Abstract
The identification of a novel β coronavirus, nCoV, as the causative agent of severe respiratory illness in humans originating in Saudi Arabia, Qatar and Jordan has raised concerns about the possibility of a coronavirus pandemic similar to that of SARS-CoV. As a definitive treatment regimen has never been thoroughly evaluated for coronavirus infections, there is an urgent need to rapidly identify potential therapeutics to address future cases of nCoV. To determine an intervention strategy, the effect of interferon-α2b and ribavirin on nCoV isolate hCoV-EMC/2012 replication in Vero and LLC-MK2 cells was evaluated. hCoV-EMC/2012 was sensitive to both interferon-α2b and ribavirin alone in Vero and LLC-MK2 cells, but only at relatively high concentrations; however, when combined, lower concentrations of interferon-α2b and ribavirin achieved comparable endpoints. Thus, a combination of interferon-α2b and ribavirin, which are already commonly used in the clinic, may be useful for patient management in the event of future nCoV infections.
Similar content being viewed by others
Introduction
Anovel β coronavirus (nCoV), has been identified as the etiological agent of 17 confirmed cases (11 deaths) of a severe respiratory illness with occasional renal failure from patients in Saudi Arabia1, Qatar2, Jordan and the United Kingdom3,4. Complete genome sequencing1,5 determined that this new virus is closely related to two Asian bat betacoronaviruses (HKU4 and HKU5)6 in lineage C. This makes nCoV the first lineage C betacoronavirus known to infect humans5. While human-to-human transmission is assumed to be less extensive as compared to SARS-CoV, three of the cases in Saudi Arabia were within one family and several healthcare workers who cared for two of the cases in Jordan have been classified as probable cases5. Moreover, in the most recent cluster of cases7, two of the three cases did not have a history of travel to the Middle East, but are suspected to have resulted from human-to-human transmission within the UK from a family member with a travel history to Saudi Arabia and Pakistan. This would suggest that human-to-human transmission can occur in close contact settings. Presumably, the sporadic nature of the apparently unlinked index cases in three different, albeit geographically proximal countries and the close relationship to Asian bat coronaviruses, suggests that the source of nCoV is zoonotic8.
Despite limited information on this new virus, it has been determined that in contrast to SARS-CoV, which uses angiotensin-converting enzyme 2 (ACE2) to gain entry into cells9,10, nCoV uses dipeptidyl peptidase 4 (DPP4 or CD26) as a functional receptor11. This finding may be important as the requirement for ACE2 was thought to be partially responsible for the pathogenicity of SARS-CoV, while also serving as one of the factors that may have limited spread from human-to-human. As the pathogenesis of nCoV could be significantly different from previously studied coronaviruses, the ability to predict whether this virus is likely to result in a larger epidemic or even pandemic, such as occurred with SARS-CoV, is unknown.
The rapid identification of therapeutics is a high priority as there is currently no specific therapy or vaccine for nCoV and the resulting disease has been severe with a high case-fatality rate. The clinical experience from SARS suggests that a number of interventions including ribavirin with and without corticosteroids12,13,14, ribavirin with protease inhibitors15,16 and interferon (alfacon-1) with corticosteroids17 may improve outcome, but a definitive treatment regimen was not clearly established18. Here we address the effectiveness in vitro of two antiviral drugs, interferon-α2b (IFN-α2b) and ribavirin, in an attempt to identify a therapeutic approach that can be immediately utilized in the clinic to benefit future cases.
Results
To determine the potential antiviral effect of IFN-α2b and ribavirin on nCoV isolate hCoV-EMC/2012 replication, susceptible cells were infected with hCoV-EMC/2012. Following 1 h absorption, infected cells were treated with either IFN-α2b or ribavirin. In Vero cells, IFN-α2b reduced the cytopathogenic effect (CPE) starting at a concentration of 250 U/ml and completely eliminated CPE at 1000 U/ml and above (Figure 1A). Ribavirin reduced CPE starting at a concentration of 100 μg/ml and completely eliminated CPE at 200 μg/ml and above (Figure 1A). Viral protein levels, as measured by nucleocapsid protein expression in cell lysates, were also reduced in the presence of increasing levels of IFN-α2b, with a reduction starting at 250 U/ml (Figure 1B). In response to ribavirin treatment, a reduction in nucleocapsid protein expression was observed at 50 μg/ml, but did not appear to be dose dependent (Figure 1B).
Interferon-α2b and/or ribavirin treatment of hCoV-EMC/2012-infected Vero cells.
Vero cells were infected with hCoV-EMC/2012 at an MOI of 0.001 for 1 h and subsequently treated with interferon-α2b (IFN-α2b) and/or ribavirin at the indicated concentration. On day 5 post-infection cells were photographed and cytopathic effect was assessed (A). Cell lysates were collected and subjected to western blotting with serum from a rabbit immunized with whole inactivated hCoV-EMC/2012 (B). β-actin was used as loading control (actin).
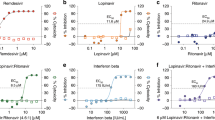
Supernatants were collected on days 1, 3 and 5 post-infection and subsequent analyses of viral loads (viral RNA) and titers were performed. Peak viral loads and infectious virus were recovered from day 3 samples; therefore, this time point was used for subsequent analysis. Supernatants collected on day 5 frequently had lower viral loads and titers than samples collected on day 3, likely due to extensive CPE. As such, day 5 samples were not included in the analyses. A dose dependent reduction in genome copies was observed for IFN-α2b treatment with a 0.53-log reduction in viral loads at 500 U/ml reaching a 1.84-log reduction at 5000 U/ml (Figure 2A). A dose dependent reduction in viral loads was also observed for ribavirin treatment, with a 0.82-log reduction at 200 μg/ml reaching a 2.04-log reduction at 2000 μg/ml (Figure 2B). Importantly, a corresponding decrease in infectious virus was also observed as a result of IFN-α2b or ribavirin treatment. A 0.57-log reduction in virus titer occurred at 500 U/ml IFN-α2b, increasing to a 1.31-log reduction at 5000 U/ml IFN-α2b (Figure 2C). For ribavirin, a 1.24-log reduction in virus titer was observed at 100 μg/ml, reaching a 4.05-log reduction at 2000 μg/ml (Figure 2D).
Replication of novel human coronavirus hCoV-EMC/2012 in response to interferon-α2b or ribavirin treatment in Vero cells.
Vero cells were infected with hCoV-EMC/2012 at an MOI of 0.001 for 1 h and subsequently treated with interferon-α2b (IFN-α2b) or ribavirin at the indicated concentration. At 1 and 3 days post-infection, supernatants were removed and subsequently analyzed for viral load by real time quantitative RT-PCR (A,B) and infectious virus titers by 50% tissue culture infectious dose (TCID50) assay (C, D). Viral loads are shown as TCID50 equivalents/ml ±SD, in response to increasing concentrations of IFN-α2b (A) or ribavirin (B). Viral titers are TCID50/ml ±SD in response to increasing concentrations of IFN-α2b (C) or ribavirin (D).
The 50% inhibitory concentration (IC50) of IFN-α2b and ribavirin was subsequently determined to be 58.08 U/ml and 41.45 μg/ml, respectively (Table 1). The IC90 (1-log reduction) and IC99 (2-log reduction) values were also calculated (Table 1). While this is a significant finding, the concentrations of IFN-α2b or ribavirin required to effectively inhibit hCoV-EMC/2012 replication are quite high and may therefore be of limited clinical application.
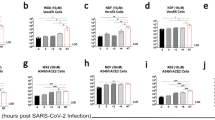
Vero cells have been described as comparatively resistant to ribavirin, as they are inefficient at converting ribavirin into its mono- and tri-phosphate forms19. Therefore, we also assessed the sensitivity of hCoV-EMC/2012 to IFN-α2b and ribavirin in LLC-MK2 cells (Figure 3A, B). Based on IC values, LLC-MK2 cells were more responsive to both IFN-α2b and ribavirin treatment (Table 1). IFN-α2b, at the maximum concentration tested (2000 U/ml), reduced infectious titers by 3.97-log (2.01-log reduction in genome copies). Ribavirin treatment, at 200 μg/ml or higher, reduced infectious virus below the detection threshold of 13.7 TCID50/ml.
Replication of novel human coronavirus hCoV-EMC/2012 in response to interferon-α2b or ribavirin treatment in LLC-MK2 cells.
LLC-MK2 cells were infected with hCoV-EMC/2012 at an MOI of 0.001 for 1 h and subsequently treated with interferon-α2b (IFN-α2b) or ribavirin at the indicated concentration. At 1 and 3 days post-infection, supernatants were removed and subsequently analyzed for viral load by real time quantitative RT-PCR (A,B) and infectious virus titers by 50% tissue culture infectious dose (TCID50) assay (C, D). Viral loads are shown as TCID50 equivalents/ml ±SD, in response to increasing concentrations of IFN-α2b (A) or ribavirin (B). Viral titers are TCID50/ml ±SD in response to increasing concentrations of IFN-α2b (C) or ribavirin (D).
Given their long history of combined use for treatment of hepatitis C20,21, we combined IFN-α2b and ribavirin treatment to determine whether one compound would augment the activity of the other. Combination treatment in Vero and LLC-MK2 cells lowered the threshold at which a decrease in CPE was noted. For Vero cells, this was reduced to 62 U/ml IFN-α2b and 12 μg/ml ribavirin with the absence of CPE at and above 125 U/ml IFN-α2b and 25 μg/ml ribavirin (Figure 1A). This represents an 8- and 16-fold decrease in the amount of IFN-α2b and ribavirin, respectively, which is required to achieve the same reduction as either treatment alone. Viral nucleocapsid protein expression was also reduced in a dose dependent manner starting at concentrations of IFN-α2b and ribavirin of 250 U/ml and 50 μg/ml, respectively (Figure 1B). The reduction in CPE and nucleocapsid protein expression also correlated with reduced virus genome copies and titers. When IFN-α2b was administered with ribavirin at 5:1 ratio, there was an additional reduction in the virus titer by 0.4- to 2.16-logs over that of IFN-α2b treatment alone (Figure 4).
Replication of novel human coronavirus hCoV-EMC/2012 in response to combined treatment with interferon-α2b and ribavirin in Vero cells.
Vero cells were infected with hCoV-EMC/2012 at an MOI of 0.001 for 1 h and subsequently treated with interferon-α2b (IFN-α2b) and/or ribavirin at the indicated concentration. At 3 days post-infection, supernatants were removed and subsequently analyzed for infectious virus titers by 50% tissue culture infectious dose (TCID50) assay. Viral titers are shown as TCID50/ml ±SE in response to increasing concentrations of IFN-α2b, ribavirin or the combination of both.
Discussion
Ongoing identification of cases of nCoV3,22 suggests continuing introduction of the virus to humans in the Middle East from an unknown source. Given the genetic relationship of hCoV-EMC/2012 to other bat coronaviruses5, one can speculate that bats may be the reservoir of this virus; however, additional host species should be considered. With documented human-to-human transmission in close contact situations and the first documented mild case22, there is a real concern that we could be observing the ‘tip of the iceberg’ and perhaps the start of an epidemic. Regardless, with a 65% case-fatality rate despite intensive medical intervention, therapeutic strategies are urgently needed. Despite the significant increase in research on coronaviruses since the discovery of SARS-CoV in 2003, there is no definitive antiviral or therapeutic treatment for coronavirus infections in humans. Pegylated interferon-α was shown to be an effective prophylactic treatment against infection with SARS-CoV in cynomolgus macaques, but was less effective when administered post exposure23. No other therapeutics have been tested for antiviral activity against SARS-CoV in a higher order animal model. In the SARS-CoV mouse model, poly IC:LC24 and mDEF201 (an adenovirus expressing mouse IFN-α)25 can protect mice from lethal disease; however, neither of these approaches yields an immediate therapeutic for use in humans. Poly IC:LC has been tested in numerous clinical trials, but is not currently approved for treatment of any human disease. Adenovirus-based therapy has multiple complicating factors, such as pre-existing immunity, that have not been adequately addressed, nor is it approved for use in humans26.
Here we identified a potential therapeutic approach against hCoV-EMC/2012 combining IFN-α2b and ribavirin. Either treatment alone reduced virus replication by at least 1-log or as much as 4-logs in susceptible cell lines. Moreover, when combined, efficacy was reached at lower concentrations. Thus, this combination may provide a benefit as a treatment in humans. Vero cells display a high level of resistance to the activity of ribavirin19,27. Thus, we also performed the same assay in LLC-MK2 cells, where sensitivity to ribavirin was observed at a much lower concentration.
Previous in vitro studies have demonstrated that SARS-CoV is sensitive to ribavirin28 and to various classes of interferon (α, β and γ)27,29,30,31,32,33,34. The sensitivity of SARS-CoV to ribavirin appears to be cell line dependent, with concentrations as low as 50 μg/ml ribavirin being reported as effective16. Unfortunately, this concentration is higher than the peak serum concentration reached in humans of approximately 24 μg/ml35. IFN-α2b was previously reported to inhibit growth of SARS-CoV starting at 1000 U/ml with a 1-log reduction at 2000 U/ml34. Following infection, only IFN-β (EC50 560 IU/ml) has shown a dose dependent antiviral effect36. In this study we report a nearly 4-log reduction in virus titers for hCoV-EMC/2012 at comparable doses.
During the outbreak of SARS-CoV, different combinations of therapeutic interventions were attempted; however, none were implemented in a manner that allowed a critical assessment of their effectiveness. The most frequently administered therapeutics were broad-spectrum antibiotics, glucocorticoids and ribavirin37,38,39. The lack of a standard dosing regimen for ribavirin makes comparisons difficult18; however, low dose ribavirin (400–600 mg/day) therapy was shown to be ineffective likely due to an insufficient plasma concentration40. In contrast, when used at higher doses other studies have found that ribavirin alone reduced viral loads in over half of the patients and when combined with the viral protease inhibitors lopinavir/ritonavir, patients had a lower incidence of adverse outcomes16. Despite being used in a large number of patients, it still remains unclear whether ribavirin alone was effective against SARS-CoV41. Alfacon-1, a synthetic IFN-α, has also been suggested to be beneficial to patients17. Unfortunately, all of these studies suffer from the confounding use of corticosteroids in doses that vary among studies making a definitive treatment elusive. It has been suggested that combination of interferon and ribavirin treatment should be evaluated18. While ribavirin can result in reversible hemolytic anemia, this complication typically occurs following longer treatment protocols35,42. This suggests that short-course ribavirin therapy for an acute infection such as nCoV may not be a significant complication as mild anemia was the most frequently reported side effect during ribavirin treatment for Lassa virus infection43.
A synergistic effect of IFN-α and ribavirin has been previously reported in vitro for both SARS-CoV27,44 and feline infectious peritonitis virus45; however, we observed an additive effect against hCoV-EMC/2012 in this study. The levels of IFNα-2b and ribavirin required for inhibition of nCoV replication must be achievable in humans in order to be relevant for clinical use. In humans, an interferon concentration of 100-750 IU/ml has been observed after intravenous injection of up to 3 × 107 U46,47, while 24 μg/ml of ribavirin is achievable following a 1000 mg intravenous dose35. Here IFN-α2b and ribavirin alone were shown to have an antiviral effect against hCoV-EMC/2012; however, in Vero cells the concentrations required to achieve a beneficial effect are likely higher than what is achievable in humans. When combined, the inhibitory concentration of both IFNα-2b and ribavirin drops to ranges that are likely achievable in humans, suggesting that the combination is a potential treatment option. Used early in the course of infection or given prophylactically to close contacts of sick individuals (close contact transmission has been documented in infection chains) this combination may improve clinical outcomes. In addition, reduced viral load would also likely translate to reduced virus shedding; thus, reducing the risk of secondary transmission. As these two drugs are currently used together in the clinic, combination therapy including IFN-α2b and ribavirin should be considered for case patient management of new nCoV cases and possibly for prophylaxis in highly exposed individuals.
Methods
Biosafety statement
All infectious work with hCoV-EMC/2012 was performed in a high containment facility at the Rocky Mountain Laboratories (RML), Division of Intramural Research (DIR), National Institute of Allergy and Infectious Diseases (NIAID), National Institutes of Health (NIH). The work was approved by the RML Institutional Biosafety Committee (IBC) at biosafety level 3 (BSL3).
Virus and cells
Human betacoronavirus EMC (hCoV-EMC/2012) was kindly provided by Erasmus Medical Center (Rotterdam, Netherlands). Vero (African green monkey kidney) and LLC-MK2 (rhesus monkey kidney) were maintained at 37°C in 5% CO2 in Dulbecco's modified Eagle's media (DMEM) supplemented with 10% fetal bovine serum (FBS), 50 U/ml penicillin and 50 μg/ml of streptomycin. HCoV-EMC was subsequently propagated on Vero cells using DMEM as above with 2% FBS (complete DMEM).
Antiviral assays
Confluent Vero and LLC-MK2 cells in 24-well culture plates (Costar, Corning, NY ) were infected in triplicate with hCoV-EMC/2012 diluted in complete DMEM at an MOI = 0.001. Following 1 h adsorption at 37°C, the inoculum was removed and the cells were washed 3 times with DMEM. Subsequently, complete DMEM containing IFN-α2b (0–5000 U/ml) (PBL Interferon Source, Piscataway, NJ) and/or ribavirin (0–2000 μg/ml) (MP Biomedicals, Solon, OH) was added to the cells. Cells were incubated for 24 h at 37°C, 5% CO2 in a humidified environment and the supernatant was removed, an aliquot was inactivated with AVL (Qiagen, Germantown, MD) for viral load quantification and the remainder was stored at −80°C for subsequent virus titration. The supernatant was replaced with fresh complete DMEM containing IFN-α2b and/or ribavirin. Supernatant was also collected at 72 h and 120 h. Five days post-infection representative wells were photographed to document cytopathic effect (CPE) and cells were subsequently collected for protein analysis in 4× SDS-PAGE loading buffer.
Genome quantification
RNA from AVL-treated supernatant was extracted with the NucleoSpin 96 Virus Core kit (Macherey-Nagel, Bethlehem, PA) on a Corbett X-tractor Gene (Valencia, CA). Quantitative real time RT-PCR using primers and probe previously described48 was performed on the RotorGene Q (Qiagen). A 10-fold dilution series of viral RNA based on TCID50 equivalents was used as a standard.
Western blot
Cell lysates were run on 10% SDS-PAGE gels and transferred to PVDF (GE Healthcare, Piscataway, NJ). Membranes were blocked with 5% non-fat milk, 0.05% Tween20 in PBS and subsequently probed with polyclonal serum diluted in blocking buffer at 1/10,000 from rabbit A691/A741 immunized with inactivated HCoV-EMC. Anti-rabbit IgG conjugated to horseradish peroxidase (KPL, Gaithersburg, MD) was used as a secondary at a 1/10,000 dilution. Western blots were developed with the Pierce ECL Plus kit (Thermo, Rockford, IL).
Infectivity assay (TCID50)
Confluent Vero cells were infected in triplicate with 10-fold dilutions of supernatants obtained from the antiviral assay. Virus was allowed to adsorb for 1 h and was then removed and replaced with complete DMEM. Cells were incubated at 37°C, 5% CO2 in a humidified environment for 5 days and then CPE was scored and TCID50 (50% tissue culture infectious dose) calculated as described by Reed and Muench49.
Data analysis
Data from the genome quantification and TCID50 assays was analyzed in Prism (GraphPad Software) and CompuSyn (combosyn.com).
References
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. & Fouchier, R. A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. The New England journal of medicine 367, 1814–1820 (2012).
Buchholz, U. et al. Contact investigation of a case of human novel coronavirus infection treated in a German hospital, October-November 2012. Euro surveillance: bulletin europeen sur les maladies transmissibles = European communicable disease bulletin 18, pii = 20406 (2013).
World Health Organization. . Novel coronavirus infection - update. (2013).<http://www.who.int/csr/don/2013_02_21/en/index.html> [Accessed March 4, 2013].
Wise, J. Patient dies from novel coronavirus in UK. BMJ 346, f1133 (2013).
van Boheemen, S. et al. Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. mBio 3, e00473–12, 10.1128/mBio.00473-12 (2012).
Woo, P. C. et al. Comparative analysis of twelve genomes of three novel group 2c and group 2d coronaviruses reveals unique group and subgroup features. Journal of virology 81, 1574–1585 (2007).
Evidence of person-to-person transmission within a family cluster of novel coronavirus infections, United Kingdom, February 2013. Euro surveillance: bulletin europeen sur les maladies transmissibles = European communicable disease bulletin 18, pii = 20427 (2013).
Holmes, K. V. & Dominguez, S. R. The new age of virus discovery: genomic analysis of a novel human betacoronavirus isolated from a fatal case of pneumonia. mBio 4, e00548–12, 10.1128/mBio.00548-12 (2013).
Muller, M. A. et al. Human Coronavirus EMC Does Not Require the SARS-Coronavirus Receptor and Maintains Broad Replicative Capability in Mammalian Cell Lines. mBio 3, e00515–12, 10.1128/mBio.00515-12 (2012).
Li, W. et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 426, 450–454 (2003).
Raj, V. S. et al. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 495, 251–254 (2013).
Peiris, J. S. et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: a prospective study. Lancet 361, 1767–1772 (2003).
Sung, J. J. et al. Severe acute respiratory syndrome: report of treatment and outcome after a major outbreak. Thorax 59, 414–420 (2004).
Booth, C. M. et al. Clinical features and short-term outcomes of 144 patients with SARS in the greater Toronto area. JAMA: the journal of the American Medical Association 289, 2801–2809 (2003).
Chan, K. S. et al. Treatment of severe acute respiratory syndrome with lopinavir/ritonavir: a multicentre retrospective matched cohort study. Hong Kong medical journal = Xianggang yi xue za zhi/Hong Kong Academy of Medicine 9, 399–406 (2003).
Chu, C. M. et al. Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax 59, 252–256 (2004).
Loutfy, M. R. et al. Interferon alfacon-1 plus corticosteroids in severe acute respiratory syndrome: a preliminary study. JAMA: the journal of the American Medical Association 290, 3222–3228 (2003).
Wong, S. S. & Yuen, K. Y. The management of coronavirus infections with particular reference to SARS. The Journal of antimicrobial chemotherapy 62, 437–441 (2008).
Shah, N. R., Sunderland, A. & Grdzelishvili, V. Z. Cell type mediated resistance of vesicular stomatitis virus and Sendai virus to ribavirin. PloS one 5, e11265, 10.1371/journal.pone.0011265 (2010).
Pearlman, B. L. Protease inhibitors for the treatment of chronic hepatitis C genotype-1 infection: the new standard of care. The Lancet infectious diseases 12, 717–728 (2012).
Paeshuyse, J., Dallmeier, K. & Neyts, J. Ribavirin for the treatment of chronic hepatitis C virus infection: a review of the proposed mechanisms of action. Current opinion in virology 1, 590–598 (2011).
Wise, J. Two more cases of novel coronavirus are confirmed in UK. BMJ 346, f1030 (2013).
Haagmans, B. L. et al. Pegylated interferon-alpha protects type 1 pneumocytes against SARS coronavirus infection in macaques. Nature medicine 10, 290–293 (2004).
Kumaki, Y. et al. Induction of interferon-gamma-inducible protein 10 by SARS-CoV infection, interferon alfacon 1 and interferon inducer in human bronchial epithelial Calu-3 cells and BALB/c mice. Antiviral chemistry & chemotherapy 20, 169–177 (2010).
Kumaki, Y. et al. Single-dose intranasal administration with mDEF201 (adenovirus vectored mouse interferon-alpha) confers protection from mortality in a lethal SARS-CoV BALB/c mouse model. Antiviral research 89, 75–82 (2011).
Gowen, B. B. et al. Extended protection against phlebovirus infection conferred by recombinant adenovirus expressing consensus interferon (DEF201). Antimicrobial agents and chemotherapy 56, 4168–4174 (2012).
Morgenstern, B., Michaelis, M., Baer, P. C., Doerr, H. W. & Cinatl, J., Jr Ribavirin and interferon-beta synergistically inhibit SARS-associated coronavirus replication in animal and human cell lines. Biochemical and biophysical research communications 326, 905–908 (2005).
Saijo, M. et al. Inhibitory effect of mizoribine and ribavirin on the replication of severe acute respiratory syndrome (SARS)-associated coronavirus. Antiviral research 66, 159–163 (2005).
Kumaki, Y. et al. Interferon alfacon 1 inhibits SARS-CoV infection in human bronchial epithelial Calu-3 cells. Biochemical and biophysical research communications 371, 110–113 (2008).
Sainz, B. Jr., Mossel, E. C., Peters, C. J. & Garry, R. F. Interferon-beta and interferon-gamma synergistically inhibit the replication of severe acute respiratory syndrome-associated coronavirus (SARS-CoV). Virology 329, 11–17 (2004).
Zheng, B. et al. Potent inhibition of SARS-associated coronavirus (SCOV) infection and replication by type I interferons (IFN-alpha/beta) but not by type II interferon (IFN-gamma). Journal of interferon & cytokine research: the official journal of the International Society for Interferon and Cytokine Research 24, 388–390 (2004).
Paragas, J., Blatt, L. M., Hartmann, C., Huggins, J. W. & Endy, T. P. Interferon alfacon1 is an inhibitor of SARS-corona virus in cell-based models. Antiviral research 66, 99–102 (2005).
Hensley, L. E. et al. Interferon-beta 1a and SARS coronavirus replication. Emerging infectious diseases 10, 317–319 (2004).
Stroher, U. et al. Severe acute respiratory syndrome-related coronavirus is inhibited by interferon- alpha. The Journal of infectious diseases 189, 1164–1167 (2004).
Koren, G., King, S., Knowles, S. & Phillips, E. Ribavirin in the treatment of SARS: A new trick for an old drug? CMAJ: Canadian Medical Association journal = journal de l'Association medicale canadienne 168, 1289–1292 (2003).
Cinatl, J. et al. Treatment of SARS with human interferons. Lancet 362, 293–294 (2003).
Lee, N. et al. A major outbreak of severe acute respiratory syndrome in Hong Kong. The New England journal of medicine 348, 1986–1994 (2003).
Poutanen, S. M. et al. Identification of severe acute respiratory syndrome in Canada. The New England journal of medicine 348, 1995–2005 (2003).
Tsang, K. W. et al. A cluster of cases of severe acute respiratory syndrome in Hong Kong. The New England journal of medicine 348, 1977–1985 (2003).
Tsang, K. & Seto, W. H. Severe acute respiratory syndrome: scientific and anecdotal evidence for drug treatment. Current opinion in investigational drugs 5, 179–185 (2004).
Groneberg, D. A. et al. Treatment and vaccines for severe acute respiratory syndrome. The Lancet infectious diseases 5, 147–155 (2005).
Brochot, E. et al. Ribavirin monitoring in chronic hepatitis C therapy: anaemia versus efficacy. Antiviral therapy 15, 687–695 (2010).
Bausch, D. G., Hadi, C. M., Khan, S. H. & Lertora, J. J. Review of the literature and proposed guidelines for the use of oral ribavirin as postexposure prophylaxis for Lassa fever. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America 51, 1435–1441 (2010).
Chen, F. et al. In vitro susceptibility of 10 clinical isolates of SARS coronavirus to selected antiviral compounds. Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology 31, 69–75 (2004).
Weiss, R. C. & Oostrom-Ram, T. Inhibitory effects of ribavirin alone or combined with human alpha interferon on feline infectious peritonitis virus replication in vitro. Veterinary microbiology 20, 255–265 (1989).
Wills, R. J. Clinical pharmacokinetics of interferons. Clinical pharmacokinetics 19, 390–399 (1990).
Hausfater, P., Cacoub, P., Assogba, U., Lebon, P. & Piette, J. C. Plasma exchange and interferon-alpha pharmacokinetics in patients with hepatitis C virus-associated systemic vasculitis. Nephron 91, 627–630 (2002).
Corman, V. M. et al. Detection of a novel human coronavirus by real-time reverse-transcription polymerase chain reaction. Euro surveillance: bulletin europeen sur les maladies transmissibles = European communicable disease bulletin 17, pii = 20285 (2012).
Reed, L. J. & Muench, H. A. A simple method for estimating fifty percent endpoints. American journal of hygiene 27, 493–497 (1938).
Acknowledgements
This work was supported by the Intramural Research Program of the National Institute of Allergy and Infectious Diseases (NIAID), National Institutes of Health (NIH). The authors would like to thank Drs. Bart Haagmans and Ron Fouchier, Erasmus Medical Center, Rotterdam, The Netherlands for providing HCoV-EMC. Anita Mora (Visual Arts, NIAID, NIH) assisted with editing the figures.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: D.F., H.F. Performed the experiments: D.F., E.d.W., C.M., J.C. Analyzed the data: D.F., E.d.W., V.J.M., H.F. Contributed essential reagents: V.J.M. Wrote the manuscript: D.F., E.d.W., V.J.M., H.F. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Falzarano, D., de Wit, E., Martellaro, C. et al. Inhibition of novel β coronavirus replication by a combination of interferon-α2b and ribavirin. Sci Rep 3, 1686 (2013). https://doi.org/10.1038/srep01686
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01686
This article is cited by
-
Targeting liquid–liquid phase separation of SARS-CoV-2 nucleocapsid protein promotes innate antiviral immunity by elevating MAVS activity
Nature Cell Biology (2021)
-
Poliovirus-nonsusceptible Vero cell line for the World Health Organization global action plan
Scientific Reports (2021)
-
COVID-19 in early 2021: current status and looking forward
Signal Transduction and Targeted Therapy (2021)
-
Can Zn Be a Critical Element in COVID-19 Treatment?
Biological Trace Element Research (2021)
-
Ribavirin Aerosol in the Treatment of SARS-CoV-2: A Case Series
Infectious Diseases and Therapy (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.