Abstract
We investigated the crystal structure of an HLA-A*2402-restricted CTL epitope in the HIV-1 nef gene (Nef134-10) before (pHLA) or after TCR docking. The wild type epitope and two escape mutants were included in the study. Y135F was an early-appearing major mutation, while F139L was a late-appearing mutation which was selected in the patients without Y135F. F139 was an eminent feature of the Nef134-10 epitope. Wild type-specific TCR was less fit to F139L mutant suggesting that F139L is an escape from the CTL against the wild type epitope. Although Y135F mutation disrupted the hydrogen bond to HLA-A*2402 His70, newly formed hydrogen bond between T138 and His70 kept the conformation of the epitope in the reconstituted pMHC. TCR from Y135F- or dually-specific CTL had unique mode of binding to the mutant epitope. Y135F has been reported as a processing mutant but CTL carrying structurally adequate TCR can be found in the patients.
Similar content being viewed by others
Introduction
Cytotoxic T lymphocytes (CTL) can exert an efficient control on HIV-1 replication if HIV-1 antigens are presented on the surface of infected cells and properly recognized by the heterodimeric T cell receptors (TCR) attached to the CTL1,2,3,4. For this to occur, viral proteins synthesized in the infected cells must first be digested to fragments, transported to the rough endoplasmic reticulum (ER) and bound within a groove formed by two α-helices of the major histocompatibility complex (HLA in human) class I molecules as a 8–10-mer peptide5,6,7. The peptide-bound HLA class I molecules (pHLA) are then exported to the cell surface for recognition by TCR on the CTL cell membranes8,9. The encounter between TCR and pHLA is the fundamental requirement and initial step that allows CTL to deliver their cytotoxicity.
The genetic hypermutability of HIV-1 can interfere with immune surveillance by CTL by introducing amino acid changes that allow the virus to “escape” immune recognition. Escape phenomena may occur at different steps during the process of antigen presentation and recognition. Mutations can result in changes in processing the viral proteins or in the way such that the processed peptides cannot bind to the HLA molecules or the pHLA cannot interact properly with TCR. For example, a CTL epitope in the HIV-1 nef gene, Nef134-10 (RYPLTFGWCF), is highly immunogenic in HLA-A*2402 (HLA-A24)-positive patients10,11. At a very early phase of primary HIV-1 infection, a Tyr-to-Phe mutation at the 2nd position of the Nef134-10 epitope (Y135F; Nef134-10(2F)) is frequently selected. HLA-A24 is the most prevalent HLA class I allele among Japanese12.
In an earlier study of pHLA/TCR interactions, we used double staining with Sendaivirus-derived pHLA-tetramers to differentiate three classes of CD8+ T cells in the peripheral mononuclear cells (PBMC) of HLA-A24-positive patients with chronic HIV-1 infection: Nef134-10(wt)-specific, Nef134-10(2F)-specific and dual-specific (reacting to both wt and 2F epitopes)13,14. Since the encounter between TCR and pHLA is critical for CTL activation, structural studies examining the interactions between pHLA and TCR should be highly relevant in understanding the immune response to HIV-1 infection, including viral escape from immune surveillance. However, very few crystal structures relevant to escape mutations have been solved to date. To gain insights into the battle between HIV-1 and cellular immune responses we established CTL clones representing each of the three classes of antigen specificity targeting the Nef134-10 epitope, reconstituted the pHLA/TCR interactions in vitro and solved the crystal structures. In the small number of patients without Y135F mutation, F139L mutation was selected. Since a wild type specific TCR showed a substantial affinity to the epitope with F139L mutation (Nef134-10(6L)), we also solved the crystal structure of the pHLA/TCR interaction between them.
Results
Characterization of escape mutations and specific T cell responses by chromium release and sequencing
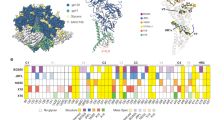
Before initiating studies on the interactions between the epitope and T cell receptors, we examined the prevalence of HLA-A24-related mutations in the Nef134-10 epitope in two independent HIV-1 cohorts: the IMSUT cohort (188 Japanese HIV-1-positive individuals in Tokyo) and the HOMER cohort (1018 HIV-1-positive individuals in British Columbia, Asian prevalence of 5%). The results confirmed the association between HLA-A24 and the Y135F mutation (Supplementary Fig. 1a, b). In both cohorts Y135F and F139L mutations were each significantly associated with the presence of HLA-A24 (p < 0.0001 and p = 0.0017, respectively). In a multicenter longitudinal acute/early infection cohort of 16 individuals expressing HLA-A24, we reconfirmed the previous finding that Y135F is selected very early after the estimated date of infection, while F139L appeared late (p = 0.0046) (Supplementary Fig. 1c).
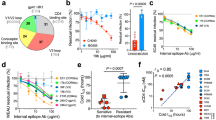
Next we established cellular clones representing dual-specific, Nef134-10(wt)-specific, Nef134-10(2F)-specific CTL from HLA-A24-positive patients in the IMSUT cohort. We showed previously that TCR repertoire of the dual-specific CD8+ T cells was highly restricted14. TRBV4-1 and TRAV8-3 gene segments were used almost exclusively as TCR β and α chains, respectively. C1-28, which used TRBV4-1 and TRAV8-3 was established and analyzed further as a CTL clone representing the dual-specific population. The TCR repertoire of the wild type specific CD8+ T cells was more diverse than the dual-specific population, but TRBV7-9 was used in more than 25% of the population. CTL clone H27-14 which used TRBV7-9 was established and represented the wild type-specific population in this study. Nef134-10(2F)-specific CD8+ T cell population was rare even after ex vivo stimulation with the cognate peptide, however, we could establish CTL clone T36-5 which represented Nef134-10(2F)-specific population. Specificities of the clones were examined by conventional chromium release assay (Fig. 1a–c).
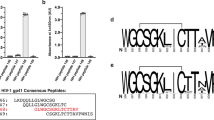
Specificities and TCR usage of the CTL clones specific to Nef134-10 epitopes.
(a) Cytolytic activities of HLA-A24-restricted Nef134-10-specific CTL clones derived from HLA-A24-positive HIV-1-infected patients. Clones H27-14 (left), T36-5 (middle) and C1-28 (right) killed HLA-A24-positive target cells pulsed with log-fold dilutions of Nef134-10 peptides, Nef134-10(wt); blue, Nef134-10(2F); red, Nef134-10(6L); orange. The effector-versus-target ratio was 5:1. (b) Vα and Vβ gene segments and CDR3 sequences of each CTL clone.
Binding characteristics of purified TCR and pHLA
The heavy chain and β2-microglobulin were expressed in E. coli and synthesized cognate peptides were refolded and pHLA was purified as previously described15. TCR derived from each CTL clone were cloned, expressed and purified from inclusion bodies of E. coli16. Refolded TCR and pHLA complexes were subjected to the surface plasmon resonance (SPR) assay (Biacore™).
Binding affinities observed between the pHLAs and TCRs are consistent with the results of the cellular killing assay (Table S3 and Supplementary Fig. 2), with the dissociation constant (KD) of the H27-14 TCR/pHLA complex increasing in the following order: A24/Nef134-10(wt) < A24/Nef134-10(6L) < A24/Nef134-10(2F). Functional TCRs against cognate epitopes have an estimated affinity ranging from 1–100 μM17. A CTL population with H27-14 TCR could recognize HIV-1-infected cells expressing the wild type A24/Nef134-10 epitope very well and could also recognize cells expressing A24/Nef134-10(6L), although less efficiently. However, H27-14 TCR affinity was too low to recognize A24/Nef134-10(2F). T36-5 TCR was highly specific against A24/Nef134-10(2F) but the TCR could recognize also with A24/Nef134-10(wt). C1-28 TCR bound almost equally with Nef134-10(wt) and Nef134-10(2F).
Kinetic analysis of H27-14 and T36-5 TCR with cognate epitopes (Table S3) yielded results that were compatible with results from equilibrium analysis. We were unable to perform kinetic analysis on C1-28 TCR due to low refolding yields.
General structures of free pHLA and TCR-pHLA
To reveal the structural differences posed by amino acid substitutions in the epitope, we determined and compared structures of A24/Nef134-10(wt), A24/Nef134-10(2F) and A24/Nef134-10(6L) before ligation to TCR (Fig. 2 and Supplementary Fig. 3a–c). The electron densities for three peptides that bound to HLA-A24 were unambiguous (Supplementary Fig. 4a–c). Large conformational changes were not seen when the Nef134-10(2F) and Nef134-10(6L) peptides in the groove of HLA-A24 were superimposed onto Nef134-10(wt) (Fig. 2a). Root mean square deviation (RMSD) values were 0.260Å and 0.259Å, respectively. Y135(P2-Y) of the Nef134-10 (wt) peptide formed a hydrogen bond with His at position 70 of the HLA-A24 (His70) and T138(P5-T) of the Nef134-10(2F) peptide formed a potential hydrogen bond with His70 (Fig. 2b).
Conformation of the three Nef134-10 peptides in the complex with HLA-A24.
(a) Overlay of three Nef134-10 peptides (Nef134-10(wt), Nef134-10(2F) and Nef134-10(6L)) bound to the HLA-A24. Nef134-10(wt), Nef134-10(2F) and Nef134-10(6L) peptides are shown blue, red and orange stick representations, respectively. HLA-A24 binding groove are not shown for clarity. (b) Interaction between the Nef134-10 peptide and HLA-A24. P2-Tyr in the Nef134-10(wt) forms a hydrogen bond with His at position 70 in the HLA-A24 (left), while P5-Thr in the Nef134-10(2F) peptide forms a hydrogen bond with His70 (right). Red dashed lines indicate hydrogen bonds. A black dashed line indicates the shortest distance between P5-Thr in the Nef134-10(wt) and His70 in the HLA-A24. (c) The side chains of P6 in the Nef134-10(wt) and Nef134-10(6L) peptides, which protrude out of the HLA-A24 binding cleft, are shown as stick and surface representations. Nef134-10(wt) (blue) and Nef134-10(6L) (orange).
Side chains of L137(P4-L) and F139(P6-F) protruded from the A24 binding groove in the A24/Nef134-10, which appeared to be the eminent feature of this epitope. The solvent-accessible surface area of the side chain of Phe(F) of the Nef134-10(wt) epitope is larger than that of Leu(L) of the Nef134-10(6L) epitope by approximately 40Å2 (188.2Å2 for P6-F, 151.9Å2 for P6-L) (Fig. 2c).
To understand how three TCRs (H27-14, T36-5 and C1-28) interact with their cognate pHLAs, we determined four TCR/pHLA complexes, including the H27-14 complexed to the agonist ligand, A24/Nef134-10(6L) (Supplementary Fig. 3d–g). The electron densities for CDR3 loops and peptide at the individual TCR/pHLA complex interfaces were unambiguous (Supplementary Fig. 4d–g).
Wild type-specific H27-14 TCR interacted with Nef134-10(wt), surrounding the F139(P6-F) residue by CDR loops (Fig. 3a, d). This complex exhibited a shape complementarity (Sc) of 0.75, which is at the higher end of the range of published TCR/pMHC17. R93α, Y98α and R31β of the H27-14 CDR rearranged to accommodate the aromatic residue of F139(P6-F) upon ligation, compared to their unligated state (Supplementary Fig. 5a, b). Y98α and R31β formed hydrogen bonds with T138(P5-T) (Supplementary Fig. 5b and Supplementary Table 3). The side chain of R93α rotated approximately 90° and formed a hydrogen bond with G100β, helping to stabilize CDRα and CDRβ at the TCR/pHLA interface.
Comparisons of interaction of TCRs with pHLAs.
Binding interfaces: (a) the H27-14 TCR-A24/Nef134-10(wt), (b) the T36-5 TCR-A24/Nef134-10(2F) and (c) the C1-28 TCR-A24/Nef134-10(2F). TCRs are shown as loop representations and the residues important for binding with the peptide are represented as stick models. Nef134-10(wt), blue; Nef134-10(2F), red; water molecules, green spheres; hydrogen bonds, red dashed lines; water-mediated hydrogen bonds, blue dashed lines. pHLA surface representations with the binding footprint of TCRs: (d) H27-14 TCR with the A24/Nef134-10(wt), (e) T36-5 TCR with the A24/Nef134-10(2F) and (f) C1-28 TCR with the A24/Nef134-10(2F). The surfaces of pHLA interacted with TCR are shown in green for Vα and yellow for Vβ. The CDRs on the pHLAs are represented as black loops. HLA-A24, grey; Nef134-10(wt), blue; Nef134-10(2F), red.
In the T36-5-A24/Nef134-10(2F) complex, the A96 residue of CDR3β formed a hydrogen bond with the W141(P8-W) residue of Nef134-10(2F) upon ligation (Fig. 3b). The T36-5 TCR interacted with A24/Nef134-10(2F) over a wide range, with a BSA approximately 200 Å2 higher than that of the other two complexes solved in this study (Fig. 3e and Supplementary Table 4) and with the highest affinity of the three TCRs in this study. The side chains of Y92α and H98β of the T36-5 TCR were packed against each other in the unligated state, but interacted with Nef134-10(2F) upon ligation (Supplementary Fig. 5c). Upon ligation the side chain of Q94α of the CDR3 loops moved 3.7 Å toward HLA-A24-α1 helix and contacted with the 135F(P2-F) carbonyl via a water molecule, while CDRβ loops (E30, A96 and H98) formed five hydrogen bonds with the amino acids (F6, G7, W8 and C9) located at the C-terminus of Nef134-10(2F) peptide.
In the C1-28-A24/Nef134-10(2F) complex, the CDR1α and 3β loops were located mainly above the Nef134-10(2F) peptide (Fig. 3c), whereas the CDR3α and 3β loops were located above the peptide in both H27-14-TCR-A24/Nef134-10(wt) and T36-5-TCR-A24/Nef134-10(2F) complexes (Fig. 3a, b). The side chains of Y32α and Y102α and the CDR3β loop surrounded the F139(P6-F) residue of the Nef134-10(2F) peptide. Water molecules acted as a “molecular glue” to stabilize the interface of the C1-28 TCR-A24/Nef134-10(2F) complex, but with only two hydrogen bonds between TCR and peptide in the C1-28-A24/Nef134-10(2F) complex, compared with four hydrogen bonds in the H27-14-A24/Nef134-10(wt) complex and five hydrogen bonds in the T36-5-A24/Nef134-10(2F) complex (Supplementary Table 3).
G28α and G98β formed hydrogen bonds with the side chains of the R134(P1-R) and C142(P9-C) residues, respectively. Surprisingly, the Vα domains, compared to Vβ domains, contributed more (~80%) in the interaction with A24/Nef134-10(2F) and were critical in the recognition of primarily the N-terminal part of the A24/Nef134-10(2F) by C1-28 TCR (Fig. 3c, f).
Wild type-specific TCR against late/minor mutant
H27-14 was slightly weaker in recognizing A24/Nef134-10(6L) than in recognizing A24/Nef134-10(wt) (Fig. 1a and Table S3). To evaluate the differences in binding ability to A24/Nef134-10(wt) or A24/Nef134-10(6L), we compared the structures of H27-14-A24/Nef134-10(wt) and H27-14-A24/Nef134-10(6L) (Fig. 4a). Conformational differences were minimal, with RMSDs of 0.485Å, 0.123Å and 0.109Å for the TCRs, Nef134-10 peptides and A24 binding clefts, respectively.
The pocket of H27-14 TCR surrounding Phe/Leu at position 6 of the Nef134-10 peptide.
(a) Overlay of the H27-14 TCR-A24/Nef134-10(wt) and H27-14 TCR-A24/Nef134-10(6L). The CDR loops in the H27-14 TCR in complex with A24/Nef134-10(wt) or A24/Nef134-10(6L) are shown in cyan or yellow, respectively. The Nef134-10(wt) or Nef134-10(6L) peptides are shown in blue or orange stick models, respectively. HLA-A24 α1 but not α2 helices are shown in grey cartoon representation for clarity. (b) The side chain of Phe at P6 (stick model in blue) is surrounded by the H27-14 TCR pocket formed by the CDR loops (cyan). (c) The side chain of Leu at P6 (stick model in orange) is surrounded by the H27-14 TCR pocket formed by the CDR loops (yellow).
In H27-14-A24/Nef134-10(wt), the aromatic side chain of F139(P6-F) protruding out of HLA-A24 binding groove was accommodated by the TCR pocket formed by the CDR1 and CDR3 loops, especially with Y31α, R93α, Y98α, R31β (Fig. 4b). Interestingly, the same binding mode was found in H27-14 TCR-A24/Nef134-10(6L) (Fig. 4c). The side chain of F139(P6-F) in the Nef134-10(wt) peptide was bulky. The 139L(P6-L) side chain in Nef134-10(6L) was much smaller, resulting in a gap in the TCR pocket surrounding the 139L(P6-L) side chain in the H27-14 TCR-A24/Nef134-10(6L) complex.
The number of van der Waals contacts (<4Å) between the TCR and P6 were 27 and 13 in H27-14-A24/Nef134-10(wt) and H27-14-A24/Nef134-10(6L), respectively. Additionally, cation-pi interactions between R93α or R31β and F139(P6-F) in H27-14-A24/Nef134-10(wt) were deformed in the case of 139L(P6L). The H27-14 TCR pocket surrounding P6 was a better fit for F139(P6-F) than for 139L(P6-L). This subtle structural difference might have been reflected in the killing assay and SPR (Fig. 1a and Table S3). Taken together, these results suggested that F139L mutation could be selected weakly but substantially as a minor escape mutant under the CTL pressure against the Nef134-10 epitope.
We were able to confirm the transformation from F to L at 6th position of Nef134-10 epitope in an HLA-A24-positive patient (Supplementary Fig. 6a). F139L appeared between Days 306 to 735 and stayed until at least 2093 days after the initial visit. Notably, the 2nd position of Nef134-10 epitope was the wild type (Y) in all IMSUT cohort patients with F139L (Supplementary Fig. 6b). Although the number of patients with F139L was limited, the emergence of the mutation at position 139 suggests that the escape mechanism of the F139L mutant could be different from that of Y135F.
Interaction between mutant- or dual-specific TCRs and A24/Nef134-10(2F)
Although Y135F has been described as an example of a CTL escape mutant in both epidemiological and immunological studies10, A24/Nef134-10(2F) was recognized by T36-5 and by C1-28 (Fig. 1a and Table S3). Therefore, we decided to analyze the structure of TCR-A24/Nef134-10(2F) complexes in both mutant-specific T36-5 and dual-specific C1-28 TCRs.
Formation of T36-5-A24/Nef134-10(2F) introduced a large TCR-induced conformational change in the peptide, with insertion of H98β into a shallow pocket formed by L137(P4-L), F139(P6-F) and W141(P8-W) (Fig. 5a). Around this pocket, Y92α and Y31β were involved in the hydrophobic interactions with L137(P4-L) and F139(P6-F). In addition, H98β and S97β formed hydrogen bonds with the main chain of L137 (P4-L) and the side chain of W141(P8-W), respectively (Supplementary Table 3). As a consequence, the side chain of W141(P8-W), which was accommodated by HLA-A24 binding pocket in unligated state, moved about 5.6 Å shift toward TCR binding surface, forming a more exposed and featured peptide conformation. Thus, Nef134-10(2F) underwent large T36-5 TCR-induced conformational change affecting the main chain hydrogen bond networks within the peptide. In the unligated state, T138(P5-T) carbonyl formed two hydrogen bonds with the nitrogen atoms of G140(P7-G) and W141(P8-W); also the W141(P8-W) nitrogen formed a bond with the F139(P6-F) carbonyl (Fig. 5b). After the TCR ligation, the hydrogen bond between T138(P5-T) carbonyl and G140(P7-G) nitrogen was lost and T138(P5-T) carbonyl reformed a single hydrogen bond with W141(P8-W) nitrogen (Fig. 5c).
Interaction of T36-5 TCR with A24/Nef134-10(2F).
(a) pHLA unligated or ligated to T36-5 TCR. Nef134-10(2F) peptide of unligated pHLA, pink; the peptide ligated T36-5 TCR, red; CDRα, blue; CDRβ, green; hydrogen bonds, red dashed lines. (b) Peptide intramolecular hydrogen bonds in unligated pHLA. Color representation are the same as (a). (c) Peptide intramolecular hydrogen bonds in T36-5 TCR-A24/Nef134-10(2F) complex. Color representation are the same as (a).
C1-28 recognized Nef134-10(2F) and Nef134-10(wt) almost equally, while T36-5 recognized Nef134-10(2F) better than Nef134-10(wt). The extensive involvement of the Vα domain in the interaction of C1-28 TCR-A24/Nef134-10(2F) was unique and quite different from typical TCR-pHLA complexes (Fig. 3f). The extended tip of the non-germline-encoded CDR3α loop lay over the A24 α1-helix (65–69 residues); by contrast, the germline-encoded CDR1α loop interacted with the N-terminus of the Nef134-10(2F) peptide (Figs. 3f, 6a and Supplementary Fig. 7a). The framework residues of Vα8-3 had multiple interactions with HLA-A24: N-terminal A1α formed a hydrogen bond with E58 and R69α formed hydrogen bonds with A158, had VDW contacts with T163 and formed a water-mediated bond and salt bridges with D166 (Fig. 6a, c). In addition, CDR1α (Y27, G28, T30) and CDR2α (F51, S52) loops made multiple contacts with HLA-A24 residues (Fig. 6a, c and Supplementary Table 3). By contrast, Vβ4-1 made relatively few contacts with HLA-A24. S97 and I99 of the CDR3β interacted with HLA-A24 residues T73 and A150, respectively (Fig. 6b, d and Supplementary Table 3). In addition, S97β formed a water-mediated bond with the main chain of F139 (P6-F).
Interaction of C1-28 TCR with A24/Nef134-10(2F).
(a) Interaction of Vα and (b) Vβ with A24/Nef134-10(2F). CDRα, cyan; Fwα (Frame work region of Vα), black; CDR3β, yellow; Nef134-10(2F), red; HLA-A24, grey, water molecules, green spheres; red dashed lines, hydrogen bonds; a green dashed line, a salt bridge. Van der Waals contact (<4.0 Å), which is represented as a black dashed line, is shown in (b), but not in (a) for clarity. (c) Interactions of Vα and (d) Vβ with A24/Nef134-10(2F). Red, blue, green and black solid lines indicate hydrogen bonds, water-mediated hydrogen bonds, a salt bridge and van der Waals contact, respectively. Fwα residues (A1 and R69) are highlighted in grey. (e) For the C1-28 TCR, interactions of the tip of the CDR3α loop (residues 96-101, cyan) with S55β and I56β residues of the Vβ4-1 segments (yellow) and G65 and A69 residues of the HLA-A24 (grey). Red dashed lines indicate a hydrogen bond.
Combination bias in the TCR variable gene segments used in dual-specific CTL
We showed previously that Vα8-3 and Vβ4-1 were public TCRs used most frequently in the dual-specific CD8+ T cell population14. Many of the interactions with HLA-A24 described above could explain the bias for Vα8-3. We looked for the molecular clues for an exclusive use of Vβ4-1. Compared to the Vβ chains of H27-14 and T36-5 TCRs, Vβ4-1 sequences coded by the germline gene segments did not have much interaction with A24/Nef134-10(2F). The CDR2β loop in Vβ4-1 was displaced outside of HLA-A24 α1-helix by the CDR3α (Supplementary Fig. 7a, b). We reported previously that the GI (Gly-Ile) motif was present at position 98-99 of CDR3β loop in most dual-specific CD8+ T cell repertoire14. Interestingly, G98β formed a hydrogen bond with C142(P9-C) of the Nef134-10(2F) peptide and hydrophobic I99β interacted with hydrophobic A150 of the HLA-A24 (Fig. 6b, d and Supplementary Table 3).
HLA class I amino acid residues 65, 69 and 155, referred to as the restriction triad, are crucial for TCR recognition18,19. Although the types of contacts differed, all three TCRs solved in this study interact with the restriction triad (Supplementary Table 3). The tips of the CDR3α loop (residues 96-101) were sandwiched between HLA-A24 (G65 and A69) and Vβ (S55 and I56) in C1-28-A24/Nef134-10(2F); G98α and S55β formed hydrogen bonds with K68 of the HLA-A24 and S101α of the CDR3α, respectively (Fig. 6e). The presence of both S55β and I56β is unique to germline-encoded TRBV4-1 gene segment and may contribute to stabilization between the CDR3α and the HLA-A24.
Although the CDR3α sequences (97–100 residues) of the public TCRs varied among dual-specific CTL clones14, the sequences were composed of small amino acids such as Gly and Ser. For insertion into an interface between G65-A69 of the HLA-A24 and S55β-I56β and formation of stable interactions between the HLA-A24 complex and CDR3α residues, small size would be beneficial. Vβ4-1 may have been co-selected by geometric constraints imposed by Vα8-3 germline and hypervariable CDR3α sequences.
The plasticity of Nef134-10 peptides
In the interactions with HLA complexes, Nef134-10 peptides assumed an M-shaped conformation similar to the structure of the cancer-related telomerase peptide presented by HLA-A24 (Fig. 2a and 7a, b)15. The side chain of T138(P5-T) faced toward the groove of HLA-A24 and functioned as the secondary anchor residue (Fig. 2b). By switching the hydrogen bond from Y135(P2-Y)/H70 to T138(P5-T)/H70, the Y135(P2-Y) to 135F(P2-F) mutation maintained the conformation of the secondary anchor to HLA-A24 α1 helix. After TCR ligation, the C-terminus shifted more than the N-terminus in H27-14 TCR-A24/Nef134-10(wt) and T36-5 TCR/A24/Nef134-10(2F) (Fig. 7c). The main chain of F139(P6-F) shifted 2.81Å in H27-14 TCR-A24/Nef134-10(wt) and G140(P7-G) shifted 2.47Å in T36-5 TCR-A24/Nef134-10(2F). By contrast, C1-28 TCR-A24/Nef134-10(2F) had less shift in the main chain of the P5-P8 (0.32-0.74 Å) residues and greater shift of the P2-P4 residues (1.06–1.68 Å) in the N-terminus.
Structural change of the Nef134-10 peptides in free and TCR-bound states.
(a) Free Nef134-10(wt) (black) and Nef134-10(wt) bound to the H27-14 TCR (blue). (b) Free Nef134-10(2F) (black), the Nef134-10(2F) bound to T36-5 TCR (red) and the Nef134-10(2F) bound to the C1-28 TCR (green). (c) The migration length of the Nef134-10 main chain after TCR ligation, when HLA-A24 binding groove is superimposed.
Discussion
Nef134-10 epitope (RYPLTFGWCF) was a highly immunogenic epitope restricted by HLA-A2410,14,20. Y135F was the major escape mutation which appeared early in the clinical course (Supplementary Fig. 1a–c). Our previous study suggesting that Y135F was a processing mutation was later confirmed by others10,21. HIV-1 with Y135F mutation has been accumulating in the population with high HLA-A24 prevalence such as Japanese. Hence, Y135F is the ultimate escape mutation which appears early in the clinical course. F139L was a minor but also HLA-A24-related mutation. Intriguingly, all 6 patients with F139L mutation kept the wild type residue at 135th position in the epitope.
According to the crystal structure of the free wild type, A24/Nef134-10 epitope took M conformation epitope. L137(P4-L) and F139(P6-F) were the eminent feature of this epitope and Y138(P5-T) was located in the valley of L137(P4-L) and F139(P6-F). N-terminal anchor, Y135(P2-Y), formed a hydrogen bond with His70 of HLA-A24 molecule. Although Y135F mutation interrupted this hydrogen bond, T138(P5-T) acted like a subanchor forming a hydrogen bond with HLA-A24 His70 keeping the M conformation very similar to the wild type. It should be noted that F is the 2nd best N-terminal anchor residue after Y for HLA-A2422,23. F139L mutation did not cause gross conformational change in the epitope, however, the solvent accessible surface area of the side chain became substantially smaller by the mutation. Namely, the epitope became slightly featureless by F139L mutation.
H27-14 TCR was highly specific to the wild type epitope (KD = 9.7 ± 0.7). Crystal structure of the complex was typical TCR/pMHC interaction in which CDR loops of α and β chains contributed almost equally to interact with the eminent feature of the epitope, F139(P6-F) residue. The binding affinity against early/major Y135F mutation diminished to nonfunctional level (KD = 291.5 ± 63.8). Although H27-14 TCR kept moderate affinity against F139L (KD = 64.8 ± 3.4), its TCR pocket was a better fit for bulky side chain of F139(P6-F) than for small side chain of 139L(P6-L). These findings may suggest that CTL with H27-14 TCR clonotype could contribute to eliminate HIV-1 with the wild type epitope but might be responsible for selection of viruses with the late/minor F139L mutation in vivo.
Nef134-10(2F)-specific T36-5 TCR had a very high affinity to A24/Nef134-10(2F) and extensive interaction shown by the BSA (approximately 2200 Å2). The side chain of 141W(P8-W) was accommodated by the HLA-A24 binding cleft in the undocked state; however, it was apparently lifted up by T36-5 TCR. Fujiwara et al. described CTL clones which recognized A24/Nef134-10(2F) more efficiently than the wild type (A24/Nef134-10(wt)). T36-5 may be one of those clones21.
In the great majority of HLA-A24-positive chronically infected patients, plasma viruses had Y135F mutation, while the patients harbored dual specific CD8+ T cell population with highly restricted TCR repertoire14. The dual-specific CD8+ T cell population expressed higher activation markers such as PD1 than the wild type-specific CD8+ T cell population (Kawana-Tachikawa, A., unpublished observation). Puzzled by the presence of dual-specific CD8+ T cell population stimulated by the epitope with the processing mutation, we wished to study the molecular interaction between the TCR and pMHC. C1-28 was chosen for the structural study as the representative of the dual-specific CD8+ T cell population. As far as we know, this is the first report of the crystal structure of Vα8-3.
According to the structural analysis of dominant public TCRs, germline residues either of the Vβ or Vα may play a major role24,25. In C1-28 TCR, Vα domains contributed predominantly (80%) in the interaction with A24/Nef134-10(2F). N-terminal A1α, R69α, residues in CDR1α (Y27, G28, T30) and CDR2α (F51, S52) had multiple interactions with HLA-A24. Nongermline CDR3α loop (residues 96-101) also contributed to interact with the two residues of restriction triad in HLA-A24, while S101α had a hydrogen bond with S55β, a germline residue of public Vβ4-1. Therefore, hypervariable CDR3α contributed to bridge the binding of HLA-A24 and Vβ4-1. In terms of the interaction with the Nef134-10(2F) peptide, germline-encoded CDR1α loop (G28α) interacted with the N-terminus of the peptide 134R(P1-R). Also, hydroxyl groups of Y32 of CDR1α and Y102 of non-germline-encoded CDR3α made water-mediated bonds with the main chain of the peptide, while both aromatic rings made hydrophobic interaction with the aromatic ring of F139 (P6-F). Thus, CDR1α and CDR3α provided a major role in interaction with the Nef134-10(2F) in addition to the interaction with HLA-A24.
Compared to the other two TCRs examined in this study, the role of Vβ4-1 of the C1-28 TCR in the recognition of HLA-A24 and the peptide was not impressive. The CDR2β loop was displaced outside of the HLA-A24 α1-helix (residues 65–69) by the CDR3α of Vα8-3. However, S55β and I56β, germline residues unique to Vβ4-1, could contribute to stabilize the tip of CDR3α, which had an important interaction with the restriction triad. Thus, the major cause of the co-selection of Vβ4-1 as a public TCR could be the bias brought by the extensive interaction of the germline Vα8-3 residues with HLA-A24. In addition, the interaction of the small residues such as Gly and Ser in CDR3α with restrictive elements in the HLA-A24 contributed to the selection of Vβ4-1. Non-germline CDR3β contributed to the interaction with HLA-A24 (T73 and A150) and C142(P9-C) of the peptide. Collectively, these structural analyses revealed that T36-5 and C1-28 TCRs have their own unique mode of binding to a mutated epitope. The three TCRs studied did not introduce a substantial conformational change in the N-terminal side of the peptide; however, two TCRs with antigen preference but not the dual-specific C1-28 TCR introduced a large conformational change in the C-terminal side.
Recently, TCR/pMHC structure of KK10 epitope restricted by HLA-B*2705 was reported26. Clone C12C was cross-reactive to both wild type and early appearing Leu268Met mutant. Although both C12C and C1-28 were dually specific to both wild type and early mutant, the character of early mutation and their TCR/pMHC structure were quite different. In the case of KK10 epitope restricted by HLA-B*2705, Leu268Met was an early mutation affecting TCR recognition and the ultimate mutations such as Arg264Lys disrupting antigen presentation follow later. However, in the case of Nef134-10 epitope restricted by HLA-A24, early Y135 mutation was the ultimate mutation. Although the number of patients are still limited, late mutation in Nef134-10 epitope, F139L, occurred only in the patients without early/ultimate Y135F mutation. It is interesting to note that HLA-B*27 is a protective but HLA-A24 is not for the disease progression after HIV-1 infection27. Although further study is needed, the different mode of mutant appearance between HLA-B*2705/KK10 and HLA-A24/Nef134-10 epitopes may be related to the role of the HLA alleles on the disease progression.
In the population with high HLA-A24 prevalence, HIV-1 with the wild type Nef134-10 epitope is replaced by the virus with Y135F mutation early after infection. Or HIV-1 with Y135F may infect HLA-A24-positive people. Whichever the case, HIV-1 with Y135F mutation replicate for many years in the presence of activated dual-specific CTL. It is tempting to speculate that the Y135F mutation could be an example of “stealth mutation.” HIV-1-infected cells might not be detected by specific- or dual-specific CTL due to processing failure or excess processing. If there is a difference in antigen processing between professional antigen presenting cells and infected T cells, CTL with functional TCR might be kept activated through cross-priming by professional antigen-presenting cells28,29. If this is the case, the immune system cannot see the infected cells as targets but may be kept activated by the spurious epitope presented by the uninfected professional antigen presenting cells. Although this hypothesis must be proven by further studies, our findings support the possibility that drugs which alter virus-peptide processing may have potential as “therapeutic vaccines.”
Methods
Approval of the study and recombinant DNA experiments in IMSUT
Plasma samples from HIV-1-positive patients attending the hospital affiliated with the Institute of Medical Science, the University of Tokyo (IMSUT) were collected and kept frozen until use. Patients provided written informed consent and the study was approved by the Institutional Review Board of the University of Tokyo (approval number 20–31). Recombinant DNA experiments used in this work were approved by the Institutional Review Board (approval number 08–30).
British columbia HOMER cohort
Founded in 1996, the British Columbia HOMER cohort is an open cohort of antiretroviral-naïve, chronically HIV-1 infected individuals. The cohort is predominantly Caucasian. Plasma HIV-1 RNA sequencing and HLA class I sequence-based typing have been performed in the HOMER cohort as described30. Here, we investigated the relationship between HLA-A*24 expression and sequence variants at Nef codons 135 and 139 in 1018 HOMER participants with HIV-1 Nef and HLA-A data available.
Longitudinal acute/early infection cohort
A total of 16 HLA-A*24 expressing individuals from a longitudinal multicenter acute/early HIV-1 infection cohort were investigated to determine the time course of selection of sequence variants at Nef codons 135 and 139 using Kaplan-Meier methods as described in11. “Time to escape” was defined as the number of days elapsed between estimated infection date and first detection of the escape variant (as a full or partial amino acid change).
Sequencing of autologous viruses
Viral RNA from EDTA-treated plasma was isolated using the QIAamp viral RNA Mini kit (QIAGEN). For heparin-treated plasma, High Pure Viral Nucleic Acid Kit (Roche) was used for RNA isolation to remove the inhibitory effect of heparin on the PCR assay. HIV-1 nef region was amplified from the RNA using the Superscript III one-step RT-PCR system with Platinum Taq DNA polymerase with High Fidelity (Invitrogen) and nef specific primers. The second-round DNA-PCR was done with EX Taq DNA polymerase Hot Start enzyme (Takara). The sequences of primers for the above PCR reactions are available upon request. Purified PCR products were directly sequenced by using BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) on an ABI 3130xl Genetic Analyzer.
Peptides
Synthetic peptides of Nef134-10(wt) [RYPLTFGWCF], Nef134-10(2F) [RFPLTFGWCF] and Nef134-10(6L) [RYPLTLGWCF] were purchased from Sigma-Genosys.
Generation of CTL clones
Nef134-10-specific CTL clones were established from peripheral mononuclear cells (PBMCs) derived from HIV-1 infected individuals carrying the HLA-A*2402, as previously described13.
Sequencing of T-cell receptor α- and β-chains
Analysis of genes encoding TCR α- and β-chains from cloned CTL were done as previously described14.
51Cr release assay
Cytotoxicity was measured by a standard 51Cr release assay as previously described10.
Protein expression and purification
The H27-14, T36-5 and C1-28 TCRs were expressed, refolded and purified essentially as described16. For the H27-14 TCR refold, 24 mg of solubilized TCR α-chains and 20 mg of β-chains were injected into 1 L of a buffer containing 5 M urea, 100 mM Tris, pH 8.5, 400 mM L-arginine-HCl, 3.7 mM cystamine, 6.6 mM cysteamine, 0.2 mM PMSF (phenylmethylsulfonyl fluoride) at 4°C. The refolding solution was dialyzed twice for 24 h against 10 vol of milli Q water and then 10 vol of 10 mM Tris, pH 8.5 at 4°C. The refolded TCR was then purified by Resource-Q column and Superdex 75 column (GE Healthcare). For the T36-5 TCR, 50 mg of α-chains and 40 mg of β-chains were refolded and purified as described above. For the C1-28 TCR refold, 20 mg of TCR α-chains and 35 mg of β-chains were injected twice into 2 L of a buffer containing 5 M urea, 100 mM Tris, pH 8.9, 400 mM L-arginine-HCl, 2 mM EDTA, 5 mM reduced glutathione, 0.5 mM oxidized glutathione, 0.2 mM PMSF at 4°C. After 24 hr incubation, the refolding solution was dialyzed twice for 24–36 h against 5 mM Tris, pH 8.5, 50 mM NaCl and then 10 mM Tris, pH 8.5, 50 mM NaCl at 4°C. The resultant was purified by Resource-Q column, Mono-Q column and Superdex 75 column (GE Healthcare).
Preparation of pHLAs
The HLA-A*2402 heavy chain, HLA-A*2402 heavy chain-BSP (BirA substrate peptide) and β2 microglobulin (β2m) were also expressed separately in E. coli, as described15. For a 1 L refold, 45 mg of solubilized HLA-A*2402 heavy chains and 15 mg of β2m were injected into a refold buffer containing 100 mM Tris, pH 8.0, 400 mM L-arginine-HCl, 2 mM EDTA, 5 mM reduced glutathione, 0.5 mM oxidized glutathione, 0.2 mM PMSF in the presence of 10 mg of HIV-1 Nef 134-10 (wt), Nef134-10(2F) or Nef134-10 (6L) peptide. The refolded protein was purified by Superdex 75 column and MonoQ column. For SPR analysis, pHLA molecules were refolded in the same way as the above, using the HLA-A*2402-BSP instead of the HLA-A*2402 heavy chain. The pHLA-BSP was biotinylated as previously described14.
Surface plasmon resonance
Surface plasmon resonance experiment was carried out at 25°C using BIAcore 2000 in a buffer containing 10 mM HEPES, pH 7.4, 150 mM NaCl, 3 mM EDTA, 0.005% Surfactant P20. Biotinylated pHLAs were immobilized to Sensor chip SA until the response was reached between 200–800 response units (RU). Each TCR was injected over the flow cells at a flow-rate of 20–30 μl/min with an indicated concentration range for equilibrium analysis. BIAevaluation software (version 4.1; GE healthcare) was used for data analysis. For kinetic analysis, 1:1 Langmuir binding model was used to calculate the Kon and Koff values.
Crystallization and data collection
All crystallizations were done by the sitting drop vapor diffusion method with a protein/reservoir drop ratio of 1:1 at 20°C. Crystallization conditions of grown crystals are shown in Supplementary Table S3. For cryoprotection, obtained crystals were soaked briefly and sequentially in reservoir solutions containing 10% and 20% ethylene glycol and then flash-frozen in liquid nitrogen. Data were collected at the beamline BL41XU in SPring 8 (Hyogo, Japan) and BL-5A, NW12A and BL1A in the PF facility (Tsukuba, Japan) and processed with HKL200031 and the CCP4 program suite32.
Structure determination and refinement
The structures were determined by molecular replacement using Molrep33. Search models used for molecular replacement were shown in Supplementary Table S3. Model building and refinement were conducted using Coot34 and CNS 1.335, respectively. The further rounds of these refinements were performed using REFMAC implemented in CCP4. For building model of the T36-5 TCR-A24/Nef134-10(2F), diffraction intensities from the various crystals exhibited twin with an estimated twinning fraction of 0.45–0.49. Therefore, this structural model was refined using CNS 1.3 as a perfect twin.
The stereochemistry of the refined models was assessed with program Rampage36. All molecular graphics representations were created with the program PyMOL (DeLano Scientific; http://www.pymol.org). Data collection and refinement statistics are shown in Supplementary Table 2.
References
Kappler, J. et al. The major histocompatibility complex-restricted antigen receptor on T cells in mouse and man: identification of constant and variable peptides. Cell 35, 295–302 (1983).
Marrack, P., Shimonkevitz, R., Hannum, C., Haskins, K. & Kappler, J. The major histocompatibility complex-restricted antigen receptor on T cells. IV. An antiidiotypic antibody predicts both antigen and I-specificity. J Exp Med 158, 1635–1646 (1983).
McMichael, A. J. & Rowland-Jones, S. L. Cellular immune responses to HIV. Nature 410, 980–987 (2001).
Walker, B. D. & Burton, D. R. Toward an AIDS vaccine. Science 320, 760–764 (2008).
de la Salle, H. et al. Homozygous human TAP peptide transporter mutation in HLA class I deficiency. Science 265, 237–241 (1994).
Fruh, K. et al. A viral inhibitor of peptide transporters for antigen presentation. Nature 375, 415–418 (1995).
Maeurer, M. J. et al. Tumor escape from immune recognition: lethal recurrent melanoma in a patient associated with downregulation of the peptide transporter protein TAP-1 and loss of expression of the immunodominant MART-1/Melan-A antigen. J Clin Invest 98, 1633–1641 (1996).
Bjorkman, P. J. et al. Structure of the human class I histocompatibility antigen, HLA-A2. Nature 329, 506–512 (1987).
Davis, M. M. & Bjorkman, P. J. T-cell antigen receptor genes and T-cell recognition. Nature 334, 395–402 (1988).
Furutsuki, T. et al. Frequent transmission of cytotoxic-T-lymphocyte escape mutants of human immunodeficiency virus type 1 in the highly HLA-A24-positive Japanese population. J Virol 78, 8437–8445 (2004).
Brumme, Z. L. et al. Marked epitope- and allele-specific differences in rates of mutation in human immunodeficiency type 1 (HIV-1) Gag, Pol and Nef cytotoxic T-lymphocyte epitopes in acute/early HIV-1 infection. J Virol 82, 9216–9227 (2008).
Itoh, Y. et al. High-throughput DNA typing of HLA-A, -B, -C and -DRB1 loci by a PCR-SSOP-Luminex method in the Japanese population. Immunogenetics 57, 717–729 (2005).
Kawana-Tachikawa, A. et al. An efficient and versatile mammalian viral vector system for major histocompatibility complex class I/peptide complexes. J Virol 76, 11982–11988 (2002).
Miyazaki, E. et al. Highly restricted T-cell receptor repertoire in the CD8+ T-cell response against an HIV-1 epitope with a stereotypic amino acid substitution. AIDS 23, 651–660 (2009).
Cole, D. K. et al. Crystal structure of HLA-A*2402 complexed with a telomerase peptide. Eur J Immunol 36, 170–179 (2006).
Boulter, J. M. et al. Stable, soluble T-cell receptor molecules for crystallization and therapeutics. Protein Eng 16, 707–711 (2003).
Rudolph, M. G., Stanfield, R. L. & Wilson, I. A. How TCRs bind MHCs, peptides and coreceptors. Annu Rev Immunol 24, 419–466 (2006).
Tynan, F. E. et al. T cell receptor recognition of a ‘super-bulged’ major histocompatibility complex class I-bound peptide. Nat Immunol 6, 1114–1122 (2005).
Burrows, S. R. et al. Hard wiring of T cell receptor specificity for the major histocompatibility complex is underpinned by TCR adaptability. Proc Natl Acad Sci U S A 107, 10608–10613 (2010).
Ikeda-Moore, Y. et al. Identification and characterization of multiple HLA-A24-restricted HIV-1 CTL epitopes: strong epitopes are derived from V regions of HIV-1. J Immunol 159, 6242–6252 (1997).
Fujiwara, M. et al. Different abilities of escape mutant-specific cytotoxic T cells to suppress replication of escape mutant and wild-type human immunodeficiency virus type 1 in new hosts. J Virol 82, 138–147 (2008).
Ibe, M. et al. Role of strong anchor residues in the effective binding of 10-mer and 11-mer peptides to HLA-A*2402 molecules. Immunogenetics 44, 233–241 (1996).
Sidney, J., Southwood, S. & Sette, A. Classification of A1- and A24-supertype molecules by analysis of their MHC-peptide binding repertoires. Immunogenetics 57, 393–408 (2005).
Kjer-Nielsen, L. et al. A structural basis for the selection of dominant alphabeta T cell receptors in antiviral immunity. Immunity 18, 53–64 (2003).
Stewart-Jones, G. B., McMichael, A. J., Bell, J. I., Stuart, D. I. & Jones, E. Y. A structural basis for immunodominant human T cell receptor recognition. Nat Immunol 4, 657–663 (2003).
Ladell, K. et al. A molecular basis for the control of preimmune escape variants by HIV-specific CD8+ T cells. Immunity 38, 425–436 (2013).
O'Brien, S. J., Gao, X. & Carrington, M. HLA and AIDS: a cautionary tale. Trends Mol Med 7, 379–381 (2001).
Lehner, P. J. & Cresswell, P. Recent developments in MHC-class-I-mediated antigen presentation. Curr Opin Immunol 16, 82–89 (2004).
Nakayama, K. et al. Imbalanced Production of Cytokines by T Cells Associates with the Activation/Exhaustion Status of Memory T Cells in Chronic HIV Type 1 Infection. AIDS Res Hum Retroviruses 28, 702–714 (2011).
Brumme, Z. L. et al. Evidence of differential HLA class I-mediated viral evolution in functional and accessory/regulatory genes of HIV-1. PLoS Pathog 3, e94 (2007).
Otwinowski, Z. & Minor, W. Processing of X-Ray Diffraction Data Collected in Oscillation Mode Macromolecular Crystallography. Methods in Enzymology 276, 307–326 (1997).
The CCP4 suite: programs for protein crystallography. Acta Crystallogr D Biol Crystallogr 50, 760–763 (1994).
Vagin, A. & Teplyakov, A. MOLREP: an Automated Program for Molecular Replacement. J Appl Cryst 30, 1022–1025 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60, 2126–2132 (2004).
Brunger, A. T. et al. Crystallography & NMR System: A New Software Suite for Macromolecular Structure Determination. Acta Cryst. D54, 905–921 (1998).
Lovell, S. C. et al. Structure validation by Calpha geometry: phi,psi and Cbeta deviation. Proteins: Structure, Function and Genetics 50, 437–450 (2003).
Acknowledgements
We thank the beam-line staffs at NW12A, BL1A and BL5A of Photon Factory (Tsukuba, Japan) and BL41XU of SPring8 (Hyogo, Japan) for technical help during data collection. We thank Drs. Richard Harrigan, Heiko Jessen, Anthony Kelleher, Martin Markowitz and Bruce Walker for specimen and/or data access. This work was supported in part by a contract research fund from the Ministry of Education, Culture, Sports, Science and Technology(MEXT)for Program of Japan Initiative for Global Research Network on Infectious Diseases (10005010)(AI); Global COE Program (Center of Education and Research for Advanced Genome-Based Medicine - For personalized medicine and the control of worldwide infectious diseases-) of MEXT (F06)(AI); JSPS KAKENHI (25293226)(AKT); Grants for AIDS research from the Ministry of Health, Labor and Welfare of Japan (H24-AIDS-IPPAN-008)(AKT); Research on international cooperation in medical science, Research on global health issues, Health and Labour Science Research Grants, the Ministry of Health, Labor and Welfare of Japan (H25-KOKUI-SITEI-001)(AI); CJB is supported by a Vanier Canada Graduate Scholarship from the Canadian Institutes of Health Research (CIHR). EM is supported by a Master's Scholarship from the Canadian Association of HIV Research and Abbott Virology. ZLB is the recipient of a CIHR New Investigator Award and a Scholar Award from the Michael Smith Foundation for Health Research.
Author information
Authors and Affiliations
Contributions
A.S. did all the experiments from protein synthesis, BIAcore assay to structural studies. A.K.-T. did all the immunological studies including establishment of CTL cell lines and characterization of CTLs. A.Y. and Y.S. did crystallographic studies. C.Y.H. and D.Z. contributed to sequencing of viruses and TCR genes and analyzed the relationship between HLA-A*2402 expression and sequence variants at Nef codons 135 and 139 in IMSUT cohort. H.N. provided the clinical data of IMSUT cohort. T.K. was responsible for the clinical management in the IMSUT hospital. J.C. analyzed the relationship between HLA-A*2402 expression and sequence variants at Nef codons 135 and 139 in longitudinal multicenter acute/early HIV-1 infection cohort. E.M. and C.J.B. analyzed the relationship between HLA-A*24 expression and sequence variants at Nef codons 135 and 139 in British Columbia HOMER cohort. Y.S. discussed the structural study described in this manuscript. G.F.G. has been collaborating with A.I. and joined the discussion for this study. Z.L.B. investigated the relationship between HLA-A*24 expression and sequence variants at Nef codons 135 and 139 in 1018 participants in British Columbia HOMER cohort and longitudinal multicenter acute/early HIV-1 infection cohort with HIV-1 Nef and HLA-A data available. S.F. managed and instructed the structural study group and provided the environment for the crystallographic study and analysis. A.I. managed all the work described and wrote the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Accession codes C1-28 TCR/Nef134-10(2F), 3VXM; A24/Nef134-10(wt), 3VXN; A24/Nef134-10(2F), 3VXO; A24/Nef134-10(6L), 3VXP; H27-14 TCR, 3VXQ; H27-14 TCR-A24/Nef134-10(wt), 3VXR; H27-14 TCR-A24/Nef134-10(6L), 3VXS; T36-5 TCR, 3VXT; T36-5 TCR-A24/Nef134-10(2F), 3VXU and 3W0W.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Shimizu, A., Kawana-Tachikawa, A., Yamagata, A. et al. Structure of TCR and antigen complexes at an immunodominant CTL epitope in HIV-1 infection. Sci Rep 3, 3097 (2013). https://doi.org/10.1038/srep03097
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03097
This article is cited by
-
Targeting of intracellular oncoproteins with peptide-centric CARs
Nature (2023)
-
Identification of TCR repertoires in functionally competent cytotoxic T cells cross-reactive to SARS-CoV-2
Communications Biology (2021)
-
RETRACTED ARTICLE: Cross-HLA targeting of intracellular oncoproteins with peptide-centric CARs
Nature (2021)
-
Computational stabilization of T cell receptors allows pairing with antibodies to form bispecifics
Nature Communications (2020)
-
Structural basis for oligoclonal T cell recognition of a shared p53 cancer neoantigen
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.