Abstract
Although attempts have been made to reveal the relationships between bacteria and human health, little is known about the species and function of the microbial community associated with oral diseases. In this study, we report the sequencing of 16 metagenomic samples collected from dental swabs and plaques representing four periodontal states. Insights into the microbial community structure and the metabolic variation associated with periodontal health and disease were obtained. We observed a strong correlation between community structure and disease status and described a core disease-associated community. A number of functional genes and metabolic pathways including bacterial chemotaxis and glycan biosynthesis were over-represented in the microbiomes of periodontal disease. A significant amount of novel species and genes were identified in the metagenomic assemblies. Our study enriches the understanding of the oral microbiome and sheds light on the contribution of microorganisms to the formation and succession of dental plaques and oral diseases.
Similar content being viewed by others
Introduction
To improve our understanding of the interactions between microbes and human hosts, large efforts have been made to characterize the composition of the human microbiome at different body sites1,2. The oral cavity consists of a complex system of tissues and organs that provide 2 primary surfaces for microbial colonization: the mucosa and teeth. To date, >200 bacterial species have been cultured from the human oral cavity and approximately 1,000 phylotypes have been detected by 16S rRNA gene sequencing of oral samples3. Recent studies have revealed that the predominant oral microbiota was largely consistent across healthy individuals4,5, which were generally nonpathogenic and considered to be commensal in the human oral cavity. However, some of the bacteria may be harmful and responsible for oral diseases, such as dental caries and periodontal disease6.
Gram-negative Porphyromonas gingivalis, Treponema denticola and Tannerella forsythia are frequently isolated from dental plaques in periodontal patients and were initially considered specific pathogens of periodontal disease7. Subsequently, a strong correlation between the proportions of several cultivable bacteria (e.g., Prevotella intermedia, Fusobacterium nucleatum, Selenomonas noxia, Actinobacillus actinomycetemcomitans and Eubacterium nodatum) and periodontal disease has been reported8,9,10. During the past 10 years, some representatives of the genera Megasphaera, Parvimonas, Desulfobulbus and Filifactor were reportedly more abundant in periodontal lesions using culture-independent molecular techniques11,12. In addition, a recent study indicated that some species present in low quantities orchestrate inflammatory periodontal disease through the commensal microbiota13; however, the microbial community composition associated with periodontal disease remains unclear.
A variety of virulence factors in periodontal bacteria, including fimbriae adhesins, lipopolysaccharides (LPSs), peptidoglycans, lipoteichoic acids, etc., are potent inducers of pro-inflammatory cytokines and the massive release of these mediators can lead to oral diseases14. For example, P. gingivalis, the most studied periodontal pathogen, can produce a number of well-characterized virulence factors. Several studies revealed that P. gingivalis LPSs stimulate osteoclasts and induce bone resorption. Butyric acid and volatile sulfur compounds cause genomic DNA damage in human gingival epithelial cells. All or part of these virulence factors were discovered in other periodontal bacteria, including A. actinomycetemcomitans15, P. intermedia16, T. denticola17 and Neisseria cinerea18. However, previous studies have only focused on several toxins of a few well-known pathogens, while virulence factors of other oral bacteria contained in the functional pools have been rarely investigated. Particularly, there are no studies that have characterized the functional divergence between oral microbiomes in healthy people and patients with periodontal disease.
Metagenomic sequencing allows screening of the genetic composition and functional potential of a microbial community. In the present study, we collected dental swabs or plaques from periodontal healthy and diseased individuals, which yielded approximately 350 million short reads using whole genome shotgun (WGS) sequencing. With the help of comparative metagenomic approaches, we provided a comprehensive view of the microbial community associated with periodontal health and chronic periodontitis. We attempted to describe the core microbiota and its metabolic functions associated with oral diseases on a genome-wide scale.
Results
Human DNA contamination and sequencing data
To avoid sequencing excessive human DNA from oral samples and to establish a pre-evaluation method, nine dental plaque samples (PY2, PY4, PY5, PZ2, PZ3, PZ8, H4, H6 and H7) were selected and examined with absolute quantification using qPCR. Contamination was assessed by measuring the levels of the human housekeeping gene β-actin. The results showed that the standard curves for β-actin were highly linear (R2 > 0.995) in the range tested by the duplicate reactions. The slopes of the standard curves were −3.36. The concentration of human DNA in samples measured by qPCR was 0.4–9.1 ng μL−1. The proportion of human DNA in samples, ranging from 2.6–53.5%, was calculated by dividing the concentration of human DNA by that of total DNA. Subsequently, 16 samples (including nine pre-evaluated plaque samples) were metagenomic sequenced and 1.5–3.1 Gbp of high-quality sequences were generated for each sample (see Supplementary Table S1). Alignments of these sequencing reads against the human genome showed that 20.5–74.4% of the total reads were of human origin. This was approximately 20% higher than that quantified by qPCR, which was probably due to the different amplification efficiencies between the pure human DNA standard and the human-microbial mixture sample. However, a linear correlation between both quantification methods was observed (see Supplementary Fig. S2), which indicated that the qPCR method can be applied to sample selection before shotgun sequencing of host-associated communities.
After screening out human DNA contaminants, 1.5–18.4 million microbial PE reads were kept in each of the 16 datasets. These PE reads were merged into 0.3–2.8 million sequences (140 ~ 194 bp each) based on the overlaps, which were then used for downstream metagenomic analyses (see Supplementary Fig. S1).
Bacterial community composition of periodontal health and disease
To explore the bacterial community composition of periodontal health and periodontitis, the PE-merged sequences were BLASTXed against the NCBI NR protein database and the MEGAN analysis pipeline was used to parse BLAST hits and to estimate bacterial abundance. Abundance differences were evaluated using the Mann-Whitney U test and Bonferroni correction for multiple comparisons. As shown in Figure 1B, the left 16 columns represented 5 subgingival plaque samples (H4, H6, H7, H9-2 and H14-2) and 2 supragingival swab samples (H9-1 and H14-1) of periodontal disease and 6 subgingival plaque samples (PY2, PY4, PY5, PZ2, PZ3 and PZ8) and 3 supragingival swab samples (Z11, Z14 and Z15) of periodontal health, which were recorded as groups H-2, H-1, PZ and Z, respectively. The most striking difference between the 7 periodontal disease and the 9 health samples was derived from the relative proportions of the four most abundant phyla, Bacteroidetes, Actinobacteria, Proteobacteria and Firmicutes (U test, P < 0.001). Bacteroidetes was the most abundant phylum (41.0–59.2%) in all samples of periodontal disease, whereas its abundance decreased to 5.6–38.0% in both swab and plaque samples of periodontal health. Instead, Actinobacteria (9.3–41.0%) and Proteobacteria (5.2–40.1%) were significantly increased in plaque of periodontal health. Firmicutes (14.8–58.3%) or Proteobacteria (9.2–46.5%) became the most abundant phylum in swab of periodontal health. Columns Z, PZ, H-1 and H-2 in the right of Figure 1B represented the bacterial community of each sampling group based on 16S rRNA gene extracted from PE-merged sequences and classified using the RDP classifier. These four columns showed similar community structures with NR-BLASTX classification of corresponding samples. Moreover, the relative proportions of the major phyla in several plaque (e.g. column PY4 and column PZ) and swab (e.g. column Z15 and column Z) samples/groups of periodontal health were consistent with two periodontally healthy samples (columns SEED and RDP) reported in previous studies, respectively19,20.
Sample collection, composition and clustering relationships of bacterial communities in periodontal health and disease.
(A) A schematic overview of sample collection. In healthy individuals, dental surfaces (Z group) were swabbed and supragingival plaques (PZ group) were collected from the gingival margin. In chronic periodontal patients, dental surfaces (H-1 group) were swabbed and subgingival plaques (H-2 group) were collected from the bottom of the periodontal pocket. (B) The distribution of major phyla in the bacterial communities of periodontal health and disease. The left 16 columns (H4-Z15) indicated the bacterial phyla classified via NR-BLAST. Columns Z, PZ, H-1 and H-2 showed the bacterial components of each group based on 16S rRNA gene sequences. Columns SEED and RDP represented the bacterial distributions of 2 reference samples (MG-RAST IDs: 4446622.3 and 4444448.3) collected from periodontally healthy volunteers. The bacterial classifications of these two reference data sets employed the SEED (www.theseed.org) and RDP classification systems, respectively. (C) Principal component analysis of 16 periodontal bacterial communities at the genus level based on the metagenomes. The first two principal components (PC1 and PC2) can explain 67% of the data variance. Different colors denote 3 distinct periodontal states (Z, PZ and H-1 and -2).
To further explore the relationship between different bacterial communities of periodontal health and disease, a PCA analysis was performed using the genus-level taxonomic profiles (see Supplementary Table S2). As shown in Figure 1C, the first two principal components, representing 67% of the variance, classified the 16 samples into three groups (H, PZ and Z), with all samples of periodontal disease forming a group (H, blue circle) apart from the plaque group (PZ, red circle) and the swab group (Z, green circle) of periodontal health. The analysis of similarity (ANOSIM) test using Bray–Curtis dissimilarity showed that the observed cluster patterns were significant (R = 0.6803, P = 0.001). Interestingly, two plaque samples (PY4 and PY5) in the PZ group were more closely related to groups Z and H, respectively, indicating that these samples may represent transition states of periodontal health.
The top 30 most abundant genera, representing 80.0–97.6% of the bacteria in each sample, are shown in Figure 2. The relative abundance of each genus is indicated by the circle area. Microbial community of the swab samples of periodontal health were dominated by Streptococcus (13.7–41.3%), Haemophilus (2.0–25.8%), Rothia (0.9–16.7%) and Capnocytophaga (3.1–13.0%). The remaining genera contributed less than 10.0% in proportion. In contrast, swab and plaque samples of periodontal disease exhibited a much different taxonomic composition, in which Prevotella formed 14.4–44.7% of the bacterial communities (U test, P < 0.001). In plaque of periodontal health, no single genus dominated the communities, but instead, Streptococcus, Capnocytophaga and several other genera exhibited similar trends in abundance.
The distribution of the major genera in the microbiomes of periodontal health and disease.
The y-axis shows the top 30 most abundant genera which constitute 80.0 ~ 97.6% of each bacterial microbiota. The relative abundance of each genus is indicated by the circle area. The 16 columns (H4-Z15) indicate the bacterial genera classified using the NR-BLASTX algorithm.
In order to visualize the bacterial shift from periodontal health to disease, we calculated the average abundance of dominant genera for each of the four sampling groups (Z, PZ, H-1 and H-2). Among these groups, swab of periodontal disease (group H-1) had the lowest equitability (Simpson's Index, SI = 0.21), followed by plaque of periodontal disease (group H-2, SI = 0.23) and plaque (group PZ, SI = 0.24) and swab (group Z, SI = 0.28) of periodontal health. As shown in the NR-BLASTX classification (see Supplementary Fig. S3A), groups of periodontal disease (H-1 and H-2) consisted of higher proportions of anaerobic and Gram-negative bacteria, such as Prevotella, Leptotrichia, Veillonella, Porphyromonas and Treponema than periodontal health (Z and PZ). There was not much difference between groups H-1 and H-2, although several known pathogens (Prevotella, Porphyromonas and Treponema) were over-represented in subgingival plaque (H-2) (see Supplementary Fig. S3A). To confirm the BLASTX-based taxonomic classification, we extracted 16S rRNA genes from the PE-merged sequences and reclassified them into various taxonomic levels using the RDP classifier, which yielded a similar taxonomic distribution (see Supplementary Fig. S3B).
Intraspecific diversity of periodontal pathogens
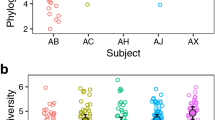
To investigate the intraspecific diversity of certain periodontal pathogens mentioned above, we aligned unassembled PE reads of all samples to currently available reference genomes and identified a large number of single nucleotide variations (SNVs) and small insertions and deletions (indels). For example, 57,479 SNVs and 931 small indels were identified from the aligned metagenomic reads when using the genomic sequence of Treponema denticola ATCC 35405 (NC_002967.9) as a reference (see Supplementary Fig. S4). It should be noted that these observed polymorphisms also occurred across different Treponema species or strains in the periodontal community. When these reads were blasted against three other closely related Treponema species (T. denticola ATTC35405, T. vincentii ATCC35580, T. phagedenis F0421 and T. pallidum ssp. pallidum Chicago), sequence similarity indicated that most reads could be assigned to T. vincentii with high sequence identities (≥90%), followed by the periodontally pathogenic bacterium T. denticola (Figure 3A). Only a small fraction of reads could be assigned to T. phagedenis and T. pallidum, but with a much lower sequence similarity (Figure 3B).
Intraspecific diversity of periodontal pathogens.
(A–B) The similarity between metagenomic reads and 4 Treponema reference genomes. The x- and y-axes indicate the sequence similarity at the amino acid level. (A) Most of the reads assigned to Treponema shared higher sequence similarity to T. vincentii than T. denticola. (B) Compared with the 2 genomes in Figure 3A, only a few reads could be assigned to T. phagedenis and T. pallidum. (C–D) Intraspecific diversities of Porphyromonas in microbiomes of periodontal disease based on FimA, Kgp and Rgp genotype classifications. (C) Porphyromonas in the microbiomes of periodontal disease consisted of 60.7% of type II, 16.4% of type Ib, 5.0% of type I and 1.5% of type IV FimA strains of P. gingivalis, as well as 16.4% of P. endodontalis. The circle size in the phylogeny is proportional to genotype frequency. (D) P. gingivalis possessing the type II (TDC60-like, 87.5%) Kgp genes were detected at a higher frequency than type I (ATCC33277-like, 12.5%). As for Rgp, the W50-like type was the most prevalent (62.2%), followed by TDC60-like type (27.3%) and ATCC33277-like type (10.5%). (E) The genetic diversity of Prevotella taxa observed in the plaque samples of periodontal health (PZ) and disease (H-2). The x-axis represents different Prevotella taxa sequenced by the Human Microbiome Project (HMP) project, whereas the y-axis represents its relative frequency in the metagenome.
P. gingivalis, another well-recognized periodontal pathogen, had been classified into several subgroups based on various virulence factors, including FimA, Kgp and Rgp cysteine proteinases21,22. With Parabacteroides johnsonii (ZP_03478485) from the family Porphyromonadaceae serving as an outgroup, a maximum likelihood (ML) phylogenetic tree (Figure 3C) was constructed using MEGA software with 1,000 replicates of amino acid sequences from 6 FimA genotypes (types I, Ib, II, III, IV and V) of P. gingivalis fimbriae (D17795, AB058848, D17797, D17801, D17802 and AB027294), P. endodontalis (ZP_04389631) and P. uenonis (ZP_04055896). Sequences assigned to 6 FimA genotypes, 2 genotypes (types I and II) of the Lys-specific cysteine proteinase Kgp gene and 3 types (TDC60-, ATCC33277- and W50-like) of the Arg-specific cysteine proteinase Rgp gene were separately extracted from unassembled PE reads of all swab and plaque samples of periodontal disease to calculate genotype frequencies. As shown in Figure 3C, the most dominant FimA genotype in P. gingivalis was type II (60.7%), followed by type Ib (16.4%) and type I (5.0%). This finding was consistent with previous studies, in which type II FimA was the most prominent among of P. gingivalis clinical isolates from a wide range of geographic locations21. Similarly, we also demonstrated that type II Kgp genes and W50-like Rgp genes in P. gingivalis strains were more prevalent than any other genotypes (Figure 3D).
Prevotella spp. was found to be widely distributed in plaque samples of both periodontally healthy (PZ) and diseased (H-2) individuals (Figure 2). To test if there was any difference in the abundance or diversity of Prevotella between both plaque groups, we assembled the PE reads for each group separately and then predicted the open reading frames (ORFs) from the assembled contigs. The putative proteins were compared against the NR database and significant hits (amino acid sequence identity ≥95%) matched to the Prevotella genus was used to determine the query sequence's taxonomic classification. As shown in Figure 3E, a much greater diversity of Prevotella taxa was observed in the H-2 group compared with the PZ group. This result was consistent to the MEGAN analysis of unassembled reads (data not shown). The most prevalent Prevotella phylotypes detected in the PZ group were P. nigrescens ATCC 33563, followed by Prevotella sp. oral taxon 472 and Prevotella sp. oral taxon 317. In contrast, the Prevotella genus in the H-2 group was highly diversified, with P. tannerae, Prevotella sp. C561, P. pallens, P. micans and P. nigrescens as the top five most abundant phylotypes, but none exclusively dominated the communities.
Functional variation between the microbiomes of periodontal health and disease
To investigate the functional divergence between the microbiomes of periodontal health and disease, we annotated the metagenomic reads using the KEGG database. Given that plaque accumulation is closely related to periodontal disease, plaque samples of periodontal health (PZ group, including PY2, PY4, PY5, PZ2, PZ3 and PZ8) and disease (H-2 group, including H4, H6, H7, H9-2 and H14-2) were compared and analyzed in detail. After comparing the amount of genes assigned to each KEGG pathway between the PZ (y-axis) and H-2 (x-axis) groups (see Supplementary Fig. S5A), we found a series of significant differences (Mann-Whitney U test with Bonferroni correction) that lead to the functional divergence between periodontal health and disease. These divergences mainly involved in membrane transport, signal transduction and cell motility (see Supplementary Fig. S5A). In detail, carbohydrate metabolism, amino acid metabolism, energy metabolism, lipid metabolism, membrane transport and signal transduction were under-represented in the H-2 group, whereas glycan biosynthesis and metabolism and cell motility were over-represented (see Supplementary Fig. S5B). Insight into these functional variations is illuminated in a schematic diagram Figure 4.
A schematic overview of the functional variations in the microbiomes associated with periodontal disease.
Briefly, genes involved in bacterial chemotaxis, flagellar assembly and biosynthesis of lipopolysaccharides and the collagenase PrtC gene were significantly over-represented in the microbiomes of plaque of periodontal disease. On the contrary, genes related to membrane transport (PTS and ABC transporters) and metabolism (e.g., H2S2 and H2O2) were underrepresented. The upward arrows in red and downward arrows in blue represented over- and under-represented pathways or genes in the microbiomes of plaque of periodontal disease, respectively.
One of the most striking observations was that almost all genes involved in bacterial chemotaxis were significantly over-represented in the microbiome of the H-2 group (Mann-Whitney U test, P < 0.001) (Figure 4). Among these genes, the expression of methyl-accepting chemotaxis protein (MCP), which encodes a transmembrane chemoreceptor, was 2–7 times greater than the chemotaxis histidine protein kinase (CheA) gene. A significant increase was also found in the coupling factor CheW, two response regulators CheY and CheB and methyltransferase CheR (U test, P < 0.001). Notably, not all bacterial chemotaxis is mediated by chemoreceptors MCP. Enzyme I (PtsI) of the phosphotransferase system (PTS) binds to and inhibits CheA when a variety of sugars are transported into the cell23. It seems that the microbiome of the H-2 group reduced the number of PTS genes to weaken the inhibition on CheA and further enhance bacterial chemotaxis (Figure 4).
An over-representation of flagellar assembly-related genes was observed in the microbiome of the H-2 group (Figure 4), which may be responsible for pathogen mobility and colonization24. Compared with the PZ group, there was a statistically significant increase (U test, P < 0.001) in synthetic genes encoding the filament, hook (FliD/C and FlgL/K/E genes) and rods and rings (FliE/F and FlgB/G/H/I) of flagella. More reads were assigned to FliG/M/N genes, which was in concert with the increase of their upstream regulator CheY. Encoding genes of the type III secretion system, FlhA/B and FliH/I/O/P/Q/R, also had a significantly higher abundance in the H-2 group than in the PZ group (U test, P < 0.001).
Bacterial endotoxins, LPS and collagenase PrtC protein are important virulence factors that elicit host immune responses and result in tissue lesions25,26. As expected, there was a significant enrichment of LPS and PrtC genes and a widespread decline in anabolism and catabolism in microbiomes of the H-2 group (Figure 4). For example, encoding genes of enzymes that mediate sulfur and peroxide metabolism were under-represented in the H-2 group. These enzymes include CysN/D/H/J/I, which deoxidizes sulfate to H2S and PatB, MetB/C and CysK/M/O, which catalyze H2S to the reductive carboxylate cycle and Cys/Met metabolism, as well as CAT, which catabolizes toxic substances (e.g., H2O2, formaldehyde, phenol and ethanol). In addition to the reduction of ATP-binding cassette (ABC) transporter genes, the genes encoding microbial membrane transport in the H-2 group were also less abundant than in the PZ group.
To clarify the bacterial contribution to the functional divergence between periodontal health and disease (Figure 4), reads assigned to bacterial chemotaxis, flagellar assembly, LPS biosynthesis and PrtC were extracted from the H-2 group and the PZ group, respectively. These reads were re-BLASTXed separately against the NR database to determine their taxonomic origin. The over-representation of bacterial chemotaxis in periodontitis samples was made up of five genera (Treponema, Selenomonas, Campylobacter, Prevotella and Fusobacterium) that were more abundant in the H-2 group than in the PZ group (see Supplementary Fig. S6A). Similarly, genera with high abundance in the PZ group were principal sources of chemotaxis genes in plaque of periodontally healthy microbiomes. Treponema, Selenomonas and Campylobacter were the first three donors that contributed to the increase of flagellar genes in the H-2 group (see Supplementary Fig. S6B). Over-representation of LPS biosynthesis genes in the H-2 group was due to the vigorous propagation of genera Prevotella, Porphyromonas and Fusobacterium (see Supplementary Fig. S6C). Another over-represented gene in the H-2 group, encoding the PrtC protein, was provided by the genera Prevotella, Treponema, Selenomonas, Porphyromonas and Fusobacterium (see Supplementary Fig. S6D). Taken together, these results suggested that the contribution to these variations (U test, P < 0.001) was generally consistent with the bacterial taxonomic composition within different periodontal status.
Novel microbial inhabitants and undiscovered functions
When comparing the sequence similarity between assigned metagenomic reads and their reference sequences (see Supplementary Fig. S7A), we found that the vast majority shared a high sequence identity with known species (≥90% at the amino acid level, based on the H4 sample for the top 30 abundant genera). Twenty-five genera, which had never been reported as members of human oral microbiota, were detected in at least three of our swabs and dental plaque samples using the NR-BLAST based approach with an identity score ≥90% (see Supplementary Table S3). Of these 25 candidates, some have been reported in the human gastrointestinal tract (e.g., genera Blautia, Bryantella, Collinsella, Holdemania, Parabacteroides and Roseburia), blood (e.g., Psychrobacter) and skin (e.g., Kytococcus); some are of animal origin, such as Basfia isolated from bovine rumen, Dichelobacter from unguis of sheep, goats and cattle and Riemerella from the blood of ducks, geese, turkeys and other birds; and some are discovered in marine or freshwater ecosystems (e.g., Citreicella, Methylobacillus and Spirochaeta). Since the NR-BLAST-based classification mainly employed complete genomes and functional sequences as references, bacteria that only had their 16S rRNA genes sequenced might be neglected. Therefore, we extracted and classified all 16S sequences from the metagenomes and identified nine new taxa that had not been reported previously in the human oral cavity (see Supplementary Table S3). Some of them (e.g., Clostridium, Parabacteroides, Eubacterium, Ruminococcus and Lactobacillus) had a relatively lower sequence identity (~75%) with known reference sequences compared with other abundant genera (see Supplementary Fig. S7B).
To estimate the level of undiscovered functions in the oral microbiota, we assembled sequencing reads of all samples and yielded approximate two million contigs with a total length of 570 Mb. After removing short contigs (<80 aa), we performed gene predictions and obtained 498,886 predicted ORFs no shorter than 80 amino acids. Only 175,080 (35.1%) of them had ≥90% sequence identities as compared to the NR database. About 8.9% of them shared no sequence similarity with any known proteins (see Supplementary Fig. S8). These findings indicated that although more than 100 oral bacterial genomes have been sequenced, there are still a large amount of novel species and functions remain undiscovered in the human oral microbiome.
Phages identified in periodontal communities
Other than bacteria, a small proportion of the PE merged reads (~0.16%) had been assigned to phages according to the result of NR BLASTX. Among them, Actinomyces and Streptococcus phages were predominant types, comprising 68.4% and 29.1% of the phage communities, respectively. Additionally, unclassified Myoviridae and Propionibacterium phage were also detected, but at very low abundance (Figure 5A). Interestingly, these identified phages were not evenly distributed among various periodontal communities. The Actinomyces phage was the most abundant type in plaque samples of periodontal health, while the Streptococcus phage was more prevalent in swab samples of periodontal health than in the others (Figure 5B). Such an uneven distribution was consistent with the abundance of their hosts, Actinomyces and Streptococcus (Figure 2 and Supplementary Fig. S3).
Phages present in the periodontal microbiomes.
(A) Actinomyces and Streptococcus phages were predominant, which comprised 68.4% and 29.1% of the phage communities, respectively. Unclassified Myoviridae (mainly Aggregatibacter or Haemophilus), Propionibacterium and some unclassified phages were also common constituents, but present at low amounts. (B) The Actinomyces phage was the most abundant type in plaque of periodontal health, while the Streptococcus phage was more abundant in swab of periodontal health than in the other groups. (C) Phylogenetic analysis of the upper collar protein homologs (YP_001333664) in the periodontal microbiomes. The proteins annotated as putative homologs of the upper collar protein (YP_001333664) in Actinomyces phage AV-1 were used to build a phylogenetic tree. (D–E) The distribution of metagenomic reads on the contigs PZ-C8421430 and PZ-scaffold426 were classified as Actinomyces phages.
From the assembled contigs no shorter than 80 amino acids, we extracted all predicted proteins, which were annotated as homologs of the upper collar protein (YP_001333664) in Actinomyces phage AV-1 and used them to build a phylogenetic tree. It clearly showed that each group of samples contained multiple distinct Actinomyces phages (Figure 5C). For example, in plaque samples of periodontal health, scaffold10190_orf67458 was nearly identical to the reference protein, whereas C8421430_orf65203 and scaffold426_orf25451 were much more divergent, which shared only 78% and 44% identity, respectively. Moreover, we used the Integrated Next-gen Genome Analysis Platform (inGAP) v2.7 bioinformatic tool27 to align paired-end reads to the contigs C8421430 and scaffold426. The alignments showed an even distribution of read coverages across the entire sequences (Figure 5D–E), which further confirmed the presence of multiple Actinomyces phages in the periodontal community.
Discussion
Metagenome sequencing has greatly facilitated the study of the human oral microbiome28. However, the current understanding of the oral microbiome mainly comes from healthy individuals29, as little is known about microbial shifts and particular functional changes associated with oral diseases. Dozens of swab and plaque samples of periodontal health and disease had been collected in our study. By using the qPCR approach, we firstly evaluated the fraction of human DNA contamination in these samples. Considering the sequencing cost and downstream bioinformatics analyses, we only selected and sequenced 16 less contaminated periodontal samples representing four different periodontal groups, Z, PZ, H-1 and H-2. Given that the sample size for each group is small, we focused on the comparison of PZ and H-2, which has 6 and 5 samples, respectively and employed statistical approaches to identify significant differences between the two groups. Our study represents the first comprehensive study of microbiomes associated with periodontal health and disease by using WGS sequencing. We re-defined a core microbiota associated with chronic periodontitis and identified 34 novel oral genera that were present in at least three individuals. Due to their relatively low abundance, we cannot rule out the possibility that these previously unreported species may be transiently present in human dental plaques. Based on comparative metagenomic analyses, we attempted to explain the mechanism of microbial community assembly of dental plaque and to characterize the bacterial agents and their functions associated with periodontitis on a genome-wide scale.
Consistent with a 16S-based study30, the WGS sequencing performed in our study further confirmed that there were significant differences in bacterial community compositions and relative abundances between patients with chronic periodontitis and healthy control subjects. The core microbiota associated with chronic periodontitis is largely consistent in both studies; however, we observed some discrepancies. First, the relative abundance of the most dominant bacterial genera differed, even within the same study30. For example, in the former study, Prevotella and Fusobacterium were the most abundant genera in subjects with chronic periodontitis using different primers and targeted regions (V1–V2 or V4 of the 16S rRNA gene, respectively). On the contrary, metagenomic shotgun sequencing used in the present study generated 16S rRNA gene sequences more randomly, which was more robust to unveil the community structure. As shown in Figure 1B, the taxonomic composition and abundance inferred from the 16S rRNA gene were roughly similar to those classified by the NR-BLAST based approach. Secondly, we detected several additional low-abundance genera associated with periodontitis, including Alistipes, Bulleidia, Butyrivibrio and Parabacteroides. Thirdly, we identified a large amount of phages in both healthy and diseased samples, which may play roles in shaping the dental microbiota. Another advantage of the shotgun metagenome sequencing approach was that it allowed us to screen for functional genes potentially associated with periodontal health and disease.
Similar to some bacterial communities in copiotrophic environments (e.g., mammalian intestines and coastal waters)31,32, functional variation of microbiomes between periodontal health and disease reveals that periodontal microbes can adapt to subgingival niches by increasing the potential for bacterial chemotaxis and flagellar assembly, which may facilitate microbial location, colonization and invasion of host tissues and thus causing periodontal inflammation. Moreover, compared to periodontal health samples, the over- or under-representation of certain pathways and genes in periodontitis samples makes them potentially prone to produce excessively intracellular toxins (e.g., LPS and PrtC). The reduced potential for decomposition (e.g., MetB/C and CAT) and transport capability (e.g., ABC transporters and PTS) may cause the accumulation of toxic substances (e.g., H2S2 and H2O2) in periodontal pockets and exacerbate tissue inflammation33,34.
In the present study, we observed a strong correlation between bacterial community structure and periodontal disease status and redefined the core disease-associated microbiota. A number of functional genes and metabolic pathways were found to be over-represented in the microbiomes in periodontal disease, which were mainly involved in bacterial chemotaxis, flagellar assembly and toxin biosynthesis. In addition, a fraction of novel phages and functional genes were identified in the assembly of metagenomic sequences. Our study enriches the understanding of the human oral microbiome and sheds light on the contribution of microorganisms in the formation and succession of human dental plaques and oral diseases. More samples are required to explore the heterogeneity of periodontal microbial communities within individuals.
Methods
Sample collection
The sample collection was approved by the Medical Ethical Committee of the Peking University and Beijing Institutes of Life Science, Chinese Academy of Sciences (Beijing, China). Participants were recruited at the Peking University School & Hospital of Stomatology and had given their informed consent. Participants were mature non-smoking females, 30 ~ 65 years of age, who were free of systemic diseases and other oral diseases except chronic periodontitis, without prosthetic dental appliances, had never received periodontal therapy and had not taken any antibiotics in the past three months. Periodontal health was defined as no probing depth or attachment loss >2 mm. Chronic periodontitis was defined as >4 sites with periodontal pockets ≥4 mm and attachment loss ≥6 mm12. Sampling was performed at least 6 h after tooth brushing and 2 h after eating.
Buccal and lingual dental surfaces were swabbed with cotton swabs per individual (Figure 1A). Dental plaques were collected using a sterile, inoculating-loop-like device to increase accumulation and to avoid gingival bleeding. In total, swabs were collected from 3 periodontally healthy, plaque-free subjects (Z11, Z14 and Z15) and 2 periodontal patients (H9-1 and H14-1). Dental plaques were collected from 6 periodontally healthy subjects (PY2, PY4, PY5, PZ2, PZ3 and PZ8) and 5 periodontal patients (H4, H6, H7, H9-2 and H14-2). The samples were categorized into 4 groups: Z, H-1, PZ and H-2. For each sample, the swab tips or plaques were placed in 1.5 mL microcentrifuge tubes and frozen at −80°C for further processing.
DNA extraction, contamination evaluation and metagenomic sequencing
Metagenomic DNA was individually extracted from swabs or plaques with the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany). The quantity and quality of isolated DNA was measured using a Nano Drop ND-1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) and agarose gel electrophoresis, respectively. Real-time quantitative polymerase chain reaction (qPCR) was employed for human DNA quantification and performed on an AB7300 Real-time PCR System (Applied Biosystems, Foster City, CA, USA) before sequencing. For each sample, 0.5 μg of purified metagenomic DNA was sheared into fragments of ~180 bp in length and a library was constructed according to a standard protocol provided by Illumina, Inc. (San Diego, CA, USA). Quantification was performed using a Qubit Fluorometer (Invitrogen, Life Technologies, Grand Island, NY, USA) and a Stratagene Mx3000P Real-time PCR Cycler (Agilent, Santa Clara, CA, USA) prior to cluster generation in a c-Bot automated sequencing system (Illumina, Inc.). Eight libraries with different indexes were pooled together and sequenced in one lane using an Illumina HiSeq 2000 high-throughput sequencing instrument with 2 × 100 bp paired-end (PE) sequencing. A total of two lanes were sequenced for the 16 libraries.
Read filtering and merging
Pre-analysis of sequencing data are described in Supplementary Fig. S1. First, low quality paired-end reads were removed before further analysis using Illumina CASAVA pipeline with default parameters. Secondly, all quality-filtered PE reads were aligned to the human genome assembly (hg19) using the Burrows-Wheeler Aligner (BWA) v0.5.9 algorithm35 to filter potential human contamination. Lastly, the remaining reads were merged into 140 ~ 194 bp sequences based on the overlap of PE reads. Briefly, we iteratively aligned a pair of reads (read1 and read2) with an overlap length ranging from 6 to 40 bp. In each iteration, the overlap score was calculated as the number of mismatches divided by the overlap length. If the score of the best overlap was smaller than the mismatch threshold (0.15), read1 and read2 were merged into a long read. Because Illumina sequencing tends to accumulate more errors at the 3′ end, the first half of the overlapped region was derived from read1 and the second half was derived from read2.
Metagenomic analysis
Identification of 16S rRNA sequences were made using the Ribosomal Database Project (RDP) classifier36 with a minimum score of 0.7 as the cutoff. Non-rRNA sequences were identified using the Basic Local Alignment Search Tool (BLASTX) algorithm and the National Center for Biotechnology Information (NCBI) non-redundant (NR) sequence database and the alignment results were further processed by the Metagenome Analyzer (MEGAN) program to statistically analyze the abundance of microorganisms in each sample37. MetaCV was used to classify unassembled reads into specific taxonomic and functional groups38. After normalizing the sequence counts of each taxon by the total number of reads, statistical analysis was performed on the bacterial composition and abundance at the phylum and genus levels. Two previously reported datasets of human oral microbiome, Metagenomics Rapid Annotation using Subsystem Technology (MG-RAST) server IDs 4446622.3 (a 454 Titanium and Illumina GAIIx sequenced metagenomic classification based on SEED)19 and 4444448.3 (an Illumina GAIIx sequenced metagenome classification based on RDP)20 were also used for phylum-level comparisons. Based on the genus-level classification, principal component analysis (PCA) was performed to evaluate the similarity among various metagenomic communities.
The merged long sequences were functionally annotated using the Kyoto Encyclopedia of Genes and Genomes (KEGG) bioinformatics database39. Functional categories and genes in the KEGG pathway were counted. The reads in pathways of LPS biosynthesis, bacterial chemotaxis and flagellar assembly, as well as PrtC genes were individually compared with the NR database and classified at the genus-level to determine their taxonomic origin. Complete genomic sequences of four Treponema species (T. denticola, T. vincentii, T. phagedenis and T. pallidum) were downloaded from the Integrated Microbial Genomes (IMG) database40 for comparative genomic analyses. Fimbriae (FimA), Lys-specific (Kgp) and Arg-specific (Rgp) cysteine proteinase gene sequences of Porphyromonas species (P. endodontalis, P. uenonis and P. gingivalis) were downloaded from GenBank as references to calculate the sequence frequency of each genotype.
Metagenome assembly and gene prediction
All non-human metagenomic PE reads were used to build a de novo assembly using the Short Oligonucleotide Analysis Package (SOAPdenovo) v1.0.5 assembly method41. The MetaGeneMark v 2.8 gene prediction tool42 was used to predict genes from the assembled contigs. The predicted proteins were compared against the NR database using BLASTP. Briefly, for each query protein, we set Sbest as the bitscore of the best BLAST hit. Then, we collected all the BLAST hits that have bitscore higher than Sbest * 0.95. Using the functional annotation of these collected BLAST hits, we applied a majority-rule consensus approach to determine the function of the query protein. For taxonomic annotation, we computed the lowest common ancestor (LCA) of all species in the collected BLAST hits and to determine its taxonomic origin.
References
Huttenhower, C. et al. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Methe, B. A. et al. A framework for human microbiome research. Nature 486, 215–221 (2012).
Dewhirst, F. E. et al. The human oral microbiome. J. Bacteriol. 192, 5002–5017 (2010).
Bik, E. M. et al. Bacterial diversity in the oral cavity of 10 healthy individuals. ISME J. 4, 962–974 (2010).
Belda-Ferre, P. et al. The oral metagenome in health and disease. ISME J. 6, 46–56 (2012).
Lamont, R. J. & Jenkinson, H. F. Oral Microbiology at a Glance. 1–85 (Wiley-Blackwell, 2010).
Zambon, J. J., Reynolds, H. S. & Slots, J. Black-pigmented Bacteroides spp. in the human oral cavity. Infect. Immun. 32, 198–203 (1981).
Slots, J. & Genco, R. J. Black-pigmented Bacteroides species, Capnocytophaga species and Actinobacillus actinomycetemcomitans in human periodontal disease: virulence factors in colonization, survival and tissue destruction. J. Dent. Res. 63, 412–421 (1984).
Kolenbrander, P. E., Andersen, R. N. & Moore, L. V. Coaggregation of Fusobacterium nucleatum, Selenomonas flueggei, Selenomonas infelix, Selenomonas noxia and Selenomonas sputigena with strains from 11 genera of oral bacteria. Infect. Immun. 57, 3194–3203 (1989).
Hill, G. B., Ayers, O. M. & Kohan, A. P. Characteristics and sites of infection of Eubacterium nodatum, Eubacterium timidum, Eubacterium brachy and other asaccharolytic eubacteria. J. Clin. Microbiol. 25, 1540–1545 (1987).
Kumar, P. S., Griffen, A. L., Moeschberger, M. L. & Leys, E. J. Identification of candidate periodontal pathogens and beneficial species by quantitative 16S clonal analysis. J. Clin. Microbiol. 43, 3944–3955 (2005).
Colombo, A. P. et al. Comparisons of subgingival microbial profiles of refractory periodontitis, severe periodontitis and periodontal health using the human oral microbe identification microarray. J. Periodontol. 80, 1421–1432 (2009).
Hajishengallis, G. et al. Low-abundance biofilm species orchestrates inflammatory periodontal disease through the commensal microbiota and complement. Cell Host Microbe. 10, 497–506 (2011).
Curtis, M. A., Zenobia, C. & Darveau, R. P. The relationship of the oral microbiotia to periodontal health and disease. Cell Host Microbe. 10, 302–306 (2011).
Wilson, M. & Henderson, B. Virulence factors of Actinobacillus actinomycetemcomitans relevant to the pathogenesis of inflammatory periodontal diseases. FEMS Microbiol. Rev. 17, 365–379 (1995).
Dorn, B. R., Leung, K. L. & Progulske-Fox, A. Invasion of human oral epithelial cells by Prevotella intermedia. Infect. Immun. 66, 6054–6057 (1998).
Dashper, S. G., Seers, C. A., Tan, K. H. & Reynolds, E. C. Virulence factors of the oral spirochete Treponema denticola. J. Dent. Res. 90, 691–703 (2011).
Stein, D. C., Miller, C. J., Bhoopalan, S. V. & Sommer, D. D. Sequence-based predictions of lipooligosaccharide diversity in the Neisseriaceae and their implication in pathogenicity. Plos One 6, e18923 (2011).
Xie, G. et al. Community and gene composition of a human dental plaque microbiota obtained by metagenomic sequencing. Mol. Oral Microbiol. 25, 391–405 (2010).
Lazarevic, V. et al. Metagenomic study of the oral microbiota by Illumina high-throughput sequencing. J. Microbiol. Methods 79, 266–271 (2009).
Yoshino, T., Laine, M. L., van Winkelhoff, A. J. & Dahlen, G. Genotype variation and capsular serotypes of Porphyromonas gingivalis from chronic periodontitis and periodontal abscesses. FEMS Microbiol. Lett. 270, 75–81 (2007).
Rylev, M. & Kilian, M. Prevalence and distribution of principal periodontal pathogens worldwide. J. Clin. Periodontol. 35, 346–361 (2008).
Grebe, T. W. & Stock, J. Bacterial chemotaxis: the five sensors of a bacterium. Curr. Biol. 8, R154–157 (1998).
Butler, S. M. & Camilli, A. Both chemotaxis and net motility greatly influence the infectivity of Vibrio cholerae. P. Natl. Acad. Sci. USA 101, 5018–5023 (2004).
Kato, T., Takahashi, N. & Kuramitsu, H. K. Sequence analysis and characterization of the Porphyromonas gingivalisPrtc gene, which expresses a novel collagenase activity. J. Bacteriol. 174, 3889–3895 (1992).
Ruiz, N., Kahne, D. & Silhavy, T. J. Transport of lipopolysaccharide across the cell envelope: the long road of discovery. Nat. Rev. Microbiol. 7, 677–683 (2009).
Qi, J., Zhao, F. Q., Buboltz, A. & Schuster, S. C. inGAP: an integrated next-generation genome analysis pipeline. Bioinformatics 26, 127–129 (2010).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Zaura, E., Keijser, B. J., Huse, S. M. & Crielaard, W. Defining the healthy "core microbiome" of oral microbial communities. BMC Microbiol. 9, 259 (2009).
Griffen, A. L. et al. Distinct and complex bacterial profiles in human periodontitis and health revealed by 16S pyrosequencing. ISME J. 6, 1176–1185 (2012).
Koch, A. L. The adaptive responses of Escherichia coli to a feast and famine existence. Adv. Microb. Physiol. 6, 147–217 (1971).
Lauro, F. M. et al. The genomic basis of trophic strategy in marine bacteria. P. Natl. Acad. Sci. USA 106, 15527–15533 (2009).
Wood, A. P. & Kelly, D. P. in Handbook of Hydrocarbon and Lipid Microbiology (ed Timmis, K. N.) 3167–31787 (Springer, 2010).
Grassi, F. et al.Oxidative stress causes bone loss in estrogen-deficient mice through enhanced bone marrow dendritic cell activation. P. Natl. Acad. Sci. USA 104, 15087–15092 (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Cole, J. R. et al. The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 37, D141–145 (2009).
Huson, D. H., Auch, A. F., Qi, J. & Schuster, S. C. MEGAN analysis of metagenomic data. Genome Res. 17, 377–386 (2007).
Liu, J. et al. Composition-based classification of short metagenomic sequences elucidates the landscapes of taxonomic and functional enrichment of microorganisms. Nucleic Acids Res. 41, e3 (2013).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36, D480–484 (2008).
Markowitz, V. M. et al. The integrated microbial genomes system: an expanding comparative analysis resource. Nucleic Acids Res. 38, D382–390 (2010).
Li, R. et al.De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010).
Acknowledgements
This study was supported by National Natural Science Foundation of China (NSFC 91131013) and CAS grant (0869011BJ5) to FZ, NSFC grant (31100094) to JQ and NSFC grant (30973317) to SW. We are grateful to Zhu BL (Institute of Microbiology, Chinese Academy of Sciences), Lu H and Mao FF (School and Hospital of Stomatology, Peking University) for their assistance in this study.
Author information
Authors and Affiliations
Contributions
J.W. conducted experiments, analyzed metagenomic data, conceived and wrote the main manuscript text. J.Q. provided algorithms, filtered and counted reads. H.Z. performed PCA analysis and prepared Figure 1. S.H. and Y.Z. recruited participants and collected samples. S.W. collected and selected samples for sequencing. F.Z. assembled sequences, analyzed metagenomic data and conceived and revised the manuscript text. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Wang, J., Qi, J., Zhao, H. et al. Metagenomic sequencing reveals microbiota and its functional potential associated with periodontal disease. Sci Rep 3, 1843 (2013). https://doi.org/10.1038/srep01843
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01843
This article is cited by
-
Influence of Environmental Factors on Salivary Microbiota and Their Metabolic Pathway: Next-Generation Sequencing Approach
Microbial Ecology (2023)
-
Functional metagenomic analysis of quorum sensing signaling in a nitrifying community
npj Biofilms and Microbiomes (2021)
-
Periodontitis associates with species-specific gene expression of the oral microbiota
npj Biofilms and Microbiomes (2021)
-
Microbiome composition comparison in oral and atherosclerotic plaque from patients with and without periodontitis
Odontology (2021)
-
Japanese subgingival microbiota in health vs disease and their roles in predicted functions associated with periodontitis
Odontology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.