Abstract
Application of PCR to multiplexing assays is not trivial; it requires multiple fluorescent labels for amplicon detection and sophisticated software for data interpretation. Alternative PCR-free methods exploiting new concepts in nanotechnology exhibit high sensitivities but require multiple labeling and/or amplification steps. Here, we propose to simplify the problem of simultaneous analysis of multiple targets in genetic assays by detecting directly the conformation, rather than mass, of target amplicons produced in the same PCR reaction. The new methodology exploits acoustic wave devices which are shown to be able to characterize in a fully quantitative manner multiple double stranded DNAs of various lengths. The generic nature of the combined acoustic/PCR platform is shown using real samples and, specifically, during the detection of SNP genotyping in Anopheles gambiae and gene expression quantification in treated mice. The method possesses significant advantages to TaqMan assay and real-time PCR regarding multiplexing capability, speed, simplicity and cost.
Similar content being viewed by others
Introduction
Research and medical laboratories normally rely on solid phase hybridization assays and real-time PCR to answer biological problems regarding gene expression profiling, determination of viral load in clinical samples, DNA and RNA quantification, bacterial identification, SNP genotyping and pharmacogenomics. Both techniques are based either on non-specific or sequence-specific fluorescent reporters that generate a signal reflecting on the amount of the PCR product; detection and quantification of fluorescently labeled targets require expensive instrumentation and sophisticated algorithms in the case of microarrays1.
Recently, the application of novel scientific and technological concepts relying on nanotechnology has resulted in the development of genetic assays of impressive performance2,3,4; such examples can be found in the wide range of nanoparticle-linked DNA “scanometric” detection strategies5,6 with detection limits in the range of few hundred- to sub-fM and, in some cases, even down to few hundred of zM7. However, these limits of detection are coupled with complicated and laborious steps including biomolecular labeling, nanoparticle functionalization, development of secondary probes and signal amplification procedures. Another promising detection scheme which offers high sensitivity but is simple to use and cost-effective includes the electronic DNA (E-DNA) sensors. These systems known as folding-based biosensors employ a surface-immobilized DNA redox-tagged probe, which, upon target hybridization, undergoes a large conformational change; this produces a strong signal as a result of the change of the redox probe distance from the electrode surface8. The E-DNA sensor can achieve detection limits in the fM range9 and without the use of amplification steps. Moreover the E-DNA sensor has been demonstrated to be able to detect nM concentrations of DNA in serum, soil and foodstuff samples10 and the gyrB gene of Salmonella typhimurium from genomic DNA11; it should be noticed, though, that in the first case the detection of a single mismatch was within the error bar of the observed signal response while in the second case, PCR was used to amplify the number of single stranded target molecules.
Our purpose here is to provide a new methodology which is generic, sensitive, selective and simple enough, so that it should be fairly easy for others to adopt in routine DNA analysis and quantification. To achieve this purpose we used conventional PCR for target amplification and acoustic wave devices for amplicon detection; PCR instrumentation is present in every research lab while acoustic devices and set-ups are commercially available by various manufacturers. The details and principles of operation of acoustic biosensors have been described before12. Briefly, the presence of an analyte at the sensor's surface affects the propagation characteristics of an acoustic wave, i.e., its velocity and energy, which in turn, are monitored as changes in frequency (ΔF) and energy dissipation (ΔD); note that frequency changes reflect on the amount of adsorbed mass and dissipation on the viscoelastic properties of the bound molecules. Theoretical treatment of experimental data in our lab revealed that energy dissipation per unit mass, ΔD/ΔF, i.e., the acoustic ratio, can be used as a direct measure of the intrinsic viscosity [η] of the surface-attached DNA molecules13,14. The particular significance of this finding lies in the fact that intrinsic viscosity is directly related to the hydrodynamic volume of the attached analyte which is a measure of its size and shape. Regarding the immobilization of DNA molecules, the acoustic ratio has three particular properties. Firstly, it is characteristic of the size and shape, of the attached DNA molecule and defines the biomolecule's acoustic signature. This has been well documented for double stranded DNA molecules varying in length and DNAs of the same length but with different degrees of curvature13 as well as the hybridization of double15 and triple16 DNAs and characterization of a Holliday nanoswitch junction17. Secondly, at equilibrium, the acoustic ratio is independent of the sample concentration loaded on the surface and thirdly, it is independent of the surface history, i.e., previous loading steps13. The practical significance of these features is great; the method possesses an inherently high selectivity determined by the conformation of each target molecule and it is possible to use the same device surface several times through the sequential loading of small amounts of various samples in a single experiment, until the surface is saturated. In this work we exploit only the length-dependence of the intrinsic viscosity and we propose the development of genetic assays where various double stranded (ds) DNA molecules, produced in the same PCR reaction, are loaded on the device surface and characterized solely by the measured acoustic ratio. As a proof of principle we apply this approach in the detection of an insecticide-resistant mutation (single nucleotide polymorphism, SNP) in the major malaria vector in Anopheles gambiae and in gene expression quantification of the ABCA1 gene in mice treated with the LXR ligand T0901317. Our results indicate that, when supported by a careful design of PCR products, acoustic wave devices can be used to achieve multiplexing, sensitive, fast and cost-effective analysis.
Results
Description of the acoustic procedure
The acoustic method involves the binding of biotinylated-DNA molecules to a neutravidin-modified device surface. In this work the commercially available Quartz Crystal Microbalance (QCM-D) setup was used to carry out acoustic measurements at 35 MHz in a flow-through system (fig. 1a). Acoustic results are expressed as the ratio of dissipation change over frequency change (ΔD/ΔF).
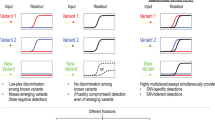
Acoustic measurements and data analysis.
(a) Real time acoustic curves. The red and the blue lines depict changes in the frequency and energy dissipation of the acoustic signal respectively. The first step (i) corresponds to the adsorption of neutravidin protein on the gold surface of the acoustic device. The following two steps correspond to the subsequent binding of a mixture of 361bp and 75bp biotinylated DNA molecules with mass ratio (ii) 1 : 3 and (iii) 1 : 4. Each sample addition is followed by buffer rinsing. (b) Plot of the measured acoustic ratio versus the number of DNA base pairs (N). A linear fit is shown (R = 0.996). (c) Plot of acoustic ratio values measured during the simultaneous presence of two different DNA molecules, as a function of their mass ratios. Mixtures of 75 bp with 361 bp DNA molecules are shown in blue and mixtures of 92 bp with 402 bp are in red. A linear fit is shown (R = 0.986).
In a previous report14 the acoustic ratios of DNA samples of various lengths were measured in a PBS buffer containing 10 mM MgCl2. The dsDNA samples applied to the surface of the acoustic devices had lengths of 20, 50, 90, 132, 167, 198, 270 and 395 and 1724 base-pairs. Here we enriched that study and further investigated the ratios from chains with 60, 75, 92, 361 and 402 bps while repeating the experiments using PBS buffer without the Mg+2 ions. The curve in fig. 1b shows our cumulative results. The acoustic ratios are identical in both buffers and a good linearity exists in the range of 20 to ~300 base pairs. The fit for this linear range is given by:

where N is the number of base-pairs in the DNA chain and ΔD/ΔF is in (x10−10/Hz) units.
It is apparent that equation (1) can be used to estimate the length of a linear DNA molecule, based on its acoustic ratio and vice versa. To go one step further, we investigated the acoustic ratio when mixtures of two different in length biotinylated-DNAs in various mass ratios were immobilized on the sensor. For that purpose we prepared separately and quantified two pure 75 bp and 361 bp DNA solutions. The 75 bp molecule was selected because it is approximately the minimum DNA length that can be produced by a PCR reaction. We prepared mixtures of 75/361 bp and measured the acoustic ratios as shown in fig. 1c. As expected, their ratios lie in the range of ~250 to 488 (x10−10/Hz), 226 being the ratio of 100% 75bp sample. It is evident that this value is nearly reached for far smaller values of the percentile 75bp content in the mixture; practically, when the mass of the 75 bp molecule is ≥ 5 times more than the mass of the 361bp component the detection suggests saturation of the signal as if this was due to a single component sample. In terms of mole ratio (instead of mass) this is equivalent to saying that when the one component is ~20–25 times more than the other the signal is dominated completely; the shorter DNA molecules are filling the available neutravidin binding sites almost exclusively. We also verified the accuracy of the calibration curve by using a combination of two different but of similar length DNA molecules of 92 and 402bp respectively. The linear fit is given by:

Application case 1
SNP genotyping of ace-1 gene in A. gambiae
The main malaria vectors in Africa are members of the Anopheles gambiae complex. Chemical insecticides are considered the most effective means to control A. gambiae populations. A glycine to serine substitution at position 119 (GGC to AGC) of the ace-1 gene, the target site of carbamates and organophosphates, confers high levels of resistance against these insecticides18. It has been recently shown that this mutation is spreading in populations of A. gambiae in West Africa19.
We developed a two-step assay for the detection of the GGC to AGC substitution in the ace-1 gene in genomic DNA. The first step involves a bidirectional PCR amplification reaction of specific alleles20,21 with genomic DNA used as the template and the second step the direct measurement of the PCR products with our acoustic sensor. Based on the sequence of the ace-1 gene18 we designed 4 different primers (fig. 2a); primers 2 and 3 were biotinylated at the 5′ end. Primers 1 and 3 could potentially produce a 179 bp biotinylated DNA fragment, primers 2 and 4 a 361 bp biotinylated fragment and primers 1 and 4 a 503 bp non-biotinylated fragment. Gel analysis of the products derived for samples of known composition showed that for wild type cases the 179 bp product is produced normally and the 361 bp is under-produced (fig. 2b). For heterozygous templates both products are almost equally produced and for homozygous mutant samples the 361 bp is normally produced while the 179 bp is significantly under-produced. Semi-quantitative analysis from agarose gels showed that for wild type samples the 179/361 bp ratio is > 2, for heterozygous ≈ 1–1.5 and for homozygous mutant < 0.25.
Design of Ace-1 genotyping assay.
(a) Relative position of the designed primers (colored arrows) on the ace-1 gene template. The orange box represents the position 119 where the adenine is substituted by a guanine in heterozygous and resistive homozygous samples. Primers 2 and 3 contain a biotin at their 5′-end. (b) A 2% agarose gel where the DNA products from a multiplex PCR with wild-type (lane 1), heterozygous (lane 2) and resistive homozygous samples (lane 3) are displayed. All products were analyzed under the same electrophoresis conditions within the same gel however, for the conciseness of the presentation only the lanes of interest are presented. Full length gel image is presented in Supplementary Fig. 1. The relative sizes of the PCR fragments are also schematically represented. The color of each strand derives from the corresponding primer. The attached yellow dots at the end of the primers 2 and 3 represent biotin. (c) Schematic representation of the gold surface of an acoustic device covered with a neutravidin layer (green spheres). Only the 361 and 179 bp fragments are immobilized while the 503 bp is washed away. The acoustic ratio measured depends on the relative number of the different size molecules that are bound.
Genomic DNA isolated from 9 A. gambiae samples were amplified during multiplexed PCR using an optimized protocol with the 4 primers. PCR products were flown over the device surface where the biotinylated fragments were immobilized while the non-biotinylated ones were washed away (fig. 2c). According to figure 2, the 179 and 361 bps DNAs were expected to give acoustic ratios equal to 356 and 600 (x10−10/Hz), respectively. Therefore, their mixtures should give acoustic ratios within this range. From the semi-quantitative gel analysis we also expect that the acoustic ratio from the heterozygous sample should be closer to that of the wild type sample. We performed triplicate acoustic experiments of the three different genotypes (fig. 3a & 3b) and found acoustic ratios equal to 380 ± 11, 430 ± 15 and 540 ± 15 (x10−10/Hz); based on the above, we attributed the first number to the wild type, the second to the heterozygous and the last to the homozygous mutant type. Our results summarized in fig. 3c were cross-checked with the TaqMan assay (fig. 3d) and gel analysis and found to be in complete agreement with both.
SNP genotyping of ace-1 gene.
(a) Real time binding curves displaying changes in the frequency attributed to a wild type (blue line), a heterozygous (black line) or a homozygous mutant (red line) sample immobilized on an acoustic biosensor. (b) Respective changes in dissipation monitored in real time. In both cases the overshoot pick at ~7 min reflects the washing off of DNA samples attached non-specifically. (c) Comparison of acoustic ratios measured for the three different types of samples with the respective calculated ratios based on equation 1. (d) Scatter plot analysis of TaqMan fluorescence data. The TaqMan assay was carried out on 47 mosquito genomic DNA samples and 1 no template negative control. Fluorescence values of the FAM labeled probe specific for the resistant allele were plotted against those of the VIC labeled probe specific for the susceptible allele. Nine of these samples were randomly selected for the development of the acoustic assay.
Application case 2
Quantification of ABCA1 gene expression
The potential of the acoustic method was further investigated by measuring differences in the expression of the ABCA1 gene between treated with T0901317 and untreated mice. The membrane protein ABCA1 (ATP Binding Cassette Transporter A1) plays an important role in the biogenesis of High Density Lipoproteins (HDL) in the liver by facilitating the efflux of membrane cholesterol and phospholipids from hepatic cells to apolipoprotein A-I6. The expression of this gene is induced by natural (oxysterols) or synthetic (T0901317) ligands of Liver X Receptors (LXRs) in many cell types22,23.
We initially prepared cDNA from mRNA derived from the liver of mice that were either treated (E48 sample) or untreated (E47 sample) with the LXR ligand T0901317. GAPDH housekeeping and ABCA1 genes were amplified in a standard PCR reaction from the prepared cDNA and semi-quantified by agarose gel electrophoresis (fig. 4a & 4b). Gel analysis revealed that in the liver the ABCA1 gene is approximately 2 times over-expressed in treated samples compared to untreated ones. In order to verify the small difference in expression in the E47-E48 liver samples we performed real-time PCR analysis. The mRNA levels of ABCA1 were measured by quantitative real-time PCR, normalized to GAPDH and showed a 2.6 ± 0.3 (fig. 4c) fold change compared to the corresponding control. The biological significance of this result was further evaluated by quantifying the gene expression levels of ABCA1 in two more treated samples (E53-E54) and one additional control sample (E52); the average difference was found to be 2.0 ± 0.7 (fig. 4d).
ABCA1 gene expression evaluation.
(a) Agarose gel images of ABCA1 and GAPDH gene fragments produced in separate PCR reactions using cDNA derived from the liver and the small intestine of mice treated with the compound T0901317 (T0) and untreated (vehicle) mice. Samples were analyzed under the same experimental conditions although the images are cropped for clarity reasons. Original image is presented in Supplementary Fig. 2. (b) ABCA1 and GAPDH fragments produced in the same tube for different number of cycles using a mixture of cDNAs from liver of treated and untreated mice. For the conciseness of the presentation the gel image was cropped. The full length image is presented in the Supplementary Fig. 3. All products were analyzed under the same experimental conditions. (c) mRNA levels of ABCA1 extracted from E48 treated sample were measured by quantitative RT-PCR, normalized to GAPDH and presented as fold change relative to control sample E47. (d) mRNA levels of ABCA1 from samples E48, E53 and E54 were measured by quantitative RT-PCR, normalized to GAPDH and presented as fold change relative to controls E47 and E52.
To apply the acoustic method in the detection of differences in gene expression levels, we took advantage of the relationship between the acoustic to mass ratio (eq. 2) of two DNA molecules varying in size (fig. 1c). We designed two sets of primers (one of the two for each set biotinylated at the 5′ end), one amplifying a 92 bp fragment corresponding to GAPDH gene and the other producing a 402 bp corresponding to the ABCA1 gene (fig. 5a). Acoustic measurements were performed as in fig. 5b & 5c. Template cDNA from either E47 or E48 samples was mixed with the four primers in a multiplexed PCR reaction and two different DNA fragments were produced during 27 cycles of amplification. This number was chosen so that the relative ratio of the two genes in the original cDNA would be maintained in the PCR products. According to equation 1, the GAPDH (92 bp) and the ABCA1 (402 bp) fragments were expected to have a ratio of 250 and 600 (x10−10/Hz), respectively and their mixtures should have intermediate values closer to one or the other end depending on their mass ratio. Each experiment was repeated six times and the acoustic ratio for the control (E47) sample was 293.3 ± 16.3 (x10−10/Hz) while for the treated (E48) was 416.7 ± 19.7 (x10−10/Hz) corresponding to mass ratios of 3.60 and 1.54 respectively (eq. 2). The differences between the ratios reflect the differences in the number of GAPDH and ABCA1 amplicons. The acoustic ratio value in each case equals to the ratio of the number of molecules produced by the PCR reaction which is directly related to the starting number of molecules in each sample since the comparison is done at 27 cycles which is below the plateau of the PCR (fig. 4b). Given that the expression of the housekeeping GAPDH gene is stable in both samples we can conclude that the differences in the measured acoustic ratios reflect the differences in the starting number of the ABCA1 molecules; this equals 2.34 ± 0.3 times (derived by dividing the two mass ratios), in good agreement with the real-time analysis data.
(a) Experimental procedure for measuring ABCA1expression changes treated and normal mice.From a multiplex PCR reaction two biotinylated fragments are produced; a short and a long one corresponding to the GAPDH and ABCA1 genes. The reaction is loaded on the neutravidin covered acoustic device surface and the acoustic ratio is directly measured reflecting the mass ratio of the two immobilized fragments. (b) Real time binding curves displaying changes in frequency during binding of a multiplexed PCR reaction derived from control (blue line) and treated (red line) sample. (c) Respective changes in dissipation of the two samples monitored in real time. Note that the data shown here were chosen so that ΔFtreated≈ΔFcontrol and the difference in the population ratio in the two samples is reflected only in the ΔD differences; this does not have to be always the case.
Regarding the mass ratio difference that can be quantified it should be noted that the curve in figure 3 allows for the discrimination of expression differences not more than five-fold limited by the maximum and the minimum acoustic ratio value for the particular DNA sizes. According to this curve a sample with more than five folds change in expression should result in the lowest or the highest possible acoustic ratio value. Indeed, measurements performed with cDNA coming from the small intestine of the same mice gave an overexpression of 5.4 times in the treated animals, as this was semi-quantified by agarose gel electrophoresis; the acoustic measurements for the same samples resulted in a 259 ± 3 (x10−10/Hz) which is at the lowest possible value, indicating that expression was equal or greater than ~5 and beyond the discrimination capability of the method. Further optimization of the operating frequency and design of alternative PCR fragments with differences in size or conformation such as dsDNA vs. ssDNA molecules, should allow measurement of higher gene-expression differences.
Real-time PCR gene expression quantification is most commonly performed by constructing standard curves for the two genes under comparison in separate tubes containing one fluorescent dye. Our method has a major advantage over this practice since we measure the relative expression ratio of two genes in two samples (control vs. test sample) and the DNA fragments are produced within the same PCR reaction. In such comparisons most unknown variables and variation among tubes cancel out24. While it is possible to have real-time measurements using two fluorescent dyes for simultaneous measurements of two genes in one tube, this would increase the assay complexity and cost significantly. The quantification accuracy of the acoustic method was found to be comparable to the one with real-time PCR while regarding cost, speed and simplicity it is proven superior.
Discussion
PCR-based DNA detection and quantification methods, currently the ultimate in terms of sensitivity, have been criticized for being complex, expensive, time consuming and difficult to apply in multiplexed field-based analysis. In this work we demonstrated a new approach for genetic testing which retains the advantages of PCR and overcomes its detection disadvantages thanks to the application of a novel acoustic approach for amplicon identification and quantification. This approach relies on a completely new detection principle, that of DNA conformation rather than mass, in a label-free manner. It also takes advantage of the detection of dsDNA molecules rather than the hybridization of a ss-target to a surface-immobilized probe, thus, eliminates problems related to the optimization of probe structure and surface density25,26. The implementation of the acoustic technique in real samples is a strong indicator that the method can be readily applied in routine DNA analysis. The multiplexing capability is another asset of the proposed scheme. Being completely label-free, the only limitation in the maximum number of detectable products is the resolution capability of acoustic ratios. Current error bars in fig. 1b allow for the safe discrimination of up to five analytes for the DNA base-pairs range depicted in fig. 1 if these were present one at a time. For simultaneous detection, we showed that we can discriminate up to two analytes (maximum of 3 acoustic ratios); the presence of three analytes, i.e., maximum 7 ratios, would be outside the resolution capability of this DNA size range. One should note though that the instrumentation applied in this work was meant for lab-based applications; the development of a purpose-made, compact measuring system that would further stabilize environmental variations is expected to improve acoustic resolution. The application of a higher frequency surface acoustic wave (SAW) device, rather than the QCM, is also expected to lead to versatile multi-analysis platforms. SAW devices, integrated with a microfluidic module have been demonstrated to be capable of performing semi-parallel analysis of cardiac markers27 in a few minutes; similar integrated systems28 hold great promise for developing a portable system for field-based applications.
In order to fully evaluate the acoustic method, issues related to the sensitivity and resolution capability should be further addressed. Quantitatively, the number of bps of the produced amplicon can be derived through measurement of the acoustic ratio and application of Eq. 1. A point of concern is how sequence-dependent Eq.1 is, i.e., how much the intrinsic viscosity of DNA molecules of same size (i.e., bps) but different sequence would differ. Results in our lab showed that the Eq. 1 holds for nearly random sequences of DNA, unless intentional insertion of sharp bend-inducing sequences takes place. In terms of the expected shape of the examined DNA range, it should be mentioned that a small (smooth) curvature is expected for molecules longer than ~150 bp because the chain length exceeds the persistence length. Finally, with the current set-up and for the DNA range investigated here, the resolution capability or error bar of detection is ~10 bases.
One should note that the high sensitivities observed with other methods9 (E-DNA, nanoparticle-linked DNA assays etc.) are not achievable with this method, which however does not use any means of signal enhancement, labels etc. In this case, the minimum amount required in order to have a measurable signal is determined by the operating frequency of the acoustic device and lies in the nM to sub-nM range using the 35 MHz QCM15 and 155 MHz SAW devices29. In the current biosensing format where a target amplification method is included, the need for achieving extremely low detection limits is obviously eliminated.
The described assay for genotyping using acoustic biosensors has the potential to evolve into a widely-used method since it is fast, sensitive, reproducible and affordable. Acoustic measurements can be combined with any standard molecular technique such as multiplexed PCR, restriction digestion or single-stranded conformational polymorphism (SSCP) in order to create various nucleic acid fragments that may differ in their size/conformation. Such examples involve determination of viral load in clinical samples, DNA and RNA quantification and bacterial identification. We believe that the simplicity and completely label-free nature of the acoustic technique has the potential to bring a paradigmatic change in genetic lab-based and point-of-care diagnostic assays.
Methods
Acoustic measurements
Neutravidin was purchased from Pierce; 200 μg/ml were sufficient to saturate the sensor surface. The acoustic method involves the binding of DNA molecules biotinylated at their 5′-end, to a neutravidin-modified device surface. Experiments are performed in a flow-through system (flow was equal to 1 μl/sec) which allows continuous additions of DNA samples and buffer in an alternating way. The DNA attachment to the device surface is considered specific since non-biotinylated DNA gives no detectable signal. Acoustic results are expressed as the ratio of dissipation change over frequency change (ΔD/ΔF).
PCR reactions for the production of the DNA molecules with the various sizes were performed in a conventional thermocycler using Taq DNA polymerase (Minotech, Heraklion Crete). Depending on the application the amplification protocol consisted of 25–30 cycles of 30 sec denaturation at 94°C, 30 sec annealing at 55 or 60°C and 30 sec extension at 72°C, in addition to a 5 min initial denaturation step and a 5 min final extension step. The reactions were purified with Nucleospin (Macherey-Nagel) and at least 300 ng of purified DNA diluted in 200 μl of 50 mM Tris pH 7.5 and 10 mM MgCl2 buffer were loaded on the sensor per acoustic measurement.
Quantitative interpretation of acoustic results
DNA molecules with a length varying from 20 to 402 bps can be quantitatively discriminated using Equation 1. Note that, for deriving fig. 1 and Eq.1, frequency measurements at 35 MHz were used without being divided by the harmonic number (i.e. ΔFraw rather than ΔF/n were used, where n = harmonic number). Regarding the solution conditions, buffers such as Tris or PBS at near neutral pH worked equally well. In addition, the ionic strength of the buffer has never been too extreme (20 to 150 mM) while Mg, if desirable, should be kept low (e.g. 10 mM) in order, for example, to avoid chain aggregation or ion-induced bending. Experiments should be carried out at a temperature well below the Tm of the particular sequences (25°C in our case).
The mass ratio calibration curve (Eq. 2) derived from acoustic measurements using two sets of DNA mixtures with similar lengths i.e., 75–361 and 92–402 bp. In case of using significantly different DNA lengths a new calibration curve should be created experimentally.
SNP genotyping using an acoustic biosensor
Adult female mosquitoes from the laboratory colonies Kisumu, a G119 (GGC) homogygous susceptible strain from Kenya, the organophosphate resistant strain ACER-KIS due to a G119S substitution (AGC) of the ace-1 gene, the target site of carbamates and organophosphates which confers high levels of resistance30 and ACER-KIS x Kisumu, a hybrid colony of the two strains progenies, all together representing a mixture of each of the three potential genotypes (wild-type, homozygous resistant and heterozygous), were used in our study.
DNA was extracted using DNAzol reagent (Molecular Research Center, Inc) at one-fifth the recommended reagent volume for each extraction and the DNAs were resuspended in TE buffer at 50 ul volumes. Genotypes of individuals for the resistance mutations G119S were determined by using a Taqman assay as previously detailed31 and confirmed by sequencing the relevant region of the ace-1 gene.
The following primers were purchased from FRIZ Biochem (Germany); primer 1: 5′-aacagctgcgtgcagatc-3′, primer 2: 5′ –biotin - tgctgtggatcttcggcggcA-3′, primer3: 5′-ggtgccggagtagaagcC -3′, primer 4: 5′- acaccgagacggcaccggcactc-3′. Amplification protocol: 1 μl of genomic DNA was initially amplified for 25 cycles using primers 1 and 4 with annealing temperature set to 60°C. The reaction was then diluted 5 times and 1 μl was re-amplified using 1 pmol of primers 2 and 4 mixed with 10 pmoles of primers 1 and 3 for 30 cycles with 55°C annealing temperature. This particular primers ratio was found to be important for the SNP discrimination assay.
For the development of an acoustic method for SNP detection using real samples we took advantage of the fact that Taq DNA polymerase lacks a 3′ -5′ exonuclease activity and that mismatched 3′ termini are extended at a lower rate than matched termini32. However, it is important to note that not all 3′-terminal mismatches affect PCR equally32,33.
The last base at the 3′ terminus of primer 2 and 3 were designed to pair with the nucleotide at position 119 of the ace-1 gene. The last base of primer 2 was adenine (A) while for primer 3 it was cytosine (C). These two particular bases were found to be responsible for causing Taq polymerase to stall significantly and amplify selectively the 361 bp or the 179 bp product depending on the template type (wild type, heterozygous, resistive). The two particular bases of primers 2 (A) and primer 3 (C) could not be replaced by (G) or (T), respectively, since in that case Taq polymerase was found to continue polymerization without stalling.
Measuring the expression levels of ABCA1 gene
Animals and animal procedures
Male 8-week-old C57BL/6 mice were obtained from the Animal House of the Institute of Molecular Biology and Biotechnology (IMBB-FORTH, Heraklion, Greece). Animals were housed in a temperature controlled facility on a 12-hour light/dark cycle, fed by standard chow diet and water ad libitum. Mice were weight matched into two groups, the control group (n = 2), samples E47, E52 and the treatment group (n = 3), samples E48, E53, E54. Mice were administered T0901317 (Cayman Chemicals,Ann Arbor, MI) at a dose of 50 mg/kg in vehicle (0.5% w/v methylcellulose/H2O) or vehicle alone (control group) by oral gavage. 18 h after treatment the mice were sacrificed by isoflurane inhalation and tissues were collected, rapidly frozen and stored at −70oC until isolation of total RNA. All animal work was conducted in accordance with the relevant national and EU ethical guidelines. All procedures described above were approved by the Veterinary Department of the Heraklion Prefecture (Heraklion, Crete, Greece).
RNA isolation and gene expression analysis
Total RNA from livers and intestines was extracted using the TRIzol reagent (Invitrogen, Carlsbad, CA) according to the manufacturer's instructions. One μg of this RNA was reverse-transcribed with random hexamers using the SuperScript II RT System (Invitrogen) and the cDNAs produced were used for PCR amplification as well as real time qPCR. PCR amplification was performed using GoTaq® Flexi DNA Polymerase (Promega, Madison, WI). The quantification of the results was performed by measuring the intensity of the bands using the Tinascan version 2 software of Raytest (Straubenhardt, Germany). Background noise was adjusted by applying the following settings: brightness -100, contrast + 34. The settings were applied equally across the entire image and were applied equally to controls. Real-time quantitative PCR was performed in an ABI 7500 Real-Time PCR System using SYBR® GreenER™ qPCR SuperMix. qPCR for each sample was carried out in triplicates. The results were analyzed using the standard curves method. All data were normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) mRNA levels. The following primers were used for quantification of mouse ABCA1 and GAPDH mRNA levels by PCR amplification: mABCA1 F: 5′ – GAA CCA ACC ACA GGC ATG GAC CCT A – 3′, mABCA1 R: 5′ – GTG GAG TCG CTT TTT GCT CTG GGA GA – 3′, mGAPDH F: 5′ – ACC ACA GTC CAT GCC ATC AC– 3′, mGAPDH R: 5′ – TCC ACC ACC CTG TTG CTG TA– 3′. The primers used in real time qPCR were: mABCA1 qF: 5′ – GGG CTG GAT ACG AAA AAC AA – 3′, mABCA1 qR: 5′ – GGG TGG TTG AAA GCA GTG AT – 3′, mGAPDH qF: 5′ – TGT TCC TAC CCC CAA TGT GT- 3′ and mGAPDH qR: 5′- CCT GCT TCA CCA CCT TCT TG - 3′.
Multiplex PCR reaction for acoustic measurements
2 μl of cDNA (liver or small intestine) were used in each multiplex PCR reaction of 25 μl total volume containing 10 pmoles of each ABCA1 reverse and forward primer and 5 pmoles each of GAPDH reverse and forward primers. The amplification protocol consisted of 27 cycles and annealing temperature was set to 60°C. At first, we run 6 identical multiplex PCR reactions (in duplicates) with this particular primer sets ratio for 18, 21, 24, 27, 31 and 36 cycles. The products were analyzed in an agarose gel and we concluded that at 27 cycles both ABCA1 and GAPDH fragments have not reached the plateau phase.
References
Drummond, T. G., Hill, M. G. & Barton, J. K. Electrochemical DNA sensors. Nature Biotech. 21, 1192–1199 (2003).
M-C Cheng, M. et al. Nanotechnologies for biomolecular detection and medical diagnostics. Curr. Opin. Chem. Biol. 10, 11–19 (2006).
Rosi, N. L. & Mirkin, C. A. Nanostructures in nanobiodiagnostics. Chem. Rev. 105, 1547–1562 (2005).
Li, Y., Cu, Y. T. H. & Luo, D. Multiplexed detection of pathogen DNA with DNA-based fluorescence nanobarcodes. Nat. Biotechnol. 23, 885–889 (2005).
Nam, J. -M., Stoeva, S. I. & Mirkin, C. A. Bio-bar-code-based DNA detection with PCR-like sensitivity. J. Am. Chem. Soc. 126, 5932–5933 (2004).
Parks, J. S., Chung, S. & Shelness, G. S. Hepatic ABC transporters and triglyceride metabolism. Curr. Opin. Lipidol. 23 (3), 196–200 (2012).
Giljohann, D. A. & Mirkin, C. A. Drivers of biodiagnostic development. Nature 462, 461–464 (2009).
Lubin, A. A. & Plaxco, K. W. Folding-based electrochemical biosensors: the case for responsive nucleic acid architectures. Accounts Chem. Res. 43, 496–505 (2010).
Fan, C., Plaxco, K. W. & Heeger, A. J. Electrochemical interrogation of conformational changes as a reagentless method for the sequence-specific detection of DNA. Proc. Natl. Acad. Sci. USA 100, 9134–9137 (2003).
Lubin, A. A., Lai, R. Y., Baker, B. R., Heeger, A. J. & Plaxco, K. W. Sequence-specific, electronic detection of oligonucleotides in blood, soil and foodstuffs with the reagentless, reusable E-DNA sensor. Anal. Chem. 78, 5671–5677 (2006).
Lai, R. Y. et al. Rapid, sequence-specific detection of unpurified PCRamplicons via a reusable, electrochemical sensor. Proc. Natl. Acad. Sci. U.S.A. 103, 4017–4021 (2006).
Gizeli, E. & Lowe, C. R. Biomolecular sensors. (Taylor & Francis, 2002).
Tsortos, A., Papadakis, G., Mitsakakis, K., Melzak, K. A. & Gizeli, E. Quantitative determination of size and shape of surface-bound DNA using an acoustic wave sensor. Biophys. J. 94, 2706–2715 (2008).
Tsortos, A., Papadakis, G. & Gizeli, E. Shear acoustic wave biosensor for detecting DNA intrinsic viscosity and conformation: A study with QCM-D. Biosens. Bioelectron. 24, 836–841 (2008).
Papadakis, G., Tsortos, A., Bender, F., Ferapontova, E. & Gizeli, E. Direct detection of DNA conformation in hybridization processes. Anal. Chem. 84, 1854−1861 (2012).
Papadakis, G., Tsortos, A. & Gizeli, E. Triple-helix DNA structural studies using a Love wave acoustic biosensor. Biosens. Bioelectron. 25, 702–707 (2009).
Papadakis, G., Tsortos, A. & Gizeli, E. Acoustic characterization of nanoswitch structures; application to the DNA Holliday Junction. Nano Lett. 10, 5093–5097 (2010).
Weill, M. et al. The unique mutation in ace-1 giving high insecticide resistance is easily detectable in mosquito vectors. Insect Mol. Biol. 13, 1–7 (2004).
Djogbénou, L. et al. Identification and geographic distribution of the ACE-1 R mutation in the malaria vector Anopheles gambiae in south-western Burkina Faso, West Africa. Am. J. Trop. Med. Hyg. 78, 298–302 (2008).
Liu, Q., Thorland, E. C., Heit, J. A. & Sommer, S. S. Overlapping PCR for bidirectional PCR amplification of specific alleles: a rapid one-tube method for simultaneously differentiating homozygotes and heterozygotes. Genome Res. 7, 389–398 (1997).
Waterfall, C. M. & Cobb, B. D. Single tube genotyping of sickle cell anaemia using PCR-based SNP analysis. Nucleic Acids Res. 29, e119 (2001).
Schmitz, G. & Langmann, T. Transcriptional regulatory networks in lipid metabolism control ABCA1 expression. Biochim. Biophys. Acta 1735 (1), 1–19 (2005).
Thymiakou, E., Zannis, V. I. & Kardassis, D. Physical and functional interactions between liver X receptor/retinoid X receptor and Sp1 modulate the transcriptional induction of the human ATP binding cassette transporter A1 gene by oxysterols and retinoids. Biochemistry 46 (41), 11473–11483 (2007).
Kubista, M. et al. The real-time polymerase chain reaction. Mol. Aspects Med. 27, 95–125 (2006).
Prigodich, A. E. et al. Tailoring DNA structure to increase target hybridization kinetics on surfaces. J. Am. Chem. Soc. 132, 10638–10641 (2010).
Wong, E. L. S., Chow, E. & Gooding, J. J. DNA recognition interfaces: the influence of interfacial design on the efficiency and kinetics of hybridization. Langmuir 21, 6957–6965 (2005).
Mitsakakis, K. & Gizeli, E. Detection of multiple cardiac markers with an integrated acoustic platform for cardiovascular risk assessment. Anal. Chim. Acta 699, 1–5 (2011).
Suna, Y., Satyanarayan, M. V. D., Nguyenb, N. T. & Kwok, Y. C. Continuous flow polymerase chain reaction using a hybrid PMMA-PC microchip with improved heat tolerance. Sens. Actuators B Chem. 130, 836–841 (2008).
Bender, F. et al. Development of a combined surface plasmon resonance/surface acoustic wave device for the characterization of biomolecules. Meas. Sci. Technol. 20, 124011 (2009).
Weill, M. et al. Comparative genomics: insecticide resistance in mosquito vectors. Nature 423, 136–137 (2003).
Bass, C., Nikou, D., Vontas, J., Williamson, M. & Field, L. Development of high-throughput real-time PCR assays for the identification of insensitive acetylcholinesterase (ace-1R) in Anopheles gambiae. Pest. Biochem. Physiol. 96, 80–85 (2010).
Black, W. C., IV & Vontas, J. Affordable assays for genotyping single nucleotide polymorphisms in insects. Insect Mol. Biol. 16, 377–387 (2007).
Kwok, S. et al. Effects of primer-template mismatches on the polymerase chain reaction: Human immunodeficiency virus type 1 model studies. Nucleic Acids Res. 18, 999–1005 (1990).
Acknowledgements
We thank Dr. Fabrice Chandre (Institut de Recherche pour le Developpement/Centre de Recherche Entomologique de Cotonou, Benin) for generously providing mosquito strains used in this study. This research was partially funded by the European Union Seventh Framework Programme FP7 (2007–2013) under agreement number 265660 (Avecnet).
Author information
Authors and Affiliations
Contributions
G.P., A.K., I.T. and E.M. carried out the experiments. E.G. and G.P. wrote the manuscript. G.P., A.T. and E.G. contributed in data interpretation. G.P., A.T., J.V. and D.K. contributed towards the design of the experimental work.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary info
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Papadakis, G., Tsortos, A., Kordas, A. et al. Acoustic detection of DNA conformation in genetic assays combined with PCR. Sci Rep 3, 2033 (2013). https://doi.org/10.1038/srep02033
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02033
This article is cited by
-
The relationship between sphingosine-1-phosphate receptor 2 and epidermal growth factor in migration and invasion of oral squamous cell carcinoma
Cancer Cell International (2023)
-
Optimized acoustic biochip integrated with microfluidics for biomarkers detection in molecular diagnostics
Biomedical Microdevices (2017)
-
DNA transducer-triggered signal switch for visual colorimetric bioanalysis
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.