Abstract
Post-translational histone modifications play key roles in gene regulation, development and differentiation, but their dynamics in living organisms remain almost completely unknown. To address this problem, we developed a genetically encoded system for tracking histone modifications by generating fluorescent modification-specific intracellular antibodies (mintbodies) that can be expressed in vivo. To demonstrate, an H3 lysine 9 acetylation specific mintbody (H3K9ac-mintbody) was engineered and stably expressed in human cells. In good agreement with the localization of its target acetylation, H3K9ac-mintbody was enriched in euchromatin and its kinetics measurably changed upon treatment with a histone deacetylase inhibitor. We also generated transgenic fruit fly and zebrafish stably expressing H3K9ac-mintbody for in vivo tracking. Dramatic changes in H3K9ac-mintbody localization during Drosophila embryogenesis could highlight enhanced acetylation at the start of zygotic transcription around mitotic cycle 7. Together, this work demonstrates the broad potential of mintbody and lays the foundation for epigenetic analysis in vivo.
Similar content being viewed by others
Introduction
Gene expression in eukaryotes is regulated at the chromatin level, including post-translational histone modifications such as acetylation and methylation on lysine residues. Whereas the epigenomic landscape of histone modifications has been uncovered in many different cell types1, the dynamic changes of modifications in single cells and living organisms remains largely unexplored due to limitations in monitoring techniques2,3. Unlike proteins, which can be tracked in living cells with genetically fused fluorescent tags, protein modifications require a separate probe for tracking. For example, Förster/fluorescence resonance energy transfer (FRET)-based sensors have been developed to track histone modification dynamics4,5,6,7. Although these sensors make it possible to monitor the modification state mediated by the balance of modification and demodification enzymes, they cannot track the spatiotemporal dynamics of endogenous histone modifications. In addition, cell lines that stably express these constructs have not been obtained3. To visualize endogenous histone modification in living cells, we previously developed a technique based on antigen binding fragments (Fabs)8,9; however, this method requires direct loading of Fabs into cells and their dilution upon cell division prevents long-term and organism-level experiments. Here we describe a genetically encoded system that overcomes the limitations of FRET sensors and Fab to enable the tracking of histone H3 lysine 9 acetylation (H3K9ac) in living cells and organisms. Specifically, we fused a single-chain variable fragment (scFv) antibody to the enhanced green fluorescent protein (EGFP) to create a modification-specific intracellular antibody, or ‘mintbody’, that retains high specificity for H3K9 acetylation with minimal interference to cell function. We anticipate this methodology will prove useful for studying a wide range of residue-specific intracellular protein modifications in vivo.
Results
Generation of ‘mintbody’
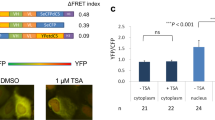
To construct H3K9ac-specific mintbodies, cDNA fragments encoding the variable regions of IgG heavy and light chains (VH and VL, respectively) were cloned from mouse hybridoma cell lines, including the one producing CMA310 that was previously used in FabLEM9 as well as two sister clones. The deduced amino acid sequences of VH and VL were highly conserved among these clones and the diversity was probably derived by somatic hypermutation (Fig. 1a). The specificity and affinity of these scFvs (designated as H3K9ac.v1, v2 and v3 for CMA310, 8A2 and 13C7, respectively) were first examined using His6-tagged proteins expressed in and purified from bacteria (Fig. 1b). Surface plasmon resonance measurements indicated that His6-scFvs are highly specific probes, binding to target H3K9ac peptides with dissociation constants at ~10−8 M, but not binding to H3K9me2 peptides (Fig. 1c). These binding affinities to H3K9ac peptide were roughly 2-orders of magnitude lower than the native IgG form of CMA310, consistent with scFv being monovalent and thus of lower affinity.
Development of scFv(H3K9ac)-EGFP probes.
(a) Amino acid sequence alignment of variable regions in heavy and light chains from three mouse monoclonal antibodies specific to histone H3K9ac. Compared to v1 (CMA310)9, v2 and v3 differ in 3 amino acids in light chain and 2 amino acids in heavy chain, respectively. (b) SDS-polyacrylamide gel analysis of purified His6-scFv(H3K9ac) proteins. Positions of size standards are shown on the left. (c) Specific binding of scFv(H3K9ac) to H3K9ac. The binding kinetics of scFv(H3K9ac) to peptides harboring H3K9ac and H3K9me2 were measured by surface plasmon resonance. The binding dissociation constants (KD; M) to H3K9ac at 25°C were shown. (d) Schematic illustration of scFv-based mintbody. VH, heavy chain variable region; VL, light chain variable region. (e) H3K9ac-mintbodies in living cells analyzed by confocal microscopy. hTERT-RPE1 cells were transfected with the expression vectors of H3K9ac-mintbodies and incubated in Hoechst33342. H3K9ac-mintbody v1 and v3 were concentrated in nuclei but excluded from nucleoli and Hoechst-dense heterochromatin. H3K9ac-mintbody v2 was distributed throughout the cell, suggesting that this probe is not functional. (f) hTERT-RPE1 cells expressing H3K9ac-mintbody v3. Cells were fixed and stained with H3K27me3 specific antibody (CMA323)9 and Hoechst33342. Arrows indicate an inactive X chromosome. Scale bars, 10 μm.
We next made mintbodies from our scFvs by fusing EGFP to their C-terminus (Fig. 1d). When expressed in human telomerase-immortalized retina pigment epithelial (hTERT-RPE1) cells (Fig. 1e,f), two of three mintbodies (v1 and v3) were concentrated in nuclei but excluded from nucleoli and Hoechst-dense heterochromatin (Fig. 1e), consistent with the euchromatic localization of H3K9ac9. This suggests that cytoplasmically expressed H3K9ac-mintbodies v1 and v3, but not v2, were functional to bind to target acetylation in living cells. Immunofluorescence with fixed cells confirmed that mintbodies were indeed excluded from hypoacetylated inactive X chromosomes, or Barr bodies, stained densely with trimethylated H3 lysine 27 (H3K27me3) and Hoechst33342 (Fig. 1f). H3K9ac-mintbody v1 and v3 behaved similarly in our preliminary experiments, suggesting both are equally useful. In what follows, v3 was used to generate stable cell lines and v1 to generate transgenic animal.
Quantitative measurement of changes in H3K9 acetylation levels in living cells
To monitor the changes in acetylation levels in living cells, we generated human osteosarcoma U2OS cells that stably express H3K9ac-mintbody. The growth rates of these cells were similar to those expressing EGFP alone (Supplementary Fig. S1), indicating that the expression of H3K9ac-mintbody at a certain level was not toxic to cells. To directly test the specific targeting of mintbodies to acetylation, we treated the cells with a histone deacetylase inhibitor, trichostatin A (TSA; 100 nM), to globally increase histone acetylation levels ~3-fold over 6 h (measured by immunoblotting; Supplementary Fig. S2a). This treatment led to a slow-down in mintbody kinetics as measured by fluorescence recovery after photobleaching (FRAP; Fig. 2a,b). Quantitative fits revealed this slowdown was due to a ~2-fold increase in the chromatin-bound fraction resulting from an enhanced binding association rate (a faster ton), as would be expected in cells with more H3K9ac target (Fig. 2b). Consistent with this, the nucleus became significantly brighter compared to the cytoplasm upon TSA treatment (Fig. 2a,c and Supplementary Video 1,2). In fact, by simply measuring the ratio of nuclear to cytoplasmic intensity, we could detect global increases in acetylation associated with as little as ~1 nM TSA, an amount not detectable by immunoblotting (Supplementary Fig. S2b). After the removal of TSA, the nuclear to cytoplasmic ratio of H3K9ac-mintbody decreased, reflecting the restoration of global acetylation levels back to the original level (Fig. 2d). Thus, H3K9ac-mintbodies accumulate reversibly in nuclei depending on the acetylation level of endogenous histone H3.
Monitoring H3K9ac levels in living cells.
(a) H3K9ac-mintbody as a measure of H3K9ac levels. The more H3K9ac levels increase, the more mintbodies bind to chromatin, resulting in more nuclear accumulation. (b) FRAP. A spot (yellow circle) was bleached in cells expressing H3K9ac-mintbody v3. FRAP curves were fitted to a reaction-diffusion model23 to obtain ton (s), toff (s) and the bound/free fraction. Averages with s.e.m. are shown (TSA, n = 19; DMSO, n = 18). (c) Nuclear concentration of H3K9ac-mintbody. Cells expressing H3K9ac-mintbody v3 were incubated in presence or absence of TSA and typical confocal images are shown on the left. Nuclear to cytoplasmic intensity ratio was measured and plotted (average with s.e.m.). The numbers of analyzed cells were 31 (vehicle), 31 (1 nM), 30 (10 nM) and 40 (100 nM). (d) After 8 hr incubation in 100 nM TSA, cells were washed and incubated in normal medium, Nuclear to cytoplasmic intensity ratio was measured and plotted (average with s.e.m; n = 32). Scale bars, 10 μm.
We next established mouse embryonic stem (ES) cells stably expressing H3K9-mintbodies (Supplementary Fig. S3a,b). A similar effect upon TSA treatment was observed in these cells (Supplementary Fig. S3c and Supplementary Video 3,4), demonstrating the potential of mintbodies for tracking H3K9 acetylation in both differentiated and undifferentiated cell types.
Visualizing H3K9ac in living organisms
To prove that mintbodies can also be used to monitor histone modification dynamics in living organisms, transgenic animals expressing H3K9ac-mintbody were generated. We first established a transgenic fly line harboring the H3K9ac-mintbody coding sequence downstream of Gal4-responsive upstream activation sequence elements (Gal4-UAS). This allowed us to control mintbody expression by crossing with a fly line harboring the driver transcription activator Gal4-VP16 under the control of tissue-specific promoters. To visualize H3K9ac dynamics during early embryogenesis, H3K9ac-mintbody was maternally supplied in eggs under the control of matα4-Gal4-VP16 driver. In very early embryos, the fluorescent signal was distributed nearly uniformly and nuclei were barely detected (Fig. 3a, 00:22; Supplementary Video 5). At around mitotic cycle 7 (00:29), at which zygotic transcription begins10,11, H3K9ac-minbodies became clearly concentrated in nuclei, consistent with a positive role of H3K9 acetylation in transcriptional activation. During later mitotic cycles, the levels of H3K9ac-mintbodies were quite homogenous in almost all nuclei up to embryonic stage 4, after which embryo morphology drastically changes (Fig. 3b; and Supplementary Video 6). Since the embryos developed normally, this data demonstrates that the expression of H3K9ac-mintbody and fluorescence imaging do not inhibit embryogenesis.
Monitoring H3K9ac during Drosophila embryogenesis.
(a) Time-laps imaging of H3K9ac-mintbody v1 during Drosophila embryo development at earlier stages. Maximum intensity projection images (the first row) are shown with z-axis optical sections (the second to the seventh rows). Nuclei become visible at 0:29 (z = 1–4). (b) Maximum intensity projection images of H3M9ac-mintbodies during Drosophila embryo development at later stages. Scale bars, 50 μm.
Besides monitoring changes in global levels of H3K9 acetylation in nuclei, H3K9ac-mintbody can also be used for imaging the enrichment of H3K9ac in discrete regions of chromatin. To demonstrate, we visualized Drosophila polytene chromosomes, where genes are organized into distinct bands. When H3K9ac-mintbodies were expressed in the salivary glands of Drosophila larvae, the banding pattern of polytene chromosomes was observed (Fig. 4a; Supplementary Fig. S4; and Supplementary Video 7). Staining with Hoechst33342 in fixed cells confirmed H3K9ac-mintbodies were generally enriched in euchromatic bands (Fig. 4b; and Supplementary Video 8), as observed by immunofluorescence using chromosome spreads12. The relative intensities were however varied in different bands and chromocenter heterochromatin was devoid of the probe. These results were again consistent with the view that histone H3 acetylation marks potentially active chromatin, even though it may not be directly correlated with ongoing transcription.
Monitoring H3K9ac in living organisms.
(a) Localization of H3K9ac-mintbody v1 on polytene chromosomes in salivary gland. Left and middle; bright-field and EGFP images of the whole salivary gland, respectively. Scale bar, 100 μm. Right; magnified image of the nucleus boxed in the middle panel. Scale bar, 10 μm. (b) Localization of H3K9ac-mintbody v1 with respect to Hoechst33342 staining in fixed salivary gland. Individual and merged confocal images are shown with magnified views (1,2). Heterochromatin is depleted in H3K9ac (red arrows). Some bands are particularly depleted or enriched in H3K9ac (red or green arrowheads). Scale bar, 10 μm. (c) H3K9ac-mintbody v1 in zebrafish. Boxed area is magnified in inset. Scale bar, 50 μm.
Finally, we also generated transgenic zebrafish, in which H3K9ac-mintbody was expressed under the control of UAS promoter in somatic cells harboring a ubiquitously expressing transcriptional activator Gal4FF [SAGFF(LF)73A]13. The fish developed normally and remained fertile (Fig. 4c; and Supplementary Video 9), indicating that mintbody can be used for in vivo imaging of H3K9ac also in vertebrates.
Discussion
We here demonstrated that H3K9ac-mintbodies can track H3K9ac acetylation in living cells and organisms. Critically, the successful generation of transgenic animals proves the H3K9-mintbody does not significantly interfere with cell division, development and differentiation, even though it specifically targets to H3K9ac. This is probably because H3K9ac-mintbodies bind their targets only for several seconds at a time, so cellular proteins can maintain access to common target modifications. We exploited this dynamic binding property to monitor changes in global H3K9 acetylation levels, both by FRAP and by measuring the nuclear-to-cytoplasmic ratio. At higher H3K9ac levels, more mintbodies bind to chromatin, resulting in more nuclear accumulation. In addition to the measurement of global H3K9ac levels, mintbodies can also detect spatial gradients in H3K9ac within a nucleus, such as the euchromatic banding patterns on Drosophila polytene chromosomes. In a future study, changes in H3K9 acetylation on specific loci may be visualized upon gene activation14.
Recently, extracellular glycan has also been visualized using a similar technique to the one developed here. Specifically, the secreted form of heparane sulfate-specific scFv tagged with GFP was used to track the glycan modification on the outer surface of cells15. A key difference between this technique and ours, however, is that mintbodies are not secreted. Thus, unlike natural antibodies, mintbodies must remain within cells, where the reducing environment can interfere with scFv folding and lead to aggregation16. Indeed, one of our H3K9ac-mintbody (v2) was not functional when cytoplasmically expressed. Fortunately, this problem can be overcome by establishing multiple hybridoma clones, as we did here, since only a few amino acid substitutions can greatly influence scFv activity. Alternatively, heavy-chain IgG from Camelidae sp might also be useful to generate mintbodies17.
In conclusion, mintbodies will be a powerful tool for studying epigenetic regulation during development, differentiation and pathogenesis in living organisms. Furthermore, they will enable high-throughput screens for the development of epigenetic drugs such as histone acetylase and deacetylase inhibitors18.
Methods
Construction of mintbody expression vectors
Total RNA was extracted from 1–5 × 106 hybridoma cells9,19 using TRIzol (Life Technologies) and reverse transcribed (1 μg in a reaction volume of 50 μl) using oligo-dT primer according to the manufacturer's protocol (SuperScriptIII; Life Technologies). The cDNAs encoding VH and VL were amplified by PCR (PrimeStar; TaKaRa), using previously reported primers20 with a slight modification; 20 and 15 nucleotides from 5′ termini were omitted from MHV.BACK primers and MKV.FOR/MLV.FOR primers, respectively. The amplified products of VH and VL were purified (QIAquick Gel Extraction Kit; Qiagen) and connected into a single chain variable fragment (scFv) by PCR using a linker sequence20. The amplified scFv fragments were purified (Qiagen) and ligated into pSC-B using a kit (StrataClone PCR Cloning Kit; Stratagene) and subjected to nucleotide sequencing. The deduced amino acid sequence agreed with the mass spectrometry profile of purified IgG peptides. scFvs were exercised from pSC-B with EcoRI and inserted into pEGFP-N2 (Clonetech) that was digested with the same enzyme. Nucleotide sequence data of scFvs (H3K9ac.v1, v2 and v3) are available in the DDBJ/EMBL/GenBank databases under the accession numbers AB793785, AB793786 and AB793787, respectively.
Bacterial expression of recombinant scFv and biochemical analysis
scFv sequence was subcloned into pET15b (Novagen) which harbors the His6 tag at the N-terminus. Each His6-scFv was expressed in Escherichia coli BL21 (DE3). The cells producing His6-scFv were collected by centrifugation and sheared by sonication in 50 mM Tris-HCl (pH 7.5) buffer containing 500 mM NaCl, 1 mM PMSF, 10% glycerol and 0.01% Triton X-100. Insoluble materials collected by centrifugation were denatured in 50 mM Tris-HCl (pH 8.0) buffer containing 7 M guanidine hydrochloride, 500 mM NaCl and 5% glycerol. After centrifugation, supernatants containing the His6-scFv were mixed with 4 ml (50% slurry) of Ni-NTA agarose (Qiagen) and rotated at 4°C for 1 h. The beads were packed into an Econo-column (Bio-Rad) and washed with 50-column volume of 50 mM Tris-HCl (pH 8.0) buffer containing 500 mM NaCl, 6 M urea, 5% glycerol and 5 mM imidazole. His6-scFv was eluted with a 50-column volume linear gradient of 5–500 mM imidazole in 50 mM Tris-HCl (pH 8.0) buffer containing 500 mM NaCl, 6 M urea and 5% glycerol. The peak fractions were dialyzed against PBS. His6-scFv was further purified by Superdex 75 gel filtration column (HiLoad 16/60 prep grade, GE Healthcare Biosciences) chromatography. The purity was checked by 16% SDS-polyacrylamide gel electrophoresis. Surface plasmon resonance (SPR) signals were measured using a Biacore X100 instrument (GE Healthcare Biosciences). Flow cells were maintained at 25°C during the measurement and the instrument was operated at the mid-flow rate (~30 μl/min). Bovine serum albumin-conjugated synthetic peptides (H3K9ac: ARTKQTAR(ac-K)STGGKAPRKQC and H3K9me2: ARTKQTAR(me2-K)STGGKAPRKQC) were conjugated to the activated surface of CM5 sensor chips (GE Healthcare Biosciences), using the standard amine coupling conditions recommended by the manufacturer. The flow cell containing a sensor chip without histone peptide was used as a reference. The running buffer was PBS containing 0.005% Tween 20. Before each binding assay, the sensor chip was regenerated by a 1-min wash with 10 mM NaOH followed by a 1-min wash with 10 mM glycine (pH 2.0). For each binding assay, different concentrations of His6-scFv proteins (6.25, 12.5, 25, 50 and 100 nM) were successively injected for 2 min without regeneration.
Cells
hTERT-RPE1 and U2OS cells were routinely grown in DME (Dulbecco's modified Eagle's medium; Nacalai Tesque) supplemented with L-glutamine/penicillin/streptomycin (2 mM L-glutamine, 100 U/ml penicillin, 0.1 mg/ml streptomycin; Sigma-Aldrich) and 10% fetal calf serum (FCS). Cells were transfected with mintbody plasmids using Fugene HD (Promega), according to the manufacturer's instruction and those stably expressing mintbody were selected in 1 mg/ml G418 (Nacalai Tesque).
Mouse embryonic stem (ES) cell line derived from C57BL/6 was grown on fibroblast feeder cells in DME (Dulbecco's modified Eagle's medium; Nacalai Tesque) supplemented with L-glutamine/penicillin/streptomycin (2 mM L-glutamine, 100 U/ml penicillin, 0.1 mg/ml streptomycin; Sigma-Aldrich), 10% FCS, non-essential amino acids (Life technologies), sodium pyruvate (Life technologies), leukemia inhibitory factor (Millipore) and 2-mercaptoethanol (Life technologies). To generate ES cells expressing mintbody, targeted homologous recombination into the ROSA locus was used21. Briefly, H3K9ac-mintbody cDNA fragment was subcloned into the NheI/SacI site of pBigT vector22, which harbors the neomycin resistant gene flanked by FRTs. The FRT-Neo-H3K9ac-mintbody-FRT cassette was exercised with PacI and AscI and subcloned into the ROSA26 vector. The targeting scheme is shown in Supplementary Fig. S3. The targeting vector was linearized with XhoI and introduced into ES cells by electroporation. After selection with 150 μg/ml G418 (Life technologies) and cloning, integrated cell lines were screened by genomic PCR using the following primers; ROSA26-SA-Fw, 5′-CCTAAAGAAGAGGCTGTGCTTTGG-3′; ROSA26-LA-Rev, 5′-GTAGTTACTCCACTTTCAAGTTCCTTATAA-3′; 19E5scFv_183as, 5′-ATTTCTCATTGTAGTCAGTACCACCACTTC-3′; and EGFP-C 5′-CATGGTCCTGCTGGAGTTCGTG-3′. After removal of the Neo cassette from the integrated locus by transient expression of FLP recombinase, mintbody was expressed stably in ES cells.
Microscopy
All fluorescence images were collected using a confocal microscope (FV1000 or FV1000D; Olympus) operated by the built-in FV1000 software (ver.3.1a), equipped with a heating (Tokai Hit) and a CO2-control (Tokken) systems. For live-cell imaging, cultured cells were grown on a glass bottom dish (Mat-Tek) and the medium was replaced to phenol red-free DME (Nacalai Tesque) containing the supplements. For Figure 1e, hTERT-RPE1 cells were incubated in 100 ng/ml Hoechst33342 (Nacalai Tesque) for 3 h and washed with DME, before colleting the images (512 × 512 pixels; pinhole 95 μm; 10× zoom; 4 μs/pixel; 6× averaging; 2.1% and 0.2% transmission of 488-nm and 405-nm lasers). For monitoring changes in H3K9ac levels in living cells, U2OS cells stably expressing H3K9ac-mintbody were imaged every 5 min (512 × 512 pixels; pinhole 300 μm; 1× zoom; 4 μs/pixel; 6× averaging; 0.1% transmission 473-nm laser ) using a 60× UPlanSApo oil-immersion lens (NA 1.35). Fluorescence intensities of the nucleus and cytoplasm were measured from maximum intensity projection (MIP) of 10 z-stacks with 0.8 μm intervals and the ratio of nucleus to cytoplasm was obtained after subtracting the background, using custom Mathematica (Wolfram Research) code, available upon request. For Supplementary Figure S3c, the time-series were collected every 10 min (640 × 640 pixels; pinhole 290 μm; 1.5× zoom; 8 μs/pixel; 4× averaging; 4% transmission 488-nm laser) using a 60× PlanApoN (NA 1.40) oil-immersion lens. Time-lapse videos were created using FV1000 built-in software (ver. 3.1a).
FRAP was performed using a 60× PlanApoN (NA 1.40) oil-immersion lens. For Figure 2b, 10 images were collected (0.2% 488-nm laser transmission; 0.375 s/frame; 2 μs/pixel; 128 × 128 pixels; pinhole 800 μm; 8× zoom), a 2 μm diameter spot was bleached (100% 488-nm laser transmission) and a further 290 images were collected. The fluorescence intensity of the bleached area was measured using Mathematica (Wolfram Research). FRAP curves and the bound/free fraction, ton (s) and toff (s), were obtained by fitting to a reaction-diffusion model23.
For immunofluorescence, cells grown on a 24-well glass bottom plate (Iwaki) were transfected and fixed with 4% formaldehyde (Electron Microscopy Sciences) in 250 mM HEPES-NaOH (pH 7.4; Wako Purechemicals) containing 0.1% Triton X-100 for 10 min at room temperature. Fixed cells were permeabilized with 1% Triton X-100 in PBS for 20 min at room temperature and washed three times with PBS. After blocking with 10% Blocking One-P (Nacalai Tesque) in PBS for 20 min at room temperature, cells were incubated in 2 μg/ml Cy3-conjugated anti-H3K27me3 (CMA323)9 for 2 h at room temperature. After washing three times with PBS, cells were incubated with 250 ng/ml Hoechst33342 in PBS for 20 minutes at room temperature. Fluorescence images of Cy3-CMA323, H3K9ac-mintbody and Hoechst33342 were sequentially collected with a 60× PlanApoN (NA 1.40) oil-immersion lens.
Immunoblotting
For immunoblotting, U2OS cells stably expressing H3K9ac-mintbody were grown on 12-well plates. After incubation with TSA (100 nM up to 6 h, or 1–100 nM for 2 hr) or DMSO, cells were washed twice with PBS and lysed with 2× SDS-gel loading buffer. Cell lysates were boiled for 5 min and separated in 15% SDS-polyacrylamide gels (Wako Purechemicals), before transfer on to polyvinylidene difluoride membranes (Pall) using a semidry blotting system (ATTO). Membranes were washed 3 times in TBST (20 mM Tris-HCl [pH 8.0], 150 mM NaCl and 0.05% Tween 20) for 5 min, blocked for 20 min in Blocking-One (Nacalai Tesque) and washed 3 times for 5 min in TBST. Membranes were then incubated for 1 h at room temperature with mouse anti-H3K9ac (CMA310; 1 μg/ml) or rabbit anti-GFP antibody (MBL; 1:1,000) in Can-get-signal (Toyobo), washed 3 times for 10 min in TBST, incubated for 1 h with peroxidase-conjugated anti-mouse or anti-rabbit IgG (1:2,000; GE Healthcare) in Can-get-signal and washed 3 times for 10 min in TBST. Signals were developed using Western Lightning Chemiluminescence Reagent Plus (PerkinElmer) and detected using a LAS-3000 (Fujifilm). Band intensity was measured using Multi Gauge, ver. 3 software (Fujifilm).
Transgenic fly and zebrafish
To generate transgenic flies, the H3K9ac-mintbody coding sequence was cloned into the pUASp vector and the resulting plasmid was injected into y w flies using a standard procedure24. H3K9ac-mintbody was expressed in early embryos and salivary gland cells under the control of matα4-Gal4-VP16 and hs-Gal4 drivers, respectively. Dechorionated embryos were placed on to a μ-dish (ibidi) in PBS and 7 z-stack images with 5 μm intervals were collected every 32.9 sec, or 10 min (512 × 512 pixels; 1.4–1.5× zoom; 2–4 μs/pixel; 4× averaging; 1% transmission of 488-nm laser) using a 20× PlanApo objective lens (NA 0.75). Salivary glands isolated from larvae were placed on to a μ-dish (ibidi) in PBS. For imaging the whole salivary gland, a 10× UPlanSApo (NA 0.40) dry lens was used. Polytene chromosomes in individual cells were imaged in live, or after fixation and Hoechst33342-staining (1 μg/ml; overnight at 4°C). For live cells, 22 z-stack images with 1 μm intervals were collected (512 × 512 pixels; 5× zoom; 8 μs/pixel; 6× averaging; 1% transmission 488-nm laser) using a 60× PlanApoN (NA 1.40) oil-immersion lens. For fixed cells, Hoechst33342 and EGFP were sequentially collected using 0.4% and 0.1% transmission of 488-nm and 405-nm lasers, respectively. Time-lapse videos with MIP and 3D-rotation videos were created using FV1000 built-in software (ver. 3.1a) and Fiji (http://fiji.sc/), respectively.
To establish double transgenic zebrafish Tg(Gal4FF;UAS:mintbody), the mintbody coding sequence was cloned into the pT2MUASMCS vector13. Both UAS:mintbody plasmids and Tol2 transposase mRNA were co-injected into one-cell stage embryos25, harboring the Gal4FF gene in the zfand5b locus with ubiquitous expression [SAGFF(LF)73A]13. For Figure 4c, a Gal4FF;UAS:mintbody double transgenic embryo at 6 days post fertilization (dpf) was placed on a glass bottom dish (Mat-Tek) in 0.05% REI-SEA marine II (Iwaki) with 0.168 mg/ml tricaine (Sigma-Aldrich) and 3% methylcellulose26. A composite image was created from four small MIP images of 12 z-stack with 10 μm intervals (512 × 512 pixels; 1× zoom; 4 μs/pixel; 4× averaging; 0.2% transmission of 488-nm laser) collected using a 10× UPlanSApo (NA 0.40) dry lens. For time-lapse imaging, the embryos at 9 h post fertilization (hpf) were dechorionated using fine forceps and mounted on a glass-bottom dish in embryo medium [0.5 μM MgCl2 and 0.5 μM CaCl2 in 0.1× Hanks' Balanced Salt Solution (HBSS; Life technologies)] containing 0.168 mg/ml tricaine and 3% methylcellulose. For Supplementary Video 9, 15 z-stack images with 15 μm intervals were collected every 10 min (512 × 512 pixels; 1× zoom; 2 μs/pixel; 4× averaging; 1% transmission of 488-nm laser). This study was carried out in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals (Fundamental Guidelines for Proper Conduct of Animal Experiment and Related Activities in Academic Research Institutions under the jurisdiction of the Ministry of Education, Culture, Sports, Science and Technology). An animal use protocol was approved by the Institutional Animal Care and Use Committee of RIKEN. Using approved anesthetics, all efforts were made to minimize discomfort and suffering during experimental procedures.
References
The ENCODE Project Consortiam. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Kimura, H., Hayashi-Takanaka, Y. & Yamagata, K. Visualization of DNA methylation and histone modifications in living cells. Curr Opin Cell Biol 22, 412–418 (2010).
Sasaki, K., Ito, A. & Yoshida, M. Development of live-cell imaging probes for monitoring histone modifications. Bioorg Med Chem 20, 1887–1892 (2012).
Lin, C. W., Jao, C. Y. & Ting, A. Y. Genetically encoded fluorescent reporters of histone methylation in living cells. J Am Chem Soc 126, 5982–5983 (2004).
Lin, C. W. & Ting, A. Y. A genetically encoded fluorescent reporter of histone phosphorylation in living cells. Angew Chem Int Ed Engl 43, 2940–2943 (2004).
Sasaki, K., Ito, T., Nishino, N., Khochbin, S. & Yoshida, M. Real-time imaging of histone H4 hyperacetylation in living cells. Proc Natl Acad Sci U S A 106, 16257–16262 (2009).
Ito, T. et al. Real-time imaging of histone H4K12-specific acetylation determines the modes of action of histone deacetylase and bromodomain inhibitors. Chem Biol 18, 495–507 (2011).
Hayashi-Takanaka, Y., Yamagata, K., Nozaki, N. & Kimura, H. Visualizing histone modifications in living cells: spatiotemporal dynamics of H3 phosphorylation during interphase. J Cell Biol 187, 781–790 (2009).
Hayashi-Takanaka, Y. et al. Tracking epigenetic histone modifications in single cells using Fab-based live endogenous modification labeling. Nucleic Acids Res 39, 6475–6488 (2011).
ten Bosch, J. R., Benavides, J. A. & Cline, T. W. The TAGteam DNA motif controls the timing of Drosophila pre-blastoderm transcription. Development 133, 1967–1977 (2006).
Tadros, W. & Lipshitz, H. D. The maternal-to-zygotic transition: a play in two acts. Development 136, 3033–3042 (2009).
Qi, D., Larsson, J. & Mannervik, M. Drosophila Ada2b is required for viability and normal histone H3 acetylation. Mol Cell Biol 24, 8080–8089 (2004).
Asakawa, K. & Kawakami, K. The Tol2-mediated Gal4-UAS method for gene and enhancer trapping in zebrafish. Methods 49, 275–281 (2009).
Yao, J., Munson, K. M., Webb, W. W. & Lis, J. T. Dynamics of heat shock factor association with native gene loci in living cells. Nature 442, 1050–1053 (2006).
Attreed, M., Desbois, M., van Kuppevelt, T. H. & Bulow, H. E. Direct visualization of specifically modified extracellular glycans in living animals. Nat Methods 9, 477–479 (2012).
Kvam, E., Sierks, M. R., Shoemaker, C. B. & Messer, A. Physico-chemical determinants of soluble intrabody expression in mammalian cell cytoplasm. Protein Eng Des Sel 23, 489–498 (2010).
Rothbauer, U. et al. Targeting and tracing antigens in live cells with fluorescent nanobodies. Nat Methods 3, 887–889 (2006).
Dancy, B. M. et al. Live-cell studies of p300/CBP histone acetyltransferase activity and inhibition. Chembiochem 13, 2113–2121 (2012).
Kimura, H., Hayashi-Takanaka, Y., Goto, Y., Takizawa, N. & Nozaki, N. The organization of histone H3 modifications as revealed by a panel of specific monoclonal antibodies. Cell Struct Funct 33, 61–73 (2008).
Zhou, H., Fisher, R. J. & Papas, T. S. Optimization of primer sequences for mouse scFv repertoire display library construction. Nucleic Acids Res 22, 888–889 (1994).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat Genet 21, 70–71 (1999).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev Biol 1, 4 (2001).
Stasevich, T. J., Mueller, F., Brown, D. T. & McNally, J. G. Dissecting the binding mechanism of the linker histone in live cells: an integrated FRAP analysis. EMBO J 29, 1225–1234 (2010).
Spradling, A. C. & Rubin, G. M. Transposition of cloned P elements into Drosophila germ line chromosomes. Science 218, 341–347 (1982).
Kitaguchi, T., Kawakami, K. & Kawahara, A. Transcriptional regulation of a myeloid-lineage specific gene lysozyme C during zebrafish myelopoiesis. Mech Dev 126, 314–323 (2009).
Westerfield, M. The Zebrafish Book 4th ed. (University of Oregon Press, Eugene, Oregon, USA, 2000).
Acknowledgements
We thank S. Takita for advice on zebrafish maintenance and F. Costantini and J. Takeda and for providing pBigT and ROSA26 vectors. SAGFF(LF)73A transgenic fish was provided by National Bioresource Project of Japan. This work was supported by grants-in-aid from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan (to H.Kimura). T.J.S. and Y.H.-T. were supported by Japan Society for the Promotion of Science fellowship. Y.H.-T. was supported by Special Coordination Funds for Promoting Science and Technology from the MEXT of Japan for the Osaka University Program for the Support of Networking among Present and Future Women Researchers.
Author information
Authors and Affiliations
Contributions
Y.S. performed most experiments and data analysis. M.Mukai, H.Kita, T.Kimura and S.H. generated transgenic fly. J.U. and K.Y. generated knock-in mouse ES cells. M.Muraki and A.K. generated transgenic zebrafish. T.J.S. performed some image analysis. N.H., T.Kujirai and H.Kurumizaka purified scFvs and measured their affinity. Y.O. contributed zebrafish embryo imaging. Y.H.-T. contributed to cloning scFv cDNA. C.O. performed mass spectrometry analysis. N.N. generated hybridoma cells. H.Kimura conceived the study. Y.S. and H.Kimura wrote the manuscript.
Ethics declarations
Competing interests
N.N. is a founder of Mab Institute Inc.
Electronic supplementary material
Supplementary Information
Supplementaly Figures
Supplementary Information
Video1. Tricostatin A (TSA) induced the nuclear accumulation of H3K9ac-mintbody in U2OS cells.
Supplementary Information
Video2. DMSO had no effects on the relative nuclear concentration of H3K9ac-mintbody in U2OS cells.
Supplementary Information
Video3. TSA induced nuclear accumulation of H3K9ac-mintbody in mouse ES cells.
Supplementary Information
Video4. H3K9ac-mintbody diffused into cytoplasm upon TSA removal in mouse ES cells.
Supplementary Information
Video5. H3K9ac-mintbody in Drosophila early embryos.
Supplementary Information
Video6. H3K9ac-mintbody in Drosophila embryos.
Supplementary Information
Video7. 3D-reconstruction of polytene chromosomes in living Drosophila salivary gland expressing H3K9ac-mintbody.
Supplementary Information
Video8. 3D-reconstruction of polytene chromosomes in fixed Drosophila salivary gland stained with Hoechst33342 (red) and H3K9ac-mintbody (green).
Supplementary Information
Video9. H3K9ac-mintbody in zebrafish embryos.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Sato, Y., Mukai, M., Ueda, J. et al. Genetically encoded system to track histone modification in vivo. Sci Rep 3, 2436 (2013). https://doi.org/10.1038/srep02436
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02436
This article is cited by
-
STREAMING-tag system reveals spatiotemporal relationships between transcriptional regulatory factors and transcriptional activity
Nature Communications (2022)
-
A live imaging system to analyze spatiotemporal dynamics of RNA polymerase II modification in Arabidopsis thaliana
Communications Biology (2021)
-
Live-cell imaging reveals the spatiotemporal organization of endogenous RNA polymerase II phosphorylation at a single gene
Nature Communications (2021)
-
Intrabody-based FRET probe to visualize endogenous histone acetylation
Scientific Reports (2019)
-
A genetically encoded probe for imaging nascent and mature HA-tagged proteins in vivo
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.