Abstract
The protein-protein interaction (PPI) networks are dynamically organized as modules and are typically described by hub dichotomy: ‘party’ hubs act as intramodule hubs and are coexpressed with their partners, yet ‘date’ hubs act as coordinators among modules and are incoherently expressed with their partners. However, there remains skepticism about the existence of hub dichotomy. Since different algorithms and data sets were used in previous studies to test the model of hub classification, the conclusions may be largely influenced by the potential inherent biases. In this study, we evaluated two data sets of yeast interactome and systematically investigated the behavior of hubs from multiple perspectives including co-expression patterns, topological roles and functional classifications. Our results revealed consistency between the two data sets, confirming the presence of hub dichotomy. Furthermore, we analyzed a human interactome data set and demonstrated that the modular architecture of the PPI networks was more complicated than hub dichotomy.
Similar content being viewed by others
Introduction
The concept of ‘date’ and ‘party’ hubs has been widely accepted in the area of protein-protein interaction (PPI) networks. Han et al first classified hubs of the yeast interactome into two classes1, by integrating PPI network information with transcriptional profiling data. Party hubs interact with most of their partners simultaneously, while date hubs bind different partners at different locations and times. In theory, date hubs preferentially connect functional modules to each other, whereas party hubs preferentially act inside functional modules1,2. The two types of hubs also display profound differences regarding topological roles and evolutionary constraint, in that party hubs posited in single modules are highly constrained, whereas date hubs connecting different modules are more plastic1,2. However, Batada et al suggested that the organization of global protein interaction network is highly interconnected in the manner that is more like the continuous dense stratus clouds than the segregated altocumulus clouds and hence argued against the classification of ‘date’ and ‘party’ hubs3. A series of subsequent papers were then involved in this debate, but there is still no definitive conclusion on it as of today4,5,6,7,8. For example, Taylor et al extended the scope of two distinct hub types from yeast to human with the evidence of a multimodal distribution of hubs co-expression in human PPI network7. However, Agarwal et al argued that the feature of multimodal distribution was not robust according to methodological changes6. In view of three-dimensional protein structures, Kim et al supported the binary partition of hubs by explaining ‘date hubs’ as single-interface hubs and ‘party hubs’ as multi-interface hubs8, while Wang et al further suggested that the number of interaction interfaces are crucial in classification of functional and topological properties associated with each hub protein9.
We proposed that the results of the PPI network analysis may be largely influenced by the potential biases in data and analytical methods. First, there was lack of consistency in the data sets used in previous analysis. Each study used different criteria or prediction methods to derive the PPI information from different public data sources, such as yeast two-hybrid assay or protein mass spectrometry, so that the number of nodes and edges are quite different between studies. For example, the network used by Han et al contains 2,491 interactions among 1,375 proteins yet the data set of Batada et al contains 3,976 interactions among 1,291 proteins. Previous study also suggested that experimental bias might play a key role in the observed properties in a given data set. Thus, it is important to construct a complete picture of cellular PPI networks and the results can be improved significantly, when more data on high-quality binary interaction information becomes available10. Second, the definitions of hubs in the network were also not consistent among different studies; for example, Han et al defined hubs with degree greater than 5 and Batada et al defined hubs as nodes whose connectivity ranked within top 5% or 10% most connected nodes in the data set. These differences in definition may also be responsible for the discordances in different studies regarding the presence of date and party hubs. This problem was also addressed by a recent study which presented three objective methods to define hub proteins in PPI networks11. Third, the principle to distinct two classes of hubs was only based on the averaged Pearson correlation coefficient (avPCC) of expression levels between hub and its interacting partners, which are not robust and may be influenced by the topological structure of the PPI network. In the paper of Han et al, they found that the avPCC of hubs (degree greater than 5) followed a bimodal distribution but the no-hubs displayed a normal distribution. Thus, hubs were split into two types: one with relatively low avPCC (date hub) and the other with relatively high avPCC (party hub). They were further inferred to play different roles in the modulization process of PPI networks. Party hubs, which highly coexpressed with their neighbors, were intramodule hubs that coordinate proteins from the same functional module, yet date hubs, which provisionally coexpressed with their neighbors, were considered as higher-level intermodule hubs that perform different functions under different conditions2,7. However, the model described above was purely based on gene expression data and ignored the network topology or protein structure information. A recent study also attempted to improve the classification of date and party hubs by features analysis12.
Modularity and community structure are important features in real complex networks13,14. In biological networks, subnetworks and functional modules are associated with certain biological process15,16,17,18,19,20. Thus, when the functional modules were identified by network topology, we can locate the hubs in functional modules and use statistic measurements to determine the intramodule and intermodule hubs based on the proportion of connections within or outside of their own module. Then, the concept that party hubs (intramodule hubs) are highly coexpressed with their neighbors while date hubs (intermodule hubs) are conditionally and temporarily coexpressed with their neighbors can be directly examined, when we take into account the modularity of PPI networks.
In this work, we used a simulated annealing method to identify modules in the networks and assign different roles to each node based on its pattern of intramodule and intermodule connections16,21. We applied this approaches on two data sets of yeast from different studies, one of which supported the ‘date’ and ‘party’ hubs concept and the other did not4,5. Our results indicated that the modularity of interactome is far more complex than the dichotomy of hubs. However, we also depicted examples of intermodule and intramodule hubs which conformed to the norm based on both topology structure and gene co-expression, suggesting that the mechanism of ‘date’ and ‘party’ hubs still played an important role in the regulation of PPI networks. In addition to yeast data sets, we also revealed an analogous dynamic organization of human interactome, indicating the universality of regulatory mechanism across species.
Results
Comparison of the detected modules between two yeast data sets
We chose to use the updated version of data sets from both sides in the debate, including the filtered high-confidence (‘filtered-HC’) data set which supported the classification of ‘date’ and ‘party’ hubs4 and the updated high-confidence (‘updated-HC’) data set which did not5. Both data sets were generated on high-throughput yeast two-hybrid system with different filter thresholds and criteria for curation. The size of the interaction network represented a major difference between the two data sets; however, the average degree of the network, a global network property to measure the connectivity of the whole network, was quite similar (Table S1). A recent study also revealed a consistent degree correlation pattern in these two data sets and suggested that protein interaction network possessed an inherent dichotomy in degree correlation22. We next calculated the largest connected component (LCC) of the network and partitioned the LCC into modules. 23 and 14 modules were identified in filtered-HC and updated-HC data sets, respectively. Although the filtered-HC was divided into more modules with relatively smaller size compared with the updated-HC, we found a high degree of consistency of the detected modules from the two data sets. 18 of 23 modules identified in filtered-HC have corresponding modules from updated-HC with an overlap coefficient above 0.5, where the overlap coefficient is defined as:  . For example, the module 4 (227 proteins) in filtered-HC has an overlap of 128 proteins with the module 12 (203 proteins) in updated-HC (Overlap coefficient = 0.63, Table S2; Fisher's Exact Test, p-value < 2.2e–16, Figure S1). Gene functional enrichment analysis confirmed that both module 4 in filtered-HC and module 12 in updated-HC were highly enriched with genes involved in ‘ribosome biogenesis and assembly’ and ‘RNA metabolism’ (Table S3). The Venn diagram (Figure 1) further showed that most of the proteins associated with ‘ribosome biogenesis and assembly’ or ‘RNA metabolism’ from module 4 in filtered-HC and module 12 in updated-HC were present in the overlapping portion of the two modules. These results implicated that biologically meaningful modules can be identified based on the topological structure, despite the large differences between these PPI data sets.
. For example, the module 4 (227 proteins) in filtered-HC has an overlap of 128 proteins with the module 12 (203 proteins) in updated-HC (Overlap coefficient = 0.63, Table S2; Fisher's Exact Test, p-value < 2.2e–16, Figure S1). Gene functional enrichment analysis confirmed that both module 4 in filtered-HC and module 12 in updated-HC were highly enriched with genes involved in ‘ribosome biogenesis and assembly’ and ‘RNA metabolism’ (Table S3). The Venn diagram (Figure 1) further showed that most of the proteins associated with ‘ribosome biogenesis and assembly’ or ‘RNA metabolism’ from module 4 in filtered-HC and module 12 in updated-HC were present in the overlapping portion of the two modules. These results implicated that biologically meaningful modules can be identified based on the topological structure, despite the large differences between these PPI data sets.
Association between node roles and avPCC
Once modules in a PPI network were identified, the role of a node can be naturally determined by how the node was located in its own module and with respect to other modules. The avPCC level for nodes can be directly calculated as the average correlation coefficient of gene expression levels between a node and all its interacting partners under various conditions or in various tissues. Using the two properties termed as within-module degree and its participation coefficient, the nodes can be assigned into seven roles based on the intramodule and intermodule connectivity (See Methods). The relationship between role assignment and avPCC was plotted in Figure 2. Similar to avPCC, another measure called expression variance (EV) can be used to evaluate the dynamic expression level of each node, which is calculated as the quantile value of the variance of its expression profile among all nodes in the network. The EV is close to 0 if the gene expression is static (with the lowest variance), yet the EV value is 1 if the gene has the most dynamic expression pattern among all genes in the genome. A high correlation was found between the value of EV and the mRNA abundance of a gene. In addition, neighbors of dynamic proteins with high EV, but not static proteins with low EV, were highly coexpressed with each other. Static protein hubs were suggested to be excluded from the date hub, since they interacted with their neighbors continuously23.
Relationship between role assignment, avPCC and EV.
(A) Updated-HC plotted by role assignment and avPCC (B) Updated-HC plotted by role assignment and EV. (C) Filtered-HC plotted by role assignment and avPCC (D) Filtered-HC plotted by role assignment and EV. Role assignments: R1, Ultra-peripheral; R2, Peripheral; R3, Non-hub connector; R4, Non-hub kinless; R5, Provincial hub; R6, Connector hub; and R7, Kinless hub. There is no clear correlation between role assignments and avPCC or EV in the plot for the two data sets.
According to the definition, R6 (Connector hub) and R7 (Kinless hub) are more likely to be the intermodule hub (date hub, the percentage is 0.63% for FHC and 1.03% for UHC), yet R5 (Provincial hub) should be the intramodule hub (party hub, the percentage is 1.97 for FHC and 0.89% for UHC). Thus, the R6 and R7 hubs are expected to have low levels of avPCC whereas R5 hubs are expected to have high values of avPCC. Figure 2 showed highly consistent patterns between the filtered-HC and updated-HC data sets, that is, all the R6 nodes had low values of avPCC. A fraction of R5 hubs showed high level of avPCC, though most of them still had low level of avPCC. The small fraction of nodes with high avPCC made the overall avPCC of R5 higher than the other roles (Figure S2). In summary, the role assignment of hubs did not correspond well with the avPCC measure, indicating that a simple dichotomy was not sufficient to interpret the diversity of hubs. Additionally, no clear correlation can be found between the role assignments and EV from Figure 2, so the dynamic levels of gene expression also cannot distinguish the topological roles of hubs.
The functional modules with high avPCC
We then focused our attention on the nodes with high values of avPCC. Strikingly, most of the nodes with avPCC above 0.5 were from the module enriched in ‘ribosome biogenesis and assembly’ and ‘RNA metabolism’ (module 4 in filtered-HC and module 12 in updated-HC, both with a p-value < 2.2e–16 by Fisher's Exact Test). Additionally, we also observed a clear bimodal distribution of the avPCC in module 4 in filtered-HC and module 12 in updated-HC. In the paper of Han et al, a bimodal distribution of avPCC of hubs suggested a natural division of the date and party hubs with a threshold of avPCC at 0.5, though the author emphasized that the bimodality was not essential evidence of the party/date hub distinction in the original report1,4. Considering that hubs were defined by 5 degrees in the initial paper of Han et al and most of the nodes with high avPCC also showed a degree larger than 5 in this module, nodes of this particular module contributed largely to the formation of the bimodal distribution of hub co-expression.
We further illustrated the topological structure of module 4 of Filtered-HC in Figure 3. A large proportion of nodes highly coexpressed with each other form a closely connected cluster in this module. Most of them belong to the ‘ribosome biogenesis and assembly’ or ‘RNA metabolism’ pathways. In all the 227 nodes of this module, 95 nodes have an avPCC above 0.5, while 64 of 99 nodes related to ‘ribosome biogenesis and assembly’ or ‘RNA metabolism’ showed an avPCC above 0.5 (p-value = 1.378 e–4, Fisher's Exact Test). Party hubs (R5, Provincial hub) situated in the center of this cluster also showed high co-expression level with their neighbors. They therefore acted as a skeleton structure of module 4.
Topological structure of module 4 in Filtered-HC.
Node representation: R1 (Ultra-peripheral), lavender; R2 (Peripheral), blue; R3 (Non-hub connector), purple; R5 (Provincial hub), orange; R6 (Connector hub), coral. Party and date hubs provided by Bertin et al are represented by rectangles and triangles, respectively4. Edge representation: black dot lines represent interactions with negative correlation of expression; solid lines represent interaction with positive correlation of expression; edge colors correspond to the Pearson Correlation Coefficient values in the legend. A small closely connected group of non-coexpressed nodes were marked with dashed oval.
In the paper of Bertin et al, the authors provided a list of updated date and party hubs based on the filtered-HC dataset. The Provincial hubs (R5) in our study were all defined as party hubs in Bertin et al. In addition, some non-hub nodes (R1, R2) in our study were also considered as party hubs by Bertin et al. In terms of biological significance, these nodes can be considered as party hubs since they also played an important role in the formation of the co-expression cluster in module 4. In contrast, there was also a small closely connected cluster with low co-expression level in Figure 3. Most of the nodes in the small cluster were classified as date hubs by Bertin et al because of the high degree and low avPCC. Intuitively, the observation is opposite to the biological interpretation of date hubs which serve as coordinators between functional modules, implying that hubs with low avPCC did not simply equate with the intermodule hubs or date hubs.
The dynamic modularity of the functional modules
Except for module 4 described above, other modules from Filtered-HC did not show a peak of avPCC distribution above 0.5 (Figure S3). However, the correlations may be impaired by integrating data sets with different conditions. When computing the correlation in certain conditions, genes in some modules became highly coexpressed. So the dynamic modularity was inferred by comparing the co-expression of modules across different conditions. The enriched functions of modules were also consistent with the conditions at which the modules reveal high co-expression. For example, module 9 was enriched with ‘cell cycle’ and ‘pseudohyphal growth’ (a pattern of cell growth occurs in conditions of nitrogen limitation) proteins and it displayed higher co-expression in the sporulation process (Figure 4), whereas module 7 was enriched with ‘protein biosynthesis and catabolism’ proteins and also showed an increasing trend of co-expression at the condition of environmental changes or DNA damage24,25,26. Module 9 contained the gene CDC28, which has the highest degree, with 202 connections in the whole network. Notably, CDC28 was listed as a date hub by Bertin et al and was also predicted as an intermodule hub (R6, connector hub) by topological structure. CDC28 was known as the catalytic subunit of the main cell cycle cyclin-dependent kinase (CDK), which can alternately associate with G1 cyclins (CLNs) and G2/M cyclins (CLBs) directing the CDK to specific substrates. Thus, this gene functions as a global regulator in yeast. As described in Figure 4, most of the genes around CDC28 were highly coexpressed with it when the sporulation process was initiated, suggesting that CDC28 played a crucial role in regulation. Additionally, CDC28 was also connected with PRE1, RPN3 and RPT1 in module 7. PRE1 was also classified as R5 (Provincial hub). In the terms of topological structure, module 7 also contained a closely connected region which included PRE1, RPN3 and RPT1. In contrast to the condition of sporulation, most of the connected partners with CDC28 showed very low correlation in the condition of DNA damage; however, the correlations of CDC28 with PRE1, RPN3 and RPT1 increased significantly: The ranks of the correlations with PRE1, RPN3, RPT1 among the 202 neighbors of CDC28 were increased from 71, 76, 78 to 12, 2, 6, respectively. The correlation between CDC28 and RPN3 was further predicted as differential co-expression with a p-value of 0.042 by the method of Cho et al27. The nodes posited in closely connected region of module 7 also displayed high co-expression accordingly under the condition of DNA damage (Figure 5). In summary, CDC28 was crucial in the regulation of sporulation process; on the other hand, CDC28 also played important function in the condition of DNA damage. Thus, CDC28 conformed to the biological interpretation of date hub and served as an intermodule connector, which was a dynamic participant in different modules.
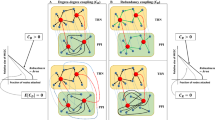
Topological structure and the co-expression change of module 9 in Filtered-HC.
(A) The co-expression pattern of module 9 and the corresponding avPCC distribution under the condition of sporulation. (B) The co-expression pattern of module 9 and the corresponding avPCC distribution under the condition in the whole compedium. Node and edge representation follows the convention in Figure 3. Party and date hubs provided by Bertin et al are expressed as rectangles and triangles respectively4. The expression correlations between connected partners in module 9 are much higher in the condition of sporulation, as most of the edges are red. In addition, the distribution of avPCC also shows that a group of nodes are highly coexpressed with their neighbors in the condition of sporulation (a peak of avPCC distribution above 0.5). Therefore, genes in module 9 are highly coexpressed in the condition of sporulation in comparison to the whole compendium.
Topological structure and the co-expression of module 7 and 9 in Filtered-HC.
(A) In the condition of DNA damage, the expression correlations between CDC28 and PRE1, RPN3, RPT1 are increased. (B) In the condition of sporulation, the correlation between CDC28 and PRE1, RPN3, RPT1 are low. Node and edge representation follows the convention in Figure 3. Party and date hubs provided by Bertin et al are expressed as rectangles and triangles respectively4.
Evolutionary constraint on proteins with high avPCC
A previous study calculated dN/dS ratio of the two hub types provided by Han et al and suggested that ‘date’ and ‘party’ hubs were under different evolutionary constraints: hubs with higher avPCC were more conserved2. However, the conclusion was still under debate since statistical bias may exist in the previous classification of the hubs5.
Our results showed a negative correlation between the dN/dS value and the avPCC in the proteins of the whole networks (Pearson correlation coefficient = −0.23, p-value < 2.2 e–16). Proteins with high avPCC (above 0.5) also showed significant lower dN/dS ratio (mean is 0.04) than the other proteins of the network whose average value of dN/dS ratio is 0.07 (two sample t-test, p-value < 1.4 e–14). Since most of the proteins with high avPCC were from module 4 as previous described, module 4 were also under a strong evolutionary constraint (two sample t-test, p-value = 8.14 e–09). Members of the most enriched biological functions in module 4, ‘ribosome biogenesis and assembly’ and ‘RNA metabolism’, were also highly constrained by purifying selection compared to members of another relatively enriched function in module 4, ‘DNA metabolism’ (Figure S4 and Figure S5). Based on the above results, we concluded that members in the co-expression modules, usually enriched in specific biological functions, were more conserved in PPI networks.
Analysis of the human interactome
The evidence of date and party hub distinctions were also elucidated in the human PPI network. Akin to the bimodal distribution of avPCC in yeast interactome, a multimodal distribution of avPCC was previously discovered in the human interactome7. However, the robustness of the evidence was questioned when changing the normalization methods for gene expression or when comparing across different interaction data sets6. We therefore applied the same strategy on the human interactome derived from HPRD (Human Protein Reference Database) to examine the existence of the binary hub classification28. We identified 16 modules with more than 20 members from the HPRD data set. These modules were also enriched in specific classification of gene ontology or pathways, suggesting the biological significance of the network partition method that solely used the topological properties (Table S4). We further assigned the roles to each node and calculated the avPCC of each node. As expected, no association was observed between different types of hubs and their avPCC as in the yeast data sets (Figure S6). We then investigated the distribution of avPCC in each module from human interactome. Unlike the yeast data sets, the distribution of avPCC in each module did not show bimodality as was observed in the module 4 of Filtered-HC. Instead, they displayed a single peak with modes ranging from 0 to 0.5. Some modules also revealed high co-expression, such as the module 15 which was enriched in function of ‘RNA transport’ (Figure S7), so hubs in module 15 were typical ‘party hubs’. The function of ‘RNA transport’ were not enriched in modules identified from yeast because the molecular machinery for RNA transport is more complex, involving much more proteins in metazoans than yeast29. As described before, genes in module enriched with ‘ribosome biogenesis and assembly’ were highly coexpressed in yeast interactome. We found a similar functional annotation termed as ‘ribonucleoprotein complex’ in human which was the second most enriched function in module 9. The genes relevant to ‘ribonucleoprotein complex’ in module 9 yielded a high co-expression correlation compared to other genes in the same module (Two Sample t-test, p-value < 5.1 e–4).
Since the avPCC described above were computed by using the entire expression compendium of different tissues, genes in a collection of more similar tissues should have higher co-expression level. We next used only the brain tissues from the human gene expression data to calculate avPCC and validated this hypothesis, especially in modules enriched in nervous system related functions (Figure S7). In tumor genesis and progression, phenotypic alternations were associated with rewiring of signaling pathways and networks30. We recalculated the avPCC from an expression data set collected from different types of cancers. The values of avPCC in each module were significantly decreased, suggesting that the normal regulation mechanisms which yielded the dynamic modularity in human interactome were disrupted in tumors tissues (Figure S7).
Discussion
In this work, we focused on the dispute about the ‘date’ and ‘party’ hub dichotomy by analyzing the roles of hubs and their dynamic modularity in PPI networks. Since the previous inference on the existence of hub dichotomy were based on indirect evidence such as the existence of a bimodal distribution of hub avPCC or the different changes of network topology by removing hubs, the established concept has been under debate for years1,3,4,5,6,7. Our analysis suggested that modularity of interactome is far more complex than the dichotomy of hubs.
We introduced a novel method to partition PPI network into several functional modules and assign roles to nodes according to the topological structure. From this perspective, we proved strong consistency between the modules identified by two different yeast interactome data sets, Filtered-HC and Updated-HC, which was previously used to draw opposite conclusion about the binary classification of hubs4,5. We further detected a module with strong co-expression which was enriched in ‘ribosome biogenesis and assembly’ and ‘RNA metabolism’. Molecular evolutionary analysis also showed that this module was highly conserved in evolution. Ribosome biogenesis is an energy intensive and complicated process to make ribosomes. Due to the importance of the process in cell growth, both the RNA and protein moieties of ribosomes and the ribosome biogenesis machinery are highly conservative from yeast to humans31,32,33. Hubs of this module satisfied the criteria of ‘party hubs’ very well1. These hubs participate in the same biological process and connect together to build up the frame of the functional module. In addition, previous evidence to support that ‘date’ and ‘party’ hubs produced distinct co-expression pattern and evolutionary rate may not be reliable, if considerable proportion of hubs were selected from this module with high conservation and high co-expression.
We also showed the dynamic modularity in PPI networks by studying the co-expression pattern of nodes in different conditions. Many modules displayed high co-expression under specific biological functions in which they were enriched. Hubs from these modules were also ‘party hubs’ according to the schematic diagram from Han et al1. In summary, ‘party hubs’ widely existed in protein interactome, accompanied by the occurrence of dynamic modularity.
In contrast to ‘party hubs’ displaying relatively high avPCC, hubs with low avPCC were far more complicated and should not be simplified defined as ‘date hubs’. Based on the module detection method, hubs were assigned as R5, R6 or R7. Among the hubs, only a fraction of R5 nodes showed high level of avPCC, yet all the others represented low-avPCC hubs (Figure 2). Therefore, the low-avPCC hubs can be R5 (intra-module hubs) or R6/R7 (inter-module hubs) in the view of topological properties. We also found low-avPCC hubs which do not show high co-expression with its neighbors or a strong evidence of participation as coordinators among modules. For example, our structural analysis of network described that members of a small closely connected cluster were all predicted as ‘date hubs’ by Bertin et al due to their low co-expression (Figure 3). Obviously, these hubs with low avPCC were not coordinators between functional modules. Similar instances widely exist in the whole proteome networks, which is why the binary classification of hubs has been argued continuously. Thus, the measure of avPCC alone was insufficient to infer them as higher-level coordinators which performed varying functions and were active at different times or under different conditions. However, we also pinpointed an intermodule hub (CDC28) which participated in a global organization of biological modules. CDC28 was annotated as a global regulator associated with G1 cyclins (CLNs) and G2/M cyclins (CLBs) alternatively to regulate the CDK to specific substrates. CDC28 also displayed low avPCC and was correctly predicted as ‘date hubs’ by Bertin et al. Therefore, hubs with low avPCC may serve diverse roles in the protein interactome, suggesting the existence of complex mechanism to modulate the protein network architecture and cell behavior.
We further expanded the analysis of dynamic modularity in protein networks from yeast to human. We demonstrated a similar dynamic organization of human interactome as yeast interactome, indicating the universality of regulatory mechanism across species. By comparing to the co-expression pattern derived from cancer tissues, we confirmed the importance of modular structure in human PPI network, since the gene co-expression of functional modules was altered in tumor tissues.
By comprehensively investigating the roles of hubs from multiple angles, we revealed that ‘party hubs’ were biologically meaningful and consistent with the role assignment of hubs from topological structures. Moreover, we confirmed the existence of ‘date hubs’ and expounded the complexity of low-avPCC hubs. Our results enhanced current understanding of the organizational principles in interactome addressing the importance of integrating multiple approaches in illustrating the biological roles of hubs. First, using the mRNA expression profiles, we can estimate temporal characteristics of hubs and their partners in the interactome networks based on static graphics. Since the PPI were static due to the experimental techniques, gene co-expression information can provide dynamic view of the interactome. Second, the mathematical methods using graph theory can exactly identify modules of highly connected nodes and the universal roles of nodes in the network, giving a comprehensive understanding of the network topology. Third, functional annotation of genes can help validate the biological significance of detected functional modules. Since the identified modules and defined node roles varied as the parameters of module detection algorithm changed, the reliability of the detected modules should be verified from an independent perspective such as gene functional enrichment analysis. We noticed that a previous study also tried to elucidate the hub dichotomy by global role assignment from topological structure, but it did not utilize the information of modules and hastily denied the concept of hub dichotomy6.
Given the rapid growth of the protein 3D structural information, future work would focus on constructing a structure-based protein-protein interaction network. A further systematic survey of the association among gene co-expression pattern, protein interaction interface and topological roles should facilitate our understanding of global organization of the proteome and provide insights to the dynamic modularity in concordance with the evolution of protein structure and interaction.
Methods
Gene expression data sets
The gene expression data on yeast were collected from 10 data sets24,25,26,34,35,36,37,38,39. The data sets were also used in previously papers debating hub dichotomy1,3,4,5. The human gene expression data were available for a panel of 79 human tissues from a previous study targeting 44775 human transcripts40. The transcripts were identified by Affymatrix (Santa Clara, CA) HG-U133A array (22,130 transcripts) and GNF1H custom array (22,645 transcripts). The gene expression data of human primary tumors were collected from a large-scale RNA profiling of 185 carcinomas including prostate, breast, lung, ovary, colorectum, kidney, liver, pancreas, bladder/ureter and gastroesophagus. The profiling experiment was performed on Affymetrix (Santa Clara, CA) U95a GeneChip, covering 12626 human transcripts41.
Protein interaction data sets
The high-confidence (‘filtered-HC’) data set containing 2,561 nodes and 5,992 edges was curated by Bertin et al to confirm the existence of ‘date’ and ‘party’ hubs4. The updated high-confidence (‘updated-HC’) data set including 4,011 nodes and 10,055 edges was released by batada et al , who did not find evidence supporting that network hubs fall into discrete classes5. Both data sets were obtained on yeast interactome.
For the human interactome, we used Human Protein Reference Database which contains scientific information of human proteins on the basis of manual curation on published literature and bioinformatics analyses of the protein sequence28.
Calculation of the average Pearson correlation coefficient
For calculation of the avPCC, Pearson correlation coefficients of gene pairs were first calculated using the expression data sets. The correlations between a specific gene ‘A’ and its connected partners from protein interaction data sets were then extracted. The avPCC of gene ‘A’ is defined as the mean of the extracted correlation values.
Module detection
The modularity M for a given partition of a network into modules is  , where NM is the number of modules, L is the number of connections in the network, ls is the number of connections between nodes in the module s and ds is the sum of the degrees of the nodes in module s. The definition of modularity is based on the notion that separating a network into modules must contain more within-module links and less possible between-module links42. We used a simulated annealing algorithm to find the optimal partition of network with the largest modularity. Details are described in the original article of the algorithm16,21.
, where NM is the number of modules, L is the number of connections in the network, ls is the number of connections between nodes in the module s and ds is the sum of the degrees of the nodes in module s. The definition of modularity is based on the notion that separating a network into modules must contain more within-module links and less possible between-module links42. We used a simulated annealing algorithm to find the optimal partition of network with the largest modularity. Details are described in the original article of the algorithm16,21.
Node role definition
The definition of the node role is based on its within-module degree and its participation coefficient16,21. The within-module degree z-score measures the connectivity of a given node to its own module and is defined as

where ki is the number of links of node i to other nodes within its own module,  is the average of k for all nodes in module Si and σSi is the standard deviation of k in module Si.
is the average of k for all nodes in module Si and σSi is the standard deviation of k in module Si.
The participation coefficient quantifies to the distribution of the links of a node among the different modules. It defined as

Where N is the number of modules, kis is the number of links of node i to other nodes in the same module S and ki is the total degree of node i. The participation coefficient Pi is close to 1 if its links are uniformly distributed among all the modules and 0 if all its links are within its own module.
At first, the nodes are classified as hubs and non-hubs according to the within-module degree (hubs: z ≥ 2.5, non-hubs: z < 2.5). Based on the participation coefficient, nodes are further subdivided into: (R1) ultra-peripheral nodes, considered as nodes with all their links within their own module (P ≤ 0.05); (R2) peripheral nodes, considered as nodes with most links within their module (0.05 < P ≤ 0.62); (R3) satellite connectors, nodes with a high fraction of their links to other modules (0.62 < P ≤ 0.80) and (R4) kinless nodes, nodes with links homogeneously distributed among all modules (P > 0.80). Hubs are divided into: (R5) provincial hubs, considered as hubs with the vast majority of links within their module (P ≤ 0.30); (R6) connector hubs, considered as hubs with many links to most of the other modules (0.30 < P ≤ 0.75) and (R7) global hubs, considered as hubs with links homogeneously distributed among all modules (P > 0.75). The threshold for within-module performs well to separate nodes with participation coefficient above 0.3, but for nodes with participation coefficient below 0.3, the role assignment for R1, R2 or R5 with a within-module degree very close to 2.5 needs to be improved.
Gene functional enrichment analysis
Gene functional enrichment analysis on yeast was based on COG functional categories43. For human, We used DAVID (http://david.abcc.ncifcrf.gov/) to test enrichment in gene sets with GO, SwissProt and InterPro terms compared with the background list of all genes44.
Evolutionary rate
The evolutionary rate (dN/dS) was estimated by a method providing adjustment of dS to correct the selection on synonymous mutations. A detailed description about the method was available in the published paper of Hirsh et al45.
References
Han, J. D. et al. Evidence for dynamically organized modularity in the yeast protein-protein interaction network. Nature 430, 88–93 (2004).
Fraser, H. B. Modularity and evolutionary constraint on proteins. Nat Genet 37, 351–352 (2005).
Batada, N. N. et al. Stratus not altocumulus: a new view of the yeast protein interaction network. PLoS Biol 4, e317 (2006).
Bertin, N. et al. Confirmation of organized modularity in the yeast interactome. PLoS Biol 5, e153 (2007).
Batada, N. N. et al. Still stratus not altocumulus: further evidence against the date/party hub distinction. PLoS Biol 5, e154 (2007).
Agarwal, S., Deane, C. M., Porter, M. A. & Jones, N. S. Revisiting date and party hubs: novel approaches to role assignment in protein interaction networks. PLoS Comput Biol 6, e1000817 (2010).
Taylor, I. W. et al. Dynamic modularity in protein interaction networks predicts breast cancer outcome. Nat Biotechnol 27, 199–204 (2009).
Kim, P. M., Lu, L. J., Xia, Y. & Gerstein, M. B. Relating three-dimensional structures to protein networks provides evolutionary insights. Science 314, 1938–1941 (2006).
Wang, H. & Zheng, H. Correlation of genomic features with dynamic modularity in the yeast interactome: a view from the structural perspective. IEEE Trans Nanobioscience 11, 244–250 (2012).
Yu, H. et al. High-quality binary protein interaction map of the yeast interactome network. Science 322, 104–110 (2008).
Vallabhajosyula, R. R., Chakravarti, D., Lutfeali, S., Ray, A. & Raval, A. Identifying hubs in protein interaction networks. PLoS One 4, e5344 (2009).
Mirzarezaee, M., Araabi, B. N. & Sadeghi, M. Features analysis for identification of date and party hubs in protein interaction network of Saccharomyces Cerevisiae. BMC Syst Biol 4, 172 (2010).
Newman, M. E. Modularity and community structure in networks. Proc Natl Acad Sci U S A 103, 8577–8582 (2006).
Girvan, M. & Newman, M. E. Community structure in social and biological networks. Proc Natl Acad Sci U S A 99, 7821–7826 (2002).
Yook, S. H., Oltvai, Z. N. & Barabasi, A. L. Functional and topological characterization of protein interaction networks. Proteomics 4, 928–942 (2004).
Guimera, R. & Nunes Amaral, L. A. Functional cartography of complex metabolic networks. Nature 433, 895–900 (2005).
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nat Genet 34, 166–176 (2003).
Chang, X., Liu, S., Yu, Y. T., Li, Y. X. & Li, Y. Y. Identifying modules of coexpressed transcript units and their organization of Saccharopolyspora erythraea from time series gene expression profiles. PLoS One 5, e12126 (2010).
Dong, J. & Horvath, S. Understanding network concepts in modules. BMC Syst Biol 1, 24 (2007).
Chang, X., Wang, Z., Hao, P., Li, Y. Y. & Li, Y. X. Exploring mitochondrial evolution and metabolism organization principles by comparative analysis of metabolic networks. Genomics 95, 339–344 (2010).
Guimera, R. & Amaral, L. A. Cartography of complex networks: modules and universal roles. J Stat Mech 2005, nihpa35573 (2005).
Hao, D. & Li, C. The dichotomy in degree correlation of biological networks. PLoS One 6, e28322 (2011).
Komurov, K. & White, M. Revealing static and dynamic modular architecture of the eukaryotic protein interaction network. Mol Syst Biol 3, 110 (2007).
Gasch, A. P. et al. Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11, 4241–4257 (2000).
Chu, S. et al. The transcriptional program of sporulation in budding yeast. Science 282, 699–705 (1998).
Gasch, A. P. et al. Genomic expression responses to DNA-damaging agents and the regulatory role of the yeast ATR homolog Mec1p. Mol Biol Cell 12, 2987–3003 (2001).
Cho, S. B., Kim, J. & Kim, J. H. Identifying set-wise differential co-expression in gene expression microarray data. BMC Bioinformatics 10, 109 (2009).
Keshava Prasad, T. S. et al. Human Protein Reference Database–2009 update. Nucleic Acids Res 37, D767–772 (2009).
Heym, R. G. & Niessing, D. Principles of mRNA transport in yeast. Cell Mol Life Sci 69, 1843–1853 (2012).
Barabasi, A. L. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nat Rev Genet 5, 101–113 (2004).
Freed, E. F., Bleichert, F., Dutca, L. M. & Baserga, S. J. When ribosomes go bad: diseases of ribosome biogenesis. Mol Biosyst 6, 481–493 (2010).
Granneman, S. & Baserga, S. J. Ribosome biogenesis: of knobs and RNA processing. Exp Cell Res 296, 43–50 (2004).
Henras, A. K. et al. The post-transcriptional steps of eukaryotic ribosome biogenesis. Cell Mol Life Sci 65, 2334–2359 (2008).
Mnaimneh, S. et al. Exploration of essential gene functions via titratable promoter alleles. Cell 118, 31–44 (2004).
Roberts, C. J. et al. Signaling and circuitry of multiple MAPK pathways revealed by a matrix of global gene expression profiles. Science 287, 873–880 (2000).
Roberts, G. G. & Hudson, A. P. Transcriptome profiling of Saccharomyces cerevisiae during a transition from fermentative to glycerol-based respiratory growth reveals extensive metabolic and structural remodeling. Mol Genet Genomics 276, 170–186 (2006).
Spellman, P. T. et al. Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol Biol Cell 9, 3273–3297 (1998).
Travers, K. J. et al. Functional and genomic analyses reveal an essential coordination between the unfolded protein response and ER-associated degradation. Cell 101, 249–258 (2000).
Yoshimoto, H. et al. Genome-wide analysis of gene expression regulated by the calcineurin/Crz1p signaling pathway in Saccharomyces cerevisiae. J Biol Chem 277, 31079–31088 (2002).
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc Natl Acad Sci U S A 101, 6062–6067 (2004).
Su, A. I. et al. Molecular classification of human carcinomas by use of gene expression signatures. Cancer Res 61, 7388–7393 (2001).
Newman, M. E. & Girvan, M. Finding and evaluating community structure in networks. Phys Rev E Stat Nonlin Soft Matter Phys 69, 026113 (2004).
Tatusov, R. L. et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 4, 41 (2003).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4, 44–57 (2009).
Hirsh, A. E., Fraser, H. B. & Wall, D. P. Adjusting for selection on synonymous sites in estimates of evolutionary distance. Mol Biol Evol 22, 174–177 (2005).
Acknowledgements
We thank members of the Wang lab for helpful comments and suggestions on the analytical strategies. The study is supported by start-up funds from the Zilkha Neurogenetic Institute and NIH grant number HG006465 from NIH/NHGRI (X.C., K.W.).
Author information
Authors and Affiliations
Contributions
X.C., K.W. conceived the project and wrote the manuscript. X.C., T.X., Y.L. analyzed the data. All authors discussed the results and gave approval to the final version of the manuscript. X.C., T.X. contributed equally to this work.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figures and Tables
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Chang, X., Xu, T., Li, Y. et al. Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of ‘date’ and ‘party’ hubs. Sci Rep 3, 1691 (2013). https://doi.org/10.1038/srep01691
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01691
This article is cited by
-
Protein-protein interaction (PPI) network analysis reveals important hub proteins and sub-network modules for root development in rice (Oryza sativa)
Journal of Genetic Engineering and Biotechnology (2023)
-
The interaction between LC8 and LCA5 reveals a novel oligomerization function of LC8 in the ciliary-centrosome system
Scientific Reports (2022)
-
Integrated network-based multiple computational analyses for identification of co-expressed candidate genes associated with neurological manifestations of COVID-19
Scientific Reports (2022)
-
Gene co-expression network analysis identifies trait-related modules in Arabidopsis thaliana
Planta (2019)
-
Rewiring of the inferred protein interactome during blood development studied with the tool PPICompare
BMC Systems Biology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.