Abstract
Centenarians exhibit extreme longevity and a remarkable compression of morbidity. They have a unique capacity to maintain homeostatic mechanisms. Since small non-coding RNAs (including microRNAs) are implicated in the regulation of gene expression, we hypothesised that longevity of centenarians may reflect alterations in small non-coding RNA expression. We report the first comparison of microRNAs expression profiles in mononuclear cells from centenarians, octogenarians and young individuals resident near Valencia, Spain. Principal Component Analysis of the expression of 15,644 mature microRNAs and, 2,334 snoRNAs and scaRNAs in centenarians revealed a significant overlap with profiles in young individuals but not with octogenarians and a significant up-regulation of 7 small non-coding RNAs in centenarians compared to young persons and notably 102 small non-coding RNAs when compared with octogenarians. We suggest that the small non-coding RNAs signature in centenarians may provide insights into the underlying molecular mechanisms endowing centenarians with extreme longevity.
Similar content being viewed by others
Introduction
The study of centenarians is important not only because they reach a very old age but also because they are an example of successful ageing. They show a remarkable compression of age associated morbidity to such an extent that their health span approximates their life span1.
A classical characteristic of (unsuccessful) ageing is that homeostatic, regulatory mechanisms are impaired or even lost2. Our aim was to determine the molecular mechanisms by which centenarians maintain a highly efficient homeostasis.
Some recent studies have searched of mutations (especially single nucleotide polymorphisms) that may be characteristic of human extreme longevity3,4,5. However, to our knowledge few studies have dealt with the regulation of RNA expression. One report analysed the whole human microRNome. This study, however compared young with middle aged people and no attempts were made to study extreme longevity6.
The small non-coding RNAs include various classes of regulatory RNAs, of which, microRNAs are the best studied. MicroRNAs (miRNAs) are small single-stranded RNAs that regulate gene expression by partial complementary base pairing to specific messenger RNAs7. The ability to regulate many targets at the same time, makes miRNAs good candidates to control many physiological processes, specially multifactorial ones like ageing8.
The aim of this work was to study miRNAs expression profiles in centenarians and compare them with octogenarians and with young individuals.
A global analysis of miRNAs expression (miRNome) indicates that there are striking similarities between their expression in centenarians and in young people. On the contrary octogenarians express miRNAs differently from centenarians and from young people.
Moreover, we have found that centenarians up-regulate the expression of miRNAs even when compared with young people. Their up-regulation of the expression of miRNAs is vastly superior that than of octogenarians. This may be a specific characteristic of centenarians that explains their striking maintenance of homeostatic mechanisms and their healthy ageing.
A consequence of these studies is that those octogenarians that up-regulate the expression of miRNAs are candidates to become centenarians. This intriguing hypothesis remains to be tested in longitudinal studies now under way in the Spanish Centenarian Study Group.
Results
Principal component analysis of small non-coding RNAs in centenarians
The principal component analysis (PCA) of all small non-coding RNAs obtained in the Genechip miRNA 2.0 Array shows that their expression in centenarians is similar to young people. Octogenarians show a very different pattern of expression of small non-coding RNA when compared with young people or with centenarians (Fig. 1).
Principal component analysis (PCA) of the small non-coding RNA profiles in mononuclear cells from centenarians, octogenarians and young people.
The small non-coding RNA expression profiles of mononuclear cells were analyzed by PCA. Each of our samples was assayed using an array (Genechip miRNA 2.0 Array) that is defined by 1,105 mature miRNAs, 1.105 pre-miRNAs, 32 scaRNA and 2,302 snoRNA from the miRBASE (v.15)). The ellipsoids (in red, octogenarian; in blue centenarian and in green young group) show a distinct directionality in different groups based on similarities and differences with age. The axes correspond to principal component 1 (PC1, x-axis), PC2 (y-axis) and PC3 (z-axis).
Up-regulation of small non-coding RNAs in centenarians
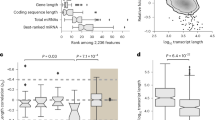
When we performed a restrictive statistical analysis with two variables: fold change |1.8| and P value ≤ 0.05 comparing the combination of the three groups (centenarians, octogenarians and young people) (see Fig. 2), we identified six miRNAs and one scaRNA that change specifically between centenarians and young people. Of those RNAs, all were up-regulated and none was down-regulated. Of the six miRNAs which we found to be relevant we did not further study one of them (miR-4281) because its expression was not different in centenarians and in octogenarians. The rest, i.e, five miRNAs and one scaRNA showed differences in centenarians versus young individuals and between centenarians versus octogenarians. Interestingly, all the RNAs that we have found are over-expressed and none is under-expressed in centenarians when compared with young people. The miRNAs found were miR-21, miR-130a, miR-494, miR-1975 and miR-1979. SCARNA17 (also called U91) was also up-regulated.
Number of small non-coding RNAs that are significantly modified when comparing the different groups, centenarians, octogenarians and young individuals.
Panel A Representation of 1-way ANOVA analysis between different contrasts: centenarians versus young individuals (C vs Y), octogenarians versus young individuals (O vs Y) and centenarians versus octogenarians (C vs O). Statiscally significant miRNAs were filtered using P value ≤ 0.05 and fold-change ≥ |1.8|. Panel B Venn diagram displaying the number of small non-coding RNAs are that commonly regulated in the different groups.
In clear contrast, when we compared the miRNome of octogenarians vs young, using the same selection criteria as for centenarians, we found that 50 small non-coding RNAs were down-regulated and only one miRNA was up-regulated. The situation was even more remarkable when we compared centenarians with octogenarians: 102 small non-coding RNAs were up-regulated in centenarians and only one was up-regulated in octogenarians. (Fig. 2). The list of all small non-coding RNAs that change in centenarians, octogenarians and young people is in Table S1, (see supplementary material).
Validation of up-regulated small non-coding RNAs in centenarians
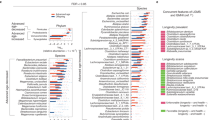
We used qPCR to validate changes of RNA expression that we had observed using microarrays and confirmed the over-expression of SCARNA17, miR-21 and miR-130a. In the case of miR-494 we found a tendency to increase but it was not statistically significant (Fig. 3).
Validation by qPCR of the small non-coding RNAs that are uniquely over-expressed in centenarians (when compared with octogenarians and with young individuals).
Expression of SCARNA17, miR-21, miR-130a and miR-494 in young individuals (white column), octogenarians (gray column) and centenarians (black column) measured by qPCR. (*) P value <0.05 and (**) P value <0.01 versus young people. (##) P value <0.01 versus octogenarians. In all cases, changes in miRNA expression observed by expression profiles were confirmed by qPCR.
The two remaining genes, i.e., miR-1975 and miR-1979 were not further studied because there is, to our knowledge, not a single reference to their biological function in vivo. Since this paper was not concerned with identifying new functions of miRNAs, we only proceeded studying the genes of which some function was known.
We further tested the validity of our microarray analysis by determining by qPCR the expression of a miR19b, a microRNA known to be down-regulated in normal aging (i.e. octogenarians)9. Figure 4 shows that miR19b expression in octogenarians was 11% of the young controls. The expression of this miRNA did not change in extreme longevity, i.e. it was the same in young individuals and in centenarians.
Validation by qPCR of the miR-19b which are uniquely under-expressed in octogenarians (when compared with centenarians and with young individuals).
Expression of miR-19b in young individuals (white column), octogenarians (gray column) and centenarians (black column) measured by qPCR. (*) P value <0.05 versus young people. (#) P value <0.01 versus centenarians.
Discussion
The incidence of centenarians in the US and Western European population is one in 10.000. Moreover, an interesting study shows that centenarians (and even more so, supercentenarians) show a compression of morbidity and thus their health span approximates their life span1.
Efforts by the scientific community are centred in finding genetic traits that may be specific for extreme human longevity. The majority of the studies, both in Europe3 and in the US4 have been focused in finding single nucleotide polymorphisms (SNPs) that may be characteristic of the centenarian population. For instance Franceschi and co-workers3 showed that there are polymorphisms in the p21 (CDKN1A) gene that correlate with longevity in an Italian population. Perls and colleagues in the New England Centenarian Study have also studied the frequency of 281 SNPs in a population of centenarians4.
Basic gerontological work2 showed that an important characteristic of unsuccessful ageing is that homestatic mechanisms are impaired. Thus one could expect that in very successful ageing homeostatic, regulatory mechanisms might be maintained.
Since small non-coding RNAs are involved in regulatory processes, we studied their expression in peripheral cells from centenarians. To our knowledge previous studies have dealt with the differential expression of the whole miRNome in relation of ageing (not in extreme longevity). In one report, authors studied the miRNome in two young people (aged 30) and two mature ones (age 65)6. Two further studies have determined that four miRNAs (miR-17, miR-19b, miR-20a and miR-106a)9 and the miR 17–92 cluster are down regulated in normal aging10.
We studied the whole miRNome in young, octogenarians and centenarians with a view of finding specific characteristics of the centenarians.
The first finding reported here is that the Principal Component Analysis of small non-coding RNAs in centenarians and in young people are almost overlapped whereas that of octogenarians is completely different from either young or centenarians (See Fig. 1). This may be taken to indicate that the regulation of genetic expression in centenarians is similar to young people but very different from that of octogenarians.
In our opinion, however, the most puzzling finding reported here is that centenarians up-regulate the expression of small non-coding RNAs whereas octogenarian down-regulate it (when compared to young people). Of course, when we compared the expression of the miRNome in centenarians and octogenarians the centenarians vastly beat octogenarians in that the former over-express 102 small non-coding RNAs and the latter only one (see Fig. 2).
Of all the small non-coding RNAs that we have found that are specifically over-expressed in centenarians, only four had a known function. These were SCARNA17, miR-21, miR-130a and miR-494. The function of two more (miR-1975 and miR-1979) is so far unknown. Thus we focused our attention on the four whose function is known and we confirmed by qPCR that their expression is up-regulated in cells from centenarians (when compared with octogenarians and with young people).
SCARNA17 is a small nuclear RNA which accumulates specifically in Cajal bodies11. Cajal bodies have been involved in RNA-related metabolic processes and in telomere maintenance12. Telomeres and telomerase are involved in ageing and we found that over-expression of telomerase increases life span of mice13.
miR-21, a mature miRNA, has been implicated in different cell processes such as cell proliferation, mitochondrial damage, cell growth, chemosensitivity, cell cycle, genome instability, response to stress, mRNA cleavage, cell invasion and translation. The over-expression of miR-21 has been related with a neuroprotective effect from ischemic death14 and cardiac protection15,16. Up-regulation of miR-21 may decrease cell death17 and inhibit mitochondrial damage18. The dysregulation of miR-21 occurs in diseases like cancer19 and in metabolic disorders20.
We have also found that miR-130a is up-regulated in centenarians. The major known function of this miRNA is the regulation of FOG-2 protein expression thus suggesting that miR-130a may play a role in the regulation of cardiac development21. Finally, over-expression of miR-494 results in an inhibition of apoptosis and cardioprotection22.
The major findings reported in this paper are that small non-coding RNAs expression in centenarians is similar to that of young people but very different from octogenarians and that centenarians up-regulate the expression of regulatory small non-coding RNAs whereas octogenarians down-regulate it.
Methods
Study population
The Spanish Centenarian Study Group at RETICEF, began in 2007 as a population-based study of all centenarians living within an area near of Valencia called La Ribera (11th Health Department of the Valencian Community, Spain), which is composed of 29 towns (240.000 inhabitants). Potential subjects were selected from the population data system of the 11th Health Department. We found 31 centenarians of whom 20 met the inclusion criteria. Then we randomly recruited 20 octogenarians of whom 16 met the inclusion criteria and 20 young people of whom 14 fulfilled the inclusion criteria. The inclusion criteria were: to be born within the dates indicated in the study (before 1908 for centenarians, between 1928 and 1938 for octogenarians and between 1968 and 1988 for young individuals), to live in the 11th Health Department for at least the last 6 years and to sign the informed consent. The exclusion criterion was to be terminally ill for any reason.
All experimental procedures were approved by the Committee for Ethics in Clinical Research of the Hospital de la Ribera, Alzira. All patients or their relatives were fully informed of the aims and scope of the research and signed an informed consent.
Peripheral blood mononuclear cells isolation
Whole blood collected in one VACUTAINER® CPT™ (Cell Preparation Tube) (BD, Franklin Lakes, NJ) containing sodium heparin as the anti-coagulant, was taken from each subject at each site. The CPT were processed at the collection site, according to the manufacturer's instructions, by centrifugation at 3000 × G for 15 minutes at room temperature within half an hour of blood collection23. After centrifugation the CPT were gently inverted several times to separate plasma, mononuclear cells and erythrocytes. We collected the white ring containing mononuclear cells. Mononuclear cells were washed twice in PBS and frozen at −80°C for subsequent RNA isolation.
Isolation of RNA from peripheral blood mononuclear cells
Total RNA containing small RNA was isolated using a mirVana miRNA Isolation Kit (Ambion, Austin, TX, USA) according to manufacturer's directions. The purity and concentration of RNA were determined from OD260/280 readings using a Genequant Pro Classic spectrophotometer (GE Healthcare). RNA integrity was determined by capillary electrophoresis using the RNA 6000 Nano Lab-on-a-Chip kit and the Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). Only RNA extracts with RNA integrity number values ≥ 6 underwent in further analysis.
Expression profiling of small non-coding RNAs
Small non-coding RNA expression profiling was performed by using GeneChip miRNA 2.0 Array (Affymetrix, Santa Clara, CA, USA). The array compromised of 15,644 mature microRNA sequences from the miRBASE (v15) encoded miRNA coverage of 131 organisms, 2,334 encompassed snoRNAs and scaRNAs and 2,202 probe sets unique to pre-miRNA hairpin sequences.
Microarray experiments were conducted according to the manufacturer's instructions. Briefly, 200 ng total RNA was labeled with FlashTag Biotin HSR RNA Labeling Kit from Genisphere. The labeling reaction was hybridized on the miRNA Array in Affymetrix Hybridization Oven 640 (Affymetrix) at 48°C for 18 h. The arrays were stained with Fluidics Station 450 using fluidics script FS450_0003 (Affymetrix) and then scanned on GeneChip Scanner 3000 7G (Affymetrix, Santa Clara, CA, USA). GeneChip® Command Console® Software supplied by Affymetrix was used to perform gene expression analysis. miRNA probe outliers were defined as per the manufacturer's instructions (Affymetrix) and further analyzed for data summarization, normalization and quality control by using the web-based miRNA QC Tool software (www.affymetrix.com).
All raw data regarding small non-coding RNA expression has been deposited in ArrayExpress, a public accessible database.
Real time PCR validation
RT was performed with random hexamers using MultiScribeTM Reverse Transcriptase (Applied Biosystems). First-strand cDNA synthesis was performed at 42°C for 30 min.
The reaction was stopped by heating the mixture at 95°C for 5 min and stored at −20°C until further use.
Pre-developed Taqman primers specific for SCARNA17, miR-21, miR-130a, miR-494 and miR19b (Hs03298712_s1, REF: 000397, REF: 00454, REF: 002365 and REF: 000396) were purchased from Applied Biosystems. The transcript levels of were detected by the 7900HT Fast Real-Time PCR System (Applied Biosystems). Each PCR reaction contains 1 μl of RT product, 5 μl Taqman Universal Maxter Mix or Gene Expression Master Mix (Applied Biosystems) and 0, 5 μl probes in a final volume of 10 μl. The PCR conditions were as follows: 50°C for 2 min, 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min. All PCR reactions were cycled in the linear region of amplification. Results were normalized according to RNU66 (housekeeping control for miRNAs, REF. 001002, Applied Biosystems) and RPLPO (housekeeping control for small RNA, SCARNA17), REF: 4333761F, Applied Biosystems) quantification in the same sample reaction. The threshold cycle (CT) was determined and then the relative miRNA and gene expression was expressed as follows:
Relative amount = 2−Δ (ΔCT),
where ΔCT = CT target − CT Housekeeping control and Δ (ΔCT) = ΔCT studied group − ΔCT baseline.
miRNA and small RNA levels in young group was chosen as the baseline.
Data analysis of microarrays
Data (.CEL files) were analyzed and statistically filtered using software Partek Genomic Suite 6.4 (Partek Inc., St. Louis, MO). Input files were normalized with the RMA algorithm for gene array on core meta probesets or miRNAs. A 1-way ANOVA was performed with the Partek Genomics Suite across all samples. Statistically significant small non-coding RNAs between different groups studied were identified using a model analysis of variance of P value ≤ 0.05. Fold-change values < |± 1.8| were removed.
The imported data were analyzed by Principal Components Analysis to determine the significant sources of variability in the data.
PCA reduces the complexity of high-dimensional data and simplifies the task of identifying patterns and sources of variability in a large data set24. The samples (fifty biological replicates, each hybridized to a separate Genechip) are represented by the spheres in the three-dimensional plot (Fig. 1). The distance between any pair of points is related to the similarity between the two samples in high-dimensional space (in this case, each variable correspond a one dimensional space). Samples that are near each other in the plot are similar in a large number of variables. Conversely, samples that are far apart in the plot are different in a large number of variables.
Finally, the selected small non-coding RNAs were imported into Pathway Studio v8 (Ariadne software) to classify the molecular function and biological processes represented by the miRNAs differentially expressed between centenarian and young samples.
Statistical analysis of validation results
All experiments were performed three times in triplicate. Data were represented by mean ± SEM. Comparison between groups was performed with a one-way ANOVA and a two-tailed t-test. P values of less than 0.05 were considered to be statistically significant.
Change history
22 March 2013
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Andersen, S. L., Sebastiani, P., Dworkis, D. A., Feldman, L. & Perls, T. T. Health span approximates life span among many supercentenarians: compression of morbidity at the approximate limit of life span. J Gerontol A Biol Sci Med Sci 67, 395–405 (2012).
Timiras, P. S. Physiological Basis of Aging and Geriatrics (Informa Heathcare, New York, 2007).
Gravina, S. et al. Identification of single nucleotide polymorphisms in the p21 (CDKN1A) gene and correlations with longevity in the Italian population. Aging (Albany NY) 1, 470–480 (2009).
Sebastiani, P. et al. Genetic signatures of exceptional longevity in humans. PLoS One 7, e29848 (2012).
Lescai, F. et al. Human longevity and 11p15.5: a study in 1321 centenarians. Eur J Hum Genet 17, 1515–1519 (2009).
Noren Hooten, N. et al. microRNA expression patterns reveal differential expression of target genes with age. PLoS One 5, e10724 (2010).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Chen, L. H., Chiou, G. Y., Chen, Y. W., Li, H. Y. & Chiou, S. H. MicroRNA and aging: a novel modulator in regulating the aging network. Ageing Res Rev 9 Suppl 1, S59–66 (2010).
Hackl, M. et al. miR-17, miR-19b, miR-20a and miR-106a are down-regulated in human aging. Aging Cell 9, 291–296 (2010).
Grillari, J., Hackl, M. & Grillari-Voglauer, R. miR-17-92 cluster: ups and downs in cancer and aging. Biogerontology 11, 501–506 (2010).
Darzacq, X. et al. Cajal body-specific small nuclear RNAs: a novel class of 2'-O-methylation and pseudouridylation guide RNAs. EMBO J 21, 2746–2756 (2002).
Zhao, Y. et al. Processive and distributive extension of human telomeres by telomerase under homeostatic and nonequilibrium conditions. Mol Cell 42, 297–307 (2011).
Tomas-Loba, A. et al. Telomerase reverse transcriptase delays aging in cancer-resistant mice. Cell 135, 609–622 (2008).
Buller, B. et al. MicroRNA-21 protects neurons from ischemic death. FEBS J 277, 4299–4307 (2010).
Dong, S. et al. MicroRNA expression signature and the role of microRNA-21 in the early phase of acute myocardial infarction. J Biol Chem 284, 29514–29525 (2009).
Cheng, Y. et al. Ischaemic preconditioning-regulated miR-21 protects heart against ischaemia/reperfusion injury via anti-apoptosis through its target PDCD4. Cardiovasc Res 87, 431–439 (2010).
Matsumoto, T. & Hwang, P. M. Resizing the genomic regulation of restenosis. Circ Res 100, 1537–1539 (2007).
Sayed, D. et al. MicroRNA-21 is a downstream effector of AKT that mediates its antiapoptotic effects via suppression of Fas ligand. J Biol Chem 285, 20281–20290 (2010).
Iorio, M. V. & Croce, C. M. Causes and consequences of microRNA dysregulation. Cancer J 18, 215–222 (2012).
Rottiers, V. & Naar, A. M. MicroRNAs in metabolism and metabolic disorders. Nat Rev Mol Cell Biol 13, 239–250 (2012).
Kim, G. H., Samant, S. A., Earley, J. U. & Svensson, E. C. Translational control of FOG-2 expression in cardiomyocytes by microRNA-130a. PLoS One 4, e6161 (2009).
Wang, X. et al. MicroRNA-494 targeting both proapoptotic and antiapoptotic proteins protects against ischemia/reperfusion-induced cardiac injury. Circulation 122, 1308–1318 (2010).
Ruitenberg, J. J., Mulder, C. B., Maino, V. C., Landay, A. L. & Ghanekar, S. A. VACUTAINER CPT and Ficoll density gradient separation perform equivalently in maintaining the quality and function of PBMC from HIV seropositive blood samples. BMC Immunol 7, 11 (2006).
I, T., J. Principal Component Analysis in Springer Series in Statistics,. Vol. XXIX (Springer, NY, 2002).
Acknowledgements
This work was supported by the following grants to JV: SAF 2010-19498 from the Spanish Ministry of Education and Science (MEC) co-financed with European FEDER funds; ISCIII2006-RED13-027 from the (RETICEF) and Prometeo 2010/074 from the Generalitat Valenciana. This study has been co-financed by FEDER funds from the European Union. ES acknowledges a 2009 PTA contract from the Ministry of Science and Innovation of Spain. We thank the following members of the Spanish Centenarian Study group for their help in this study: Ms Jesica Portero (Unit for Multigenic Analysis, UCIM, INCLIVA), Dr F.J. Tarazona; Dr J.R. Domenech, Dr Carmen Valldecabres, Dr. David Cuesta, Dr Ricardo Bou (Hospital de la Ribera, Alzira). Special thanks go to the nurses: Rosa Carrasco, Maria Jesús Martínez, Miriam Hoya and Alicia Pérez for their excellent care of the people involved in this study.
Author information
Authors and Affiliations
Contributions
E.S., J.G., K.M. performed experimental work; A.B., P.S. and J.A.A. performed clinical work, C.B. and L.R.M. co- directed research and J.V. designed research and directed the project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplemantary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Serna, E., Gambini, J., Borras, C. et al. Centenarians, but not octogenarians, up-regulate the expression of microRNAs. Sci Rep 2, 961 (2012). https://doi.org/10.1038/srep00961
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00961
This article is cited by
-
Early downregulation of hsa-miR-144-3p in serum from drug-naïve Parkinson’s disease patients
Scientific Reports (2022)
-
Epigenetic changes during ageing and their underlying mechanisms
Biogerontology (2020)
-
Genetic background, epigenetic factors and dietary interventions which influence human longevity
Biogerontology (2019)
-
Functional similarities of microRNAs across different types of tissue stem cells in aging
Inflammation and Regeneration (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.