Abstract
Large expansions of a non-coding GGGGCC-repeat in the first intron of the C9orf72 gene are a common cause of both amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). G-rich sequences have a propensity for forming highly stable quadruplex structures in both RNA and DNA termed G-quadruplexes. G-quadruplexes have been shown to be involved in a range of processes including telomere stability and RNA transcription, splicing, translation and transport. Here we show using NMR and CD spectroscopy that the C9orf72 hexanucleotide expansion can form a stable G-quadruplex, which has profound implications for disease mechanism in ALS and FTD.
Similar content being viewed by others
Introduction
Amyotrophic lateral sclerosis (ALS) is a devastating neurodegenerative disorder in which the loss of motor neurons in brain and spinal cord causes progressive weakness and paralysis, ultimately leading to death from respiratory failure1. Frontotemporal dementia (FTD) is one of the most common forms of young-onset dementia and is characterized by the progressive degeneration of the frontal and anterior temporal lobes, leading to changes in personality or language impairment2,3. ALS and FTD share numerous similarities at the genetic and neuropathological level, clinically they can co-occur and they have been proposed to be part of the same spectrum of disease4.
Large expansions of the non-coding GGGGCC-repeat in the first intron of the C9orf72 gene have been recently demonstrated to cause ALS and FTD5,6. Whilst the unaffected normal control population carries <30 repeats of this hexanucleotide, approximately 8–10% of FTD and ALS European patients carry very large expansions which have been reported to range between 700 and 1,600 repeats5. The introns containing these large expansions are transcribed and indeed in patients the GGGGCC-repeat expansion is detectable by in situ hybridization in nuclear RNA foci5.
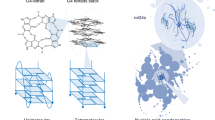
Guanine (G)-quadruplexes are highly stable nucleic acid secondary structures formed from short tracts of G-rich sequence associating together. These can occur in both DNA and RNA and consist of stacks of planar layers of G-tetrad units, named G-quartets in which the G bases are arranged in a square cyclic pattern held together by eight hydrogen bonds ( Fig. 1a ). When stacked the tetrads form a central cavity formed from guanine O6 atoms, where they can interact with metal cations. Monovalent metal cations bound in this central channel significantly affect the stability (K+ > Na+ > Li+) and topology of the folded G-quadruplex structure7.
Schematic representation of the parallel stranded GGGGCC RNA G-quadruplex.
(a) Tetrads are stabilized by eight hydrogen bonds (black lines), with metal ions sitting centrally (orange circle) and phosphate backbones laterally (blue squares). (b) Four stacked G-tetrads, anti-glycosidic torsion angles, with phosphate backbones connected through a propeller loop arrangement, comprised of the two cytosines, ensures a parallel topology.
The presence of RNA G-quadruplexes has been demonstrated in vitro and in vivo and they have been found in the transcripts of diverse organisms, ranging from viruses to humans8,9. Computational algorithms used to predict G-quadruplex forming sequences suggest that there are approximately 197,000 G-quadruplex forming sequences in the human genome9. Interestingly their presence is particularly enriched in the regulatory 5′ UTR, first intron and 3′ UTR regions of transcripts10,11. It has been shown that RNA G-quadruplexes have been recognized and bound by specific proteins and implicated in a wide range of biological processes including alternative splicing, RNA transport, translation regulation, RNA degradation and telomere stability12,13,14.
The GGGGCC expansion of C9orf72 forms runs of adjacent G-repeats and raises the possibility these could form G-quadruplexes. The secondary structure of the hexanucleotide repeat is very likely to be involved in determining the proteins it interacts with. These interactions may play a critical role in disease – as has been shown in Myotonic Dystrophy, where similar non-coding RNA repeat expansions form nuclear foci which sequester key RNA-binding proteins thereby causing functional defects15.
Thus, given the importance of expanded non-coding repeats in other RNA repeat expansion diseases15, we have investigated the secondary structure of the C9orf72 hexanucleotide repeat by biophysical methods to determine whether it forms RNA G-quadruplexes.
Results
In silico analysis predicts C9orf72 GGGGCC-repeats form G-quadruplexes
The GGGGCC-expansion lies in the 5′ region of C9orf72 intron 1 ( Fig. 2b ). Control individuals are reported to have <30 repeats, but the majority of controls have 2 repeats5,6,16,17. Nonetheless, using the EuQuad database18, which identifies G-quadruplex forming sequences in eukaryotic genomes, the three GGGGCC-repeats and adjacent GGGGC nucleotides present in the human reference C9orf72 gene sequence, are recognized as a G-quadruplex forming sequence. We therefore termed this sequence the minimal C9orf72 G-quadruplex repeat unit (C9Gru). The prediction is supported by the G-quadruplex analysis tool QGRS Mapper9: the full C9orf72 genomic sequence containing the C9Gru sequence shows one predicted four stacked G-quadruplex with a high predictive score ( Fig. 2a ). When inputting a pathogenic sequence containing 800 hexamer (GGGGCC) repeats the number of predicted 4 stacked G-quadruplexes increases accordingly ( Fig. 2b ).
The C9orf72 hexanucleotide repeat is predicted to form a G-quadruplex structure.
The G-quadruplex prediction tool GQRS mapper was used to identify potential G-quadruplex forming sequences in the entire C9orf72 genomic DNA sequence, which is shown with exons highlighted in red and the GGGGCC repeats in green. GQRS mapper provides a G-score (plotted in blue) which indicates the likelihood of G-quadruplex formation. (a) The highest G-score in the C9orf72 reference sequence from Ensembl GRCh37 corresponds to the three GGGGCC repeats and adjacent GGGGC which we have termed the minimal C9orf72 G-quadruplex repeat unit (C9Gru). (b) 800 GGGGCC repeats were added, which is representative of the disease-causing expansions, leading to an extended region of high G-score. Image is adapted from QGRS mapper (http://bioinformatics.ramapo.edu/QGRS/index.php). E1a = exon 1a; E1b = exon 1b.
GGGGCC-repeats form RNA G-quadruplexes
1D 1H NMR is a key technique in providing unequivocal experimental evidence of G-tetrad and quadruplex formation. We used NMR to investigate the structure of an RNA oligonucleotide consisting of the C9Gru sequence. NMR analysis in 10 mM potassium phosphate buffer, pH 7.0, prior to refolding in the presence of quadruplex stabilizing cations, revealed the presence of imino proton peaks between 12 to 14 ppm, a region of the spectrum characteristic of Watson-Crick (W-C) hydrogen bonding ( Supplementary Fig. S1 ). Raising the potassium chloride concentration to 40 mM and annealing the RNA allowed the C9Gru oligonucleotide to be refolded in the presence of stabilizing cations. At room temperature the 1D 1H NMR spectrum showed the presence of a broad envelope of peaks between 10 and 11.5 ppm and several distinct peaks characteristic of G-tetrad formation ( Fig. 3 ). The appearance of peaks consistent with tetrad formations coincides with the loss of W-C peaks between 12 to 13 ppm. The buffered C9Gru RNA oligonucleotide was then heated, stepwise (+5 K) from 273 K to 333 K, but without a significant change observed in the spectra, indicating the stability of the folded quadruplex and retention of the G-tetrads. As clear and distinct peaks could not be differentiated within the imino region, further structural analysis was not pursued.
NMR analysis of the C9orf72 GGGGCC RNA hexanucleotide repeat shows formation of G-quadruplexes.
The 1D proton spectrum of the C9Gru RNA oligonucleotide annealed in 10 mM K2PO4 40 mM KCl buffer, pH 7.0, 298 K. Peaks in the imino proton region (arrow) between 10 and 11.5 ppm correspond to quadruplex formation.
GGGGCC G-quadruplexes are highly stable and parallel oriented
G-quadruplexes can adopt three possible topologies according to the directions of the strands, these can be parallel, anti-parallel or mixed. Riboguanosine has a strong propensity to adopt the ‘anti’ conformation and therefore to give rise to parallel G-quadruplex structures. Circular dichroism (CD) spectroscopy is a standard method used to analyse structural features of G-quadruplexes19,20. We used this technique to characterize the structural conformation of the C9Gru RNA sequence in a 40 mM KCl buffer. This showed a positive peak at 262 nm and a negative peak at 237 nm ( Fig. 4a ), which are the hallmarks of a parallel-oriented G-quadruplex structure19.
CD analysis shows the GGGGCC RNA G-quadruplex structures are very stable, cation dependent and parallel oriented.
CD spectra of the C9Gru RNA oligonucleotide (4.6 μm) show a positive peak at 262 nm and a negative peak at 237 nm, which is characteristic of parallel-oriented G-quadruplex structures. (a) and (b) represent the temperature unfold and refold spectra respectively, with the RNA oligonucleotide in KCl 40 mM, K2PO4 10 mM buffer. The peak at 262 nm only decreases at 85°C indicating a very stable structure. Temperature unfold (c) and refold (d) spectra of identical RNA in LiCl 50 mM, Na2PO4 10 mM buffer. The characteristic cation dependence of G-quadruplex structures was confirmed by the observed reduced stability in the presence of Li+ ions.
To study the stability of the G-quadruplex motif found in the C9Gru G-quadruplex structure, we performed melting experiments by increasing the temperature from 15°C to 95°C using a 1°C/min gradient. The CD spectra were measured with 10°C intervals and showed a strong overlay from 15°C to 75°C with a reduction of the 262 nm peak only starting at 85°C, indicating a strong stability of the formed structure ( Fig. 4a ). Folding was fully reversible, but lagged behind the unfolding indicating either a slow refolding process or intermolecular quadruplex formation ( Fig. 4b ).
To gain information on whether the G-quadruplexes are formed intermolecularly or intramolecularly, we analysed the melting profile temperature by diluting the C9Gru RNA oligonucleotide 10-fold (0.46 μM) in buffer containing 40 mM KCl. The spectrum displayed the same characteristic maximum and minimum around 262 and 237 nm, respectively. Furthermore, the structure remained highly stable and retained the same amount of structure at 95°C as seen in the concentrated sample ( Supplementary Fig. S2 ). This result indicates the formation of an intramolecular G-quadruplex19.
GGGGCC G-quadruplex stability is cation dependent
The stability of the G-quadruplex structure is strongly influenced by the presence of monovalent cations between the G-quartet stacks. K+ promotes stable folding over Na+ and Li+ ions. We therefore compared the C9Gru RNA structure in the presence of either K+ or Li+. CD results showed a reduction of the formed structure in the presence of Li+, with a reduction of the 262 nm peak by ~30% and a reduction in stability of the quadruplex and the cooperativity of the fold ( Fig. 4c ).
Together, these CD data show that GGGGCC RNA repeats fold into a very stable, parallel intramolecular G-quadruplex structure.
Discussion
Our investigations show the C9orf72 GGGGCC-hexanucleotide repeat forms G-quadruplexes and these adopt a parallel topology as illustrated in the model in Figure 1b . Our structural data confirm in silico predictions that attribute a high probability of forming G-quadruplexes to the GGGGCC-repeat.
With regards to quadruplex topology, in contrast to DNA G-quadruplexes, RNA G-quadruplexes generally disfavour folded topologies with phosphate backbones running antiparallel containing mixed syn and anti glycosidic bonds. A preference of the ribose sugar puckers for the C3’-endo conformation favours the anti-conformation and so an all-parallel topology21. Structural studies have suggested that stability is derived through the 2′-OH groups and their intra-and inter molecular interactions with the ribose O4′ atom for C3′-endo pucker and the N2 guanine amine for C2′-endo puckers22. However, this preference is not absolute and examples of RNA adopting the anti-conformation exist23. Our results show that similar to most other RNA G-quadruplexes, the C9orf72 GGGGCC-repeat G-quadruplex adopts a parallel conformation. The GGGGCC-repeat G-quadruplexes, similar to other RNA G-quadruplexes, are very thermodynamically stable at physiological intracellular potassium ion concentrations. Our CD data obtained from the RNA oligomer dilution also suggests that quadruplexes form intramolecularly. In the context of the C9orf72 expansion, where the hexanucleotide repeats are >700, it remains to be assessed whether quadruplexes form by association of adjacent repeats or from distant tracts of the sequence.
The ALS-FTD causing expansions in C9orf72 are formed by a very large number of repeats, which is thought to range between 700 and 1,400 hexamers5. Although we have shown the propensity of the basic repeats composing this expansion to form G-quadruplexes, the extended structure of this very long RNA sequence is not yet known. Evidence of ternary structure from RNA transcripts of telomeric repeat (TERRA) sequences containing several hundred G-quadruplex-forming hexamers, indicates a “bead on a string model” with stacked quadruplex dimers, using 3′ or 5′ interfaces, connected by the intervening linkers to the next pairing24,25. Therefore, we would predict that many quadruplexes would be formed within the C9orf72 expansion and that quadruplexes would stack as dimers.
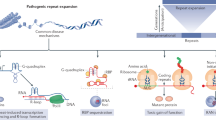
Three transcript variants (V1, V2, V3) have been described for the C9orf72 gene: V2 and V3 utilize exon 1a and therefore include the hexanucleotide repeat, while V1 utilizes the alternative exon 1b therefore excluding the hexanucleotide repeat, which is located upstream of the transcription start site5 ( Fig. 2a ). Real time RT-PCR data from patient brain and cell lines has shown that the presence of the expanded repeat causes a reduction in V1 transcription5,16. V2 and V3 were shown to be transcribed in patient frontal cortex cDNA, which was confirmed by the detection of the hexanucleotide repeats in nuclear foci in patients' brains by in situ hybridization5. A second study used real-time PCR to show that V2 levels were reduced in patient frontal cortex16. Therefore further studies are required to clarify the extent of transcription of the expanded repeat in V2 and V3 in relevant patient tissues. As GGGGCC RNA repeats form G-quadruplexes, it is likely that the DNA sequence also forms these structures. There is evidence that DNA G-quadruplexes on the template strand (also known as the non-coding strand) can inhibit transcription26. It has also been suggested that G-quadruplexes forming on the coding or non-template strand, as is the case with the C9orf72 repeats, could enhance transcription by keeping the template strand single stranded26. This would be consistent with transcription through the repeat and the identification of nuclear RNA foci containing the repeat in patient tissue5. Further work will be required to determine the extent and mechanism of transcription of the C9orf72 repeat sequence. Whether disease is caused by a toxic effect of the GGGGCC-repeat RNA, by a loss of function of C9orf72 or by both mechanisms is yet to be determined.
Repeat expansions in the 3′ UTR of DMPK and in intron 1 of ZNF9 were shown to cause Myotonic Dystrophy type 1 and 2 respectively (DM1 and DM2)27,28. These mutations are similar to C9orf72 expansions in being located in non-coding regions of the respective genes and in forming nuclear RNA foci in patient tissue and cells. Although loss of transcript has been suggested to play a role in both DM1 and DM229,30, the fact that two repeat sequences located in entirely different genes can cause such similar disease features, implies a potential common pathogenic mechanism by RNA gain-of-function31. Indeed, it has been shown that RNA binding proteins, such as MBNL1, are sequestered away from their normal RNA targets by interaction with the expanded repeats32. This leads to aberrant functional downstream effects which directly cause aspects of the myotonic dystrophy phenotype33,34,35. The expanded repeats that cause DM1 fold into a stable hairpin structure that mimics the normal MBNL1 RNA-binding site36,37. These findings suggest it is possible that the expanded GGGGCC-repeat may act in a similar fashion. In this light, the secondary structure of the RNA repeat expansion is crucial in determining which proteins it binds to.
G-quadruplexes have been shown to play a role in a variety of biological processes13,14. A number of these functions are carried out by the interaction with G-quadruplex binding proteins and the sequestration of these proteins could lead to downstream effects that play a role in the disease pathogenesis. While many DNA G-quadruplex binding proteins have been identified, relatively few proteins have been confirmed to bind RNA G-quadruplexes38. A prominent example of an RNA G-quadruplex binding protein is the fragile X mental retardation protein FMRP39, but it is currently unknown whether FMRP is sequestered by the C9orf72 expanded GGGGCC-repeats. Further work will be required to determine whether specific proteins bind to the repeats and what role they might play in disease pathogenesis. If RNA G-quadruplex binding proteins were sequestered by the expanded repeats, thousands of potential target genes with predicted G-quadruplex structures in their regulatory regions could be affected40.
An intriguing finding, which carries relevance for neurons, is that G-quadruplexes in the 3′ UTR of mRNAs are necessary and sufficient for the localization of these mRNAs to dendrites, through interactions with proteins such as FMRP12. Whether the accumulation of the C9orf72 expanded RNA causes sequestration of proteins relevant in mRNA transport needs to be addressed, however, this highlights the possibility of RNA alterations in C9orf72 ALS/FTD that are not limited to RNA quantity and splicing defects. Therefore the study of C9orf72 ALS/FTD may need to couple quantitative RNA sequencing studies with RNA localization investigations in order to fully understand disease pathogenesis. Finally, small molecules have been identified that interact with G-quadruplexes13 and our data suggests that they may have use as potential therapeutics in ALS and FTD caused by C9orf72 repeat expansions.
Methods
RNA oligonucleotide sample preparation and annealing
An HPLC purified RNA oligonucleotide of sequence GGGGCCGGGGCCGGGGCCGGGGC was purchased from Integrated DNA Technologies and supplied as a lyophilized powder and reconstituted in ultrapure water to a stock concentration of 2.5 mM. We termed this sequence the C9orf72 minimal G-quadruplex repeat unit (C9Gru). Annealing was carried out by heating the samples to 90°C and allowing them to cool overnight to 20°C.
NMR spectroscopy
NMR data was acquired using a Bruker AVANCE 500 NMR spectrometer operating at a proton resonance frequency of 500.13 MHz and equipped with a QNP cryoprobe. 1D 1H NMR spectra of the RNA sample in 90%H2O/10%D2O were acquired and processed using Topspin (version 2.1, Bruker Biospin, Karlsruhe) and excitation sculpting pulse sequence (Bruker pulse program zgesp). Spectra of the RNA sample were acquired before annealing in the presence of 10 mM potassium phosphate buffer, pH 7.0 and after annealing in the presence of 40 mM of KCl and 10 mM potassium phosphate buffer, pH 7.0. Samples were equilibrated in both cases at a calibrated temperature of 298 K. Data was accumulated centered at the solvent resonance with 1024 transients over a frequency width of 10.33 kHz into 32 K data points for an acquisition time aq = 1.58 s with a relaxation delay d1 = 2 s between transients using a 90° radiofrequency (r.f.) pulse (p1 = 13 μs at a power level of 2 dB).
Circular dichroism
RNA concentrations ranged from 4.6 to 0.46 μM and are indicated in the results section. The following buffers were used: 40 mM KCl, 10 mM potassium phosphate, pH 7.0; 50 mM LiCl, 10 mM sodium phosphate, 0.35 mM KCl, pH 7.0. CD experiments were performed at temperatures between 15°C and 95°C, with a 1°C/min temperature gradient, using a Jasco J715 spectropolarimeter (Jasco Hachioji, Tokyo, Japan) equipped with a Jasco peltier temperature control system. A CD spectrum of the buffer was recorded and subtracted from the spectrum obtained for the RNA-containing solution. Data were zero-corrected between 340–350 nm.
References
Haverkamp, L. J., Appel, V. & Appel, S. H. Natural history of amyotrophic lateral sclerosis in a database population. Validation of a scoring system and a model for survival prediction. Brain 118 (Pt 3), 707–719 (1995).
McKhann, G. M. et al. Clinical and pathological diagnosis of frontotemporal dementia: report of the Work Group on Frontotemporal Dementia and Pick's Disease. Arch. Neurol. 58, 1803–1809 (2001).
Neary, D. et al. Frontotemporal lobar degeneration: a consensus on clinical diagnostic criteria. Neurology 51, 1546–1554 (1998).
Lillo, P. & Hodges, J. R. Frontotemporal dementia and motor neurone disease: overlapping clinic-pathological disorders. J Clin Neurosci 16, 1131–1135 (2009).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
Campbell, N. H. & Neidle, S. G-quadruplexes and metal ions. Met Ions Life Sci 10, 119–134 (2012).
Xu, Y., Suzuki, Y., Ito, K. & Komiyama, M. Telomeric repeat-containing RNA structure in living cells. Proc. Natl. Acad. Sci. U.S.A. 107, 14579–14584 (2010).
Kikin, O., Zappala, Z., D'Antonio, L. & Bagga, P. S. GRSDB2 and GRS_UTRdb: databases of quadruplex forming G-rich sequences in pre-mRNAs and mRNAs. Nucleic Acids Res 36, D141–D148 (2008).
Huppert, J. L., Bugaut, A., Kumari, S. & Balasubramanian, S. G-quadruplexes: the beginning and end of UTRs. Nucleic Acids Res. 36, 6260–6268 (2008).
Eddy, J. & Maizels, N. Conserved elements with potential to form polymorphic G-quadruplex structures in the first intron of human genes. Nucleic Acids Res. 36, 1321–1333 (2008).
Subramanian, M. et al. G-quadruplex RNA structure as a signal for neurite mRNA targeting. EMBO Rep. 12, 697–704 (2011).
Collie, G. W. & Parkinson, G. N. The application of DNA and RNA G-quadruplexes to therapeutic medicines. Chem Soc Rev 40, 5867–5892 (2011).
Ji, X. et al. Research progress of RNA quadruplex. Nucleic Acid Ther 21, 185–200 (2011).
Cooper, T. A., Wan, L. & Dreyfuss, G. RNA and Disease. Cell 136, 777–793 (2009).
Gijselinck, I. et al. A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol 11, 54–65 (2012).
Rutherford, N. J. et al. Length of normal alleles of C9ORF72 GGGGCC repeat do not influence disease phenotype. Neurobiol. Aging 33, 2950.e5–7 (2012).
Wong, H. M., Stegle, O., Rodgers, S. & Huppert, J. L. A Toolbox for Predicting G-Quadruplex Formation and Stability. J Nucleic Acids 2010 (2010).
Kumari, S., Bugaut, A., Huppert, J. L. & Balasubramanian, S. An RNA G-quadruplex in the 5′ UTR of the NRAS proto-oncogene modulates translation. Nat Chem Biol 3, 218–221 (2007).
Wieland, M. & Hartig, J. S. RNA quadruplex-based modulation of gene expression. Chem. Biol. 14, 757–763 (2007).
Tran, P. L. T., Cian, A. D., Gros, J., Moriyama, R. & Mergny, J.-L. Tetramolecular Quadruplex Stability and Assembly. Topics in current chemistry (2012).
Collie, G. W., Haider, S. M., Neidle, S. & Parkinson, G. N. A crystallographic and modelling study of a human telomeric RNA (TERRA) quadruplex. Nucleic Acids Res. 38, 5569–5580 (2010).
Tang, C.-F. & Shafer, R. H. Engineering the quadruplex fold: nucleoside conformation determines both folding topology and molecularity in guanine quadruplexes. J. Am. Chem. Soc. 128, 5966–5973 (2006).
Martadinata, H., Heddi, B., Lim, K. W. & Phan, A. T. Structure of long human telomeric RNA (TERRA): G-quadruplexes formed by four and eight UUAGGG repeats are stable building blocks. Biochemistry 50, 6455–6461 (2011).
Collie, G. W. et al. Electrospray mass spectrometry of telomeric RNA (TERRA) reveals the formation of stable multimeric G-quadruplex structures. J. Am. Chem. Soc. 132, 9328–9334 (2010).
Bochman, M. L., Paeschke, K. & Zakian, V. A. DNA secondary structures: stability and function of G-quadruplex structures. Nat. Rev. Genet. 13, 770–780 (2012).
Ranum, L. P., Rasmussen, P. F., Benzow, K. A., Koob, M. D. & Day, J. W. Genetic mapping of a second myotonic dystrophy locus. Nat. Genet. 19, 196–198 (1998).
Harley, H. G. et al. Expansion of an unstable DNA region and phenotypic variation in myotonic dystrophy. Nature 355, 545–546 (1992).
Reddy, S. et al. Mice lacking the myotonic dystrophy protein kinase develop a late onset progressive myopathy. Nat. Genet. 13, 325–335 (1996).
Chen, W. et al. Haploinsuffciency for Znf9 in Znf9+/− mice is associated with multiorgan abnormalities resembling myotonic dystrophy. J. Mol. Biol. 368, 8–17 (2007).
Lee, J. E. & Cooper, T. A. Pathogenic mechanisms of myotonic dystrophy. Biochem. Soc. Trans. 37, 1281–1286 (2009).
Miller, J. W. et al. Recruitment of human muscleblind proteins to (CUG)(n) expansions associated with myotonic dystrophy. EMBO J. 19, 4439–4448 (2000).
Kanadia, R. N. et al. Reversal of RNA missplicing and myotonia after muscleblind overexpression in a mouse poly(CUG) model for myotonic dystrophy. Proc. Natl. Acad. Sci. U.S.A. 103, 11748–11753 (2006).
Lin, X. et al. Failure of MBNL1-dependent post-natal splicing transitions in myotonic dystrophy. Hum. Mol. Genet. 15, 2087–2097 (2006).
Du, H. et al. Aberrant alternative splicing and extracellular matrix gene expression in mouse models of myotonic dystrophy. Nat. Struct. Mol. Biol. 17, 187–193 (2010).
Yuan, Y. et al. Muscleblind-like 1 interacts with RNA hairpins in splicing target and pathogenic RNAs. Nucleic Acids Res. 35, 5474–5486 (2007).
Warf, M. B. & Berglund, J. A. MBNL binds similar RNA structures in the CUG repeats of myotonic dystrophy and its pre-mRNA substrate cardiac troponin T. RNA. 13, 2238–2251 (2007).
Bugaut, A. & Balasubramanian, S. 5′-UTR RNA G-quadruplexes: translation regulation and targeting. Nucleic Acids Res. 40, 4727–4741 (2012).
Melko, M. & Bardoni, B. The role of G-quadruplex in RNA metabolism: involvement of FMRP and FMR2P. Biochimie 92, 919–926 (2010).
Beaudoin, J.-D. & Perreault, J.-P. 5′-UTR G-quadruplex structures acting as translational repressors. Nucleic Acids Res. 38, 7022–7036 (2010).
Acknowledgements
We thank the UK Medical Research Council (E.M.C.F., P.F., A.N.), the Motor Neurone Disease Association (E.M.C.F., P.F.), Alzheimer's Research UK and MHMS General Charitable Trust (A.M.I.) for funding. PF is funded by a Medical Research Council/Motor Neurone Disease Association Lady Edith Wolfson Fellowship.
Author information
Authors and Affiliations
Contributions
P.F., E.M.C.F., G.P., A.M.I. conceived the project and wrote the manuscript; P.F., S.M., A.N., M.Z., G.P. planned and performed the experiments and analysed the data. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Fratta, P., Mizielinska, S., Nicoll, A. et al. C9orf72 hexanucleotide repeat associated with amyotrophic lateral sclerosis and frontotemporal dementia forms RNA G-quadruplexes. Sci Rep 2, 1016 (2012). https://doi.org/10.1038/srep01016
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01016
This article is cited by
-
Mutant GGGGCC RNA prevents YY1 from binding to Fuzzy promoter which stimulates Wnt/β-catenin pathway in C9ALS/FTD
Nature Communications (2023)
-
The ALS/FTD-related C9orf72 hexanucleotide repeat expansion forms RNA condensates through multimolecular G-quadruplexes
Nature Communications (2023)
-
Leukocyte telomere length and amyotrophic lateral sclerosis: a Mendelian randomization study
Orphanet Journal of Rare Diseases (2021)
-
Molecular mechanisms underlying nucleotide repeat expansion disorders
Nature Reviews Molecular Cell Biology (2021)
-
MeCP2 duplication causes hyperandrogenism by upregulating LHCGR and downregulating RORα
Cell Death & Disease (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.