Abstract
Time-resolved quantitative analysis of auxin-mediated processes in plant cells is as of yet limited. By applying a synergistic mammalian and plant synthetic biology approach, we have developed a novel ratiometric luminescent biosensor with wide applicability in the study of auxin metabolism, transport and signalling. The sensitivity and kinetic properties of our genetically encoded biosensor open new perspectives for the analysis of highly complex auxin dynamics in plant growth and development.
Similar content being viewed by others
Introduction
Auxin is a key regulator of plant growth as well as of physiological and developmental processes from embryogenesis to maturity1,2. In the analysis of these processes it is of central importance to sensitively quantify the hormone at high spatial and temporal resolution. Auxin levels are typically analysed as average concentrations over whole tissues with biochemical or physicochemical methods such as isotope labelling and GC-MS3,4. Though sensitive and robust, these methods require disruption of tissues, thus preventing in vivo dynamic assays. Alternatively, the expression of reporter genes under control of hormone-responsive promoters enables the semi-quantitative monitoring of auxin homeostasis and signalling5,6,7. This approach, however, is still unsuitable for rapid analysis and may not reflect auxin levels directly but also the influence of other interfering genetic regulatory networks8. The conserved mechanism of auxin perception and signal transduction into transcriptional programs9 has recently been used for the development of genetically encoded biosensors8,10. Auxin-dependent formation of a co-receptor complex between TIR1/AFB (F-box proteins, constituents of an SCF E3 ubiquitin-ligase complex) and Aux/IAA (family of negative regulators of the auxin response) leads to the ubiquitylation and proteolysis of Aux/IAA, thus relieving the repression of auxin responsive genes11. Sensors relying either on the auxin-dependent formation of the co-receptor complex directly10 or on the further ubiquitylation and degradation of fluorescent proteins fused to Aux/IAA8,12, have been successfully applied to the study of signalling components and the mapping of relative auxin distribution in plant tissues at high spatial resolution. Despite these properties and a wide range of applications covered by the above-described methods, there is still an unmet need for tools that enable the time-resolved quantitative monitoring of auxin dynamics. Development of such a tool would lead to a major breakthrough for research on hormone metabolic, transport and signalling processes.
To meet these demands we have developed a chemiluminescent ratiometric sensor making use of the auxin-mediated interaction between TIR1 and Aux/IAA. By applying the degradation-based sensor in a transient gene expression cell system, e.g. protoplasts, a high temporal resolution is expected. Furthermore, luminescent reporters provide with large signal-to-noise ratios and hence high sensitivity as well as a necessary wide dynamic range13,14. Lastly, performing ratiometric quantifications contributes to the robustness of the tool by correcting for cell specific differences or the variability typical of transient assays15.
Results
Sensor development, evaluation and optimisation, in a mammalian cell system
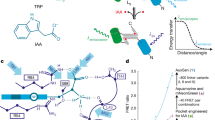
To fulfil the described specifications for a ratiometric auxin sensor, we designed a synthetic construct comprising firefly luciferase fused to an auxin-dependent degradation sequence/element of Aux/IAA proteins (sensor module) and renilla luciferase (normalisation element) (sensor design, Fig. 1a). Both components are linked by a 2A peptide16 that leads to their stoichiometrical co-expression, thereby allowing auxin-dependent degradation of the sensor to be monitored as a decrease in firefly relative to renilla luminescence (F/R) (Fig. 1a).
Sensor design principle and evaluation in a mammalian cell system.
(a) Ratiometric luminescent auxin biosensor. The sensor construct comprises two components: a sensor module (SM), fused to firefly luciferase and renilla luciferase. Both components are linked by a 2A peptide. 13-amino-acid minimal degradation sequences of three selected AtAux/IAA family members (AtIAA17, AtIAA28 and AtIAA31) that confer auxin-dependent degradation, were used as sensor modules (See also Supplementary Table 1). The 2A peptide allows for the stoichometrical co-expression of SM-firefly and renilla luciferase. Auxin concentration-dependent degradation of the sensor could be monitored as a decrease in firefly relative to renilla luminescence (F/R). (b) Characterisation of sensor variants in a mammalian cells system. HEK-293T cells were co-transfected with the min17-Luc, min28-Luc and min31-Luc sensor constructs and rice TIR1. 24 h after transfection, the cell culture medium was supplemented with 0, 1, 10 or 100 μM IAA and the cells were further incubated for 60 (upper panel) or 240 min (lower panel) prior to luciferase activity determination. Results are means ± s.e.m. (n = 4). Statistical significances for each sensor are indicated with lower case letters (one-way ANOVA, P < 0.01).
For the development of the sensor module of the auxin sensor we took into consideration essential features of the Aux/IAA proteins. A. thaliana has 29 Aux/IAA family members, most of them containing four conserved domains: domain I binds to the transcriptional repressor TOPLESS, domains III and IV are responsible for dimerisation and interaction with ARF transcription factors and domain II mediates auxin binding and degradation17. A multiple sequence alignment of Aux/IAA proteins identified a highly conserved 13-amino-acid consensus sequence within domain II, that proved to be sufficient for conferring auxin-dependent degradation18. In a pioneering work involving yeast-two-hybrid and quantitative in vitro pull down assays, Calderon-Villalobos et al were able to systematically determine that different Aux/IAA proteins form co-receptor complexes with TIR1/AFB F-box proteins with varying auxin-binding affinities, which are mainly determined by the Aux/IAA protein10. Moreover, the auxin-dependent stability of the Aux/IAA protein inversely correlates with the strength of the co-receptor complex10,19.
We thus profited on the described characteristics of the Aux/IAA proteins for the rational design of sensor modules: i) a series of synthetic sensor modules were engineered based on different Aux/IAA proteins to obtain sensors with different dynamic ranges of auxin sensitivity; ii) to minimise potential interferences of the sensor with endogenous auxin signalling components, only the 13-amino-acid minimal degron sequences of the different Aux/IAA proteins were used.
Accordingly, we used as sensor modules the 13-amino-acid degron sequences from three different Aux/IAAs spanning a wide range of sensitivities, namely AtIAA17, AtIAA28 and AtIAA31, with reported auxin binding affinities of 33 nM, 75 nM and >1,000 nM, respectively10. The minimal 13-amino-acid degron sequence of AtIAA17 coincides with the auxin-sensitive 13-amino-acids consensus sequence of 22 Aux/IAA proteins18 (QVVGWPPVRSYRK, sensor min17-Luc, Supplementary Table 1), while the ones from AtIAA28 and AtIAA31 deviate considerably from it (PVVGWPPVRSSRR and ARQDWPPIKSRLR, yielding the sensors min28-Luc and min31-Luc, respectively, Supplementary Table 1). A construct lacking the sensor module served as a negative control of the system (Ctrl-Luc, control construct, Supplementary Table 1).
The characterisation of the resulting sensors was first performed in a heterologous expression system to avoid possible interference from plant endogenous systems. Based on the high degree of conservation of the SCF-dependent proteasomal degradation machinery in eukaryotes20,21, we evaluated the sensors in human embryonic kidney 293T cells (HEK-293T). For this purpose, the different sensor constructs were co-transfected with rice TIR1, as the necessary co-receptor is absent from mammalian cells20. After 24 h, the culture medium was supplemented with 1, 10 or 100 μM indole-3-acetic acid (IAA) and after 60 and 240 min the response of the sensors was evaluated (Fig. 1b). The results show the functionality of the min17-Luc and min28-Luc sensors, which were able to detect IAA in the range of 1- to 10 μM after 240 min (Fig. 1b, lower panel). The min17-Luc sensor showed a higher sensitivity, namely 47% decrease when incubated with 10 μM IAA in comparison to 25% for min28-Luc (Fig. 1b, lower panel) and a faster response, detecting IAA already after 60 min (Fig. 1b, upper panel). On the other hand, the min31-Luc sensor did not show auxin-dependent degradation in the range of concentrations used, similarly to the negative control (Ctrl-Luc, Fig. 1b). The results indicate a clear correlation between the auxin-detection capabilities of the different sensors (min17-Luc > min28-Luc > min31-Luc) and the nature of the sensor module (minimal degron derived from AtIAA17, AtIAA28 and AtIAA31, respectively), in agreement with the experimental data on auxin-binding affinities for Aux/IAA family members10.
We selected the min17-Luc sensor, which showed the highest auxin sensitivity among tested sensors, for further characterisation and optimisation prior to its use in plant cells. In the first place, to address possible sterical effects at the N-terminus of the sensor module, additional versions of the min17-Luc sensor were constructed which include linkers varying in length and composition between the consensus and the 2A peptide sequence (L1min17-Luc, L2min17-Luc, L3min17-Luc, full sequences in Supplementary Table 1). Moreover, we also constructed a sensor containing full length AtIAA17 as sensor module (FL-Luc), to assess and to compare the auxin-sensing capabilities and consequent destabilization of the whole protein, in relation to the minimal degron module of the min17-Luc variants. Accordingly, we transfected HEK-293T cells with the different sensors along with rice TIR1. After 24 h, the culture medium was supplemented with 1, 10 or 100 μM IAA and after 5, 30 or 240 min incubation, the time- and dose-dependent response of the sensors was evaluated (Fig. 2 and Supplementary Fig. 1). All sensor variants exhibited quantitative responses in the range of IAA concentrations assayed after incubation for 240 min (Fig. 2, lower panel). In particular, the L2min17-Luc sensor showed the fastest response, detecting IAA in the 1- to 10 μM range already after 30 min of treatment (Fig. 2, upper panel). This response was abolished in the absence of TIR1 (Supplementary Fig. 2) and inhibited by MG132, thereby confirming the proteosome-dependent auxin-induced degradation of the sensor (Supplementary Fig. 3).
Characterisation of sensor variants in a mammalian cell system.
(a) HEK-293T cells were co-transfected with the indicated sensor constructs (FL-Luc, min17-Luc, L1min17-Luc, L2min17-Luc, L3min17-Luc and Ctrl-Luc) and rice TIR1 (see also Supplementary Table 1). 24 h after transfection, the cell culture medium was supplemented with 0, 1, 10 or 100 μM IAA and the cells were further incubated for 30 (upper panel) or 240 min (lower panel) prior to luciferase activity determination. Results are means ± s.e.m. (n = 4). Statistical significances for each sensor are indicated with lower case letters (one-way ANOVA, P < 0.001).
Characterisation and application of the ratiometric auxin sensor in plant cells
After having characterised and identified functional sensors in the orthogonal mammalian cell system, we evaluated whether their sensing responses were maintained upon expression in plant cells. For this purpose and based on its suitable biochemical, genetic and physiological characteristics22, we decided to use Arabidopsis leaf protoplasts as the transient gene expression system. We transformed the L2min17-Luc and FL-Luc constructs into protoplasts and after 24 h incubation in hormone free medium, IAA was added to the samples. The ratiometric determinations were performed following 5, 15 and 45 min treatments (Fig. 3a). The L2min17-Luc and FL-Luc sensors proved also functional in the plant system, showing ca. 70 and 40% F/R ratio decrease, respectively, after merely 15 min of treatment with IAA (Fig. 3a). The activity of Ctrl-Luc did not change in response to auxin, thereby confirming its function as a negative control of the system (Fig. 3a). The sensitivity of the sensors was higher in plant cells than in the orthogonal system. This observation could be related to the fact that, despite the high degree of conservation of the SCF-type E3 ubiquitin ligase components in mammalian and plant cells20, the efficiency of the SCFTIR1 complex formed in mammalian cells with the plant F-box protein TIR1, could be suboptimal. This aspect should be considered when working with approaches involving the use of heterologous systems. Nevertheless, the results obtained in the plant cells concur with the insight obtained with the mammalian cell system, validating the orthogonal approach as a versatile and efficient tool for the design, preliminary characterisation and optimization of plant biosensors.
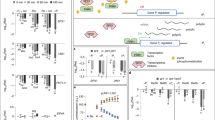
Characterisation and application of the ratiometric luminescent auxin sensor in plant cells.
(a) Initial characterisation of selected sensor variants in A. thaliana protoplasts. 24 h after transformation, protoplasts expressing FL-Luc, L2min17-Luc or Ctrl-Luc were incubated for 5, 15 or 45 min in either hormone-free or auxin-supplemented medium (1 μM IAA). Ratiometric luciferase activities are shown as means ± s.e.m. (n = 6). Statistical significances are indicated (two-sided t-test, * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001, n.s. non significant). (b) Sensitivity of the sensor towards different auxins. Protoplasts were transiently transformed with the L2min17-Luc sensor construct and after 24 h incubation, the culture medium was supplemented with either IAA or 1-naphtylacetic acid (NAA) in a pM to μM range. Luciferase activities were determined after the indicated periods of time. Results are means ± s.e.m. (n = 6). Statistical significances at each point in time are indicated with lower case letters (one-way ANOVA, P < 0.001). (c) Rapid determination of changes in auxin homeostasis. The sensor was co-expressed in protoplasts with either AtAUX1 or AtPIN1, auxin influx and efflux carriers, respectively. GFP was used as a negative control. 24 h after transformation, IAA in a pM to μM range was supplemented to the cell culture medium. After the indicated time periods, ratiometric luciferase determinations were performed. Results are means ± s.e.m. (n = 6).
In the absence of exogenously added auxin, the FL-Luc sensor exhibited considerably lower F/R values in comparison to L2min17-Luc (<20%), suggesting a higher rate of degradation of the full length AtIAA17 sensor module towards endogenous auxin levels in the plant cells (Fig. 3a). The engineered L2min17-Luc sensor on the other hand exhibited optimal characteristics in terms of dynamic range, sensitivity and temporal responses and was therefore selected for further characterization and applications. Additionally, the use of L2min17-Luc, which is based on the minimal degron sequence and thus depleted of other domains present in the FL-Luc sensor, minimises undesirable interactions with the endogenous components.
We then explored the capability of the L2min17-Luc sensor to monitor the bioactivity of natural or synthetic auxin analogues. For this purpose we compared the sensitivity of the sensor towards the natural auxin IAA, the synthetic analogues, 1-naphtylacetic acid (NAA) and 2,4-Dichlorophenoxyacetic acid (2,4-D) and the inactive auxin analogue 2-NAA. Protoplasts expressing the sensor were incubated with either compound at various concentrations for different time periods. The sensor could rapidly detect exogenously added IAA in the 10 nM to 1 μM range already after 5 min of treatment (Fig. 3b). Moreover, longer incubation times revealed a high sensitivity of the tool, namely threshold values in the low nM range. In comparison to IAA, the sensor showed a delayed and less sensitive response to NAA (Fig. 3b) and 2,4-D, the latter being detected only after 45 min treatment and at high concentrations (Supplementary Fig. 4). The non-active analogue 2-NAA, did not lead to sensor degradation in the whole range of concentrations and time points analysed (Supplementary Fig. 4). These results are in agreement with reported data indicating that the synthetic auxins NAA and 2,4-D though promoting TIR1/AFB-Aux/IAA interaction in vitro and in vivo, exhibit considerably lower binding affinities compared to IAA (2,4-D < NAA < IAA)10,23. Remarkably, in addition to its high sensitivity and responsiveness, the consensus sequence-based sensor is able to discriminate between different auxin analogues, unravelling its potential as a tool to study the metabolism, transport and signalling of physiologically active compounds.
The genetically encoded sensor was further applied to monitor the directed modulation of intracellular auxin levels upon modification of the transport machinery in transient expression assays. As a proof of principle, the sensor was co-expressed with either AtAUX1 or AtPIN1, plasma membrane localised auxin influx and efflux carriers, respectively24,25. As a control, the sensor was co-expressed with GFP, a protein not involved in auxin metabolism, in place of the transporters (expression controls, Supplementary Fig. 5). The results showed that in the presence of the influx carrier, AtAUX1, the sensor detected exogenously added IAA at lower concentrations (100 pM–1 nM) and at shorter treatment periods than the control (Fig. 3c). On the contrary, expression of the efflux carrier, AtPIN1, delayed the response and increased the concentration threshold of detection (Fig. 3c). The shift in sensor sensitivity correlates with the expected transport activity of the two proteins thus the results besides confirming the high sensitivity of the sensor, i.e. exogenously added auxin in the picomolar range, they depict its prospects for the study of genes involved in auxin-mediated processes.
Discussion
Quantitative monitoring of auxin in living cells has proven challenging. By utilising a ratiometric luminescent reporter system, we were able to develop a degradation-based biosensor to be used in single cell systems, fulfilling the requirements of high sensitivity and time resolution needed for studies of complex auxin-mediated processes. For this purpose, we applied a synergistic synthetic biology approach comprising the engineering and preliminary characterisation of the sensor in an interference-free heterologous mammalian cell system, followed by its successful application in plant cells.
We showed in developed proof of principle applications the suitability of the sensor for rapid, robust and highly sensitive auxin analysis. The sensor enabled quantitative time-resolved monitoring of intracellular auxin changes upon exogenous application of natural and synthetic auxins (chemical assay prototype) or carrier-mediated modulation of auxin levels in cells (genetic assay prototype). It is of note, that the range of sensitivity (dynamic range) of the sensor to exogenous IAA is within the physiological auxin concentrations and remarkably close to the dissociation constants of TIR1/AFB-Aux/IAA co-receptor complexes determined by in vitro quantitative IAA binding assays, as previously reported by Calderon-Villalobos et al10. Furthermore the differential sensitivity of the sensor towards synthetic auxins (2,4-D, NAA) correlates with the varying binding affinities reported for these compounds10,23. As shown in this work, the design principles applied allow for the engineering of synthetic sensor modules with tuneable dynamic ranges or even distinct specificities. The capacity of the sensor to perceive variations in intracellular auxin levels upon expression of the efflux carrier AtPIN1 or the influx carrier AtAUX1 provides an alternative to the more elaborate radioactive labelling transport assays26,27. Notably, to our knowledge this is the first report on AUX1 mediated auxin uptake assay in a plant cell system.
A series of indirect and direct analytical methods ranging from radiolabelling, microelectrode-based analysis to GC-MS in combination with fluorescent activated cell sorting, are currently available for quantitative auxin determinations3,4. These approaches are highly sensitive and accurate, but at the same time they present intrinsic drawbacks such as the necessity of disrupting cells or tissues, often complex and time-demanding preparation procedures, discontinuity, or high maintenance and running costs for the equipments. In addition to the high sensitivity and selectivity of these tools, the genetically encoded luminescent auxin biosensor here described allows for the time-resolved quantitative monitoring of auxin dynamics in living cells. Moreover, the sensor-based assay platform is technically simple, requires only a relatively low cost luminescence plate reader and thus this approach is amenable for straightforward establishment in an ordinary plant research laboratory.
The sensor here presented shares its functional principle with the recently reported DII-Venus auxin biosensor. Both sensors rely on the auxin-dependent formation of a TIR1/AFB-Aux/IAA-reporter like complex. While the DII-Venus sensor provides with unmatched high-resolution spatio-temporal assessment of auxin responses in planta8, our tool is optimised for the use in single cell systems. It combines a luminescent reporter with an internal normalization element, for performing highly sensitive and more quantitatively accurate determinations, as described in this work. The complementary features of both approaches could therefore synergistically be used in benefit of more comprehensive studies on auxin dynamics in plants.
We illustrate here the greater potential of this novel auxin sensor that, in combination with other available methods and molecular/genetic tools such as mutants, RNAi approaches, analogues and inhibitors, will generate quantitative datasets essential for comprehensive understanding of auxin homeostasis and signalling networks. Moreover, the luminescent biosensor in combination with the plant cell platform is suitable for multi-well plate format assays allowing for mid-to high-throughput dynamic analysis in genetic and chemical screenings.
The remarkable kinetic and sensitivity properties of our genetically encoded biosensor, together with its wide applicability and versatility, not only represent an improvement to currently available methods for auxin determination but also provide with novel perspectives. Furthermore, an adaptation of this system to the analysis of other hormones sharing the hormone-induced proteolytic mechanism, such as jasmonic acid and gibberellins1, opens up new research prospects for the plant hormone field.
Methods
Plasmid construction
The different sensor constructs were obtained by fusion of PCR-generated amplicons into the mammalian expression vector TOPO-pEF5 (Invitrogen, US). Renilla and firefly luciferases were amplified from the plasmids pRL-TK (Promega, US) and pGL3-basic (Promega, US), respectively. The minimal degradation consensus sequences and all linkers were introduced into PCR primers. The pNHK6020 plasmid comprising AtAUX/IAA17 (GeneID839568) and OsTIR1 (GeneID3435696) cDNA were kindly provided by M. Kanemaki, Osaka University, Japan. The corresponding amino acid sequences of all components of the sensors are provided in Supplementary Table 1. For construction of the plant expression vectors, the different sensor constructs were recombined into the Gateway28 compatible binary vector pMIR, a derivative of the pAM-PAT expression vector (GenBank accession AY436765.1). For co-expression of AtAUX1 (GeneID818390), AtPIN1 (GeneID843693) or GFP, the corresponding cDNA sequences were introduced by Gateway cloning into pMIR vectors containing the L2min17-Luc construct. Gene expression in all cases was driven by the CaMV 35S promoter.
Mammalian cell culture, transfection and auxin treatment
Human embryonic kidney 293-T cells (HEK-293T) were cultivated in DMEM (DMEM, PAN Biotech GmbH, Aidenbach, Germany, cat. No. P03-0710), 10% FCS (PAN Biotech GmbH, Aidenbach, Germany, cat. no.1502, batch P123002) and 1% (v/v) penicillin/streptomycin (PAN Biotech GmbH, Aidenbach, Germany, cat. no. P06-07100). HEK-293T cells were cultivated at 37°C in a humidified atmosphere containing 5% CO2 and transfected using an optimised polyethylenimine-based protocol29 (PEI, linear, MW: 25 kDa, Polyscience, US). In brief, 1 mg/ml PEI was dissolved in H2O and adjusted to pH 7.0 with HCl, sterile filtered and stored in aliquots at −80°C. 60,000 cells per well were seeded in a 24-well plate and cultivated for 24 h. For transfection of one well, 0.75 μg of DNA was diluted in 50 μl of OptiMEM (Life Technologies, US) and subsequently mixed with 2.5 μl of PEI solution while vortexing. After 15 min at room temperature this mixture was added to the cells. To induce auxin-mediated protein degradation, appropriate auxin dilution series were prepared in DMEM and added to the cells 24 h post transfection and incubated for designated periods. MG132 was prepared as 20 mM stock in DMSO and added to cell culture medium to a final concentration of 20 μM.
Plant material
A. thaliana (Col-0) seeds were plated in a line on autoclaved filter paper stripes (200–300 seeds/stripe) placed on 12 cm square plates (Greiner Bio-One, Germany) containing SCA culture medium30. After 24 h incubation at 4°C, the plates were placed in a growth chamber, with a 16 h light regime at 21°C. 2 to 3-week old plantlets were used for protoplast isolation.
Protoplast isolation, transformation and auxin treatment
Plant shoots were dissected with a sterile scalpel and cut into small pieces in 2 ml of MMC medium30. Tissue pre-plasmolysis, digestion and isolation prior the flotation step were performed according to Dovzhenko et al. (2003), with centrifugation for 15 min at 100 g (Eppendorf 5810R Centrifuge, Eppendorf, Germany). After flotation, protoplasts were transferred to a 12 ml tube (Greiner Bio-One, Germany) and the total volume adjusted to 10 ml with W5 medium31. After cell number evaluation, protoplasts were sedimented for 10 min at 100 g. After discarding the supernatant, protoplast density was adjusted to 107 cells ml−1 with the transformation medium (TM: 5 mM MES, 15 mM MgCl2, 600 mOsm with mannitol, pH 5.8). The following modifications were implemented for PEG-mediated DNA uptake procedure: 10 μg of plasmid DNA were resuspended in TM (final volume of 50 μl) and mixed with 100 μl of protoplast suspension with subsequent 5 min incubation at room temperature (23–25°C) in 3.5 cm culture dishes or 6-well plates (Greiner Bio-One, Germany). 100 μl of 40% PEG1500 were added drop-wise for 8 min treatment. Afterwards, the total volume was adjusted to 2.5 ml with PCA-0 (hormone free PCA medium30). Protoplasts were incubated in the dark at 25°C for 20 to 24 h prior to auxin treatment. 100 μl aliquots were transferred to 2 ml deep-well storage plates (ABgene, Germany), mixed in a 1:1 ratio with PCA-M medium (PCA salts, 600mOsm mannitol, pH 5.8) containing appropriate auxin dilutions and incubated for the designated time periods.
Analysis of transgene expression
20 h. after transformation, 1*106 protoplasts for each construct were collected by centrifugation and used for RNA isolation with a RNeasy plant mini kit (Qiagen, Germany), according to the manufacture instructions, with an additional DNAse treatment. For the RT-PCR, equal amount of purified RNA was used for cDNA synthesis with Maxima Reverse Transcriptase (Fermentas, Germany) with oligodT primers and then for a PCR reaction with the primers 5′-ggtggtggtcggaactctaa and 5′-tagcaggaccaccgtcttct for AtPIN1, 5′- ttgggttcggtggatgggct and 5′-aagcggcgaccggggcatgt for AtAUX1, and 5′-tggaccgctcttatcaaagg and 5′-cttgaggaggttgcaaagga for AtPEX4. Equal amounts of each reactions were loaded on a 2% agarose gel.
Inducers
Indole-3-acetic acid (IAA) (Sigma Aldrich, St. Louis, MO) was prepared as 57 mM stock in 95% ethanol. 1-naphtylacetic acid (NAA) was purchased as 5.37 mM sterile stock solutions (Sigma Aldrich, St. Louis, MO). 2,4-Dichlorophenoxyacetic acid (2,4-D) (Duchefa Biochemie, Netherlands) was prepared as a 200 mM stock in 95% ethanol. 2-naphtylacetic acid (2-NAA) (Sigma Aldrich, St. Louis, MO) was prepared as a 200 mM stock in 1 M NaOH.
Luminescence analysis
In order to determine luciferase activity, mammalian cells were first lysed by addition of 250 μl luciferase lysis buffer (25 mM Tris-HCl pH 7.8, 1% Triton X-100, 15 mM MgSO4, 4 mM EGTA, 1 mM DTT) per well and incubation at 4°C for 10 min. 80 μl mammalian cell lysate were transferred to Costar® 96-well flat-bottom white plate (Corning Incorporated, Germany). After indicated incubation times, 80 μl of protoplast suspensions were directly used for luminescence determinations. Firefly and Renilla luminescence was directly monitored using either a Synergy 4 multimode microplate reader (BioTek Instruments Inc., Winooski, VT) or a Mithras LB940 luminescence reader with integrated injectors (Bertold Technologies, Germany) after addition of 20 μl of either firefly luciferase substrate (20 mM Tricine, 2.67 mM MgSO4, 0.1 mM EDTA, 33.3 mM DTT, 0.52 mM ATP, 0.27 mM Acetyl-CoA, 5 mM NaOH, 50 mM MgCO3, 0.47 mM luciferin) or renilla luciferase substrate (472 μM coelenterazine stock solution in methanol; diluted directly before use, 1:15 in PBS).
Statistical analysis
One-way ANOVA or t-test analysis, were performed with Matlab 7.120.635 (R2011a). Corresponding p-values are indicated in the figure legends.
References
Santner, A., Calderon-Villalobos, L. I. & Estelle, M. Plant hormones are versatile chemical regulators of plant growth. Nat Chem Biol 5, 301–307 (2009).
Kieffer, M., Neve, J. & Kepinski, S. Defining auxin response contexts in plant development. Curr. Opin. Plant Biol. 13, 12–20 (2010).
Petersson, S. V. et al. An auxin gradient and maximum in the Arabidopsis root apex shown by high-resolution cell-specific analysis of IAA distribution and synthesis. Plant Cell 21, 1659–1668 (2009).
Barkawi, L. S., Tam, Y. Y., Tillman, J. A., Normanly, J. & Cohen, J. D. A high-throughput method for the quantitative analysis of auxins. Nat Protoc 5, 1609–1618 (2010).
Ulmasov, T., Murfett, J., Hagen, G. & Guilfoyle, T. J. Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9, 1963–1971 (1997).
Sorefan, K. et al. A regulated auxin minimum is required for seed dispersal in Arabidopsis. Nature 459, 583–586 (2009).
Barbez, E. et al. Single-cell-based system to monitor carrier driven cellular auxin homeostasis. BMC Plant Biol 13, 20 (2013).
Brunoud, G. et al. A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482, 103–106 (2012).
Vanneste, S. & Friml, J. Plant signaling: Deconstructing auxin sensing. Nat Chem Biol 8, 415–416 (2012).
Calderon Villalobos, L. I. et al. A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nat Chem Biol 8, 477–485 (2012).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Havens, K. A. et al. A synthetic approach reveals extensive tunability of auxin signaling. Plant Physiol. 160, 135–142 (2012).
Stefan, E. et al. Quantification of dynamic protein complexes using Renilla luciferase fragment complementation applied to protein kinase A activities in vivo. Proc. Natl. Acad. Sci. U. S. A. 104, 16916–16921 (2007).
Fan, F. et al. Novel genetically encoded biosensors using firefly luciferase. ACS Chem Biol 3, 346–351 (2008).
Jacobs, J. L. & Dinman, J. D. Systematic analysis of bicistronic reporter assay data. Nucleic Acids Res 32, e160 (2004).
de Felipe, P. et al. E unum pluribus: multiple proteins from a self-processing polyprotein. Trends Biotechnol. 24, 68–75 (2006).
Overvoorde, P. J. et al. Functional genomic analysis of the AUXIN/INDOLE-3-ACETIC ACID gene family members in Arabidopsis thaliana. Plant Cell 17, 3282–3300 (2005).
Ramos, J. A., Zenser, N., Leyser, O. & Callis, J. Rapid degradation of auxin/indoleacetic acid proteins requires conserved amino acids of domain II and is proteasome dependent. Plant Cell 13, 2349–2360 (2001).
Gray, W. M., Kepinski, S., Rouse, D., Leyser, O. & Estelle, M. Auxin regulates SCF(TIR1)-dependent degradation of AUX/IAA proteins. Nature 414, 271–276 (2001).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat Methods 6, 917–922 (2009).
Kämpf, M. M. et al. Rewiring and dosing of systems modules as a design approach for synthetic mammalian signaling networks. Mol Biosyst 8, 1824–1832 (2012).
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2, 1565–1572 (2007).
Kepinski, S. & Leyser, O. The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435, 446–451 (2005).
Gälweiler, L. et al. Regulation of polar auxin transport by AtPIN1 in Arabidopsis vascular tissue. Science 282, 2226–2230 (1998).
Swarup, R. et al. Localization of the auxin permease AUX1 suggests two functionally distinct hormone transport pathways operate in the Arabidopsis root apex. Genes Dev. 15, 2648–2653 (2001).
Yang, H. & Murphy, A. S. Functional expression and characterization of Arabidopsis ABCB, AUX 1 and PIN auxin transporters in Schizosaccharomyces pombe. Plant J. 59, 179–191 (2009).
Petrasek, J. et al. PIN proteins perform a rate-limiting function in cellular auxin efflux. Science 312, 914–918 (2006).
Karimi, M., Inze, D. & Depicker, A. GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7, 193–195 (2002).
Müller, K. et al. A red/far-red light-responsive bi-stable toggle switch to control gene expression in mammalian cells. Nucleic Acids Res (2013).
Dovzhenko, A., Dal Bosco, C., Meurer, J. & Koop, H. U. Efficient regeneration from cotyledon protoplasts in Arabidopsis thaliana. Protoplasma 222, 107–111 (2003).
Menczel, L., Galiba, G., Nagy, F. & Maliga, P. Effect of radiation dosage on efficiency of chloroplast transfer by protoplast fusion in Nicotiana. Genetics 100, 487–495 (1982).
Acknowledgements
This work was supported by the Excellence Initiative of the German Federal and State Governments (EXC 294 and GSC-4, Spemann Graduate School), the Initiating and Networking Fund (IVF) of the Helmholtz Association within the Helmholtz Initiative on Synthetic Biology (SO-078), the Bundesministerium für Bildung und Forschung (Sysbra, Systec), the Baden-Württemberg Stiftung, the German Research Foundation (DFG) grants SPP1395 Inkombio and SFB746 and the Alexander von Humboldt Foundation Research Grant, no. 1141629. We are indebted to Dr. Claude Becker for vector development and to Sophia Samodelov for careful reading of the manuscript.
Author information
Authors and Affiliations
Contributions
S.W., C.D.B., M.M.K., F.R., K.P., W.W., A.D. and M.D.Z. designed experiments. S.W., C.D.B., M.M.K., F.R., A.D. and M.D.Z. performed experiments and analysed data. S.W., C.D.B., A.D. and M.D.Z. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Info
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Wend, S., Bosco, C., Kämpf, M. et al. A quantitative ratiometric sensor for time-resolved analysis of auxin dynamics. Sci Rep 3, 2052 (2013). https://doi.org/10.1038/srep02052
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02052
This article is cited by
-
NERNST: a genetically-encoded ratiometric non-destructive sensing tool to estimate NADP(H) redox status in bacterial, plant and animal systems
Nature Communications (2023)
-
Design principles for synthetic control systems to engineer plants
Plant Cell Reports (2023)
-
Variation in auxin sensing guides AUX/IAA transcriptional repressor ubiquitylation and destruction
Nature Communications (2017)
-
Regulation of seedling growth by ethylene and the ethylene–auxin crosstalk
Planta (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.