Abstract
The anti-hyperglycaemic drug metformin has important anticancer properties as shown by the direct inhibition of cancer cells proliferation. Tumor cells avidly use glucose as a source for energy production and cell building blocks. Critical to this phenotype is the production of glucose-6-phosphate (G6P), catalysed by hexokinases (HK) I and II, whose role in glucose retention and metabolism is highly advantageous for cell survival and proliferation. Here we show that metformin impairs the enzymatic function of HKI and II in Calu-1 cells. This inhibition virtually abolishes cell glucose uptake and phosphorylation as documented by the reduced entrapment of 18F-fluorodeoxyglucose. In-silico models indicate that this action is due to metformin capability to mimic G6P features by steadily binding its pocket in HKII. The impairment of this energy source results in mitochondrial depolarization and subsequent cell death. These results could represent a starting point to open effective strategies in cancer prevention and treatment.
Similar content being viewed by others
Introduction
One of the primary metabolic changes observed in malignant transformation is an increased catabolic glucose metabolism characterized by high rates of anaerobic glycolysis regardless of oxygen concentration1. Critical to this phenotype is glucose cellular entrapment by its conversion to glucose-6-phosphate (G6P). In normal tissues, this basic process is regulated by four different hexokinase (HK) isoforms indicating that regulation of glucose phosphorylation can vary in different tissues under different condition2. In cancer cells, this reaction is mainly catalysed by HK II whose glucose affinity and mitochondrial localization are highly advantageous for cancer survival and growth3. Inhibition of HKII enzymatic activity and its mitochondrial localization, are associated with cancer cells death4,5.
The relevance of the so-called “Warburg effect” could be complementary with recent evidence documenting that metformin can inhibit cancer growth at least partially by an enhanced phosphorylation of AMP-activated protein kinase (pAMPK)6. This highly conserved Ser/Thr protein kinase complex is activated by reduction in fuel supply and directs nutrients toward catabolic processes, switching off anabolic/growth-promoting pathways7. In normal tissues, biguanides activate this energy checkpoint by lowering ATP via a direct inhibition of respiratory complex-18. However other mechanisms could participate to this inhibition as shown by the finding that metformin decreases 18F-fluorodeoxyglycose (FDG) uptake in experimental model9.
This tracer enters the cytosol via the same facilitative transporters of glucose (GLUT), while its entrapment is mediated by phosphorylation to FDG6P through a reaction catalysed by the same HKs responsible for the conversion of glucose into G6P10,11. Accordingly, these data seem to indicate that metformin might reduce cancer fuel supply by directly hampering glucose trans-membrane transport and/or phosphorylation.
To test this hypothesis, we studied the effect of metformin on the first steps of glucose metabolism in Calu-1 cells, derived from human non-small cell lung cancer. This approach documented that metformin directly inhibits HK I and HK II function in a dose and time dependent fashion. The impairment of these important promoters of cell proliferation contributes to explain the anticancer properties of this biguanide and represents a starting point for effective strategies in cancer prevention and treatment.
Results
Metformin effect on glucose metabolism in cancer cells
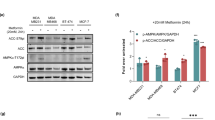
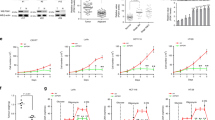
Metformin effect on cancer metabolism was evaluated by estimating Calu-1 cells capability to retain FDG. Metformin treatment decreased tracer uptake in a dose and time dependent manner up to its virtual abolition after 24 hours exposure to 10 mM drug concentration (32.7 ± 1.0% in controls vs 3.1 ± 0.4% in treated cells, p < 0.0001) (Figure 1A).
Effect of metformin on Calu-1 cells glucose consumption and HKs activity.
(A) Cell uptake of FDG was expressed as percentage of total tracer availability according to different metformin concentrations and exposure times. 1 mM metformin did not produce any significant modification, while tracer uptake decreased after 24 hrs exposure to metformin 5 mM. Highest drug concentration (10 mM) caused a significant reduction and virtually abolished glucose consumption at 6 and 24 hr. p values are shown for each comparison that was performed by one way analysis of variance. (B) Calu-1 HKs activity (expressed as percentage of control) is represented as function of metformin concentrations. The reaction was switched on after 10 minutes of metformin pre-incubation with Calu-1 total cell lysate (Lysate) or plus ATP 0.8 mM (Lysate + ATP) or Glucose 100 mM (Lysate + Glu). The reaction was switched on by adding to the solution respectively ATP + Glu (Lysate), Glucose (Lysate + ATP) and ATP (Lysate + Glu). Pre-incubation with metformin and glucose (Lysate + Glu) caused an inhibition of the HK I and II enzymatic activity that was dependent upon metformin concentration. This finding was not observed when the enzymes were pre-exposed to metformin alone (Lysate) nor to metformin and ATP (Lysate + ATP). (C) Enzymatic activity (expressed as percentage of control) of human purified HK I, HK II and HK IV observed after pre-incubation with glucose and different metformin concentrations. The reaction was switched on after 10 minutes by adding to the solution 0.8 mM ATP. Metformin induced a dose-dependent inhibition of catalytic activity of HK I and HK II. By contrast, it did not affect enzymatic activity of HK IV. (D) Dose dependent interference of ATP on human purified HK II inhibition caused by metformin. Phosphorylation rate is expressed as percentage of HKs activity measured after ten minutes pre incubation with glucose (100 mM) and different metformin concentrations and starting the reaction with ATP (0.4–1.2 mM). ATP 0.8 mM, was considered as reference value. Starting the reaction with ATP concentrations ≥1.2 mM fully abolished metformin effect. On the contrary, ATP levels below 1.2 mM reduced the extent of metformin inhibitory action in a dose dependent fashion. * = p < 0.05; ** = p < 0.01:*** = p < 0.001. Error bars indicate standard error.
Theoretically, a reduced FDG uptake might reflect an accumulation of G6P caused by a block in downstream glycolytic reaction. To verify this hypothesis, we measured the effect of metformin on mRNA levels encoding for thyoredoxin inhibiting protein (TXNIP). This pathway has been recently found to regulate glucose metabolism by inhibiting glycolysis rate and being activated by G6P12. Prolonged biguanide treatment silenced TXNIP expression (Figure S1A) indicating that the reduction in FDG retention was not caused by a block in glycolytic flux and rather reflected decreased rates of glucose transport and/or phosphorylation.
We thus investigated whether metformin reduces the availability of membrane GLUT. In agreement with the malignant phenotype13,14, Calu-1 cells were found to mostly express GLUT1 (Figure S1B). Metformin did not affect gene expression, total protein amount or plasma membrane localization of this carrier (Figure S1B, C and D), thus excluding drug effects on trans-membrane glucose transport.
Selective effect of metformin on HK I and HK II enzymatic activity
Since the mechanism underlying glucose uptake was not significantly impaired, we hypothesized an alteration in tracer retention. To this purpose, we verified gene expression of the four different HK isoforms. Real time PCR showed that Calu-1 cells preferentially express HK I and II, while mRNA encoding for HK III and IV are almost undetectable (Figure S1E). This expression pattern predicts a poor prognosis, playing a key role in cancer growth and survival3. Again, metformin exposure did not modify gene expression nor protein amount of HK I and II isoforms (Figure S1C and E).
We thus tested whether metformin interferes with HK I and II enzymatic function. To this purpose, we first suspended cell lysates in a solution containing MgCl2 (5 mM) and different concentrations of metformin (from 0.0375 mM to 10 mM) alone or in the presence of either glucose (100 mM) or ATP (0.8 mM). After ten minutes pre-incubation, the reaction was started providing the full availability of all substrates. HK I and II activity remained unchanged after pre-incubation with metformin alone or metformin plus ATP. On the contrary, pre-incubation of cell lysates with glucose elicited a dose dependent drug inhibition of enzymatic function (Figure 1B). Experiments on human purified HKs provided the same results: pre-incubation with glucose and metformin inhibited phosphorylating activity of I and II isoforms without affecting HK IV (Figure 1C). These data indicate that the conformational changes caused by prior exposure to glucose are needed to allow the interference of metformin on the catalytic site of HK I and II. Although, the inhibitory effect of metformin remained partial, even when drug concentration was increased beyond saturating values (Figure 1C).
Molecular modelling of HK II and metformin interaction
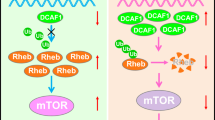
HK I and II are very similar in structure, with the latter accounting for the highly glycolytic phenotype in cancer cells3,15. Accordingly, to elucidate the molecular action of metformin, we modelled the conformational states of HK II catalytic domain with in silico techniques (Table S1–3 and Figure S2–3). We found that the open conformation, which is characteristic of the substrate-free HK II (Figure 2A), cannot accommodate metformin with a stable binding mode (Table S4 and Figure S4–5). On the contrary, the closed conformation, induced upon glucose binding, can steadily harbour different orientation of metformin (Table S4 and Figure S4–5, Media Video1–2).
Molecular mechanism of HK II inhibition by metformin.
(A) Open conformation of HK II in complex with ATP highlighted with the red surface (Table S1 and Figure S2). (B) Closed conformation of HK II in complex with glucose and glucose-6-phosphate highlighted with green and orange surfaces, respectively (Table S2 and Figure S2). (C) Closed conformation of HK II in complex with metformin as suggested by Induced Fit Docking calculation and molecular dynamic simulations. (D) Closed conformation of HK II in complex with glucose, ATP and Mg2+ highlighted with green, red and blue surfaces, respectively. (E) Chemical structure of metformin (tautomeric and protonation forms are reported in Figure S3). (F) Chemical structure of glucose-6-phosphate. (G) Interactions map of the most stable binding mode of metformin in the closed HK II conformation. (H) Interactions map glucose-6-phosphate in the closed HK II conformation.
While biochemical results suggest that metformin alone is not able to elicit the open-closed conformational change, we found that the pharmacological action of metformin on the closed conformation occurs in a region interposed between substrate and ATP location (Figure 2D). In physiological condition, this region is critical for the fine-tuning of the catalytic activity of HK II. In fact, two players compete for this site: Mg2+, that catalyse the phosphorylation reaction (Figure 2D) and G6P, the product of the reaction that inhibit the enzyme (Figure 2B). Showing a similar interaction pattern with respect to G6P (Figure 2E–H), metformin is thus prefigured as an uncompetitive (Figure S1F) and allosteric inhibitor of HK II as only the enzyme-substrate complex can be bound. The mechanism of metformin inhibition, however, implies a competition with ATP (Figure 1D). This is consistent with the stable binding modes of metformin in the closed conformation that overlaps with the β- and γ-phosphoryl groups of ATP (Figure 2C and D). Finally, metformin binding does not affect the closed conformation HK II (Figure S6), suggesting that this binding region may be conveniently used for the rational design of other small-molecule inhibitors16. Altogether, these data document that the reduced FDG uptake reflects a selective metformin induced impairment of glucose phosphorylation.
Effect of metformin on HK II mitochondrial localization and cell death
HK I and II are tightly bound to mitochondria, thereby gaining a preferential access to ATP11,15,17,18. This interaction is relatively stable for HK I, while it is dependent upon G6P levels for HK II2,17,19. Immunofluorescence analysis showed that metformin inhibition of HK enzymatic activity was associated with a preferential cytosolic localization of isoform II (Figure 3). The ability of metformin to mimic and occupy G6P binding site could explain the reduced HKII localization in mitochondrial compartment.
Metformin displaces HK II from Mitochondria.
(A) HK I (green color) and mitochondria (red color) co-localization under control condition and after treatment with 5 mM metformin for 24 hr. Images obtained by immunofluorescence labelling HK I and mitochondria (Mit) show that metformin did not affect HK I interaction with mitochondria. (B) Images display the corresponding finding for HK II staining. HK II binding to mitochondria is impaired by treatment with 5 mM metformin. (C, D) Quantitative measurement of HK I and HK II and mitochondria co-localization under control condition and after metformin treatment. Metformin does not affect the binding of HK I to mitochondria (78% ± 4.7 vs 80% ± 5.2, p = ns) while it decreases the binding of HK II to mitochondria (43% ± 3.8 vs 66% ± 5, p < 0.05). * = p < 0.05. Error bars indicate standard error.
Metformin effects resulted in a marked and dose-dependent increase in the AMP/ATP ratio (Figure 4A). The altered energy asset was accompanied by a mitochondrial membrane depolarization as shown by JC-1 staining (Figure 4B). Flow-cytometry analysis confirmed confocal microscopy data demonstrating an increase in number of depolarized mitochondria from 6% under control condition up to 50% in cells treated with 10 mM metformin (Figure 4B).
Metformin treatment impairs mitochondrial function inducing cell death.
(A) Calu-1 cells were treated for 1 hr with different doses of metformin; ATP and AMP concentrations (μMol/mg of proteins) were determined by enzymatic assay. Measured AMP/ATP ratio increased as a function of metformin concentration. * = p < 0.05, ** = p < 0.01. Error bars indicate standard error. (B) 10 mM metformin exposure of Calu-1 cells for 24 hr affect mitochondrial potential (ΔΨm). Upper pannels: Bivariate JC-1 analysis of mitochondrial membrane potential by flow cytometry. Botton pannels: Immunofluorescence analysis of Calu-1 cells treated with JC-1.Metformin treatment induced an increase of depolarized regions as indicated by the disappearance of red and increase of green fluorescence. (C) Apoptosis was evaluated after treating Calu-1 cells with metformin and staining with Annexin-V and propidium iodide (PI) at 24 hr. The values indicate the percentage of death (Annexin-V−, PI+), early apoptotic (Annexin-V+, PI−) and necrotic/advanced apoptotic stage (Annexin-V+, PI+) cells.
The combined impairment of glucose phosphorylation and mitochondrial function preceded the expected cell damage. Indeed, the decrease of glucose consumption already occurred at 6 and 24 hours when Calu-1 cell viability was almost all preserved. By contrast, measurable numbers of dead cells occurred after 48 hours of drug exposure (Figure 4C) confirming previous reports4,20,21. The effect on energy metabolism was followed by cell death without evidence of early apoptosis by annexin V staining (Figure 4C).
These data indicate that a prolonged exposure to metformin impairs the energy asset of cancer cells. This effect is explained by the capability of metformin to blunt fuel availability through a direct inhibition of HK I and II enzymatic function thus hampering cell capability to retain and metabolize glucose2.
This effect is further worsened by the increased cytosol localization of HK II decreasing ATP availability and impairing polarization of mitochondrial membranes. This finding suggests the possibility to develop small molecules with greater affinity for HK enzymatic pocket to improve effectiveness of metabolic cancer treatment22.
Discussion
Recent evidence indicates that the widely used anti-hyperglycemic drug metformin has important anticancer properties. Its best-characterized action is the respiratory chain complex I inhibition, resulting in ATP depletion and AMPK activation. However, significant evidences suggest that metformin can affect glucose metabolism by alternative independent mechanisms. The present study demonstrates that metformin decreases FDG uptake by directly inhibiting enzymatic activity of HKII and HKI and mimicking G6P in NSCLC Calu-1 cells.
The first evidence of this new metformin action is represented by the decreased FDG entrapment, which occurred at short exposure time condition. FDG enters the cytosol via the same facilitative transporters of glucose (GLUT), it remains phosphorylated and trapped within the cell, being a false substrate for all further reactions channeling G6P to alternative pathways10,11. Accordingly, the reduced retention of FDG might reflect a direct alteration in glucose transport and phosphorylation mechanisms as well as a block in G6P utilization. Metformin reduces FDG uptake to a critical level in a dose and time dependent manner, without altering membrane GLUT availability. The ability of metformin to interfere with cancer cells glucose uptake was showed also by others and in different cellular models9,23. It remains to explain how metformin could negatively modulate this mechanism.
Metformin induces AMPK phosphorylation, a cellular energy sensor which downregulates energy consuming activity of cells23. This finding appears counterintuitive if we consider that AMPK activation induces Glut transporter externalization favoring glucose uptake24,25. In the same time there are several cellular targets, which can drive metformin effect independently from AMPK. It has been demonstrated that inhibition of glucose phosphorylation could be explained by an AMPK-independent mechanism26. Hexokinases-2 (HKII) is over-expressed in most malignant tumors, regulates cancer cell energy disposition and represents an important attractive target for anticancer drug development15. Active HKII is strategically bound to the outer mitochondrial membrane where it helps couple ATP production in mitochondria to glucose phosphorylation conferring a highly glycolytic phenotype to cancer cells15. We show for the first time that metformin directly inhibits phosphorylating activity of both HKI and HKII in Calu-1 cells, impairing their enzymatic activity in a dose dependent fashion. Our results demonstrate that the inhibition of metformin on HK activity was uncompetitive and thus present only in the presence of the complex formed by HKII and its substrate glucose. As showed by Lineweaver-Burk plot of the hexokinase activity (Supplementary Figure 1F), in the presence of different concentration of glucose and metformin, HKII Vmax and the Km decrease by the same factor, so the ratio of Km/Vmax does not change the value of Ki which is not relevant in this kind of inhibition.
The determination of the binding mode (position, orientation and conformation in addition to protonation and tautomeric states) of small molecules in protein-ligand complexes is highly challenging. In silico predictions, are useful to determine binding modes of small molecules like metformin that, due to their small size and chemical nature, have the capacity to adapt protonation forms, tautomeric forms and orientations depending on the dynamic behavior of the target binding site27,28. Our data show that only the HKII closed conformation, induced upon glucose binding, can steadily harbour different orientations of metformin. The region occupied by metformin corresponds to the binding site of G6P, inhibitor of the enzymatic reaction catalyzed by HKI and HKII but not by HKIV. In silico data corroborate with experimental evidences showing that metformin does not inhibit glucokinase (HKIV), which is the only hexokinase isoform that does not show any auto-inhibition by it product G6P. In the same way of G6P, metformin shows a competition with ATP as documented by the increase inhibitory effect on HKII activity decreasing ATP concentration. Thus ATP depletion induced by metformin treatment should be considered largely advantageous to the inhibitory effect of metformin on HKII. In this scenario metformin is proposed to mimic G6P.
G6P influences HKII release from mitochondrial compartment19. The ability of metformin to mimic and occupy G6P binding site could explain the reduced HKII ability to be localized in mitochondrial compartment.
Beneficial anticancer effects of metformin are principally attributed to complex I inhibition mitochondrial dysfunction. Here we provide evidence of a new effect of metformin associated with mitochondria compartment strengthening metformin as a mitochondrial inhibitor in anticancer strategy. The mechanism by which metformin inhibits complex I and the oxidative phosphorylation remained unclear. Addition of metformin decreases oxygen consumption and mitochondrial membrane potential only in intact cells but not in permeabilized cells or isolated mitochondria suggest the existence of novel cellular signaling pathway, targeted to the respiratory chain complex I. In this contest HKII could represent a good upstream candidate target of metformin action.
In conclusion the key finding of the present study is that metformin inhibits HK II in Calu-1 cells through an allosteric modification of its molecular structure blocking the synthesis of G6P. Moreover, our results demonstrate that HK II inhibition by metformin causes release of this enzyme from the outer membrane of mitochondria, thus leading to the activation of apoptotic signals. Once extended to other cancer models the blockage of HKII may be therapeutically useful for the management of cancer cells metabolism. These results open new strategies not only toward metformin therapy but also for the study of new metformin related compounds.
Methods
Reagents
Metformin was provided by Sigma-Aldrich (St. Louis, MO, USA). FDG was produced according to standard methodology. Daily quality controls always documented adequate standards and, in particular, a radiochemical purity ≥98%.
Cell culture
Calu-1 cells were obtained from Interlab Cell Line Collection (ICLC) and cultured in low Glucose Dulbecco's modified Eagle's medium supplemented with 2 mM glutamine, 10% Fetal Bovine Serum (FBS), 100 U/ml penicillin and 100 mg/ml streptomycin sulphate at 37°C in 5% CO2.
FDG uptake evaluation
Labelling was performed by incubating 1 × 106 cultured cells with FDG, according to a modified procedure developed and validated in our laboratory29. On the day of the experiment, uptake medium was added with PBS and FDG so to obtain a tracer concentration of 1 μCi/mL in each well. Glucose-free medium was used for FDG uptake. Tracer exposure was maintained for 60 minutes at 37°C. Preliminary studies indicated that FDG uptake is linear over this period of time for all cell lines and that there is no depletion of the radioactive substrates from the culture media. Cells then rinsed twice in cold PBS and counted to determine cell associated fluorine-18 radioactivity, using a Packard Cobra II gamma counter (Packard, Meriden, CT). Washing did not induce significant efflux of FDG, as confirmed by preliminary studies comparing FDG uptake of cells washed for different periods of time in the presence or absence of the glucose transport inhibitors, cytochalasin B and phloretin. This indicates that after an incubation period of 60 minutes, almost all FDG was trapped intra cells as FDG6P. Radioactivity uptake values were normalized to the number of viable (trypan blue-negative) cells.
Real-time PCR analysis
Total RNAs were isolated using the RNeasy Mini Kit (Qiagen, Hilden, Germany). Oligo dT primer cDNAs were synthesized using the SuperScript® III First-Strand Synthesis SuperMix (Invitrogen, Irvine, CA). Expression analysis was performed by quantitative real-time RT-PCR on a LightCycler 480 (Roche Applied Science, Monza, Italy). Expression data were normalized on the mean of housekeeping gene GAPDH gene expression data obtained in parallel. Relative expression values with standard errors and statistical comparisons (one way analysis of variance) were obtained using Q-gene software.
Western blotting analysis
Calu-1 cell extracts were resolved by SDS-PAGE and transferred to a nitrocellulose membrane as previously described (FASEB). Membranes were incubated overnight at 4°C and probed with the following antibodies: anti human HK I (C35C4) and HK II (C64G5) purchased from Cell Signaling Technologies (Boston, MA), anti human Actin and Glut1 (SPM498) purchased from Abcam (Cambridge, UK), anti human Glut1 purchase from Scbt (CA, USA). Primary antibodies were detected using a goat anti-mouse or goat anti-rabbit IgG HRP conjugated and developed with ECL substrate (Thermo Fisher Scientific, IL, USA).
Immunofluorescence analysis
Calu-1 cells, cultured on glass coverslips, were incubated with MitoTracker probe (Life Technologies) and treated as previously described (FASEB). Coverslips were incubated O/N with rabbit anti-HK I (C35C4) or HK II (C64G5) primary antibodies (Cell Signaling Technologies). Specific staining was visualized with a goat anti-Rabbit Alexa Fluor 488 secondary antibody (Molecular Probes, Eugene, OR, USA) washed and mounted using Prolong Gold antifade reagent (Life Technologies Ltd, Paisley, UK). The images were collected using a three-channel TCS SP2 laser-scanning confocal microscope (Leica Microsystems, Wetzlar, Germany). Sequential acquisition was performed to avoid cross-talk between color signals. Spatial co-localization was analysed by ImageJ 1.34f software (Wayne Rasband, National Institutes of Health, Bethesda, MD, USA).
To measure GLUT1 plasma membrane expression Calu-1 cells were fixed with methanol and incubated with the anti-GLUT1 antibody (AbCam, Cambridge, MA, USA). A FITC-conjugated goat anti-mouse Ig (LiStarFish, Milano, Italy) was used as second step reagent. Samples were analysed with a FACScan (BD Pharmigen, San Jose, CA, USA).
Hexokinase assay
To assay the HK activity, the reduction of NADP+ was followed at 340 nm, through a coupled reaction with glucose-6-phosphate dehydrogenase (G6PD). The assay mix contained: 100 mM Tris HCl pH 8.5 mM MgCl2, 100 Mm Glucose, 0.8 mM ATP, 1 mM NADP, 3 Units of G6PD.
The assay was made on human purified HK II (ProSpec-Tany TechnoGene Ltd. Israel). Several metformin concentrations (between 0.375 mM and 10 mM) were used. The enzyme was incubated with metformin for ten minute before the assay, in the absence or in presence of one of the HK substrates. No metformin interference with G6PD activity was observed (data not shown).
Molecular modelling
A complete description of in silico methodologies used in this study is available in Supplementary Information
ATP/AMP ratio assay
Cells were washed twice with PBS, lysed with 2.5% Percloric Acid (PCA) and sonicated in ice three times for 10 seconds. After centrifugation, supernatants containing PCA were collected and neutralized with K2CO3. ATP and AMP were assayed enzymatically.
Mitochondrial potential detection
Mitochondrial membrane potential (Δψm) was determined using the cationic dye JC-1 (5,5,6,6-tetrachloro-1,1,3,3-tetraethylbenzimidazolcarbocyanine iodide) (Life Technologies Ltd, Paisley,UK). Samples were analyzed by a FACScan (BD Pharmigen) or by three-channel TCS SP2 laser-scanning confocal microscope (Leica Microsystems, Wetzlar, Germany) according to the manufacturer's procedures.
Cell survival and apoptosis assay
Apoptosis was determined by annexin V-FITC that was associated with propidium iodide (PI) staining (as an index of cell death) double staining (eBioscience, San Diego, CA, USA) according to the manufacturer's instructions. Analysis was performed using FACScan (BD Pharmigen).
Statistical analysis
All experiments were performed >3 times. Statistical differences were assessed by ANOVA (SPSS 16.0; SPSS, Chicago, IL). Values of p < 0.05 were considered statistically significant.
References
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–33 (2009).
Wilson, J. E. Isoenzymes of mammalian hexokinase: structure, subcellular localization and metabolic function. J. Exp. Biol. 206, 2049–2057 (2003).
Mathupala, S. J., Ko, Y. H. & Pedersen, P. L. Hexokinase-2 bound to mitochondria: Cancer's stygian link to the “Warburg effect” and a pivotal target for effective therapy. Semin. Cancer. Biol. 19, 17–24 (2009).
Kim, W. et al. Apoptosis-inducing antitumor efficacy of hexokinase II inhibitor in hepatocellular carcinoma. Mol. Cancer Ther. 6, 2554–2562 (2007).
Chiara, F. et al. Hexokinase II detachment from mitochondria triggers apoptosis through the permeability transition pore independent of voltage-dependent anion channels. Plos One. 19, 3(3), e1852 (2008).
Faubert, B. et al. AMPK is a negative regulator of the Warburg effect and suppresses tumor growth in vivo. Cell Metabolism 17, 113–124 (2013).
Hardie, D. G. Why starving cells eat themselves. Science 331, 410–411 (2011).
Zhou, G. et al. Role of AMP-activated protein kinase in mechanism of metformin action. J. Clin. Invest 108(8), 1105–7 (2001).
Mashhedi, H. et al. Metformin abolishes increased tumor (18)F-2-fluoro-2-deoxy-D-glucose uptake associated with a high energy diet. Cell Cycle. 10(16), 2770–8 (2011).
Aloj, L., Caraco', C., Jagoda, E., Eckelman, W. C. & Neumann, R. D. Glut-1 and hexokinase expression: relationship with 2-Fluoro-2-deoxy-D-glucose uptake in A431 and T47D cells in culture. Cancer Res. 59, 4709–4714 (1999).
Ahn, K. J. et al. Evaluation of the role of hexokinase type II in cellular proliferation and apoptosis using human hepatocellular carcinoma cell lines. J. Nucl. Med. 50, 1525–1532 (2009).
Stoltzman, C. A. et al. Glucose sensing by MondoA:Mlx complexes: a role for hexokinases and direct regulation of thioredoxin-interacting protein expression. Proc. Natl. Acad. Sci. U S A. 105, 6912–6917 (2008).
Macheda, M. L., Rogers, S. & Best, J. D. Molecular and cellular regulation of glucose transporter (GLUT) proteins in cancer. J. Cell Physiol. 202, 654–662 (2005).
Kaira, K. et al. Correlation between 18F-FDG uptake on PET and molecular biology in metastatic pulmonary tumors. J. Nucl. Med. 52, 705–711 (2011).
Pedersen, P. L. Warburg, me and hexokinase 2: multiple discoveries of key molecular events underlying one of cancers'most common phenotypes, the “Warburg effect”, i.e., elevated glycolysis in the presence of oxygen. J. Bioenerg. Biomembr. 39, 211–222 (2007).
Barbosa, A. J. M. & Del Rio, A. Freely accessible databases of commercial compounds for high- throughput virtual screenings. Curr. Top. Med. Chem. 12, 866–77 (2012).
Bustamante, E. & Pedersen, P. L. High aerobic glycolysis of rat hepatoma cells in culture: Role of mitochondrial hexokinase. Proc. Natl. Acad. Sci. 74, 3735–3739 (1977).
Kroemer, G., Galluzzi, L. & Brenner, C. Mitochondrial membrane permeabilization and cell death. Physiol. Rev. 87, 99–163 (2007).
John, S., Weiss, J. N. & Ribalet, N. Subcellular localization of Hexokinases I and II directs the metabolic fate of glucose. PLoS One 6(3) (2011).
Alimova, I. N. et al. Metformin inhibits breast cancer cell growth, colony formation and induces cell cycle arrest in vitro. Cell cycle. 8, 909–915 (2009).
Song, C. W. et al. Metformin kills and radiosensitizes cancer cells and preferentially kills cancer stem cells. Sci. Rep. 362, 2–9 (2012).
Pollak, M. N. Investigating metformin for cancer prevention and treatment: the end of the beginning. Cancer Discov. 2, 778–790 (2012).
Habibollahi, P., Van Den Berg, N. S., Kuruppu, D., Loda, M. & Mahmood, U. Metformin--anadjunct antineoplastic therapy--divergently modulates tumor metabolism and proliferation, interfering with early response prediction by 18F-FDG PET imaging. J Nucl Med. Feb 54(2), 252–8 (2013).
Winder, W. W. & Hardie, D. G. Inactivation of acetyl-CoA carboxylase and activation of AMP-activated protein kinase in muscle during exercise. Am J Physiol. 270, E299–E304 (1996).
Kudo, N. et al. Characterization of 5′AMP-activated protein kinase activity in the heart and its role in inhibiting acetyl-CoA carboxylase during reperfusion following ischemia. Biochim Biophys Acta. 1301, 67–75 (1996).
Guigas, B. et al. 5-Aminoimidazole-4-carboxamide-1-beta-D-ribofuranoside and metformin inhibit hepatic glucose phosphorylation by an AMP-activated protein kinase-independent effect on glucokinase translocation. Diabetes. 55(4), 865–74 (2006).
Cooper, D. R., Porebski, P. J., Chruszcz, M. & Minor, W. X-ray crystallography: Assessment and validation of protein small molecule complexes for drug discovery. Expert opinion on drug discovery 6, 771–782 (2011).
Malde, A. K. & Mark, A. E. Challenges in the determination of the binding modes of non-standard ligands in X-ray crystal complexes. Journal of computer-aided molecular design 25, 1–12. 2. 1 (2011).
Su, H. et al. Monitoring tumor glucose utilization by positron emission tomography for the prediction of treatment response to epidermal growth factor receptor kinase inhibitors. Clin. Cancer. Res. 12, 5659–5667 (2006).
Acknowledgements
This work was supported by MIUR (PRIN 2010JS3PMZ_009), the Italian Ministry of Health, Grant program 2008 (GMS), the Italian Association for Cancer Research (AIRC Start Up grant N. 6266).We thank Prof De Flora for his helpful discussion and suggestions.
Author information
Authors and Affiliations
Contributions
BS designed and performed most of the experiments, discussed the data, analysed immunofluorescence, western blot and biochemical experiments; CM produced, performed and analysed FDG uptake experiments; AD produced molecular modelling data, participated in the design and analysis of biochemical experiments; SR designed, produced and analysed biochemical data; MM produced FDG uptake experiments; AMO produced, analysed FACS experiments; AA performed RT-PCR experiments; MP performed immunofluorescence experiments; SM participated to perform western blot experiments; UP designed, produced and analysed RT-PCR data; RC, DM and GS initiated, conceived and directed the project. All authors discussed the results and collaborated to write the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Video pose1
Supplementary Information
Video pose2
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Salani, B., Marini, C., Rio, A. et al. Metformin Impairs Glucose Consumption and Survival in Calu-1 Cells by Direct Inhibition of Hexokinase-II. Sci Rep 3, 2070 (2013). https://doi.org/10.1038/srep02070
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02070
This article is cited by
-
Research Progress on the Use of Metformin in Leukemia Treatment
Current Treatment Options in Oncology (2024)
-
Early effects of LPS-induced neuroinflammation on the rat hippocampal glycolytic pathway
Journal of Neuroinflammation (2022)
-
Metformin alters therapeutic effects in the BALB/c tumor therapy model
BMC Cancer (2021)
-
Loss of hexokinase 1 sensitizes ovarian cancer to high-dose metformin
Cancer & Metabolism (2021)
-
Metabolic reprogramming by driver mutation-tumor microenvironment interplay in pancreatic cancer: new therapeutic targets
Cancer and Metastasis Reviews (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.