Abstract
Macrophage derived foam cells are actively involved in the initial phase of atherosclerosis. Uptake of modified lipoprotein such as oxidized LDL (oxLDL) is a critical step for foam cell formation. CD36 is the major receptor mediating oxLDL uptake by macrophage. However, the molecular mechanism underlying CD36 mediated oxLDL uptake remains unclear. Here we reported that IRGM1 (IRGM in human), a member of immunity-related small GTPase family, is essential for the actin-dependent CD36 mediated oxLDL uptake by macrophage. IRGM/IRGM1 was highly expressed by macrophage around the atherosclerotic plaque and was up-regulated by oxLDL both in vitro and in vivo. Moreover loss of IRGM/IRMG1 significantly decreased oxLDL uptake in both mouse and human. Furthermore, the IRGM1 knock-out mice displayed impaired CD36 internalization in macrophage, which was associated with the deficiency of F-actin polymerization. These results revealed a novel function of IRGM1 in regulating oxLDL uptake by macrophage during atherosclerosis.
Similar content being viewed by others
Introduction
Atherosclerosis is one of the most common chronic inflammatory vascular diseases in human1. The exact etiology of atherosclerosis remains unclear. Studies have shown that hyperlipidemia, one of the most important risk factors, is strongly associated with atherosclerosis2. The abnormal accumulation of lipoproteins, especially low density lipoprotein (LDL) in the arterial intima can be modified by factors such as oxygen free radicals from resident macrophage, which causes further recruitment of monocyte-derived macrophage3. These macrophages together with smooth muscle cells engulf the modified LDL, which in turn, transforms these cells into foam cells. Foam cells can release various pro-inflammatory cytokines and trigger an inflammatory cascade leading to atherosclerotic plaque formation4. Studies indicated that macrophage derived foam cells are important for the initial phase of the plaque formation during atherosclerosis5.
OxLDL accounts for large proportion of modified LDL that can be recognized by a variety of receptors, including the class A scavenger receptor SR-A and the class B scavenger receptor CD36 6. CD36 is thought to be responsible for more than 50% of oxLDL uptake by mouse and human macrophages7. Upon exposure to oxLDL, CD36 together with oxLDL is internalized into punctuate organelles in the cytoplasm8. Further study has revealed that CD36 diffusion within linear confinement regions promoted unengaged receptor clustering, which is dependent on the cytoskeleton-mediated organization of receptor diffusion in the membrane and enhances CD36 responsiveness to ligand9. The internalization of CD36 together with the oxLDL is not only important for oxLDL uptake but is also crucial for signal transduction. However, little is known about how the actin-dependent CD36 internalization being regulated upon oxLDL activation.
IRGM1 (IRGM in human) belongs to the immunity-related small GTPase family. Studies have revealed the multi-function of IRGM1 in diseases, including intracellular pathogen infection10,11,12,13 and autoimmune diseases14,15 and stroke16. Recent study also has shown that IRGM1 can facilitate macrophage migration through regulating actin polymerization17. Since actin also involved in oxLDL uptake, we asked whether IRGM1 is involved in oxLDL uptake by macrophage through actin-dependent CD36 internalization. To test this possibility, we tested the expression of IRGM/IRGM1 in atherosclerotic lesion and investigated how IRGM1/IRGM expression is regulated by oxLDL during atherosclerosis. The data demonstrated that IRGM1 plays a potent role in regulating the oxLDL uptake by macrophage via modulating the actin-dependent CD36 receptor-ligand internalization during atherosclerosis.
Results
IRGM/IRGM1 was highly expressed by foam cells in the lesion of atherosclerosis
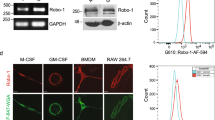
To test whether IRGM1 was associated with the pathogenesis of atherosclerosis, we first examined the IRGM1 expression in the atherosclerotic plaque. ApoE−/− mice were fed with western diet to induce the mouse model of atherosclerosis. The expression of IRGM1 in the aortic arch was detected by western blot. Compared with the control mice, IRGM1 was highly expressed in the western diet-fed mice (Figure 1a, 1b and 1c). Co-staining of IRGM1 with macrophage marker CD68 suggested that IRGM1 was expressed in the CD68+ macrophage (Figure 1d). Consistent with mice, IRGM was also highly expressed by the macrophage around the human AS plaque (Figure 1e).
IRGM/IRGM1 is highly expressed by foam cells in the lesion of atherosclerosis.
(a) AS plaque of ApoE-/- mice was validated by oil-red staining. IRGM1 expression in aortic arch of the mice with or without western diet was detected by western blot (b, c) (p = 0.0003, n = 5) and located by immuno-fluorescence staining (IRGM1-red, CD68-green, DAPI-blue) (d). Human IRGM expression in autopsy AS lesion was detected by immunohistochemistry staining (e). Data are represented as mean ± SEM.
OxLDL up-regulated IRGM/IRGM1 expression in macrophage
As oxLDL is an important factor that regulates macrophage response during atherosclerosis, also because IRGM1 is highly expressed by macrophages in atherosclerotic lesion, it is intriguing to assume that the IRGM1 expression can be regulated by oxLDL in macrophages during atherosclerosis. To test this hypothesis, peritoneal macrophages (PMΦ) and bone marrow derived macrophages (BMMΦ) were isolated from C57/B6 mice and treated with or without oxLDL for 48 hours. As shown in figure 2, IRGM1 was strongly up-regulated in both BMMΦ (Figure 2a) and PMΦ (Figure S1a) after oxLDL treatment compared with the nil condition (Figure 2a and S1a). Consistently, we further confirmed this finding by using human THP1 cell derived macrophages (Figure S1b and S1c) and human primary monocyte-derived macrophages (HMMΦ, Figure 2b). Interestingly, using primary monocytes, we also found a positive correlation between IRGM expression and serum LDL level (Figure 2c), indicating that the potential impact of LDL on IRGM expression in vivo. MiR196 has been reported that can direct regulate human IRGM expression15. We found that oxLDL strongly decreased miR196 expression (Figure 2d), which is consistent with a recent miRNAs profiling study using whole atherosclerotic plaque materials18. To further confirm that oxLDL regulates human IRGM expression through miR196, we transfected human macrophage with miR196 mimics. Compared to the control condition, miR196 mimics significantly decreased oxLDL induced IRGM expression in human macrophage (Figure 2e and Figure S2), indicating that oxLDL may regulate IRGM expression through down-regulation of miR196. These data together indicated that oxLDL induced IRGM/IRGM1 expression both in vitro and in vivo.
OxLDL up-regulates IRGM/IRGM1 expression in macrophage.
Bone marrow macrophages (BMMΦ) were recruited from C57/B6 mice. Cells were then treated with or without oxLDL for 48 hours. IRGM1 expression was tested by western blot ( a, n = 4, p = 0.01). (b) IRGM expression of human monocyte derived macrophage (HMMΦ) was detected by western blot (n = 3, p = 0.01). (c) Correlation of IRGM1 expression and LDL level in human (n = 17, p = 0.003). (d) Human macrophage was treated with oxLDL for 48 hrs. MiR196 expression was detected by real-time PCR (n = 6, p = 0.001). (e) Human primary monocyte derived macrophage was either untransected or transfected with either control miRNA or miR196 mimics. Western blot was used to detected IRGM expression (n = 3). Data are represented as mean ± SEM.
Loss of IRGM/IRGM1 decreased oxLDL uptake by macrophage both in vivo and in vitro
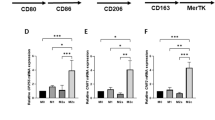
We next asked whether IRGM is involved in atherosclerosis pathogenesis in vivo. We fed Irgm+/+ApoE−/− and Irgm+/−ApoE−/− with western diet and compared the area of the aortic lesion. We showed a significant decrease of red-oil stained atherosclerotic lesion in the Irgm+/−ApoE−/− (Figure 3a). Moreover, after treating either Irgm1+/+ or Irgm1−/− BMMΦ with oxLDL for 48 hours, we found a decrease of total cellular cholesterol (Figure 3b). Together, these data suggested that loss of IRGM1 decreased the foam cell formation in vivo and in vitro. Since the influx and efflux of the oxLDL together determine the formation of the foam cells, the decrease of foam cell formation may be due to the deficiency of oxLDL uptake or the increase of degraded oxLDL efflux or both. To address that question, we first compared oxLDL uptake by BMMΦ and PMΦ from either Irgm1+/+ or Irgm1−/− mice using confocol microscopy and flow cytometry. Dil labeled oxLDL was used to trace the oxLDL up-take. To avoid the contamination of surface sticking oxLDL, cells were washed with ice-cold acid buffer before detection. Results from both BMMΦ (Figure 3c and 3d) and PMΦ (Figure S3a and S3b) suggested that oxLDL uptake by macrophages was significantly impaired in the Irgm1−/− mice. To confirm this finding in human, small-interfering RNA (siRNA) specifically targeting human IRGM was used to knock down its expression in human primary monocyte-derived macrophages. Consistent with the finding in mice, oxLDL uptake by macrophages was lower in the IRGM-siRNA group, compared with control-siRNA treated group (Figure 3e and 3f). We also measured the expression of the receptors related to oxLDL efflux such as ABCA1 and ABCG1. Interestingly, the natural expression of both ABCA1 and ABCG1 was significantly higher in the Irgm1−/− mice (Figure S4). However, such differences were lost after oxLDL treatment. It has been reported that oxLDL can increase both ABCA1 and ABCG1 expression19. But in order to do that, the proper uptake of oxLDL is required for the signal transduction. So the inability to further increase ABCA1 and ABCG1 expression may due to the significant decrease of oxLDL uptake in the Irgm1−/− mice. These data indicated that IRGM1/IRGM is involved in the atherosclerosis pathogenesis and part of reason is through regulating oxLDL uptake by macrophage in both mouse and human.
Loss of IRGM/IRGM1 decreases oxLDL uptake by macrophage.
Irgm1+/+ or Irgm1−/− mice were fed with western diet for 3 months. Aorta was isolated and stained with red oil. (a) The representative and the quantified data of atherosclerotic lesion (n = 6/group, p = 0.04). BMMΦ were isolated from either Irgm1+/+ or Irgm1−/− mice. Ox-LDL or Dil-oxLDL was added for 48 hrs after isolation or induction. (b) Biochemical lipid quantitive assay was used to measure total cellular lipid. Dil-oxLDL was detected by either (c) confocal microscope (n = 4/group, p = 0.0004) or (d) flow cytometry (n = 8/group, p = 0.001). Human monocyte derived macrophage (HMMΦ) was transfected with either control-siRNA or IRGM-siRNA. IRGM-siRNA was validated by real-time PCR (, n = 3, p = 0.02). Dil-oxLDL uptake was measured by flow cytometry (f, n = 3, p = 0.04). Data are represented as mean ± SEM.
IRGM1 regulates CD36 internalization via modulating F-actin polymerization
CD36 and scavenger receptor A are important for oxLDL uptake by macrophage. We next compared the expression of CD36 and SRA expression between Irgm1+/+ and Irgm1−/− mice. We found that Irgm1+/+ and Irgm1−/− mice have similar basic expression of CD36 and SRA (Figure S5), which indicated that the impaired of oxLDL uptake in the Irgm1−/− mice is not due to the lower expression of these receptors. A recent study has demonstrated that CD36 linear diffusion and its receptor and signaling function is controlled by actin cytoskeleton9. Interestingly, studies have also revealed a role of IRGM1 in modulating cytoskeleton remodeling and membrane dynamics of macrophage17, which raised the possibility that IRGM1 may regulate the CD36 function via modulating actin cytoskeleton. To address this possibility, we first determined whether the function of CD36 in regulating the ligand recognition and internalization is impaired in the absence of IRGM1. As shown in figure 3, upon binding to CD36 cross-linking antibody, CD36 internalization was remarkably suppressed in the Irgm1−/− mice as compared to Irgm1+/+ mice (Figure 4a and 4b). We then asked whether IRGM1 control the CD36 internalization via actin cytoskeleton. As shown in figure 4c and 4d, F-actin polymerization was significantly reduced in Irgm1−/− mice with or without oxLDL. As expected, in the wild type mice, oxLDL enhanced F-actin polymerization (Figure 4c and 4d). Interestingly, unlike wild type mice, macrophage from Irgm1−/− mice was failed to increase F-actin polymerization after oxLDL treatment as we compared the ratio of actin MFI with or without oxLDL (Figure 4d). These data demonstrated that IRGM1 regulates CD36 function via controlling the F-actin polymerization and, possibly, the CD36 diffusion and clustering on the membrane.
IRGM1 controls CD36 internalization by regulating actin polymerization.
BMMΦ from either Irgm1+/+ or Irgm1−/− mice was labeled with CD36 cross-linking antibody. Then, they were either left in 37°C to cross-link CD36 or on ice. Cells were then washed by cold acid wash buffer to deplete surface CD36. Confocal microscope and flow cytometry were used to detect and quantify CD36 internalization respectively. (a and b n = 8/group, p = 0.004). F-actin polymerization was detected by phalloidin-FITC in BMMΦ from either Irgm1+/+ or Irgm1−/− mice with or without oxLDL. The fluorescence was measured by confocal microscope and quantified by flow cytometry (c and d, n = 3, p = 0.02). Data are represented as mean ± SEM.
Discussion
Although the uptake of oxLDL by macrophage via scavenger receptors has been well established to be a critical step to foam cell formation and atherosclerosis, it remains poorly understood how this process being regulated. Our study has uncovered a novel function of IRGM/IRGM1 as a regulator for the CD36-mediated oxLDL uptake by macrophage. During atherosclerosis, oxLDL up-regulates IRGM/IRGM1 in macrophage, which, in turn modulates the CD36 internalization of receptor-ligand complex and promotes the oxLDL uptake. This positive feedback process involves the modulation of F-actin polymerization by IRGM1.
IRGM/IRGM1 has been implicated in a wide variety of biological functions, including host resistance to infections by intracellular bacterial and protozoan pathogens (e.g Mycobacteria and Toxoplasma gondii), stem cell development and maturation, T cell and neuronal autophagy, etc10,11,12,13,14,15,16. Further studies have demonstrated that Irgm1 is critically required for normal motility of activated macrophages and is pivotal in controlling the innate response to infection in vivo17. However, these observations have been confined to infectious conditions. Our study reveals a novel mechanism for Irgm1 in macrophages involved in atherosclerotic pathology.
It has been shown that the reciprocal interaction between oxLDL-CD36 and actin is important for oxLDL induced cellular and molecular events. On the one hand, through CD36, oxLDL can induce actin polymerization and control macrophage migration, polarity and locomotion20,21. One the other hand, uptake of oxLDL by CD36 occurs by an actin-dependent pathway distinct from macropinocytosis8. OxLDL or antibody-induced clustering of CD36 triggers internalization of the receptor-ligand complex. Cytoskeleton perturbations that inhibited diffusion in linear confinement regions reduced receptor clustering in the absence of ligand and, following ligand addition, suppressed CD36-mediated signaling and internalization. Our results suggested that IRGM1 could be one of the important regulators for the actin-driven CD36 internalization in macrophage. Although the precise mechanism connecting IRGM1 to the cytoskeleton remains to be determined, Taylor's group showed that IRGM1 can influence the activity of Ras-related C3 botulinun toxin subsetrate 1 (Rac1) in macrophages during activated condition, which is an important modulator for F-actin polymerization17. Further studies are needed to verify this possible mechanism.
In conclusion, for the first time we demonstrated a pathogenic role of IRGM/IRGM1 in atherosclerosis. The data expand our vision of IRGM1 as a potent regulator for macrophage motility and activity in both infectious and non-infectious conditions and have important implications for developing novel strategies for anti-atherosclerotic therapies.
Methods
Cells
Peritoneal macrophages were recruited from Irgm1+/+ and Irgm1−/− mice by injecting 2 ml 10% thioglycollic acid broth 3 days before collecting the cells with ice-cold RPMI. Cells were then rested on 24 well plates in RPMI with 10% fetal bovine serum for 24 hrs before experiments. For bone marrow-derived macrophages (BMM), femurs were dissected from Irgm1+/+ and Irgm1−/− mice, BMMs were cultured in DMEM supplemented with conditioned medium collected from B16-GM-CSF cells for 5 days. Human THP1 cells were activated with 50 ng/ml PMA for 48 hrs before experiments. Healthy subjects were recruited from healthy volunteers. All subjects provided informed consent as approved by the HMU ethics review board. Human peripheral blood monocytes were purified by using CD14 beads (Miltenyi bio) based on manufacture's protocol and activated with 20 ng/ml hGMCSF (R&D system) for 4 days before experiments. OxLDL used in the experiments is from human peripheral blood LDL oxidized using Cu2SO4(oxidant) in PBS;Dil-oxLDL is from oxLDL labled with 1,1′-dioctadecyl - 3,3,3′,3′ – tetramethyl - indocarbocyanine perchlorate (Dil). The concentration of Dil-oxLDL is 50 μg/ml and the concentration of oxLDL is 25 μg/ml.
Mice
C57BL/6 (B6) mice were purchased from HFK BIOSCIENCE (Beijing). Irgm1 knock-out mice (Irgm1−/−) were a gift by Dr. Gregory A. Taylor (Department of Medicine, Division of Geriatrics and Immunology, Duke University, USA) and were backcrossed to B6 background for 10 generations. ApoE−/− mice under the background of C57BL/6 were feed with western diet (TD.88137 Adjusted Calories Diet, from HFK BIOSCIENCE) for 12 weeks before experiments. All mice were housed in the animal facilities of the Harbin Medical University. All experimental procedures involved were performed according to protocols approved by the Institutional Animal Care and use Committee at the Institute of Genetics and Developmental Biology.
Antibodies
Monoclonal mouse anti-mouse IRGM1 (Clone:1c11) and monoclonal mouse anti-human IRGM (Clone:1G9) from AbMart; Polyclonal rabbit anti-IRGM from HuaAn Biotechnology Company; Monoclonal rat anti-mouse CD68; Donkey anti-mouse IgG-DyLight550 (ab98767); Donkey anti-rat IgG-DyLight488 (ab102232); Donkey anti-rabbit IgG-DyLight488 (ab98488) and Goat-anti-mouse IgA FITC (ab97234) from Abcam; Monoclonal mouse anti-β-actin from sigma; Rat anti-mouse CD36 IgA (Clone: No.63, 1956928) from Millipore; Rat anti-mouse CD36-APC (Clone: No.72-1); Mouse anti-human CD36 IgM-eFluor660 (50-0369); Rat anti-mouse F4/80-APC (Clone: BM8) from eBioscience; Rat anti-mouse CD11b-FITC (553310) from BD Pharmingen; Goat anti-mouse IgG, HRP-linked antibody from Cell Signaling; Goat anti-rabbit IgG, biotin linked antibody from Cell Signaling; anti-biotin, HRP-linked antibody from Cell Signaling.
Oil-red staining
The aortic arch from ApoE−/− mice feed with or without western diet was fixed for 24 hours in ice-cold 4% buffered paraform, After washed by PBS for 2 times, samples were stained with 1% of oil Red O solution (Sigma, Aldrich, USA) for 2hours. Samples were then washed for three times with PBS. Atherosclerostic lesions were measured by Image-Pro Plus 6.0 software with double-blinded assessment.
Western blot and Quantitative analysis
The protein concentration was measured by using BCA assay (P0012, Beyotime). Protein lysates were separated by SDS-PAGE and transferred to NC membranes. The membranes were blocked in non-fat-milk in TBS-T and were incubated with primary antibody overnight at 4°C. Then, after several washes, the membranes were incubated with secondary antibodies. After final washes, the signals were detected by chemiluminescence. The bands were quantified by ImageJ and values were normalized to β-actin.
Immunocytochemistry Staining
The sections were washed with PBS and then blocked with normal goat serum, 0.3% BSA, 0.05% saponin and 0.3% Triton X-100 in PBS for 1 hour. Then they were incubated at 4°C overnight with the primary antibody. The sections were later washed with PBS and incubated for 1 hour with secondary antibodies. The sections were then rinsed three times in PBS and mounted with GEL/Mounting Medium (MO1, Biomeda, USA) after nuclear staining. Fluorescence microscope (Nikon, Japan) and confocal microscope (Zeiss, Germany) were used to capture the images.
Real-time PCR
Total RNA was extracted by using TRIzol reagent (15596-026, Invitrogen) according to the manufacturer's protocol. cDNA synthesis was performed using random hexamer primers and the TaqMan reverse transcription kit (4366596, Applied Biosystems). Samples were subjected to real-time PCR analysis on an ABI Step-one Sequence Detection System under standard conditions. The sequence of pre-designed primer for human IRGM: forward primer: GGACTCTGGCAATGGGATGT, reverse primer: CCCTCATGTCCTGTGTTTCGA, probe primer: FAM – ACCTTCATCAGTGCCC - MGB (Invitrogen). Expression of IRGM/IRGM1 was normalized with the expression of actin. Other primers including: mABCA1: forward primer:5′CGTTTCCGGGAAGTGTCCTA3′ Reverse primer:5′ GCTAGAGATGACAAGGAGGATGGA3′ mABCG1: forward primer:5′ GGGAAGTTGATAAAGGATGT3′ Reverse primer:5′ GATTCGGGCTATGTATGG3′ MiR.196b: forward primer: CGCCGCTACTAGGTAGTTTCC; Reverse primer: CGTATCCAGTGCGTGTCGTG; U6: forward primer: GCTTCGGCAGCA -CATATACTAAAAT; Reverse primer: CGCTTCACGAATTTGCGTGTCAT; mCD36: Forward: 5′-GATGACGTGGCAAAGAACAG-3′ ; Reverse: 5′-TCCTCGGGGT -CCTGAGTTAT-3′mSRA:Forward: 5′-TAGGCACTTGGGATGTCTGA-3′; Reverse : 5′-GTCCTCAATTTGTATTGGTGCT-3′
Small interfere RNA experiment for human IRGM
IRGM gene silencing was performed in THP1 cells and human primary monocyte derived macrophages with siGENOME SMART pool reagent (Dharmacon). Control Non-targeting siRNA pool (Dharmacon) was used to assess the silence specificity of IRGM siRNA. The transfection was performed 24 hours before any experiment.
MiR196 mimic transfection
Human primary macrophage was induced with 20 ng/ml hGM-CSF for 48 hours and was transfected with miR196 mimic (40 nM) and miRNA negative control (40 nM) using lipofectamine RNAiMAX (Invitrogen) for 24 hours and changed with fresh medium.
Cholesterol collection and measurement in macrophage derived foam cell
Mice bone marrow derived macrophage in 24-well plate was treatment with 50 ug/ml oxLDL for 48 hours, removed the medium, washed with ice 0.01 M PBS for 2 times, intracelluar cholesterol was distracted by added 0.5 ml hexane : isopropanol (3:2), collected in vials and dried in fume hood and the total cholesterol in foam cells were detected with Amplex Red Cholesterol Assay Kit (Invitrogen,A12216). The protein assay was performed using the BCA assay kit from Pierce.
Detection of Dil-OxLDL uptake by flow cytometry
Dil-labeled oxLDL (YB-0010, Yiyuan Biotech.) was used to trace the oxLDL uptake. Cells were washed with cold PBS for 2 times before any staining. Cold acid washing buffer (0.5 M glacial acetic acid, 150 mM sodium chloride, pH 2.5) was used to wash surface adherent ox-LDL. Cells were then detached from the plate with 0.05 mM EDTA in PBS. Cell surface markers were stained on ice for 20 min. After 2 times washes by FACS washing buffer (2% FBS+PBS), samples were analyzed by either FACS Calibur (BD) or Accuri 6 cytometry (BD). Data was further quantified by FlowJo (Tree star Inc.).
CD36 cross-linking and internalization assay
Cells were washed with ice-cold RPMI for two times. And then they were incubated with anti-CD36 IgA for 10 min on ice, after two times washes with cold RPMI, cells were later incubated with anti-mouse IgA-FITC for another 10 min. Then the control group will be left on ice and the cross-linking group will be transferred to the 37°C incubator for 30 min in RPMI. Then cells were washed twice with cold PBS and incubated with cold acid wash buffer (as above) for 2 min to wash out the surface CD36, followed by recovery in ice-cold RPMI for 2 min. Samples were analyzed by Accuri 6 cytometry (BD).
Phalloidin staining
Cells were fixed with cold 4% paraform in PBS for 5 min on ice and then perforated with 0.1% trion-X100 in PBS for other 5 min. After one time wash with cold PBS, cells were then stained for F-actin with phalloidin-FITC (sigma) for 30 mim in 4°C. After two times washes with PBS, cells were analyzed by Accuri 6 cytometery (BD).
Statistical Analysis
The statistical analyses were performed using SPSS 19.0 for Windows. All of the values were presented as the mean ± SEM and were analyzed by using either student's t-test or a one-way analysis of variance (ANOVA). Statistical significance was defined as p < 0.05.
References
Hansson, G. K. & Hermansson, A. The immune system in atherosclerosis. Nat Immunol. 12, 204–12 (2011).
Weber, C. & Noels, H. Atherosclerosis: current pathogenesis and therapeutic options. Nat Med. 17, 1410–22 (2011).
Moore, K. J. & Tabas, I. Macrophages in the pathogenesis of atherosclerosis. Cell. 145, 341–55 (2011).
Allahverdian, S., Pannu, P. S. & Francis, G. A. Contribution of monocyte-derived macrophages and smooth muscle cells to arterial foam cell formation. Cardiovasc Res. 95, 165–72 (2012).
Tabas, I. Macrophage death and defective inflammation resolution in atherosclerosis. Nat Rev Immunol 10, 36–46 (2010).
Endemann, G. et al. CD36 is a receptor for oxidized low density lipoprotein. J Biol Chem. 268, 11811–6 (1993).
Collot-Teixeira, S. et al. CD36 and macrophages in atherosclerosis. Cardiovasc Res. 75, 468–77 (2007).
Collins, R. F. et al. Uptake of oxidized low density lipoprotein by CD36 occurs by an actin-dependent pathway distinct from macropinocytosis. J Biol Chem. 284, 30288–97 (2009).
Jaqaman, K. et al. Cytoskeletal control of CD36 diffusion promotes its receptor and signaling function. Cell. 146, 593–606 (2011).
MacMicking, J. D., Taylor, G. A. & McKinney, J. D. Immune control of tuberculosis by IFN-gamma-inducible LRG-47. Science. 302, 654–9 (2003).
Hunn, J. P. et al. Regulatory interactions between IRG resistance GTPases in the cellular response to Toxoplasma gondii. EMBO J. 27, 2495–509. (2008).
King, K. Y. et al. Irgm1 protects hematopoietic stem cells by negative regulation of IFN signaling. Blood. 118, 1525–33 (2011).
Feng, C. G. et al. The immunity-related GTPase Irgm1 promotes the expansion of activated CD4+ T cell populations by preventing interferon-gamma-induced cell death. Nat Immunol. 9, 1279–87 (2008).
Xu, H. et al. Genetic deficiency of Irgm1 (LRG-47) suppresses induction of experimental autoimmune encephalomyelitis by promoting apoptosis of activated CD4+ T cells. FASEB J. 24(5), 1583–92 (2010).
Brest, P. et al. A synonymous variant in IRGM alters a binding site for miR-196 and causes deregulation of IRGM-dependent xenophagy in Crohn's disease. Nat Genet. 43, 242–5 (2011).
He, S. Y. et al. Immune-related GTPase M (IRGM1) regulates neuronal autophagy in a mouse model of stroke. Autophagy. 8, 1621–1627 (2012).
Henry, S. C. et al. Regulation of macrophage motility by Irgm1. J Leukoc Biol. 87, 333–43 (2010).
Bidzhekov, K. et al. MicroRNA expression signatures and parallels between monocyte subsets and atherosclerotic plaque in humans. Thromb Haemost. 107, 619–25 (2012).
Xu, M. et al. ABCG1 mediated oxidized LDL-derived oxysterol efflux from macrophages,. Biochem. Biophys. Res. Commun. 390, 1349–1354 (2009).
Park, Y. M., Febbraio, M. & Silverstein, R. L. CD36 modulates migration of mouse and human macrophages in response to oxidized LDL and may contribute to macrophage trapping in the arterial intima. J Clin Invest. 119,136–45 (2009).
Park, Y. M. et al. Oxidized LDL/CD36 interaction induces loss of cell polarity and inhibits macrophage locomotion. Mol Biol Cell. 23, 3057–68 (2012).
Acknowledgements
We thank Dr. Gregory A. Taylor for providing the Irgm1−/− mice. We also thank Dr. Zhiheng Xu for project discussion. These studies were funded by National Natural Science Foundation Projects to Dr. Hongwei Xu (No. 81071949), Dr. Chaodong Wang (No. 81271249) and Dr. Shaohong Fang (No. C080102); and by grants from department of education, Heilongjiang province funding to Dr. Shaohong Fang (No. 12511301).
Author information
Authors and Affiliations
Contributions
X.F.C. performed the majority of the experiments and wrote the manuscript; L.R. developed concept of this study, revised the manuscript and involved in the human part of this study; W.C.D. revised the manuscript and provided the human AS samples; Y.S. & F.S.H. collected human blood samples; T.L.L., D.H.Y., P.C.Y., H.S.Y. & J.P.Y. are involved in the mouse part of the study and project discussion; H.R.C. was involved in the animal model induction and project discussion; L.H.L. & X.H.W. raised the initial idea of this study, revised the manuscript and analyzed the data.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Suppementary Figure 1 - 5
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Xia, F., Li, R., Wang, C. et al. IRGM1 regulates oxidized LDL uptake by macrophage via actin-dependent receptor internalization during atherosclerosis. Sci Rep 3, 1867 (2013). https://doi.org/10.1038/srep01867
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01867
This article is cited by
-
The neurorepellent, Slit2, prevents macrophage lipid loading by inhibiting CD36-dependent binding and internalization of oxidized low-density lipoprotein
Scientific Reports (2021)
-
CD36 in Atherosclerosis: Pathophysiological Mechanisms and Therapeutic Implications
Current Atherosclerosis Reports (2020)
-
Macrophagic CD146 promotes foam cell formation and retention during atherosclerosis
Cell Research (2017)
-
IRGM1 enhances B16 melanoma cell metastasis through PI3K-Rac1 mediated epithelial mesenchymal transition
Scientific Reports (2015)
-
IFNg-induced Irgm1 promotes tumorigenesis of melanoma via dual regulation of apoptosis and Bif-1-dependent autophagy
Oncogene (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.