Abstract
The c-Myc oncogenic transcription factor is known to regulate microRNA (miRNA) expression at the transcriptional level. However, little is known about the function of c-Myc in miRNA processing. Here, we report that Drosha, one of the most important components of the miRNA processing machinery, is a c-Myc target gene. c-Myc transactivates drosha mRNA expression, thus upregulating the Drosha protein level. A chromatin immunoprecipitation (ChIP) experiment revealed that c-Myc binds directly to the E-box of the drosha promoter. Both in vitro and in vivo microRNA processing assays demonstrated that c-Myc promotes miRNA processing by upregulating the Drosha expression level. Overall, our study reveals a previously unrecognised function of c-Myc in miRNA processing and provides valuable insight into a new aspect of how c-Myc regulates microRNA expression.
Similar content being viewed by others
Introduction
The c-myc oncogene encodes a conserved basic helix-loop-helix (HLH) leucine zipper transcription factor that binds to approximately 10–15% of genes in the genome and regulates genes encoding both proteins and non-coding RNAs1,2,3. MicroRNAs (miRNAs) are a set of small, non-protein-coding RNAs that regulate gene expression at the post-transcriptional level. Pri-miRNA precursors (pri-miRNAs) are transcribed from the genome by RNA polymerase II4 and fold into hairpin structures, substrates that are processed by the RNA-processing Drosha5 – DGCR86,7,8 complex into 60–70 nt miRNA precursors (pre-miRNAs). These precursors are subsequently transported to the cytoplasm where they are further processed by Dicer9,10, generating small RNA duplexes containing the mature miRNA11,12. The biogenesis of miRNA is regulated at both the transcriptional and post-transcriptional levels12,13,14 and many transcription factors regulate miRNA expression positively or negatively in a tissue-specific or developmental-specific manner15,16,17,18,19,20. Recent studies have also demonstrated that several activators and repressors regulate microRNA (miRNA) biogenesis through either protein-protein or protein-RNA interactions21,22,23,24,25.
Emerging evidence has demonstrated that c-Myc, a transcription factor that is reported to regulate cellular proliferation and tumourigenesis, is known to directly regulate miRNA expression at the transcriptional level15,16,17,26,27,28. Moreover, microRNA is also recognised as one of the important components of the c-Myc target gene network; however, less is known with regard to how c-Myc regulates miRNA biogenesis.
In this study, we show that drosha is a c-Myc target gene and that c-Myc directly binds to the E-box of the drosha promoter to activate the expression of Drosha. Through in vitro and in vivo miRNA processing assays, we demonstrate that c-Myc promotes miRNA processing by upregulating Drosha expression. Our study reveals a previously unrecognised function of c-Myc in miRNA processing.
Results
Expression profiles of c-Myc-targeted pri-, pre- and mature miRNAs
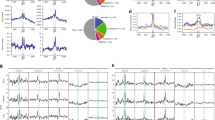
As a starting point in uncovering the molecular details of c-Myc in miRNA biogenesis, we investigated the mature, precursor and primary miRNA expression levels of c-Myc target miRNAs, including c-Myc-activated miRNAs (miR-17, miR-20a, miR-18a, miR-19a and miR-92a1/215,27) and c-Myc-repressed miRNAs (miR-16, miR-26a and miR-34a17) (Figure 1a). The P493-6 human B cell line, which carries a tetracycline-repressible c-Myc transgene, was used as our test system29. Specifically, quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) was performed to compare the levels of pri-, pre- and mature c-Myc target miRNAs in tetracycline-treated (Tet, low c-Myc) and untreated (Mock, high c-Myc) cells. As expected, the pri-miRNAs levels of the three c-Myc-repressed miRNAs were found to be upregulated under the low-c-Myc conditions (Figure 1c). Unexpectedly, four out of six primary miRNAs of the c-Myc-activated miRNAs were found to be upregulated in the low-c-Myc cells. This unexpected result may be explained by the fact that the previously defined c-Myc-activated miRNA only considered mature miRNA levels, disregarding the levels of primary microRNA transcripts. Indeed, the expression levels of the primary transcripts of c-Myc-activated miRNAs are not necessarily lower in low-c-Myc cells than in high-c-Myc cells. A similar experiment was repeated in A549 cells in which c-Myc expression was silenced by short interfering RNA (siRNA), with an increase in the pri-miRNA levels of the c-Myc-activated miRNAs in the low-c-Myc cells when compared to the high-c-Myc cells (Figure 1f). If c-Myc merely regulates c-Myc target miRNA expression via transcription, the levels of pri-miRNA for those c-Myc-activated miRNAs would be expected to be downregulated in low-c-Myc cells. Obviously, the discrepancy between the levels of pri-miRNA and mature miRNA cannot be explained simply by the transcriptional capability of c-Myc; rather, some intermediate events, for example, primary RNA processing, might be involved. We then speculated whether the apparent higher level of pri-miRNA of c-Myc-activated miRNAs in low-c-Myc cells could be the result of less miRNA processing due to the low level of c-Myc. Accordingly, we further addressed whether c-Myc affects the expression of some of the processing components (e.g., Drosha), which, in turn, modulates pri-miRNA biogenesis.
Expression profiles of c-Myc target mature, pre- and pri-miRNAs.
(a), (b) and (c). The expression levels of the mature (a), precursor (b) and primary (c) miRNAs of the indicated c-Myc target miRNAs were examined by qRT-PCR analysis in P493-6 cells treated with tetracycline (Tet) to inhibit c-Myc or left untreated to induce c-Myc. U6 snRNA was used for normalisation. The data represent the mean ± SD of three independent experiments. *indicates P < 0.05. (d), (e) and (f). qRT-PCR analysis showing the mature (a), precursor (b) and primary (c) miRNA expression levels of the indicated c-Myc target miRNAs in A549 cells treated with c-Myc siRNA or control siRNA. U6 snRNA was used for normalisation. The data represent the mean ± SD of three independent experiments. *indicates P < 0.05.
c-Myc enhances the expression of Drosha
All available evidence implicates that Drosha, DGCR8 and Dicer are the most important enzymes involved in miRNA processing. Thus, to investigate the role of c-Myc in microRNA processing, we searched for a possible link between c-Myc and one of these components of the miRNA-processing machinery. First, we examined whether the expression of c-Myc was correlated with Drosha in response to serum stimulation and found that c-Myc expression was rapidly induced after serum (10%) stimulation, reaching its peak at 2 hours, followed by a decrease in its level. As observed in Figure 2a, Drosha exhibited similar expression profiles upon serum stimulation. Moreover, using the Tet-off system, we found that the suppression of c-Myc expression was correlated with Drosha suppression (Figure 2b) and that the levels of both c-Myc and Drosha recovered rapidly after tetracycline was removed from the cell culture. At the same time, we have checked some other cell lines including HepG2 (liver hepatocellular cells) and HeLa (cervical cancer cells) cells (Supplemental Figure a) and the conclusion that positive correlation between Drosha and c-Myc upon serum stimulation is also true in these cell lines. In addition, we expressed Flag-c-Myc in 293 T and H1299 cells (Supplemental Figure b) and Drosha is indeed upregulated with increasing concentration of Flag-c-Myc. These results indicate that the finding that c-Myc upregulates Drosha appears to be a general phenomenon.
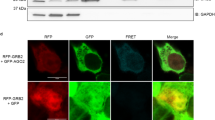
c-Myc enhances both the protein and mRNA Drosha expression levels.
(a) Western blot showing Drosha and c-Myc expression in A549 cells that were re-stimulated with serum for the indicated times. GAPDH was used as a loading control. (b) Western blot showing Drosha and c-Myc expression in P493-6 B cells treated with tetracycline (Tet) to inhibit c-Myc expression for the indicated times or first treated with tetracycline for 48 hours, washed (Wash) to remove the tetracycline and then incubated for the indicated times (48 and 72 h) after washing. Actin was used as a loading control. (c) Real-time PCR and western blotting showing that both Drosha mRNA and protein levels decrease markedly in A549 cells with c-Myc knockdown. 18 s rRNA was used for normalisation. GAPDH was used as a loading control. The data represent the mean ± SD of three independent experiments. **indicates P < 0.01. (d) Real-time PCR and western blotting showing that both Drosha mRNA and protein levels decrease markedly in LO2 cells with c-Myc siRNA knockdown. 18 s rRNA was used for normalisation. Actin was used as a loading control. The data represent the mean ± SD of three independent experiments. **indicates P < 0.01. (e) Real-time PCR and western blotting showing that both Drosha mRNA and protein levels decrease markedly in A549 cells that were serum-starved for 24 hours and then stimulated with serum (10%) for 2 hours. 18 s rRNA was used for normalisation. GAPDH was used as a loading control. The data represent the mean ± SD of three independent experiments. **indicates P < 0.01. (f) Real-time PCR showing the Drosha mRNA levels in P493 cells either treated with tetracycline for the indicated times or first treated with tetracycline for 48 hours, washed (Wash) to remove the tetracycline and then incubated for the indicated times (24 and 48 h) after washing. 18 s rRNA was used for normalisation.
Because of the positive correlation between c-Myc and Drosha, we addressed whether the transcription factor c-Myc upregulates Drosha at both the transcriptional and protein levels. To this end, c-Myc expression was silenced by siRNA in A549 (Figure 2c) and LO2 (Figure 2d) cells, which led to decreases in both Drosha mRNA and protein levels (Figure 2c, d) in each of these cell lines. Furthermore, when c-Myc expression was induced by serum stimulation, the levels of Drosha mRNA and protein were upregulated accordingly (Figure 2e). Additionally, we examined the Drosha mRNA levels in P493-6 B cells at different time points following tetracycline treatment. Consistent with our western blotting results (Figure 2b), the mRNA levels of Drosha decreased gradually over a 72-hour time period with a corresponding suppression of c-Myc (Figure 2f). Similarly, the Drosha mRNA levels were readily recovered after repeated washing (Figure 2f). Taken together, these results indicate that c-Myc enhances the expression of Drosha at both the mRNA and protein levels.
drosha is a direct target gene of c-Myc
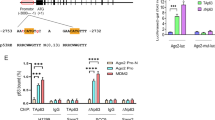
Next, we inspected the drosha promoter region for putative c-Myc binding sites. Our sequence analysis revealed the presence of a canonical E-box sequence, 5′-CACGTG-3′ (4133 bp upstream of the Drosha translational start site), which could be recognised by c-Myc (Figure 3a). To determine whether c-Myc truly binds to this region in vivo in response to serum stimulation, we performed a chromatin immunoprecipitation (ChIP) experiment. As shown in Figure 3b, c-Myc indeed bound to the E-box of the drosha promoter and this binding was further enhanced when c-Myc was upregulated by serum stimulation.
Drosha is a direct target gene of c-Myc.
(a). Schematic representation of the drosha genomic locus. The E-box (c-Myc binding site) is indicated. The deleted 6 bp of the E-box are shown (c-Myc BS mut). The exons are represented as dark boxes. The number represents the nucleotides upstream of the initiation codon (ATG) of the drosha gene. Arrows indicate the primers for the PCR reaction in the chromatin immunoprecipitation assay and luciferase constructs. (b). Chromatin immunoprecipitation was performed using A549 cells first serum-starved for 24 hours and then re-stimulated for 2 hours. The chromatin from A549 cells was fragmented and incubated with an antibody against c-Myc. An irrelevant IgG antibody was used as a negative control. (c). 293 T cells were co-transfected with Drosha-luciferase constructs (pGL3-BS) and increasing amounts of Flag-c-Myc; luciferase activity was then measured. The pGL3-vector and pGL3-BS mutant were used as controls. The error bars indicate the standard deviations. GAPDH was used as a loading control.
We then evaluated whether the E-box within the drosha promoter region confers c-Myc-dependent transcriptional activation. DNA fragments containing the wild-type E-box or a mutant E-box were inserted into the promoter region of a firefly luciferase reporter plasmid. c-Myc was able to induce luciferase expression in a dose-dependent manner from the wild-type but not the mutant reporter plasmid (Figure 3c).
c-Myc promotes pri-miRNA processing in vitro and in vivo
To understand the mechanism of c-Myc in miRNA processing, we performed an in vitro pri-miRNA processing assay by separately incubating radiolabelled pri-let7a-1 and pri-miR30a substrates with the immunoprecipitated Drosha complex from P493-6 B cells. In particular, we examined the immunoprecipitated Drosha complex from P493-6 B cells that were either treated with tetracycline (Tet) to inhibit c-Myc expression or that had been washed (Wash) to remove the tetracycline and re-induce c-Myc expression. As shown in Figure 4a, b and c, the processing activities for both the pri-let7a-1 and pri-miR30a primary transcripts were greatly reduced when c-Myc expression was turned off (Tet) in comparison to the Mock samples. Furthermore, higher processing activity was recovered (Wash) after tetracycline was removed. Additionally, we performed a similar pri-miRNA processing assay in A549 cells in which the expression of c-Myc, Drosha, or both was silenced using siRNA. As shown in Figure 4d, e and f, knockdown of either c-Myc, Drosha, or both greatly diminished the processing activity for both pri-let7a1 and pri-miR30a when compared to the control siRNA.
c-Myc accelerates pri-miRNA processing in vitro and in vivo.
(a) and (b). In vitro pri-miRNA processing assays of pri-let-7a-1 and pri-miR-30a in P493-6 B cells. The amount of pre-miRNA resulting from pri-miRNA processing was determined by a densitometric analysis of a western blot and is indicated by Arabic numbers. (c). Western blot analysis of c-Myc and Drosha expression in Tet- and Wash-treated cells. GAPDH was used as a loading control. (d) and (e). In vitro pri-miRNA processing assays of pri-let-7a-1 and pri-miR-30a in A549 cells treated with siRNA knockdown against c-Myc and Drosha separately and in combination. The amount of pre-miRNA resulting from pri-miRNA processing was determined by a densitometric analysis of a western blot and is indicated by Arabic numbers. (f). Western blot showing the siRNA gene-silencing efficiency for c-Myc and Drosha in A549 cells. Actin was used as a loading control. (g). A schematic diagram of the in vivo monitoring of pri-miRNA processing. (h). Luciferase activity was measured in A549 cells transfected with psiCHECK-pri-miR30a to monitor pri-miRNA processing in vivo. c-Myc knockdown increased the luciferase activity. The intensities were normalised by Renilla luciferase. The data represent the mean ± SD of three independent experiments. *indicates P < 0.05.
To reinforce this conclusion, we utilised a highly reliable system for the in vivo monitoring of pri-miRNA processing. A549 cells were transfected with a luciferase plasmid carrying a segment of pri-miR30a located between the luciferase-coding region and the polyadenylation signal. We speculated that the luciferase transcripts, once transcribed in the transfected cells, would lose their polyadenylation tail after the cleavage of pri-miR30a, resulting in a decrease in reporter expression (luciferase) activity (Figure 4g). Using this assay system, we observed that the knockdown of c-Myc expression caused an increase in the luciferase activity of psiCHECK2-miR30a (Figure 4h), indicating that c-Myc enhances Drosha-mediated pri-miRNA processing in vivo. In addition, we measured the pri-miRNA expression levels for a group of selected miRNAs; the pri-, pre- and mature expression forms of these miRNAs have been previously reported in different cell lines by the Schmittgen TD group30. Specifically, after knocking down the expression of c-Myc using siRNA in A549 cells, the expression levels of the majority of the pri-miRNAs (91.9%) were increased (Supplemental Figure c). Similar results were obtained in P493-6 B cells treated with tetracycline (Tet) to inhibit the expression of c-Myc (81.7%) (Supplemental Figure d). It has been reported that c-Myc actively contributes to tumourigenesis by both positively and negatively regulating the expression of diverse miRNAs; moreover, the downregulation of a subset of miRNAs is an important mechanism of c-Myc-mediated tumourigenesis. In fact, our pri-miRNA expression qRT-PCR data (Supplemental Figures c and d) revealed that for c-Myc-induced miRNAs, their pri-miRNA levels are generally high under low-c-Myc conditions and low under high-c-Myc conditions due to c-Myc-mediated miRNA processing, regardless of whether they are c-Myc activated or repressed. These results clearly suggested that when we suppressed the expression of c-Myc either by siRNA knockdown or by tetracycline induction, pri-miRNA processing was blocked at the first step, a process that is regulated by Drosha, resulting in increased pri-miRNA levels. Taken together, these results demonstrate that c-Myc modulates the Drosha-mediated processing of miRNAs both in vivo and in vitro.
Discussion
Although c-Myc has been shown to be a general negative regulator of miRNA expression17, there are important miRNAs for which expression is activated by c-Myc15,16,27. Our study provides evidence that c-Myc functions as a modulator of miRNA expression through the regulation of miRNA processing. c-Myc is currently considered a transcription factor that can directly regulate miRNA expression and our data show that c-Myc can affect miRNA processing by controlling Drosha levels through the transactivation of Drosha expression.
Recent studies have demonstrated that the regulation of miRNA biosynthesis can be mediated by many important factors, including SMAD21,31, p5332, ATM33, MutLα34 and BRCA135, which modulate miRNA processing through direct or indirect interactions with the microprocessor complex. However, a direct interaction between c-Myc and Drosha was not detected in our study (data not shown). Instead, we show that c-Myc transcriptionally regulates Drosha, whereby c-Myc indirectly regulates the activity of the pri-miRNA processing complex by controlling the amount of Drosha (Figure 5). As reported previously, DGCR8 (also known as Pasha) is also required for miRNA processing6,7,8 and the Drosha protein cleaves the hairpins on the DGCR8 mRNA, thus destabilising it36. We also examined the level of DGCR8 in response to c-Myc expression. Different from c-Myc, we found that the level of DGCR8 mRNA increased rather than decreased in P493-6 B cells upon tetracycline (Tet) treatment (low c-Myc) (Supplemental Figure e). Moreover, we found that the DGCR8 protein level did not present a corresponding marked increase (data not shown). Using a ChIP assay, we also determined that c-Myc is unlikely to directly bind to the dgcr8 promoter region, even though there is a putative c-Myc binding motif present (data not shown). These results indicated that c-Myc does not appear to regulate DGCR8 directly, but rather, affects DGCR8 mRNA via Drosha and that the c-Myc-Drosha-DGCR8 axis maintains the microprocessor activity for processing pri-miRNA to pre-miRNA.
A model for the role of c-Myc in miRNA biogenesis.
c-Myc is known to directly regulate miRNA transcription. We demonstrate here that c-Myc transactivates drosha mRNA expression, thus upregulating the Drosha protein level. As a key component of the miRNA processing machinery, once upregulated by c-Myc, Drosha is able to accelerate pri-miRNA processing. Therefore, c-Myc modulates miRNA biogenesis not only by transcriptional regulation but also by post-transcriptional modification.
Previous studies have indicated that c-Myc directly activates the transcription of the miR-17-92 cluster15, which acts as a bone fide oncogene37. Furthermore, this cluster promotes tumourigenesis in a mouse model27,38. Previous studies have also found that the activation of c-Myc causes the widespread repression of miRNA expression17, which contributes to tumourigenesis. Overall, the work reported here introduces a possible new approach to tumour treatment by modulating the c-Myc and miRNA pathways. The transactivation of Drosha by c-Myc may add another layer of complexity to Myc-mediated miRNA regulation.
Methods
Antibodies and reagents
Antibodies against the following proteins were used in this study: Drosha (Abcam; 1:1,000); c-Myc (Cell Signaling; 1:1,000); Flag (Sigma; 1:4,000); GAPDH (Cell Signaling; 1:1,000); and Actin (Abmart; 1:4,000).
Cell culture and gene knockdown with siRNA
P493-6 human B lymphoma cells were maintained in RPMI 1640 with 10% foetal bovine serum (FBS) and A549 cells were maintained in Dulbecco's modified Eagle's medium (DMEM, Invitrogen) with 10% FBS. The targeted siRNA sequences are as follows: siDrosha, 5′-AACGAGUAGGCUUCGUGACUU-3′36; c-Myc siRNA, 5′-ACGGAACUCUUGUGCGUAAUU-3′, 5′-GAACACACAACGUCUUGGAUU-3′, 5′-AACGUUAGCUUCACCAACAUU-3′ and 5′-CGAUGUUGUUUCUGUGGAAUU-3′26; and control siRNA, 5′-UUCUCCGAACGUGUCACGUTT-3′. The siRNAs were transfected into cells using Lipofectamine 2000 (Invitrogen).
Luciferase assays
A DNA fragment containing the c-Myc-binding sequence (CACGTG) was amplified by PCR with the following primers: 5′-CCTCTCGAGGCTTTCTCATTCTCTCAAGGTC-3′ and 5′-CGCAAGCTTCGATTCTTCTGTCTCAGCCT-3′. The PCR products were digested with XhoI and HindIII and cloned into the pGL3-Basic vector (Promega). To construct a mutant vector, the CAAGTG sequences were deleted by PCR-mediated site-directed mutagenesis. The luciferase assay was performed according to the manufacturer's instructions (Promega).
Chromatin immunoprecipitation
The chromatin immunoprecipitation assay was performed using the Chromatin Immunoprecipitation Kit (Upstate, USA) according to the manufacturer's instructions. The PCR primers used for the chromatin immunoprecipitation assays were 5′-GCTTTCTCATTCTCTCAAGGTC-3′ and 5′-CGATTCTTCTGTCTCAGCCT-3′.
In vitro pri-miRNA processing analysis
An in vitro pri-miRNA processing assay was performed as described previously5,6,7, with the following modifications. The PCR primers used to generate the pri-miRNA were as follows: 5′-TTTCTATCAGACCGCCTGGATGCAGACTTT-3′ was used for first-strand synthesis and 5′-TAATACGACTCACTATAGGGGATTCCTTTTCACCATTCACCCTGGATGTT-3′ and 5′-TTTCTATCAGACCGCCTGGATGCAGACTTT-3′ were used for let-7a-1 PCR amplification; and 5′-CCCTTGAAGTCCGAGGCA-3′ was used for first-strand synthesis and 5′-TAATACGACTCACTATAGGGTTGCTGTTGACAGTGAGCGA-3′ and 5′-CCCTTGAAGTCCGAGGCA-3′ were used for miR-30a PCR amplification. The PCR product was directly used as the template for in vitro transcription using a T7 Transcription Kit (Fermentas) according to the manufacturer's instructions. A total cell extract was prepared for immunoprecipitation using an anti-Drosha antibody (Abcam). After washing, beads were used for in vitro processing. The processing reaction mixture contained 10 μl of beads, 6.4 mM MgCl2, 1 unit of RNase Inhibitor (Takara, Japan) and the labelled pri-miRNA transcripts. The reaction mixture was incubated at 37°C for 60–90 min.
qRT-PCR assays
qRT-PCR assays were performed for determining the expression levels of the primary, precursor and mature miRNAs, as described previously21,32. In brief, total RNA was extracted from cells using TRIzol reagent and cDNA was synthesised from the total RNA using a PrimeScript RT reagent kit (Takara, Japan) with oligo dT primers, according to the manufacturer's instructions. For detection of the precursor and mature miRNAs, as an alternative approach, the small RNA fraction was purified with the miRvana miRNA isolation kit (Ambion, USA) and was subjected to reverse transcription and qRT-PCR analyses (normalisation using U6 snRNA); similar results were obtained using this method. qRT-PCR was performed in a real-time PCR machine (Agilent Technologies Stratagene MX3000P). The PCR cycling parameters were as follows: 94°C for 15 s and 40 cycles of 94C for 5 s and 60°C for 34 s. The data analysis was performed using the comparative Ct method. The primers used were as follows6,26,30,32,39: Drosha, 5′-TAGGCTGTGGGAAAGGACCAAG-3′ and 5′-GTTCGATGAACCGCTTCTGATG-3′; 18 s rRNA, 5′-CGGCGACGACCCATTCGAAC-3′ and 5′-GAATCGAACCCTGATTCCCCGTC-3′; pri-miR-30a, 5′-ATTGCTGTTTGAATGAGGCTTCAGTACTTT-3′ and 5′-TTCAGCTTTGTAAAAATGTATCAAAGAGAT-3′; pri-miR-34a, 5′-CCTCCAAGCCAGCTCAGTTG-3′ and 5′-TGACTTTGGTCCAATTCCTGTTG-3′; pri-miR-15a, 5′-CACACATTCGCGCCTAAAGAA-3′ and 5′-CCTATAGCACTGTGCTGGGCA-3′; pri-miR-16, 5′-GCAATTACAGTATTTTAAGAGATGAT-3′ and 5′-CATACTCTACAGTTGTGTTTTAATGT-3′; pri-miR-23a, 5′-TCTCATATGCAGGAGCCACCA-3′ and 5′-GCAAGTTGCTGTAGCCTCCTTG-3′; pri-miR-17, 5′-GCAGGAAAAAAGAGAACATCACC-3′ and 5′-TGGCTTCCCGAGGCAG-3′; pri-mirR18, 5′-TAAGGTGCATCTAGTGCAGATAG-3′ and 5′-GAAGGAGCACTTAGGGCAGT-3′; pri-miR-19a, 5′-CCAATAATTCAAGCCAAGCA-3′ and 5′-CAGGCAGATTCTACATCGACA-3′; pri-miR-20a, 5′-GCACTAAAGTGCTTATAGTGCAG-3′ and 5′-GTACTTTAAGTGCTCATAATGCA-3′; pri-miR-92a-1, 5′-TCTACACAGGTTGGGATCGG-3′ and 5′-CGGGACAAGTGCAATACCATA-3′; pri-miR-92a-2, 5′-ATGCGTATCTCCAGCACTCA-3′ and 5′-CCACCCGACAACAGCAA-3′; and U6 snRNA, 5′-CTCGCTTCGGCAGCACA-3′ and 5′-AACGCTTCACGAATTTGCGT-3′.
In vivo monitoring of pri-miRNA processing
Plasmid constructs with pri-miRNA at the 3′ untranslated region of firefly luciferase cDNA (psiCHECK2-miR30a) and expression vectors or siRNAs were transfected into A549 cells. Cell extracts were prepared at 48 h after transfection and the ratio of firefly and Renilla luciferases was measured using a Dual-Luciferase Reporter Assay system according to the manufacturer's instructions (Promega, USA).
References
Patel, J. H., Loboda, A. P., Showe, M. K., Showe, L. C. & McMahon, S. B. Analysis of genomic targets reveals complex functions of MYC. Nat Rev Cancer 4, 562–568 (2004).
Dang, C. V. et al. The c-Myc target gene network. Semin Cancer Biol 16, 253–264 (2006).
Meyer, N. & Penn, L. Z. Reflecting on 25 years with MYC. Nat Rev Cancer 8, 976–990 (2008).
Lee, Y. et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23, 4051–4060 (2004).
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Han, J. et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev 18, 3016–3027 (2004).
Gregory, R. I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Denli, A. M., Tops, B. B., Plasterk, R. H., Ketting, R. F. & Hannon, G. J. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004).
Bernstein, E., Caudy, A. A., Hammond, S. M. & Hannon, G. J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Hutvagner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834–838 (2001).
Lee, Y., Jeon, K., Lee, J. T., Kim, S. & Kim, V. N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J 21, 4663–4670 (2002).
Kim, V. N. MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 6, 376–385 (2005).
Winter, J., Jung, S., Keller, S., Gregory, R. I. & Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol 11, 228–234 (2009).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nature reviews. Genetics 11, 597–610 (2010).
O'Donnell, K. A., Wentzel, E. A., Zeller, K. I., Dang, C. V. & Mendell, J. T. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 435, 839–843 (2005).
Ma, L. et al. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat Cell Biol 12, 247–256 (2010).
Chang, T. C. et al. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat Genet 40, 43–50 (2008).
He, L. et al. A microRNA component of the p53 tumour suppressor network. Nature 447, 1130–1134 (2007).
Conaco, C., Otto, S., Han, J. J. & Mandel, G. Reciprocal actions of REST and a microRNA promote neuronal identity. Proc Natl Acad Sci U S A 103, 2422–2427 (2006).
Han, L., Witmer, P. D., Casey, E., Valle, D. & Sukumar, S. DNA methylation regulates MicroRNA expression. Cancer biology & therapy 6, 1284–1288 (2007).
Davis, B. N., Hilyard, A. C., Lagna, G. & Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454, 56–61 (2008).
Sabin, L. R. et al. Ars2 regulates both miRNA- and siRNA- dependent silencing and suppresses RNA virus infection in Drosophila. Cell 138, 340–351 (2009).
Trabucchi, M. et al. The RNA-binding protein KSRP promotes the biogenesis of a subset of microRNAs. Nature 459, 1010–1014 (2009).
Wu, H. et al. A splicing-independent function of SF2/ASF in microRNA processing. Molecular cell 38, 67–77 (2010).
Xhemalce, B., Robson, S. C. & Kouzarides, T. Human RNA methyltransferase BCDIN3D regulates microRNA processing. Cell 151, 278–288 (2012).
Gao, P. et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature 458, 762–765 (2009).
Dews, M. et al. Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat Genet 38, 1060–1065 (2006).
Bui, T. V. & Mendell, J. T. Myc: Maestro of MicroRNAs. Genes Cancer 1, 568–575 (2010).
Pajic, A. et al. Cell cycle activation by c-myc in a burkitt lymphoma model cell line. Int J Cancer 87, 787–793 (2000).
Jiang, J., Lee, E. J., Gusev, Y. & Schmittgen, T. D. Real-time expression profiling of microRNA precursors in human cancer cell lines. Nucleic Acids Res 33, 5394–5403 (2005).
Davis, B. N., Hilyard, A. C., Nguyen, P. H., Lagna, G. & Hata, A. Smad proteins bind a conserved RNA sequence to promote microRNA maturation by Drosha. Molecular cell 39, 373–384 (2010).
Suzuki, H. I. et al. Modulation of microRNA processing by p53. Nature 460, 529–533 (2009).
Zhang, X., Wan, G., Berger, F. G., He, X. & Lu, X. The ATM kinase induces microRNA biogenesis in the DNA damage response. Molecular cell 41, 371–383 (2011).
Mao, G., Lee, S., Ortega, J., Gu, L. & Li, G. M. Modulation of microRNA processing by mismatch repair protein MutLalpha. Cell research (2012).
Kawai, S. & Amano, A. BRCA1 regulates microRNA biogenesis via the DROSHA microprocessor complex. The Journal of cell biology 197, 201–208 (2012).
Han, J. et al. Posttranscriptional crossregulation between Drosha and DGCR8. Cell 136, 75–84 (2009).
Mendell, J. T. miRiad roles for the miR-17-92 cluster in development and disease. Cell 133, 217–222 (2008).
Uziel, T. et al. The miR-17 ~ 92 cluster collaborates with the Sonic Hedgehog pathway in medulloblastoma. Proc Natl Acad Sci U S A 106, 2812–2817 (2009).
Takamizawa, J. et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 64, 3753–3756 (2004).
Acknowledgements
We thank Dr. P. Jiang (University of Pennsylvania School of Medicine) for providing valuable suggestions for this project and Anthony Partridge (Pharmaron, Beijing) and Zhongren Wu (Pharmaron, Beijing) for editing the manuscript. This work was supported by grants from the Ministry of Science and Technology of China (2011CB966302 and 2010CB912804), National Natural Science Foundation of China (31030046) and the Fundamental Research Funds for Central Universities (USTC, WK2060190018).
Author information
Authors and Affiliations
Contributions
X.W. and X.Z. conceived and designed the project. X.W. and X.Z. performed all the experiments with the help of P.G. M.W. wrote the manuscript with the help of X.W. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
supplement
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Wang, X., Zhao, X., Gao, P. et al. c-Myc modulates microRNA processing via the transcriptional regulation of Drosha. Sci Rep 3, 1942 (2013). https://doi.org/10.1038/srep01942
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01942
This article is cited by
-
microRNA-184 in the landscape of human malignancies: a review to roles and clinical significance
Cell Death Discovery (2023)
-
Oncogenic AURKA-enhanced N6-methyladenosine modification increases DROSHA mRNA stability to transactivate STC1 in breast cancer stem-like cells
Cell Research (2021)
-
MicroRNA-27a-5p regulation by promoter methylation and MYC signaling in prostate carcinogenesis
Cell Death & Disease (2018)
-
MicroRNA therapeutics: towards a new era for the management of cancer and other diseases
Nature Reviews Drug Discovery (2017)
-
Interaction of RNA-binding protein HuR and miR-466i regulates GM-CSF expression
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.