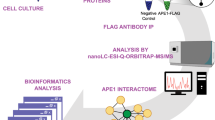
Abstract
IDPs, while structurally poor, are functionally rich by virtue of their flexibility and modularity. However, how mutations in IDPs elicit diseases, remain elusive. Herein, we have identified tumor suppressor PTEN as an intrinsically disordered protein (IDP) and elucidated the molecular principles by which its intrinsically disordered region (IDR) at the carboxyl-terminus (C-tail) executes its functions. Post-translational modifications, conserved eukaryotic linear motifs and molecular recognition features present in the C-tail IDR enhance PTEN's protein-protein interactions that are required for its myriad cellular functions. PTEN primary and secondary interactomes are also enriched in IDPs, most being cancer related, revealing that PTEN functions emanate from and are nucleated by the C-tail IDR, which form pliable network-hubs. Together, PTEN higher order functional networks operate via multiple IDP-IDP interactions facilitated by its C-tail IDR. Targeting PTEN IDR and its interaction hubs emerges as a new paradigm for treatment of PTEN related pathologies.
Similar content being viewed by others
Introduction
The concept of “Intrinsic Disorder” in proteins has rapidly gained attention as the preponderance and functional roles of IDPs are increasingly being identified in eukaryotic proteomes1,2. Structured proteins adopt energetically stable three-dimensional conformations with minimum free energy. In contrast, IDPs, due to their unique amino acid sequence arrangements, cannot adopt energetically favorable conformations and, thus, lack stable tertiary structure in vitro3. This structural plasticity allows IDPs to operate within numerous functional pathways, conferring multiple regulatory functions4,5,6. Indeed, mutations in and dysregulation of IDPs are associated with many diseases including cancer1,6,7, signifying that IDPs play vital roles in functional pathways. Evidence suggests that ~80% of proteins participating in processes driving cancer contain IDRs6. For example, tumor suppressor p53 as an IDP, functions via its C-terminal IDR, which simultaneously exists in different conformations, each of which function differently1. Since PTEN is the second most frequently mutated tumor suppressor with versatile functions8, we hypothesized that PTEN may contain IDR(s) that can be exploited for therapeutic targeting in cancers and diseases associated with pathogenic PI3K/Akt/mTOR (Phosphoinositide 3-Kinase/Akt/ mammalian Target of Rapamycin) signaling9,10,11.
PTEN (phosphatase and tensin homolog), a 403 amino acid dual protein/lipid phosphatase converts phosphatidylinositol(3,4,5)-triphosphate (PIP3) to phosphatidylinositol(4,5)-bisphosphate (PIP2), thereby regulating the PI3K/Akt/mTOR pathway involved in oncogenic signaling, cell proliferation, survival and apoptosis12. PTEN, as a protein phosphatase, autodephosphorylates itself13. Deficiency or dysregulation of PTEN drives endometrial, prostate, brain and lung cancers and causes neurological defects14,15. PTEN is activated after membrane association16, providing conformational accessibility to the catalytic phosphatase domain (PD) that converts PIP3 to PIP216 (Figure 1a). Because PTEN reduces PIP3 levels and inhibits pathogenic PI3K signaling, therapeutically targeting PTEN to the membrane to enhance its activity is of significance in treating several pathologies including cancer.
PTEN: A newly identified IDP.
(a) Diagrammatic representation of PTEN structure. PTEN, a 403 amino acid protein, comprises of PBM: PIP2 Binding Module (AA 1–13; in green), a phosphatase Domain (AA 14–185; in pink), C2 Domain (AA 190–350; in blue), C-terminal region or Tail (AA 351–400; in orange) and a PDZ binding domain (AA 401–403; in dark blue). The PDZ-binding motif is considered as a part of the C-terminal region. *Figure not to scale. (b) Crystal structure of PTEN. Only the phosphatase (in pink) and C2 domain (in blue) are amenable to crystallization. The first seven residues and the last 50 residues represent unstructured/loosely-folded regions that are yet to be crystallized. These regions represent the N- and C-termini of PTEN, respectively. (Source: RCSB Protein Data Bank). (c) Disorder analysis of PTEN. PONDR-VLXT and PONDR-FIT prediction tools were used to determine the disorder score of PTEN. Any value above 0.5 indicates intrinsic disorder. There are several disordered stretches within the PTEN protein, however, the most prominent of these disordered regions is a 50 amino-acid stretch located at the C-terminus of the PTEN protein. (d) IDPs are enriched in polar (R, Q, S, T, E, K, D, H) and structure breaking (G, P) amino acids and are depleted in hydrophobic (I, L, V, M, A), aromatic (Y, W, F) and cysteine (C) and asparagine (N) residues. The amino acid sequence of PTEN highlights these classes of residues with their relative distribution. (e) Composition profiling for full-length PTEN (in green), its ordered domain (in yellow) and its IDR (in red). The tool used is Composition Profiler (Vacic et al, 2007). As shown in the graph, the disordered region in PTEN is enriched in polar residues (specifically H, T, D, S and E), structure breaking residues (specifically P) and is depleted in all hydrophobic residues, cysteine and all aromatic residues. (f) Histogram representing the percentage of hydrophobic, polar, aromatic, structure breaking, cysteine and asparagines residues in ordered vs. disordered regions. The disordered region has an amino acid composition in line with the definition of IDPs.
PTEN crystal structure revealed that the PD and membrane-binding C2 domains are ordered (Figure 1b); however, the structures of the N-terminus, the CBR3 loop and the 50 amino-acid C-tail remain undetermined17. The C-tail is of particular significance due to its ability to regulate PTEN membrane association, activity, function, stability18,19,20,21. Herein, we identify PTEN as an IDP with its C-tail being intrinsically disordered. The PTEN C-tail IDR is heavily phosphorylated by a number of kinases and regulates the majority of PTEN functions, including a large number of PPIs that forms the PTEN primary and secondary interactomes, comprising critical functional protein hubs, most of which are related to cancer. Our analysis provides a mechanistic insight into the functioning of the PTEN C-tail IDR at the systems level, including inter- and intra-molecular interactions that will aid in designing drugs to enhance the lipid phosphatase activity of PTEN for the pharmacotherapy of cancers and pathological conditions driven by hyperactive PI3K-signaling.
Results
PTEN is an IDP
Utilizing two disorder prediction software programs, PONDR-VLXT and PONDR-FIT22,23, we have identified PTEN as a bona fide IDP. PTEN has a highly disordered, functionally versatile, C-tail encompassing amino acids 351–403 (Figure 1a and 1c). A PDZ-binding motif (amino acids 401–403) is part of the disordered region. Thus, the PTEN C-tail IDR facilitates interactions with a vast repertoire of PDZ domain-containing proteins (Figs. 1a and 2d). The unique amino acid composition of IDRs dictates their structural plasticity3,23,24. IDRs are enriched in polar and structure-breaking amino acid residues, depleted in hydrophobic and aromatic residues and, rarely, contain Cys and Asn residues1,23,24. The ordered region of PTEN (AA 1–350) has 25% hydrophobic, 43% polar, 9% structure breaking, 13% aromatic and 9% Cys and Asn residues. In contrast, the PTEN C-tail (AA 351–403) is enriched in polar (66%) and structure breaking (11%) residues and is depleted in hydrophobic (11%), aromatic (6%) and Cys and Asn residues (6%), indicating an ideal profile for the IDR (Figs. 1d and 1f ). Further, compositional analysis of PTEN using the Composition Profiler24 reveals that the disordered region in PTEN is enriched in polar residues (specifically H, T, D, S and E) and structure breaking residues (specifically P) but is depleted in all aromatic and hydrophobic residues in addition to cysteine. (Figure 1e), again exhibiting universal characteristics of IDPs. Taken together, we establish the PTEN C-tail as a functional IDR and classify PTEN as a new IDP.
The functional relevance of the PTEN IDR.
(a) The number of mutations observed in PTEN over its 403 amino-acid stretch is plotted. Fewer mutations are observed in the tail region (in red) possibly indicating the deleterious nature of mutations in the functionally critical C-terminal region. [Source: Sanger Institute Catalogue of Somatic Mutations in Cancer (COSMIC), Human Gene Mutation Database (HGMD)]. (b) Number of mutations in every successive 50 amino-acid stretch of the PTEN protein. The last 50 amino-acid stretch, representing the tail region has at least one-eighth the number of mutations seen in any other 50 amino-acid stretch along PTEN, pointing to its critical function in cell homeostasis. (c) Correlation of mutations with the amino acid composition of PTEN. The ratio of mutations in specific residues in the disordered vs. ordered region are represented in this graph. The residues considered here are those used to define IDRs: hydrophobic, polar, aromatic, structure-breaking, cysteine and asparagine residues. Compared to the other classes of residues, mutations in aromatic residues are much higher in the disordered region when compared to the ordered region. (d) The PTEN primary interactome. Forty proteins interact with known regions of PTEN. There are approximately 340 more proteins that interact with PTEN at sites that are yet to be determined (see Supplementary Table S2). Proteins shown in pink interact with the phosphatase domain, those in blue interact with the C2 domain and those in orange interact with the disordered tail. (Visualization tool: Cytoscape). (e) The PTEN C-tail has a higher propensity for PPIs. Of the 40 mapped proteins, 60% interact with the disordered indicating a strong correlation between degree of disorder and the number of protein interactions. (f) Most proteins within the PTEN interactome are highly disordered. Approximately 80% of PTEN-interacting proteins within the primary interactome are disordered, as indicated in red. The proteins within the interactome that are ordered are indicated in blue.
Low mutability of PTEN IDR suggests critical biological functions
Mutations in PTEN are associated with several types of cancers14. To correlate PTEN mutations to its structure, we analyzed all human PTEN mutations deposited in the COSMIC Database (http://www.sanger.ac.uk/genetics/CGP/cosmic/). The disordered PTEN C-tail IDR shows unusually low mutability (~8-fold less) compared to any other 50 amino-acid stretch of PTEN (Figure 2a and 2b). To confirm our finding of the low mutability of the C-tail region, we also analyzed all human PTEN mutations deposited in the Human Gene Mutation Database (HGMD, http://www.hgmd.cf.ac.uk/ac/index.php)25 (Figure 2a), cBioPortal for Cancer Genomics26,27 (Supplementary Figure S1) and the Roche Cancer Genome Database28 (Supplementary Figure S1) which was consistent with the COSMIC database mutational data. It is likely that evolutionary pressure maintains a survival advantage and ipso facto abrogates progeny with mutations in highly functional protein sequences29,30,31. Thus, the functionally versatile PTEN C-tail IDR cannot afford mutations, hence showing least number of mutations. It is equally likely that mutations in individual residues within the IDR are well tolerated, as the evolutionary pressure may have shifted to maintaining global biophysical properties and structural malleability of the IDR to safeguard the critical protein function29. In either case, on a global scale, the versatile structural pliability of the PTEN IDR dictates functional diversity and biological activities29. Thus, the slightest functional perturbation in the PTEN IDR due to mutations, either within the IDR or in domains interacting with it, could disrupt cellular homeostasis as seen in cancers and neurodegenerative disorders associated with PTEN mutations. This is supported by our data indicating that PTEN, as an IDP when mutated, causes several cancers14.
Moreover, the PTEN C-tail IDR exhibits preferential mutations in aromatic residues compared to the ordered region (Figure 2c). The ratio of mutations in aromatic residues in the disordered to ordered region is much higher than any other class of residues (structure breaking, hydrophobic, polar, Cys and Asn), likely attributed to the structure-imparting property of aromatic residue32. Specifically, aromatic residues within IDRs engage in stacking interactions, enhancing nucleation between distinct residues at functional protein-protein interaction interfaces32. Thus loss of this critical structural and functional property imparted by aromatic residues is associated with a disease phenotype. In summary, the disordered PTEN C-tail IDR has functionally evolved to contain a combination of peptides that cannot tolerate mutations.
Disorderliness in PTEN primary interactome drives functional networks
Protein-Protein Interactions (PPIs) typically occur between conserved, structurally rigid regions of two or more proteins, particularly ordered proteins that display energetically favorable, highly-folded conformations. Intriguingly, IDPs lack tertiary structure, yet engage in PPIs, albeit with lower affinities but high specificity1. The lack of structure within IDPs enhances their biophysical landscape, conferring them with the ability to attain structural complementarities required for PPIs. Since IDPs do not conform to a stable structure, they are less compact, providing a larger physical interface and energetic adaptability to interact with multiple proteins1,7. Thus, conditional folding within IDPs is effectively utilized for interaction with a multitude of binding partners, enabling them to shuttle between several signaling cascades as efficient “cogs”, mediating and regulating PPIs4,7,33,34,35,36. Indeed, we discovered that PTEN, being an IDP, interacted with more than 400 proteins (Supplementary Table S1) when a combination of online software, literature search and database mining tools were used. Proteins with known PTEN interaction domains were classified as “mapped” (Figure 2d and Supplementary Table S1), whereas those with uncharacterized/predicted interactions were designated as “unmapped” proteins (Supplementary Table S1). Derivation of PTEN primary interactome from the mapped proteins using Cytoscape (http://www.cytoscape.org/) indicated that PTEN disorderliness is efficiently used for interaction with 40 proteins, most existing in distinct functional pathways (Figure 2d, 2e and Supplementary Table S2).
Interestingly, within the PTEN primary interactome, 60% of interactions occurred within the disordered C-tail region. Furthermore, disorder analysis on the primary interactome revealed that 33 proteins (>82%) were IDPs, of which two-thirds interacted with the C-tail IDR (Figure 2e, 2f and Supplementary Table S3), indicating a high propensity for disorder-disorder (D-D)-type interactions.
In order to study evolutionary conservation of the PTEN C-tail and its interactions across species, several sequence alignments were performed (Figure 3a). Sequence alignment of the entire PTEN protein from different animal species shows a good conservation of the catalytic phosphatase domain between vertebrates and invertebrates with 100% sequence conservation for the dual specificity phosphatase catalytic motif HCKAGKGR8 (Supplementary Figure S2). The C-tail shows good conservation in the vertebrate species, likely indicating the recent emergence of the function of PTEN C-tail region in regulating PTEN activity and enriching its PPI potential, translating to its versatile functions. In order to examine the conservation across species for the PTEN C-tail interacting proteins, a literature search was conducted to identify experimentally verified domains/motifs involved in interaction with the C-tail. The domains involved in these interactions with the C-tail for 13 proteins with relevant literature sources for these interactions are part of Supplementary Figure S3. Subsequent sequence alignments for these thirteen proteins (Supplementary Figure S3) shows good sequence homology for the domains/motifs involved in interaction with the PTEN C-tail. These findings support the concept that the PTEN C-tail has evolved in vertebrates to incorporate features that allow it to interact with these proteins.
Sequence conservation in PTEN and its interacting partners reflects functionality.
(a) Sequence alignment of the PTEN protein for vertebrate and invertebrate animals. Green color indicates sequence similarity while red indicates sequence dissimilar amino acid residues. All comparisons are made with respect to the human PTEN protein. (b) Network analysis for PTEN was performed to assess its potential as a network hub. The network shows multiple secondary interactions within the 40 mapped proteins, indicating their role in multiple signaling cascades mediated via PTEN. The proteins SMAD2/3, AR, PCAF, ANAPC7, B-arrestin 1 and p53 appear to be critical within these signaling cascades and also happen to be intrinsically disordered (Supplementary Table S3), reinforcing the concept of preferential interactions between disordered proteins. (Analysis Tool: Metacore by GeneGo).
Further, to assess whether PTEN acts as a functional hub protein and regulates pathways through its protein-binding partners, we performed functional network analysis using the Analyze Network option from MetaCore (GeneGo Inc, Thomson Reuters, 2011) (Figure 3b). The PTEN primary interactome was used as input with PTEN as the central node. We identified multiple interactions not only between PTEN (node) and SMAD2/3, AR, PCAF, ANAPC3, ANAPC4, Caveolin, β-arrestin 1 and p53 (edges), but also amongst the edge proteins themselves (Figure 3b). Interestingly, all the edge proteins are themselves highly disordered (Supplementary Table S3). Further supporting this finding, our functional enrichment revealed that 13 proteins (one-third) of the PTEN primary interactome were cancer-related and highly disordered (Figure 4a, Supplementary Table S3 and S4).
Derivation and disorder analysis of the PTEN cancer interactome.
(a) Derivation of the PTEN Cancer Interactome. Functional enrichment of the PTEN primary interactome identified 13 cancer-related proteins which are also intrinsically disordered. Subsequently, the PTEN secondary interactome was derived from the primary PTEN interacting proteins. A subset of the secondary interactome was designated as the PTEN Cancer Interactome and it represents the proteins that interact with the 13 cancer-related proteins of the primary interactome. (b) PTEN Cancer Interactome. PTEN is the primary node that interacts with the 13 cancer-related proteins representing the partial primary interactome. Proteins that interact with each of the 13 cancer-related proteins comprise the secondary interactome. Disordered proteins are represented in red while ordered proteins are shown in blue. Cancer-related proteins in the PTEN primary interactome were identified using IPA (Ingenuity® Systems, www.ingenuity.com). (c) We identified 40 proteins that are part of the PTEN primary interactome of which 13 are highly disordered (IDP) and identified as potential cancer network hubs based on functional network analysis. We further identify 299 IDPS from the secondary PTEN interactome. A filter for cancer-related proteins revealed that approximately two-thirds of the IDPs that form the secondary interactome (193 out of 299) are involved in oncogenesis, suggesting a high degree of functional enrichment. (Functional network analysis was performed using IPA (Ingenuity® Systems, www.ingenuity.com).
Pliant PTEN secondary interactome relays function of the primary network
The disorderliness of the PTEN primary interactome prompted us to investigate the possibility that PTEN radiates its function via a malleable network of IDPs that extends beyond the primary interactome. Therefore, we derived the PTEN secondary interactome (Supplementary Table S5) and ascertained the interaction of 13 cancer-related proteins identified in the primary interactome (Figure 4a). The entire PTEN secondary interactome consisted of 299 IDPs, of which 193 IDPs (two-thirds) were associated with the 13 cancer-related proteins, generating a “PTEN-Cancer Interactome” (Figure 4, Supplementary Table S5 and S6). Thus, two-third of the IDPs within the PTEN secondary interactome associates with one-third of the cancer related IDPs within the PTEN primary interactome, indicating that cancer-related functions are driven by IDPs in the PTEN interactome and that the flexibility of IDP-IDP interactions modulates diverse functions; dysregulation of which causes cancers.
Functional network analysis of the 193 cancer-related IDPs identified 31 proteins that shared multiple nodes (Figure 5a and Supplementary Table S6). We overlaid this network with the cancer-related IDPs of the primary interactome to predict functionally critical protein hubs (indicated in yellow circles in Figure 5a and b). Our analysis revealed 16 proteins as highly populated hubs, most enriched in disordered regions, again demonstrating that a high degree of structural and functional association between the hubs required IDP-IDP interactions (Figure 5b). The involvement of these hubs in multiple, critical oncogenic signaling pathways make them attractive drug targets in the field of clinical oncology. Our bioinformatic analysis resonates well with observed biological phenomena as seen in the case of MDM2 protein, which is a major PPI hub regulating p53. Interaction of the human androgen receptor (AR) protein and MDM2 influences prostate cell growth and apoptosis37. Mdm2-Daxx interaction activates p53 following DNA damage38 and Daxx binds and inhibits AR function39. Conversely, the breast cancer susceptibility gene 1 (BRCA1) interacts directly with AR and enhances AR target genes, such as p21(WAF1/CIP1), that may result in the increase of androgen-induced cell death in prostate cancer cells40. Further, BRCA1 complexes with Smad3 and is inactivated, leading to early-onset familial breast and ovarian cancer41. Within the same network, MDM2 inhibits the transcriptional activity of SMAD proteins including SMAD342, thereby, emerging as a major player in prostrate, breast and ovarian cancer. Loss of PTEN, on the other hand, results in resistance to apoptosis by activating the MDM2-mediated antiapoptotic mechanism. We also identified proteins like NCL, DAXX and SUMO that play critical roles in mediating cancers as being a part of the PTEN centric cancer interactome (Figure 5b). Interestingly, all of the 16 predicted hubs can be traced back to PTEN (either directly or through other signaling adaptors) reinforcing our analysis (Figure 5c). These findings support the prevailing concept of preferential interaction between disordered regions of two distinct proteins; with PTEN being the common disordered interacting hub, giving functional centrality to PTEN in many critical cellular pathways.
Predicting functionally relevant network hubs in the PTEN cancer interactome.
(a) Methodology to identify functional hubs within the PTEN Cancer Interactome. The PTEN Cancer Interactome contains 193 IDPs that are potential hubs. Over-represented IDPs (or IDPs with multiple occurrences) in the PTEN Cancer Interactome would have a greater propensity to function as hubs. Upon sorting for over-represented IDPs the list of 193 proteins is brought down to 31 proteins. In order to assess the possibility of these 31 proteins as functional hubs a network analysis is warranted. (b) We identified 31 potential hubs based on multiple associations from within the 193 cancer-associated IDPs of the PTEN secondary interactome. Regulatory networks derived from these 31 proteins were overlaid with a similar network from the 13 cancer-related proteins. Based on the number of associations within the network, we identify 16 potential functional hubs in the PTEN cancer interactome (indicated in yellow). Regulatory interactions were generated using the Transcriptome Browser tool (Lopez et al, 2008). (c) Functional network analysis of the 16 predicted hubs. In order to assess the functional association of the 16 predicted hubs with PTEN – a network analysis with PTEN as a central node was done. The analysis identifies MDM2 protein, a major regulator of p53, as one of the major PPI hubs in the PTEN cancer interactome. A number of other critical cancer-related proteins, such as AR, SMAD2/3 and PDGFRB that are part of the PTEN primary interactome, feature prominently in the PTEN cancer interactome. We also identified proteins like NCL, DAXX and SUMO that play critical roles in mediating cancers as being a part of the PTEN centric cancer interactome. Interestingly, all of the 16 predicted hubs can be traced back to PTEN (either directly or through other signaling adaptors) reinforcing our analysis. (Functional network analysis was performed using IPA (Ingenuity® Systems, www.ingenuity.com).
To further validate our methodology in using intrinsic disorder and cancer as filters to identify key signaling hubs, we compared our data sets with a previously published cancer signaling data set. We derived 7 common hubs (Supplementary Table S7), which were extended using the expansive human signaling network described previously43,44,45,46 to obtain the PTEN associated cancer interactome (Figure 6a). An extensive disease associated network analysis using IPA validated our predictions as all the seven predicted hubs had an extensive cross-talk across multiple cancer disease types (Figure 6b).
Derivation of PTEN associated cancer hubs.
(a) A PTEN linked cancer network was derived using seven of the 16 predicted cancer hubs that were common with the human cancer associated gene set. The associated partners of the seven hubs were extracted from the human signaling network (Cui et al, 2007, Awan et al, 2007, Li et al, 2012 and Newman et al, 2013). Red color denotes the potential cancer hubs and blue color are their associated partners. Topological analysis identifies p53 as the most significant network hub in the PTEN linked cancer network (Supplementary Table S7). (b) Disease associated network of PTEN cancer hubs. A functional network was constructed with the seven topologically relevant hubs identified previously using the Core Analysis function from the IPA suite to derive the primary network (denoted as MP). A disease network was constructed using the Path Designer option and disease associated biological functions were overlaid on the primary network. Fx denotes the different functions associated with the members of the networks.
Modulation of PTEN PPIs by linear binding motifs
Recent evidence has shown that IDPs mediate PPIs via short linear amino acid sequences (~20 residues) called Molecular Recognition Elements (MoREs) or Molecular Recognition Features (MoRFs)35,47. MoRFs undergo disorder-to-order transitions upon binding and adopt thermodynamically stable well-defined structures47, increasing the propensity of IDPs to interact with a vast repertoire of proteins. MoRFs also display molecular recognition elements that capture the binding partner proteins with high specificity. These partner-dependent conformational differences are critical to imparting versatile binding properties to IDRs35.
Since the PTEN IDR engages in multiple PPIs, we tested the possibility for the existence of MoRFs. The MORFPred algorithm48 revealed that PTEN contains major MoRF sites at amino acids 273–279 (part of the disordered CBR3 loop of the C2 domain), amino acids 339–347 (in close vicinity of the disordered C-tail) and amino acids 395–403 (part of the disordered C-tail) (Figure 7a and Supplementary Figure S4). The primary restriction of MoRFs to the PTEN C-tail IDR or adjacent regions indicates that these MoRFs directly participate in modulating PPI functions (Figure 7a). However, mutational analysis within MoRFs is required to establish their active role in functional PPIs.
Biochemical features modulating PTEN PPIs.
(a) MoRFs in the PTEN C-tail IDR. MoRFpred (Disfani et al, 2012), a computational tool, was used to identify MoRF regions within the PTEN protein (Supplementary Figure S4). The MoRFs in the vicinity of the C-tail IDR are highlighted in red. Interestingly, all of the major MoRFs (with a length greater than 5 residues) are observed in the vicinity of disordered regions (either part of the disordered CBR3 loop of the C2 domain or the C-tail IDR) indicating a positive correlation between intrinsic disorder and PPIs. (b) ELMs in PTEN C-tail IDR. Eukaryotic Linear Motifs (or ELMs) are 3–11 amino acid long sequences that mediate PPIs. IDRs are particularly enriched in ELMs (Dinkel et al, 2012). The linear motifs occurring in the disordered segment of PTEN (tail + PDZ domain) have been highlighted. The motifs with a high conservation score (>0.75) are indicated in red. Interestingly, all of the motifs with a high conservation score are restricted to the C-tail IDR. (c) Phosphorylation sites in the C-tail IDR. Phosphorylation of PTEN, particularly on serine and threonine residues in the disordered region, regulates the function and stability of PTEN. Phosphorylation occurs at Ser 362, Thr 366, Ser 370, Ser 380, Thr 382, Thr 383, Ser 385 by various enzymes such as Casein Kinase II, Glycogen synthase kinase 3-B and Polo-like kinase 3. Each of these phosphorylation events helps regulate the availability and stability of the PTEN molecule within the cell.
Protein-protein interactions are also facilitated by very short motifs (3–10 amino acids) called Short Linear Motifs (SLiMs) or Eukaryotic Linear Motifs (ELMs)49,50. Because of their short sequences, ELMs arise/disappear by simple point mutations, providing the evolutionary plasticity that the ordered protein domains lack. Thus, ELMs easily adapt to novel interactions in signaling pathways, where rapid assembly/disassembly of multi-protein complexes is a prerequisite. The frequent occurrence of ELMs in a typical proteome indicates their critical cellular functions. Consistent with this notion, a higher density of ELMs are observed in hub proteins and IDPs50. Since ELMs have short sequences, they interact with low-affinity, however, they engage in highly cooperative binding in protein complexes, triggering productive signaling50. Therefore, at increased intracellular local concentrations they competitively bind to mutually overlapping physiological targets of each other as seen with PDZ, SH2 and PTB interaction domains found in cancer-associated proteins and in IDRs49,50. As PTEN contains a PDZ-binding motif within the IDR (Figure 1a and c), we probed for the existence and features of ELMs in PTEN using The Eukaryotic Linear Motif Resource (http://elm.eu.org). We identified 34 different classes of ELMs in PTEN that mediate PPIs (Supplementary Figure S5). Interestingly, the four ELMs that are most conserved (conservation score>0.75) occurred within the PTEN C-tail IDR, indicating its high level of functional/biological significance (Figure 7b). ELM functions are further modulated by post-translational modifications, mainly by phosphorylation50. Indeed, the PTEN IDR possesses nine phosphorylation sites51,52 (Figure 7c).
PTEN phosphorylation modulates intramolecular association and PPI function
Post-translational Modifications (PTMs) in IDPs facilitate PPIs5. Modifying enzymes readily dock on structurally flexible IDRs, making them a hot spot for PTMs4,7,53,54. Consistent with this notion, regulatory cancer-associated proteins have twice as much disorder and undergo more frequent phosphorylation/dephosphorylation than other cellular proteins as predicted by DISPHOS (a DISorder-enhanced PHOSphorylation prediction software)54, implicating a tight interconnection between protein phosphorylation and disorder. Consistent with the function of PTM in IDRs, clustering of Ser and Thr phosphorylation sites (Figure 7c) in the C-tail IDR regulates PTEN stability, membrane association and activity19,20. Phosphorylation in the PEST [proline (P), glutamic acid (E), serine (S) and threonine (T)] domain within the C-tail IDR (amino acids 352 to 399) inhibits degradation of PTEN51. Casein kinase II (CK II), Glycogen synthase kinase 3-beta (GSK3-β) and PLK3 (Polo-like kinase 3) phosphorylate Ser and Thr residues within the IDR, each providing a distinct function51 (Figure 7c). The microtubule-associated serine/threonine (MAST), serine/threonine kinase 11(STK11) or LKB1 and casein kinase I (CKI) kinases have also been implicated in PTEN phosphorylation. STK11/LKB1 modifies T383, while CKI modifies T366, S370 and S38552. Indeed, our DISPHOS prediction for C-tail IDRs supports these experimental observations (Supplementary Figure S6).
Substrate-kinase interactions are typically of the disordered-ordered (D-O) type and are stabilized by hydrogen bonding (Figure 7c), a hallmark of IDRs54. Indeed, computational analysis revealed that large ordered regions comprising the catalytic domains of CKII, GSK3B, PLK3, Rak and Src kinases interact with the C-tail IDR (Supplementary Table S8), indicating that PTEN engages in D-O type intermolecular interactions with the modifying kinases.
At the intramolecular level, phosphorylation at C-tail residues triggers a conformational change in PTEN, inhibiting its membrane association and, therefore, its lipid phosphatase activity18,19,21,55. The phosphorylated C-tail IDR folds onto the PD and C2 domains giving rise to the “closed-closed” conformation of PTEN (Figure 8a) that is incapable of interaction with the membrane18,20. The “closed- closed” form of PTEN is enzymatically inactive and cannot convert PIP3 to PIP2. The identification of the exact resides involved in this intramolecular interaction remains an active area of research18,20,56.
Targeting PTEN C-tail IDR.
Most PTEN functions emanate from the C-tail IDR, including aberrant PPIs that hyper-activate oncogenic pathways. (a) Phosphorylation mediates an intramolecular interaction in the PTEN molecule. Phosphorylation causes a conformational change in PTEN converting it to the enzymatically inactive “closed closed ” form wherein the flexible tail folds onto residues in the C2 and phosphatase domain, thereby making it incapable of interacting with the membrane. Dephosphorylation (by an unknown phosphatase or via auto-dephosphorylation) converts PTEN to the “open-closed” form. Electrostatic interactions, mediated by the PBM, further convert PTEN to the “open-open” form wherein it binds to the membrane and acts as a lipid phosphatase converting PIP3 to PIP2, thereby, abrogating signaling via the PI3K/Akt/mTOR pathways. Subsequent to membrane binding, several E3 ubiquitin ligases polyubiquitinate PTEN marking it for proteasomal degradation. Phosphorylation, by inducing the intramolecular interaction, masks the ubiquitination sites thereby increasing the half-life of the PTEN protein within the cell. Therefore, phosphorylation negatively regulates PTEN function but positively regulates its stability. (b) PTEN IDR engages in PPIs of the disorder:order type (D-O type). As revealed in the present study, this occurs via the use of a MoRF or SLiM region. Therefore, designing a peptidomimetic drug molecule that competes with the PTEN MoRF/SLiM binding to the ordered protein will abrogate PTEN binding, therefore PTEN function. PTEN IDR is highly accessible to multiple kinases that phosphorylate and modulate PTEN function, mainly its inhibition via intra-molecular interactions. PTEN inhibition hyper-activates the PI3K/AKT/mTOR pathway, which increase the oncogenic potential of the cell and drives cancer growth. Therefore, targeting the PTEN C-tail IDR with small molecules that bind and sterically hinder PTEN phosphorylation and/or intra-molecular interactions will be an ideally adjunctive therapy to multiple inhibitor therapy targeting of the PI3/AKT/mTOR pathway.
It was recently shown that the phosphorylation events of PTEN occur in two independent cascades of ordered events, with the S380–S385 cluster being modified prior to the S361–S70 cluster52. Even within the two clusters, the phosphorylation events follow a specific pattern with a distributive kinetic mechanism. Not surprisingly, distributive kinetics is energetically favorable on protein domains that are highly disordered with multiple ensembles of flexible structures52. Thus the dynamic nature of these phosphorylation events is contingent to the inherent flexibility in the PTEN structure driven by intrinsically disordered C-tail crucial for PTEN stability and localization within the cell (Figure 8a).
Targeting intrinsic disorder in PTEN and its interactome
Drug targeting to critical protein regions can mitigate aberrant cellular processes driving oncogenesis57. However, despite numerous clinical trials with molecularly targeted therapies, failure rates for cancer treatments remain high. Conventional therapies targeting pathway-specific kinases suffer from “off-target effects” and often fail due to the emergence of compensatory and alternative pathways58. As a novel approach, facile drug targeting to IDRs within critical signaling hub proteins is highly plausible59,60,61. Moreover, as IDRs undergo extensive PTMs53 and engage in PPIs4,34,36, the multitude of resulting protein interactions (normal and aberrant) can be targeted concomitantly with a cocktail of distinct inhibitors, which dampens oncogenic signaling60.
Indeed, targeting PPIs is a more selective treatment strategy over conventional enzyme inhibitors60. However, disruption of multiple ordered interfaces within PPIs by small molecule inhibitors remains challenging62. The advantage of targeting IDPs engaged in PPIs is that, unlike ordered proteins, they engage in PPIs via MoRFs or ELMs, which are small peptide regions that bind with low affinity and thus are susceptible to disruption by small molecule inhibitors59. Consistent with this notion, small molecules disrupted highly disordered complexes of p53-Mdm2 and c-Myc-Max interactions by inducing order upon binding60,63. Likewise, targeting the PTEN C-tail IDR may reduce its intra- and inter-molecular interactions and limit accessibility to enzymes mediating PTMs (Figure 8b), providing a means to increase PTEN activity. Our analysis shows that since the C-tail IDR is rich in conserved MoRFs/SLiMS, targeting these regions will prove to be a rational therapeutic modality for a large number of cancers that show compromised PTEN activity or hyperactivation of the oncogenic PI3K/AKT/mTOR pathway9,10,11. Since reductions in the levels and activity of PTEN are sufficient to drive oncogenesis11,14,15, increasing PTEN activity is an ideal therapy for cancers associated with hyperactive PI3K-signaling.
Discussion
Recent studies on genome- and proteome-wide molecular alterations in diseases indicate that pathological conditions are caused by perturbations in complex, highly interconnected biological networks64. Thus, current reductionist approach of studying structure-function relationship in diseases has limited our abilities to discover effective targeted therapeutics. In an attempt to overcome these limitations, in the current study, we have undertaken a novel approach to drug discovery that exploits systems and network biology at the structural, topological and functional level. Using PTEN, a tumor suppressor, we have applied computational and systems biology approaches and integrated extensive data-mining and biochemical properties of IDP interactions to reach a finer understanding of PTEN function. These results have identified PTEN C-tail IDR and several hub proteins in PTEN-driven molecular network implicated in human diseases as therapeutic targets, enhancing the repertoire of clinically relevant biological targets for pharmacotherapy.
Our derivation and analysis of PTEN primary and secondary interactome indicates that altered levels or interactions of IDPs perturb myriad cellular signaling pathways, leading to pathological conditions including cancer. IDPs have the propensity to aggregate and cause cellular toxicity65. Therefore, PTEN as an IDP has evolved a mechanism, wherein, the level of active PTEN, its cellular localization and PTEN-PPIs are regulated via phosphorylation of the C-tail IDR. Furthermore, evolutionarily conserved ELMs and MoRFs that we have identified within the C-tail IDR may play a critical role in orchestrating the formation and function of the PTEN interactome.
Increase in complexity of PPIs is either directed by the number and type of proteins or by increasing the number of interactions required to execute cellular functions66. To delineate how PTEN executes myriad functions, we first derived the PTEN primary interactome. We found 40 proteins to directly interact on the PTEN molecule, out of which 25 were associated with the C-tail IDR, consistent with the concept that disorderliness within PTEN executes its myriad functions. To enhance our understanding of PTEN functions in the context of multiple distinct pathways at the systems-level, we delineated functional networks operating within the primary interactome. Our findings showed a high degree of cross-talk between edges, implying that shared regulatory modules, comprised of multiple signaling cascades, operate via PTEN-mediated interaction networks. When these networks are altered, diseases ensue with extreme functional penalties. We also found that the edge proteins were themselves highly disordered indicating that disorderliness within the PTEN primary interactome confers functional versatility. Supporting this notion, 13 proteins that were functionally classified as cancer-related were also highly disordered forming a pliable “PTEN-Cancer Interactome”. Thus, PTEN lesions influence the flexibility of IDP-IDP interactions modulating diverse functions, likely causing cancer.
Owing to the inherent ability of PPIs to be flexible while being complex, specific cellular functions are readily fine-tuned as per the biological demands. Emerging evidence suggests that certain features on the IDRs are recognized as a way of conferring plasticity to protein interaction networks. Consistent with this concept, our data suggest that PTEN, a hub protein containing an IDR, likely utilizes MoRFs and ELMs, gets differentially modified via PTMs, acquiring complementary structures to engage and modulate PPI activity by facilitating adaptive binding to multiple protein partners in many cellular pathways. Thus, our present work provide a novel entrée in targeting intrinsic disorder in PTEN and its interactome to dampen the aberrant PI3K-signaling that drives many cancers. First, imparting order to the PTEN structure may help dampen multiple oncogenic signaling pathways mediated via the 16 hub proteins identified in the present study, by limiting their affinity for PPIs. Second, targeting intrinsic disorder in PTEN and its interactome can become an adjunctive or alternative approach to the use of various kinase inhibitors, which are toxic and have many off-target effects when used to mitigate the aberrant hyperactivation of PI3K/AKT/mTOR oncogenic signaling pathway. Taken together, the present findings provide a novel entrée to design strategies for drug discovery and may become a logical intervention in the pharmacotherapy of cancer and other PTEN-associated disease treatment modalities.
Methods
Disorder analysis
Disorder analysis for PTEN, its primary interactome, secondary interactome and kinases that phosphorylate it were performed using the PONDR-FIT software22. The software assigns a disorder score to each amino acid of the protein. Residues with disorder scores > 0.5 were considered to be disordered. For the PTEN primary interactome, proteins having long disordered regions (i.e a stretch of 30 or more contiguous disordered residues) are considered to be disordered. Percent disorderliness for each protein in the primary interactome is calculated as [(No. of disordered residues/Total no. of amino acids in the protein)*100]. A plot of percent disorderliness is part of Figure 2. For kinases that phosphorylate PTEN, those having long disordered regions (i.e a stretch of 30 or more contiguous disordered residues) are considered to be disordered. The exact residues that constitute the kinase domain were obtained from UniProtKB (http://www.uniprot.org/).
Compositional analysis
Compositional Analysis for PTEN was carried out using the Compositional Profiler Tool24. Comparative analysis is performed for full-length PTEN, its ordered and disordered regions respectively. The analysis makes use of PDB Select 25 and DisProt as a database for ordered and disordered proteins respectively. Any change in amino acid levels for the query protein (i.e enrichment or depletion) over the average value (obtained from the databases) is expressed as (C-Corder)/Corder . C is the level of a particular amino acid in the query protein, which in this case is full-length PTEN, its ordered domain and the intrinsically disordered PTEN tail. Corder corresponds to the level of the same amino acid obtained from a database of ordered proteins (PDB Select 25).A similar profile for typical IDPs is obtained using the DisProt database.
Further, a comparative compositional analysis was performed for the disordered and ordered regions of PTEN. Percentage frequency of a specific class of residues (i.e hydrophobic, polar, aromatic, structure breaking, cysteine and asparagine) is calculated for the ordered and disordered regions respectively. The percentage frequency for the ordered region is calculated as [(No. of a given class of residues in the ordered region/Total number of amino acids in the ordered region)*100]. Similarly, the percentage frequencies are calculated for the disordered region.
Mutational analysis
Mutation data for the PTEN protein was compiled from Sanger Institute Catalogue of Somatic Mutations in Cancer (COSMIC), Human Gene Mutation Database (HGMD), cBioPortal for Cancer Genomics and the Roche Cancer Genome Database (RCGDB). The number of observed mutations at every amino acid position is determined.
In order to correlate the mutations in PTEN with its amino acid composition, we calculated the ratio of mutations in the different classes of residues used to define IDRs (hydrophobic, polar, aromatic, structure breaking, cysteine and asparagine) in the disordered vs ordered region. The ratio is calculated as:

Where, Rd is calculated as:

Similarly, Ro is calculated as:

Sequence alignments for PTEN and the PTEN C-tail interacting proteins
FASTA sequences for PTEN and PTEN C-tail interacting proteins in various animal species was retrieved from UniProtKB. Sequence alignments were performed using Clustal Omega tool (http://www.ebi.ac.uk/Tools/msa/clustalo/).
Derivation of primary and secondary interactomes and network analysis
The PTEN primary interactome was compiled using manual data curation and various softwares including Database of Interacting Proteins (DIP), Interologous Interaction (I2D) Database, InnateDB, IntAct, MatrixDB, The Molecular INTeraction Database (MINT), Molcon, The Microbial Protein Interaction Database (MPID), Uniprot, Simons Foundation Autism Research Initiative (SFARI). The PTEN primary interactome is divided into the mapped and unmapped categories, those proteins that interact with known regions of PTEN are classified as “mapped proteins” whereas those with uncharacterized or computationally predicted interactions are termed as “unmapped proteins”. The PTEN secondary interactome was compiled using various softwares listed above. Network Analysis for the PTEN primary interactome was performed using the Metacore software suite (GeneGo) using PTEN as the node.
Thirteen cancer-related proteins in the PTEN primary interactome were identified by functional enrichment analysis done using the IPA software (Ingenuity® Systems, www.ingenuity.com).
Subsequent network analysis is performed using the IPA software (Ingenuity® Systems, www.ingenuity.com) and Transcriptome Browser67.
Deriving the PTEN associated cancer network
The 16 hubs that were predicted to be cancer related hubs were compared with the human cancer associated gene list43. Seven potential cancer associated hubs were identified from the cancer-associated gene list containing 2128 genes. A PTEN centric cancer network was derived from these seven genes as potential hubs using the human signaling network43,44,45,46. The human signaling network contains ~6,300 proteins and ~63,000 signaling relations. In order to make the network PTEN-centric, we added the PTEN protein-protein interaction data to the original human signaling network. The PTEN associated network was obtained from this updated data set by selecting all the links associated with PTEN and the seven potential cancer associated hubs. The network was visualized using Cytoscape. Network analysis was performed to identify topologically significant hubs using the Network Analysis Plug in tool.
MoRF and ELM prediction
The MoRFPred algorithm48 was used to identify MoRF regions within the PTEN protein while the ELM software50 was used to scan the PTEN protein for annotated ELMs.
References
Uversky, V. N., Oldfield, C. J. & Dunker, A. K. Intrinsically disordered proteins in human diseases: introducing the D2 concept. Annu Rev Biophys 37, 215–246 (2008).
Ward, J. J., Sodhi, J. S., McGuffin, L. J., Buxton, B. F. & Jones, D. T. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. Journal of Molecular Biology 337, 635–645 (2004).
Dunker, A. K. et al. Intrinsically disordered protein. J Mol Graph Model 19, 26–59 (2001).
Fong, J. H., Shoemaker, B. A. & Panchenko, A. R. Intrinsic protein disorder in human pathways. Mol Biosyst 8, 320–326 (2012).
Gsponer, J. & Babu, M. M. The rules of disorder or why disorder rules. Prog Biophys Mol Biol 99, 94–103 (2009).
Iakoucheva, L. M., Brown, C. J., Lawson, J. D., Obradovic, Z. & Dunker, A. K. Intrinsic disorder in cell-signaling and cancer-associated proteins. Journal of Molecular Biology 323, 573–584 (2002).
Babu, M. M., van der Lee, R., de Groot, N. S. & Gsponer, J. Intrinsically disordered proteins: regulation and disease. Curr Opin Struct Biol 21, 432–440 (2011).
Song, M. S., Salmena, L. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol advance online publication (2012).
Zoncu, R., Efeyan, A. & Sabatini, D. M. mTOR: from growth signal integration to cancer, diabetes and ageing. Nat Rev Mol Cell Biol 12, 21–35 (2011).
Vivanco, I. & Sawyers, C. L. The phosphatidylinositol 3-Kinase AKT pathway in human cancer. Nat Rev Cancer 2, 489–501 (2002).
Sansal, I. & Sellers, W. R. The biology and clinical relevance of the PTEN tumor suppressor pathway. J Clin Oncol 22, 2954–2963 (2004).
Maehama, T. & Dixon, J. E. The tumor suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J Biol Chem 273, 13375–13378 (1998).
Zhang, X. C., Piccini, A., Myers, M. P., Van Aelst, L. & Tonks, N. K. Functional Analysis of the Protein Phosphatase Activity of PTEN. Biochem J (2012).
Hollander, M. C., Blumenthal, G. M. & Dennis, P. A. PTEN loss in the continuum of common cancers, rare syndromes and mouse models. Nat Rev Cancer 11, 289–301 (2011).
Leslie, N. R. & Foti, M. Non-genomic loss of PTEN function in cancer: not in my genes. Trends in Pharmacological Sciences 32, 131–140 (2011).
Das, S., Dixon, J. E. & Cho, W. Membrane-binding and activation mechanism of PTEN. Proc Natl Acad Sci USA 100, 7491–7496 (2003).
Lee, J. O. et al. Crystal structure of the PTEN tumor suppressor: Implications for its phosphoinositide phosphatase activity and membrane association. Cell 99, 323–334 (1999).
Rahdar, M. et al. A phosphorylation-dependent intramolecular interaction regulates the membrane association and activity of the tumor suppressor PTEN. Proc Natl Acad Sci USA 106, 480–485 (2009).
Vazquez, F., Ramaswamy, S., Nakamura, N. & Sellers, W. R. Phosphorylation of the PTEN tail regulates protein stability and function. Mol Cell Biol 20, 5010–5018 (2000).
Ross, A. H. & Gericke, A. Phosphorylation keeps PTEN phosphatase closed for business. Proc Natl Acad Sci USA 106, 1297–1298 (2009).
Odriozola, L., Singh, G., Hoang, T. & Chan, A. M. Regulation of PTEN activity by its carboxyl-terminal autoinhibitory domain. J Biol Chem 282, 23306–23315 (2007).
Xue, B., Dunbrack, R. L., Williams, R. W., Dunker, A. K. & Uversky, V. N. PONDR-FIT: a meta-predictor of intrinsically disordered amino acids. Biochim Biophys Acta 1804, 996–1010 (2010).
Romero, P. et al. Sequence complexity of disordered protein. Proteins 42, 38–48 (2001).
Vacic, V., Uversky, V. N., Dunker, A. K. & Lonardi, S. Composition Profiler: a tool for discovery and visualization of amino acid composition differences. BMC Bioinformatics 8, 211 (2007).
Stenson, P. D. et al. The Human Gene Mutation Database: 2008 update. Genome Med 1, 13 (2009).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2, 401–404 (2012).
Gao, J. J. et al. Integrative Analysis of Complex Cancer Genomics and Clinical Profiles Using the cBioPortal. Science Signaling 6, (2013).
Kuntzer, J., Eggle, D., Lenhof, H. P., Burtscher, H. & Klostermann, S. The Roche Cancer Genome Database (RCGDB). Hum Mutat 31, 407–413 (2010).
Mosca, R., Pache, R. A. & Aloy, P. The Role of Structural Disorder in the Rewiring of Protein Interactions through Evolution. Mol Cell Proteomics 11, M111 014969 (2012).
Liu, J., Tan, H. & Rost, B. Loopy proteins appear conserved in evolution. J Mol Biol 322, 53–64 (2002).
Schlessinger, A. et al. Protein disorder--a breakthrough invention of evolution? Curr Opin Struct Biol 21, 412–418 (2011).
Espinoza-Fonseca, L. M. Aromatic residues link binding and function of intrinsically disordered proteins. Molecular BioSystems 8, 237–246 (2012).
Dyson, H. J. & Wright, P. E. Coupling of folding and binding for unstructured proteins. Current Opinion in Structural Biology 12, 54–60 (2002).
Fong, J. H. et al. Intrinsic Disorder in Protein Interactions: Insights From a Comprehensive Structural Analysis. PLoS Comput Biol 5, e1000316 (2009).
Dunker, A. K., Cortese, M. S., Romero, P., Iakoucheva, L. M. & Uversky, V. N. Flexible nets. FEBS Journal 272, 5129–5148 (2005).
Shimizu, K. & Toh, H. Interaction between Intrinsically Disordered Proteins Frequently Occurs in a Human Protein–Protein Interaction Network. Journal of Molecular Biology 392, 1253–1265 (2009).
HK, L., L, W., YC, H., S, A. & C, C. Phosphorylation-dependent ubiquitylation and degradation of androgen receptor by Akt require Mdm2 E3 ligase. EMBO J 21, 4037–4048 (2002).
Tang, J. et al. Critical role for Daxx in regulating Mdm2. Nat Cell Biol 8, 855–862 (2006).
Lin, D.-Y. et al. Negative Modulation of Androgen Receptor Transcriptional Activity by Daxx. Molecular and Cellular Biology 24, 10529–10541 (2004).
Yeh, S. et al. Increase of androgen-induced cell death and androgen receptor transactivation by BRCA1 in prostate cancer cells. Proceedings of the National Academy of Sciences 97, 11256–11261 (2000).
Dubrovska, A. et al. TGF[beta]1//Smad3 counteracts BRCA1-dependent repair of DNA damage. 24, 2289–2297 (2005).
Yam, C. H. et al. MDM2 and MDMX Inhibit the Transcriptional Activity of Ectopically Expressed SMAD Proteins. Cancer Research 59, 5075–5078 (1999).
Cui, Q. et al. A map of human cancer signaling. Mol Syst Biol 3, 152 (2007).
Awan, A. et al. Regulatory network motifs and hotspots of cancer genes in a mammalian cellular signalling network. IET Syst Biol 1, 292–297 (2007).
Li, L. et al. The human phosphotyrosine signaling network: evolution and hotspots of hijacking in cancer. Genome Res 22, 1222–1230 (2012).
Newman, R. H. et al. Construction of human activity-based phosphorylation networks. Mol Syst Biol 9, 655 (2013).
Mohan, A. et al. Analysis of molecular recognition features (MoRFs). J Mol Biol 362, 1043–1059 (2006).
Disfani, F. M. et al. MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in proteins. Bioinformatics 28, i75–i83 (2012).
Diella, F. et al. Understanding eukaryotic linear motifs and their role in cell signaling and regulation. Frontiers in Bioscience 13, 6580–6603 (2008).
Dinkel, H. et al. ELM-the database of eukaryotic linear motifs. Nucleic Acids Res 40, D242–D251 (2012).
Singh, G. & Chan, A. M. Post-translational modifications of PTEN and their potential therapeutic implications. Curr Cancer Drug Targets 11, 536–547 (2011).
Cordier, F. et al. Ordered phosphorylation events in two independent cascades of the PTEN C-tail revealed by NMR. J Am Chem Soc (2012).
Gao, J. X. D. Correlation between posttranslational modification and intrinsic disorder in protein. Pac Symp Biocomput. 94–103 (2012).
Iakoucheva, L. M. et al. The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Res 32, 1037–1049 (2004).
Salmena, L., Carracedo, A. & Pandolfi, P. P. Tenets of PTEN tumor suppression. Cell 133, 403–414 (2008).
Rahdar, M. Johns Hopkins University School of Medicine. (2011).
Rotea, W., Jr & Saad, E. D. Targeted drugs in oncology: new names, new mechanisms, new paradigm. Am J Health Syst Pharm 60, 1233–1243; quiz 1244–1235 (2003).
Pearson, M. A. & Fabbro, D. Targeting protein kinases in cancer therapy: a success? Expert Rev Anticancer Ther 4, 1113–1124 (2004).
Metallo, S. J. Intrinsically disordered proteins are potential drug targets. Curr Opin Chem Biol 14, 481–488 (2010).
Uversky, V. N. Intrinsically disordered proteins and novel strategies for drug discovery. Expert Opinion on Drug Discovery 7, 475–488 (2012).
Tzakos, A. G. Intrinsic Protein Disorder as a Drug Target in Oncology: Designing Drugs Targeting Plasticity. Biochemistry and Pharmacology (2012).
Blundell, T. L. et al. Protein-protein interactions in receptor activation and intracellular signalling. Biol Chem 381, 955–959 (2000).
Chene, P. Inhibition of the p53-MDM2 interaction: Targeting a protein-protein interface. Molecular Cancer Research 2, 20–28 (2004).
del Sol, A., Balling, R., Hood, L. & Galas, D. Diseases as network perturbations. Curr Opin Biotech 21, 566–571 (2010).
Gsponer, J., Futschik, M. E., Teichmann, S. A. & Babu, M. M. Tight regulation of unstructured proteins: from transcript synthesis to protein degradation. Science 322, 1365–1368 (2008).
Deeds, E. J., Krivine, J., Feret, J., Danos, V. & Fontana, W. Combinatorial complexity and compositional drift in protein interaction networks. PLoS One 7, e32032 (2012).
Lopez, F. et al. TranscriptomeBrowser: A Powerful and Flexible Toolbox to Explore Productively the Transcriptional Landscape of the Gene Expression Omnibus Database. PLoS One 3, e4001 (2008).
Acknowledgements
This work was supported by American Heart Association Grant. SDG-155-N (V.D.), Moffitt Cancer Center Lung SPORE Career Development grant (V.D.) and by the Program of the Russian Academy of Sciences for the "Molecular and Cellular Biology" (V.N.U.). The authors would like to thank Dr Robert Deschenes for helpful discussions and Jaymin Kathiriya for assistance with the sequence alignments.
Author information
Authors and Affiliations
Contributions
P.M. analyzed the properties of PTEN IDR, derived the PTEN primary interactomes and performed the sequence alignments. R.R.P. performed network analysis and obtained the PTEN secondary interactome. P.M. and R.R.P. identified ELMs and MoRFs in PTEN. B.X. and V.N.U. performed the disorder analysis and obtained amino acid composition profile of the PTEN molecule. The project was conceived and designed by V.D. and written by V.D. and P.M. with input from all authors.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Complete dataset
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Malaney, P., Pathak, R., Xue, B. et al. Intrinsic Disorder in PTEN and its Interactome Confers Structural Plasticity and Functional Versatility. Sci Rep 3, 2035 (2013). https://doi.org/10.1038/srep02035
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02035
This article is cited by
-
Precise definition of PTEN C-terminal epitopes and its implications in clinical oncology
npj Precision Oncology (2019)
-
A growth hormone receptor SNP promotes lung cancer by impairment of SOCS2-mediated degradation
Oncogene (2018)
-
PTEN proteoforms in biology and disease
Cellular and Molecular Life Sciences (2017)
-
Functions of intrinsic disorder in transmembrane proteins
Cellular and Molecular Life Sciences (2017)
-
Tick receptor for outer surface protein A from Ixodes ricinus — the first intrinsically disordered protein involved in vector-microbe recognition
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.