Abstract
Humoral immunity to Plasmodium falciparum circumsporozoite protein is partly mediated by a polymorphic NANP tetra-amino acid repeat. Antibody response to these repeats is the best correlate of protective immunity to the RTS,S malaria vaccine, but few descriptions of the natural variation of these repeats exist. Using capillary electrophoresis to determine the distribution of NANP repeat size polymorphisms among 98 isolates from Lilongwe, Malawi, we characterised the diversity of P. falciparum infection by several ecological indices. Infection by multiple distinct variants was common and 20 distinct repeat sizes were identified. Diversity of P. falciparum appeared greater in children (18 variants) than adults (12 variants). There was evidence of genetic distance between different geographic regions by Nei's Standard Genetic Distance, suggesting parasite populations vary locally. We show that P. falciparum is very diverse with respect to NANP repeat length even on a local level and that diversity appears higher in children.
Similar content being viewed by others
Introduction
Worldwide about 219 million people became sick and 660,000 died from falciparum malaria in 20101. The currently used control strategies – insecticide treated bed nets, indoor residual spraying and chemotherapy – are unlikely to lead to elimination or eradication without new tools. A malaria vaccine is a high priority and would provide an important weapon in the armamentarium of malaria control. Historically, efficacy trials of vaccines against Plasmodium falciparum have demonstrated incomplete protection and preliminary results of a recent phase 3 trial of RTS,S/AS01E have demonstrated only modest efficacy. The first results of this trial showed 50.4% efficacy against clinical malaria in children 5 to 17 months of age2 but only 30.1% in children 6 to 12 weeks old after 12 months3. Unfortunately, follow up studies have shown that vaccine efficacy gradually disappears over four years and is less in areas of high transmission4. Reasons for the suboptimal efficacy of this vaccine are not known. One possible explanation for poor vaccine performance might be the lack of cross-protection against genetic variants not included in the vaccine.
The RTS,S vaccine consists of components of the P. falciparum circumsporozoite protein (CSP) joined to hepatitis B surface antigen. CSP is a surface protein highly expressed on the pre-erythrocytic stage of P. falciparum and contains the NANP repeat region, a B cell epitope and several T cell epitopes. The NANP repeat region is an important target of antibodies isolated from people with naturally-acquired immunity to malaria5. This repeat evolves by concerted evolution, whereby repeat arrays expand and contract by internal duplications and deletions6. To date, there have been few systematic reports of the natural variation in repeat size6,7. The RTS,S vaccine contains an NANP repeat of 19 repeats and the C-terminal T cell epitopes8; however, a much wider range of sizes of the NANP repeat have been documented in different parasite isolates6,9. In addition, the sequence encoding the repeat may be degenerate and different repeats including NVDP and NPDP may be found; however, the repeat length is consistently 4 amino acids. Studies into the effect of NANP repeat size on vaccine efficacy are being conducted in conjunction with the phase 3 RTS,S vaccine trial, but currently it remains unclear how variations in the size of this repeat in natural parasite populations may affect vaccine efficacy.
In addition to the diversity within a parasite population, individual malaria infections are often composed of diverse populations of genetically distinct parasites. A person may be infected by multiple strains of malaria concurrently10,11. While the effect of different CSP variants on vaccine performance is not well-characterized, with no strain selection based on T cell epitopes seen in Phase II trials but a decrease in multiplicity of infection (MOI) seen in vaccine recipients12,13,14, vaccines against other P. falciparum surface proteins have shown strain specificity15. Thus further characterization of the genetic diversity of immune epitopes of vaccine antigens is important. Because children are the target population for malaria vaccines and the most vulnerable to P. falciparum infection, the role of parasite diversity is of particular interest in children.
In order to understand the genetic diversity of these antigens, characterisation of the antigenic diversity of P. falciparum in individual patients, among populations and across space is a necessary starting point. In this report, we describe the in-host and local spatial diversity of P. falciparum in Lilongwe, Malawi with respect to the NANP repeat length of CSP. Using samples from 98 parasitemic patients (49 children and 49 adults) at outpatient clinics at Kamuzu Central Hospital in Lilongwe, Malawi, we describe the genetic diversity of P. falciparum with respect to the number of repeats in the NANP repeat region determined by capillary electrophoresis using an ecological approach. Treating each patient as an individual population, we calculated within-host and between-host measures of diversity and estimated the total number of variants present in the study area. Given that children are particularly susceptible to severe malaria, we examined the differences in parasite diversity between children and adults. We also mapped the study area to characterize the spatial distribution of malaria variants and the spatial variability in parasite diversity. We have been able to detect differences in genetic diversity of P. falciparum on a smaller geographic scale than previously reported, which have previously been reported only on a regional scale16,17,18,19.
Results
Fifty children (21 male, 29 female) and 50 adults (27 male, 23 female) were recruited into the study and underwent interviews and blood draws. Mean age was 3.0 years for children and 32.1 years for adults; median age was 3 years in children and 27.5 years in adults. There was no PCR amplification of 2 samples (one male child and one female adult) and these samples were excluded from analyses.
There were no differences in intensity of parasitemia (p = 0.4) between children and adults. For this study, multiplicity of infection (MOI) refers to the number of NANP repeat variants determined in the sample. There was no difference in MOI between adults and children (p = 0.6). Children had a mean of 1.29 variants per infected person while adults had a mean of 1.22 variants per infected person. As expected due to the limitations of capillary electrophoresis, the MOI was lower than that estimated by the number of T-cell epitope haplotypes (TH2 and TH3) determined in the same samples using ultra deep sequencing as previously reported (mean MOI 2.31)20. There was no significant relationship between MOI and age for the entire population (p = 0.8); results were similar when the population was stratified into children (p = 0.5) and adults (p = 0.8).
Within-host diversity
We found 20 different alleles containing between 25 and 49 NANP repeats within our study population of 98 patients. Figure 1 shows the frequency of NANP repeat variants among the 98 patients, demonstrating that variants were not represented equally in the population (24 degrees of freedom, p = 0.00009). A total of 123 variants were detected in the 98 samples with mean MOI of 1.26 (range: 1–4), with individual variants isolated from a range of 1 to 24 patients. We found 18 variants in children and 12 variants in adults, of which 10 were shared between populations. NANP repeat lengths at the extremes of the curve appeared to be more common in children than in adults (Figure 1), but Kolmogorov-Smirnov test for different distributions did not reach significance (p = 0.08). Males had higher mean MOI than females (1.32 vs. 1.19., p = 0.2) and mean intensity of infection was slightly higher in females than in males (3.2 vs. 2.9, p = 0.06), but these differences were not statistically significant.
Indices of α diversity, which measure species richness (the number of species in a population), were calculated for children, adults and the combined population. A summary of ecological analyses is shown in Table 1. The species richness analyses showed differences in the number of variants seen in the population between adults (fewer variants) and children (more variants), but confidence intervals overlapped likely due to the small sample size. In order to estimate the true number of variants and account for adequacy and depth of sampling, we employed capture-recapture analysis and plotted variant accumulation/rarefaction curves. Capture-recapture analysis, which accounts for variants missed by sampling, predicted 15 total variants in adults, 32 in children and 34 in the entire population. If a population has been sampled thoroughly, the variant accumulation curve should begin to approach the asymptote. Our variant accumulation curves (Figure 2A–C) for all participants, adults and children appeared to be nearing their asymptotes but had not reached them yet. This data was used to develop smoothed rarefaction curves for our population, which estimate the number of haplotypes that would be detected at any given sampling level. Figure 2A–C shows the rarefaction curves for all participants, adults and children, respectively. The curves for adults and children suggest the parasite populations may be different, but the confidence intervals do overlap and have not begun to collapse.
Variant species accumulation and rarefaction curves with 95% confidence intervals.
Variant accumulation and rarefaction curves for all participants (a), adults (b) and children (c) are shown. Variant accumulation curves of the raw data are shown in dark blue. Computed smoothed rarefaction curves determined by simulation are shown as thick red lines and 95% confidence intervals for the rarefaction curves are demonstrated by thin red lines. The computed rarefaction curve represents the expected average rate of variant accumulation that would be produced by repeated sampling of the same population.
Simpson's and Shannon's indices are measures of species diversity that take into account both species richness and evenness (i.e. frequency and distribution of variants)21. These indices suggest that children probably harbour a richer and more evenly distributed parasite population than adults, although confidence intervals overlapped (Table 1). This difference in evenness between populations is suggested visually by the histogram of variant distribution (Figure 1) as well as by the fact that 8 variants uniquely occurred in children while only 2 variants uniquely occurred in adults.
Spatial diversity
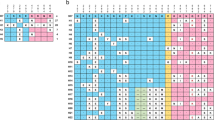
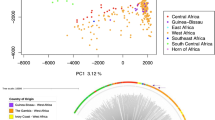
Patients reported residing in 21 of the city's 58 urban districts and 5 nearby Traditional Authorities. There was geographic data for 92 individuals in which we could locate a place of residence within the defined regions. Because very few participants lived in each individual Area, we grouped neighbouring Areas with similar geographic (east vs. west side of the river) and demographic (urban vs. periurban) features together into Regions, as shown in Figure 3A. Multiplicity of infection did not differ among the four regions (p = 0.8) nor between residents of urban and periurban/rural Areas (p = 0.6). The East Urban was the most represented region (48 individuals, 61 variants), followed by the West Urban (25 individuals, 32 variants), West Peri-urban (10 individuals, 11 variants) and East Peri-urban (9 individuals, 11 variants). The urban Regions showed a low Nei's Standard Genetic distance between the populations (0.11). Comparisons between the urban Regions and periurban Regions showed higher pairwise genetic distance (range: 0.28–1.09). All regions showed high levels of heterozygosity (range: 0.78–0.86). More alleles were seen in the urban Areas (Figure 4), though there were more participants living in these Areas than in periurban Traditional Authorities. However, the frequency of specific variants differed between Regions (Figure 4). For example, the variant with 40 NANP repeats was the single most common allele size in the total population, but it was completely absent in samples from the east periurban Region. Separation between urban and rural populations was confirmed by principle coordinate analysis (PCoA) of genetic distance between locations (Figure 3B). Assessing the significance of PCoA results using Tracy-Widon distribution was not applicable here because of the small number of clusters. Over 97% of the variance was predicted by the first two coordinates (78.92% in the first principle coordinate and 18.75% in the second principle coordinate).
Analysis of genetic similarity between different parts of the city of Lilongwe.
Panel A shows the geographic regions within and around Lilongwe used for the spatial analysis. Arrows show the pairwise comparison between each region and Nei's standard genetic distance between parasites identified in each population. The East Urban Region was the most diverse and had the highest number of participants (48 individuals, 61 variants), followed by the West Urban (25 individuals, 32 variants), West Peri-urban (10 individuals, 11 variants) and East Peri-urban (9 individuals, 11 variants). The map was prepared using ArcGIS (ESRI, Redlands, CA) and Microsoft Powerpoint (Microsoft, Seattle, WA). Panel B shows a principle coordinate analysis of genetic relatedness between regions. East and west urban regions cluster more closely to each other than either periurban region. Coordinate 1 explains 78.9% of the variation and coordinate 2 explains 18.8% of the variation.
Discussion
This cross-sectional study of parasitemic patients at a clinic in Lilongwe, Malawi reveals a high degree of genetic diversity in P. falciparum on a small scale – within individual hosts and in different neighbourhoods of the same city. In individual patients, simultaneous infection with multiple strains of malaria is common, with 22% of all patients harbouring more than one parasite variant. Using an ecological approach, we identified a non-significant trend towards greater diversity of NANP repeat length in children than adults. However, all statistics suggested this same trend, which likely would have been stronger with more samples. Interestingly, no differences were seen in MOI between the adults and children, so this could not account for the differences in diversity.
Currently, it is unclear what is driving this difference in diversity. Our data suggest that children are infected with a broader, more evenly distributed population of P. falciparum variants, a hypothesis supported visually by the distribution of variants and ecologically by the species diversity indices. One potential explanation is that adults have acquired strain-specific immunity to NANP variants. The lower diversity seen in adults could be due to pre-existing immunity to some of the variants found in the children. This cannot be confirmed in this study as we do not know what strains individuals were previously exposed to and do not have a measure of specificity of immune response. Ancillary studies to the phase 3 RTS,S study, however, will specifically address this question in relation to NANP repeat size.
There are limitations to the approach used in this study. First, as we used capillary electrophoresis to genotype infections, we can only determine the size of the allele and not the nucleotide or amino acid sequence. This approach has long been used to study diversity within malaria infections at other alleles. Size polymorphisms of the genes glurp, msp1 and msp2 are the standard means for determining multiplicity of infection by World Health Organization recommended genotyping protocols22. Thus, there may be an underestimation of alleles from this approach; however, at the amino acid level, NANP is by far the most common repeat, often representing over 90% of repeats in a population7. In addition, it has been shown that sequence alignment in repeat regions is difficult and often highly uncertain23. Thus the capillary electrophoresis approach, though it likely underestimates the number of alleles, is likely robust and will likely closely represent the number of NANP repeats due to the frequency of that amino acid repeat type. Second, selection bias may have led to undersampling of some variants. All study participants were ill enough to seek medical attention and parasites isolated from these patients may not be representative of the population, particularly of parasites causing asymptomatic parasitemia. Third, environmental, behavioural, parasitological (such as the impacts from other antigens or mixed species infections) or vector-related factors may also influence the distribution of P. falciparum variants and could not be assessed in this study. Lastly, based on capture-recapture analysis (Table 1) and our rarefaction curves (Figure 2), we predicted that we did not isolate all existing variants in Lilongwe. The impact of this under-sampling on the differences in diversity seen is unclear.
There were no significant differences in multiplicity of infection for patients living within the city limits compared to those who lived in more rural or periurban regions. This finding was somewhat unexpected, but much of “urban” Lilongwe is not very developed; furthermore, transmission intensity is very high in Malawi and urban residents may remain at very high risk. Despite the lack of difference in MOI, we were able to detect genetic distance between parasite populations in different regions within and around Lilongwe. We measured a much smaller genetic distance between parasite populations in the two urban Regions than between urban and periurban Regions or between periurban Regions. Spatial analysis was limited by the lack of address-level data, allowing us only to draw conclusions about data aggregated by district. In addition, the rural regions did have fewer participants than the urban regions, which may bias the results form under sampling those regions. Some of the periurban districts were quite large in area, so the periurban Regions may have contained substantial heterogeneity in parasite diversity that we could not detect in aggregate centroid-based analyses. The frequency of specific alleles did not change did not change over the course of enrolment.
To date, there are no reports documenting genetic distances between parasite populations within a single trial site. A few studies have shown that within relatively close distances (less than1 km), variations in exposure may lead to differences in malaria incidence and potentially immunity. A cohort of 558 children in a single parish of Kampala, Uganda, demonstrated a strong “dose-response” relationship between risk of malaria and distance of a subject's residence from a local swamp. Children living in the swamp had almost 4 times the risk of malaria than did children living more than 200 meters away24. This translated in another study to decreased efficacy of antimalarial treatment, which is known to be impacted by a patient's immunity, the further a child lived from the swamp25. Thus, differences in parasite populations on a small geographic scale are not unexpected.
The parasite's genetic diversity may be an important deterrent to the development of a malaria vaccine and is currently a topic of intense research. Design of an effective vaccine against malaria has proved elusive. There have been high hopes for the RTS,S/AS01 vaccine, but early results have been mixed. While results for older children showed greater than 50% efficacy against clinical malaria2, the most recent result in very young infants demonstrated a disappointing 30% efficacy3. Furthermore, vaccine efficacy was only 16% in high-transmission areas4. The components of the CSP antigen incorporated into the vaccine are derived from a well-established laboratory strain8 and it is not known how well the vaccine will protect against strains of malaria that are different from the vaccine strain at the NANP and T cell epitopes. Initial studies from phase 2 trials suggest that protection is not strain-specific12; however, another study has shown that the vaccine reduces multiplicity of infection with respect to msp-1 and msp-2 as well as allelic variation in the Th2 and Th3 regions of CSP in patients with breakthrough malaria14. Other candidate vaccines such as FMP2.1/AS02A against the apical membrane antigen 1 have shown poor activity against strains different from the vaccine strain15. In addition, there is evidence of selection occurring in CSP that suggests allele specific immunity, primarily in the C-terminal region20. Understanding and monitoring the diversity of the antigens in any malaria vaccine should be part of any vaccine monitoring in order to detect strains tolerant to the vaccine, even if allele specific immunity is weak to the antigen.
Our study took place within the catchment area of the phase 3 RTS,S trial site in Malawi. We showed that there is a high degree of parasite diversity present in at least one of the vaccine trial sites at baseline, so forthcoming results from these trials will shed light on the vaccine's performance against a very diverse malaria population. The RTS,S data in infants showed poor performance of the vaccine3, which is probably related at least in part to infants' immature immune system and minimal prior exposure to malaria. If our results are extrapolated to this population, we would guess that parasite diversity is very high in these infants and future studies should investigate the role of this phenomenon in vaccine efficacy.
If, as our data suggests, parasite populations may not be the same across trial sites, in the presence of strain specific immunity this may alter how we need to interpret the results of malaria vaccine trials. Traditional vaccine trials assumed that the effect of the vaccine is the same throughout the trial area, but, this view is changing. In the case of other vaccines for infectious diseases, differences in the local ecology have had an important impact on estimates of vaccine efficacy, leading to the development of “ecological” analysis of vaccine trials26,27,28,29. In depth analyses of the RTS,S trials are needed to determine if this will be the case for malaria.
This study shows that there is a large amount of genetic diversity with respect to P. falciparum circumsporozoite antigen found in the RTS,S/AS01 vaccine even in a small geographic area. This high level of diversity with respect to a single gene has unknown implications; however, highly diverse malaria populations on a local or host level may modify the protective effect of vaccine candidates. This is especially concerning in light of our findings that parasite diversity is highest in young children, the primary target population for vaccination against malaria.
Methods
Study population
A convenience sample of patients (50 children and 50 adults) at outpatient clinics at the Kamuzu Central Hospital in Lilongwe, Malawi were enrolled between 19 March and 6 May 2010. Peak malaria transmission occurs in Lilongwe between February and June. The population has been described in detail elsewhere20. Diagnostic blood smears were independently reviewed by 2 microscopists on site and smear positivity was defined as identification of P. falciparum monoinfection with parasites at a density of ≥ 2+ on a scale of 0 to 4+ (0 is not infected; 1+, 1–9 rings/100 microscope fields; 2+, 10–100 rings/100 fields; 3+, 1–10 rings/field; 4+, > 10 rings/field). Patients with severe malaria were excluded from the study. After consent was obtained, dried bloodspots were collected from each participant and stored with desiccant in individual packets at room temperature until shipment to UNC-Chapel Hill. Basic demographic and malaria-related behavioural data were collected from each patient by study personnel fluent in the patient's native language, Chichewa. All parasitemic patients were treated immediately for malaria per Malawian and WHO guidelines.
Study site
Lilongwe is the capital city of Malawi with a population of 647,448 in the urban centre and 1,230,834 in the entire district30. The city lies at 13° 59′ south, 33° 47′ east at an elevation of 1050 metres above sea level. It is divided into 58 districts and the surrounding land is broken into Traditional Authorities. Mean annual temperature is 10–22.5°C and mean rainfall is 800–1000 mm, falling almost exclusively in the rainy season between November and April. Malaria is endemic to Malawi with year-round transmission, though transmission is highest in the warm wet season. With an estimated 6 million cases per year nationally, malaria is the largest cause of morbidity and mortality in children and pregnant women in Malawi31. P. falciparum is the predominant species accounting for more than 95% of parasitemias by PCR. In a recent cross-sectional population survey, 43% of the population was parasitemic32.
Laboratory procedure
P. falciparum DNA was extracted from filter paper using the Invitrogen Pro 96 DNA kit (Invitrogen, Carlsbad, CA). We designed our PCR primers using the csp gene (Pfc0210c, PlasmoDB) using Primer3 software (http://primer3.sourceforge.net). The samples were amplified using an Eppendorf Master cycler (Eppendorf, Haupaugge, NJ) with the following reaction mix: 300 nM forward primer (GSGGATGRTAATCCTGATCC), 300 nM reverse primer (CATTTCGGTTTGGGTCATTT), 5 μl of extracted P. falciparum DNA, 1 μl 10 mM dNTP mix, 36 μl water, 5 μl 10X PCR buffer and 0.5 μl of Qiagen HotStar Taq (Qiagen, Inc., Valencia, CA) with the following PCR conditions: 95°C for 15 minutes; 40 cycles of 94°C for 30 seconds, 55°C for 45 seconds and 72°C for 2 minutes; followed by 72°C for 10 minutes for final extension and 4°C hold. The NANP region in each sample was sized by capillary electrophoresis on an Agilent 2100 Bioanalyzer using the DNA 1000 kit (Agilent Technologies, Santa Clara, CA). Variants were binned in 12 bp bins based upon the expected size of the NANP repeat region of the 3d7 strain (Pfc0210c), which contains 42 repeats (38 NANP and 4 NVDP). Ten percent of samples, including samples containing variants at the far ranges of size, were repeated to ensure reproducibility. All peaks representing more than 5% of the signal in the sample (based upon the total peak area of all peaks in the sample) and less than 50 repeats were reproducible. Peaks not meeting these criteria were excluded.
Data collection and statistical analysis
Patient data and blood spots on filter paper preparation were collected during a single patient visit to the clinic. Filter paper specimens were shipped at ambient temperature to UNC-Chapel Hill where laboratory work was performed. Linear or Poisson regression as indicated and Student's t-test were used to compare continuous variables and intensity of infection (an ordinal variable). Kruskal-Wallis test was used for categorical variables. Binary variables were analyzed with chi-squared test. Kolmogorov-Smirnov test was used to compare distributions. Two-tailed p-values were used where applicable, with statistical significance defined as p < 0.05.
Ecological analysis
Ecological measures were calculated for parasite variants in adults, children and the total population using EstimateS v8.233, using 1000 randomizations with replacement. Chao 2 statistics were calculated using the classical technique because bias-corrected values can be inaccurate with small sample sizes. Rarefaction curves were created using individual-based curves, calculated with replacement34. Simpson's index reflects the probability that two individuals in the population belong to the same species. Since mean proportional abundance of each variant increases with decreasing number of variants, Simpson's Index would have small values in datasets of high diversity and high values in data sets of low diversity. This is counterintuitive for reporting diversity, thus traditionally the reciprocal Simpson's index (1/λ) is reported. Shannon's index incorporates species richness and evenness and increases as diversity increases. The Shannon Evenness Index (SEI) is calculated by dividing the Shannon mean diversity statistic by its potential maximum [h(m)], thus limiting the range between 0 and 1 allowing for easier interpretation.
To estimate the total number of malaria variants in the population we used capture-recapture techniques. Each individual patient was treated as one sample to capture parasite variants and estimators developed by Chao were used to calculate estimates of the total number of parasite variants35. Computations for capture-recapture measurements were calculated with Chao's formula

where N is the total number of variants, S is the number of observed variants, fi is the number of distinct variants captured exactly i times and Zi is the number of distinct variants captured only in the ith sample. Statistical and ecological analyses were performed with Stata 12 (StataCorp LP, College Station, TX), Microsoft Excel (Microsoft, Seattle, WA) and EstimateS v8.233.
Spatial analysis
Since we did not have physical addresses or geographic coordinates of our study subjects, we mapped each participant to the centroid of their area of residence, using an existing geographic information system (GIS) of the study site representing urban districts and periurban/suburban Traditional Authorities. When participant's provided data about area of residence did not correspond to a governmental district or Traditional Authority (i.e., village or neighbourhood names), the location of their neighbourhood was identified with Google Earth (Google, Inc., Mountainview, CA). Area level frequency and prevalence of malaria variants were integrated into the GIS and with district-level prevalence data for P. falciparum parasitemia collected as part of an on-going study at our study site. In order to obtain adequate sample size in each geographic area, Lilongwe was divided into 4 regions: west urban, east urban, west peri-urban and east peri-urban. The west and east urban regions are naturally divided by the Lilongwe River which runs through Lilongwe. West and east peri-urban regions consisted of Traditional Authorities on the outskirts of the city. Nei's standard genetic distance was calculated using Genalex, v.6.4136,37. Nei's standard genetic distance was used rather than the unbiased correction which may give spurious results when homozygosity and sample size are small38. Principle coordinate analysis (PCoA) was performed in Genalex, v6.4136,37. Mapping and spatial analysis was performed with ArcGIS (ESRI, Redlands, CA) and Stata 12 (StataCorp LP, College Station, TX).
Ethical considerations
This study was approved by the Institutional Review Boards at the University of North Carolina and the Malawi National Health Sciences Review. All study participants provided written informed consent at enrolment.
References
World Health Organization. World Malaria Report 2012. WHO Publications: Geneva, (2012).
RTS,S Clinical Trials Partnership. First results of phase 3 trial of RTS,S/AS01 malaria vaccine in African children. New Engl J Med. 365, 1863–1875 (2011).
RTS,S Clinical Trials Partnership. A phase 3 trial of RTS,S/AS01 malaria vaccine in African infants. New Engl J Med. 367, 2284–2295 (2012).
Olutu, A., Fegan, G. & Wambua, J. et al. Four-year efficacy of RTS,S/AS01E and its interaction with malaria exposure. New Engl J Med. 368, 1111–1120 (2013).
Calvo-Calle, J. M. et al. Immunogenecity of multiple antigen peptides containing B and non-repeat T cell epitopes of the circumsporozoite protein of Plasmodium falciparum. J Immunol. 150, 1403–1412 (1993).
Zeeshan, M. et al. Genetic variation in the Plasmodium falciparum circumsporozoite protein in India and its relevance to RTS,S malaria vaccine. PLoS One. 7, e43430. 10.1371/journal.pone.0043430 (2012).
Putaporntip, C., Jongwutiwes, S. & Hughes, A. L. Natural selection maintains a stable polymorphism at the circumsporozoite protein locus of Plasmodium falciparum in a low endemic area. Infect Genet Evol. 9, 567–573 (2009).
Casares, S., Brumeanu, T. D. & Richie, T. L. The RTS,S malaria vaccine. Vaccine. 28, 4880–4894 (2010).
Rich, S. M., Ferreira, M. U. & Ayala, F. J. The origin of antigenic diversity in Plasmodium falciparum. Parasitol Today. 16, 390–396 (2000).
Kyabayinze, K. D. et al. Multiplicity of Plasmodium falciparum infection predicts antimalarial treatment outcome in Ugandan children. African Health Sci. 8, 200–205 (2008).
Lee, S. A. et al. Complexity of Plasmodium falciparum infections and antimalarial drug efficacy at 7 sites in Uganda. J Infect Dis. 193, 1160–1163 (2006).
Alloueche, A. et al. Protective efficacy of the RTS,S/AS02 Plasmodium falciparum malaria vaccine is not strain specific. Am J Trop Med Hygiene 68, 97–101 (2003).
Enosse, S. et al. RTS,S/AS02A malaria vaccine does not induce parasite CSP T cell epitope selection and reduces multiplicity of infection. PLoS Clin Trials. 1, e5. 10.1371/journal.pctr.0010005 (2006).
Waitumbi, J. N. et al. Impact of RTS,S/AS02A and RTS,S/AS01B on genotypes of P. falciparum in adults participating in a malaria vaccine clinical trial. PLoS One. 4, e7849 (2009).
Thera, M. A. et al. A field trial to assess a blood-stage malaria vaccine. N Engl J Med 365, 1004–1013 (2011).
Alam, M. T. et al. Tracking origins and spread of sulfadoxine-resistant Plasmodium falciparum dhps alleles in Thailand. Antimicrob Agents Chemother 55, 155–164 (2011).
Griffing, S. M. et al. South American Plasmodiumfalciparum after the Malaria Eradication Era: clonal population expansion and survival of the fittest hybrids. PloS one 6, e23486 (2011).
Rebaudet, S. et al. Genetic structure of Plasmodium falciparum and elimination of malaria, Comoros archipelago. Emerg Infect Dis 16, 1686–1694 (2010).
Schultz, L. et al. Multilocus haplotypes reveal variable levels of diversity and population structure of Plasmodium falciparum in Papua New Guinea, a region of intense perennial transmission. Malaria J 9, 336–348 (2010).
Bailey, J. A. et al. Diversity of and selection on Plasmodium falciparum csp T cell epitopes determined by massively parallel pyrosequencing in Lilongwe, Malawi. J Infect Dis 206, 580–587 (2012).
Magurran, A. E. Measuring Biological Diversity. Blackwell Publishing: Malden MA (2007).
World Health Organization. Recommended genotyping procedures (RPGs) to identify parasite populations. Available online at http://www.who.int/malaria/publications/atoz/rgptext_sti/en/index.html (2007).
Jongwutiwes, S. et al. Allelic variation in the circumsporozoite protein of Plasmodium falciparum from Thai field isolates. Am J Trop Med Hyg. 51, 659–668 (1994).
Clark, T. D. et al. Factors determining the heterogeneity of malaria incidence in children in Kampala, Uganda. J Infect Dis. 198, 393–400 (2008).
Greenhouse, B. et al. Decreasing efficacy of antimalarial combination therapy in Uganda is explained by decreasing host immunity rather than increasing drug resistance. J Infect Dis. 199, 758–65 (2009).
Ali, M. et al. Vaccination of adult women against cholera protects infants and young children in rural Bangladesh. Ped Infect Dis J 27, 33–37 (2008).
Ali, M. et al. Herd immunity conferred by killed oral cholera vaccines in Bangladesh. Lancet 366, 44–49 (2005).
Emch, M., Ali, M., Yunus, M., Sack, D., Acosta, C. & Clemens, J. D. Efficacy calculation in randomized vaccine trials: global or local measures? Health and Place 13, 238–248 (2007).
Emch, M., Ali, M., Yunus, P., Park, J. K., Yunus, M., Sack, D. & Clemens, J. D. Int J Epidemiol 35, 1044–1050 (2006).
Malawi Census. Available online at http://www.maseda.mw/censusinfomw/.
National Statistical Office (NSO) and ICF Macro. Malawi Demographic and Health Survey 2010. Zomba, Malawi and Calverton, MarylandUSA: NSO and ICF Macro (2011).
Malawi Ministry of Health. Malawi National Malaria Indicator Survey 2010. Lilongwe, Malawi: Malawi Ministry of Health (2010).
Colwell, R. K. EstimateS: Statistical estimation of species richness and shared species from samples. Version 8.2.User's Guide and application published at: http://purl.oclc.org/estimates (2009).
Gotelli, N. J. & Colwell, R. K. Quantifying biodiversity: procedures and pitfalls in the measurement and comparison of species richness. Ecology Letters. 4, 379–391 (2001).
Chao, A. Estimating population size for sparse data in capture-recapture experiments. Biometrics. 45, 427–438 (1989).
Peakall, R. & Smouse, P. E. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes. 6, 288–295 (2006).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research – an update. Bioinformatics. 10.1093/bioinformatics/bts460 (2012).
Hedrick, P. W. Genetics of Populations. Third edition. Sudbury, Massachusetts: Jones and Bartlett. (2005).
Acknowledgements
We would like to thank John Bruno and Feng Chang for their helpful suggestions and critical review of the analyses. We also thank the staff of the KCH pediatric and adult outpatient clinics, the staff of the UNC Project laboratory and the study participants who were generous with their time and the hope for a malaria free Malawi. Natalie Bowman is supported by Ruth Kirschstein NRSA T32 715134. This work was supported by the National Institutes of Health grants 1RO1AI089819, KL2RR025746 and UL1RR025747.
Author information
Authors and Affiliations
Contributions
S.C., T.M., F.M. and I.H. recruited patients and collected data onsite in Malawi. S.C. and J.J.J. performed the laboratory studies. N.M.B., J.C.P., S.M. and J.J.J. performed statistical analyses. V.E. and M.E. prepared spatial data and advised on spatial analysis. N.M.B. and J.J.J. prepared the manuscript and figures. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Bowman, N., Congdon, S., Mvalo, T. et al. Comparative population structure of Plasmodium falciparum circumsporozoite protein NANP repeat lengths in Lilongwe, Malawi. Sci Rep 3, 1990 (2013). https://doi.org/10.1038/srep01990
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01990
This article is cited by
-
Restricted valency (NPNA)n repeats and junctional epitope-based circumsporozoite protein vaccines against Plasmodium falciparum
npj Vaccines (2022)
-
Prevalence and risk factors associated with asymptomatic malaria among school children: repeated cross-sectional surveys of school children in two ecological zones in Ghana
BMC Public Health (2021)
-
Subclass and avidity of circumsporozoite protein specific antibodies associate with protection status against malaria infection
npj Vaccines (2021)
-
Structural and biophysical correlation of anti-NANP antibodies with in vivo protection against P. falciparum
Nature Communications (2021)
-
Epitope-coated polymer particles elicit neutralising antibodies against Plasmodium falciparum sporozoites
npj Vaccines (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.