Abstract
Biologists have devoted much attention to assortative mating or homogamy, the tendency for sexual species to mate with similar others. In contrast, there has been little theoretical work on the broader phenomenon of homophily, the tendency for individuals to interact with similar others. Yet this behaviour is also widely observed in nature. Here, we model how natural selection can give rise to homophily when individuals engage in social interaction in a population with multiple observable phenotypes. Payoffs to interactions depend on whether or not individuals have the same or different phenotypes and each individual has a preference that determines how likely they are to interact with others of their own phenotype (homophily) or of opposite phenotypes (heterophily). The results show that homophily tends to evolve under a wide variety of conditions, helping to explain its ubiquity in nature.
Similar content being viewed by others
Introduction
Homophily, the tendency to interact with others of similar type, is widely observed in nature. Sex- and age-related homophily, for example, shapes the formation of clusters of preferred companionships in zebras1, dolphins2 and predicts both the quantity and quality of many primate interactions3,4. Meerkats tend to assortatively associate with other group members of similar attributes in dominance and foraging networks5. And across many dimensions of phenotypes, humans exhibit high levels of homophily in social tie formation6,7,8. In fact, recent evidence suggests that humans may even exhibit genotypic homophily, meaning that individuals with a certain genotype are more likely to be friends with others of the same genotype9.
Heterophily, the tendency to interact with others of different type, also exists in nature at both the cellular10,11,12 and organismic levels. For example, research on collaboration networks suggests that people are likely to form heterophilic task-related ties with those who are complementary to their own skill sets8. Analogously, hunter-gatherer life is characterised by long-term imbalances in productivity and consumption and by the division of labour13; hence, one might possibly expect that social interactions would, at least in part, be heterophilic, offering complementary advantages to interacting parties; but they are not7.
Indeed, heterophily is far less common than homophily. Much effort has focused on examining the functional role homophily plays in a wide range of domains, including social segregation14, cultural polarization15, friendship formation16, social contagion17 and the evolution of cooperation18,19. And the ubiquity of homophily suggests that natural selection may favour it. Yet, to our knowledge, there have to date been no attempts to understand the possibly evolutionary origin of this phenomenon.
Here, we conceptualise the benefits to homophily and heterophily as the results of a simple coordination game. For example, homophily may yield fitness advantages because individuals using the same mode of communication may be able to act together more effectively. These advantages are sometimes called synergy. On the other hand, heterophily may be beneficial because it gives rise to specialisation or gains from trade, such as when a farmer interacts with a baker20, when different scientists collaborate21, or when individuals at different stages in the life cycle interact22.
We elaborate a simple model that assigns benefits to interactions and allows individuals to have preferences to interact with others with similar or different phenotypes. We then let these preferences co-evolve with the set of available phenotypes. This model shows that homophily emerges as the dominant preference under a wide variety of conditions.
For simplicity, suppose there is a haploid asexual population of size N. There are M possible phenotypes (size, colour, behaviour, etc.) and each individual i has an observable phenotype denoted by Gi ∈ {1, 2, …, M}. To be sure the phenotypes do not drive the results, we assume that none of these phenotypes alone make individuals more or less fit; only combinations of phenotypes between individuals determine fitness. When individuals i and j interact, they obtain a homophilic interaction payoff, a, when they are of the same phenotype (Gi = Gj). We can think of this as the payoff to synergy. When i and j are of different phenotypes (Gi ≠ Gj), they receive a heterophilic interaction payoff, b; we can think of this as the payoff to specialisation. We assume that when individuals do not interact, their payoff is 0.
We can describe three kinds of environments from this basic set up. When a > b there is an advantage to homophilic interactions. When a < b there is an advantage to heterophilic interactions. Finally, when a = b there is no advantage to either homophily or heterophily.
At each time period, we assume there is a process that allows each individual to interact with another individual. This process proceeds in three stages, 1) choose, 2) meet, 3) interact. With probability pi ∈ [0, 1], individuals will choose to interact with individuals of the same phenotype (homophily) and with probability 1 − pi, they will choose to interact with individuals of different phenotypes (heterophily). Note that pi = 1 means that individual i always chooses individuals of the same phenotype and never of the opposite phenotype (perfect homophily). Conversely, pi = 0 means that individual i always chooses individuals of different phenotypes and never individuals of the same phenotype (perfect heterophily). And for intermediate values 0 < pi < 1, individuals show a tendency to favour one kind of interaction over another. When pi > 0.5, then individual i tends to be homophilic and when pi < 0.5, individual i tends to be heterophilic.
Next, individuals meet. For simplicity, we assume they are randomly paired with other members of the population (one can imagine more complex assumptions about meeting – we consider this possibility below). Once they meet, they interact, but only if both individuals have chosen to initiate an interaction with a partner that is compatible with their preferences and phenotypes. Hence, the probability of a successful interaction between two individuals i and j is pipj if they are of the same phenotype (Gi = Gj) and (1 − pi)(1 − pj) if they are of different phenotypes (Gi ≠ Gj).
Results
One advantage of conceptualising homophily in this way is that we can estimate the average homophilic preference p for several species and phenotypes in a number of available data sets (Table 1; see also SI) assuming that observed interactions are successful matches. Note that these real world observations show that homophily (p > 0.5) is apparent in all cases, with the estimated value ranging between 0.53 and 0.68.
Returning to the model, if we let δij = 1 when Gi = Gj and 0 otherwise, then the expected payoff πij for each interaction is  . And letting
. And letting  be the proportion of individuals in the population with preference i and phenotype l, the likelihood of each encounter qij is
be the proportion of individuals in the population with preference i and phenotype l, the likelihood of each encounter qij is  . If we let each individual in the population initiate an interaction, the average expected payoff to individual i is then
. If we let each individual in the population initiate an interaction, the average expected payoff to individual i is then  .
.
We assume fitness is an exponential function of payoffs23 and individuals reproduce proportional to their fitness according to a frequency-dependent Moran process24, which can occur either via natural selection (the less fit die and are replaced by the more fit), or learning (the less fit copy the preferences of the more fit)25,26,27,28,29. We also allow for mutation. The probability that an offspring changes at random to one of the M phenotypes (each with equal likelihood) is v and in expectation ν = vN offspring will change. Similarly, the probability that an offspring changes to a preference drawn from a uniform distribution with support [0, 1] is u, with µ = uN offspring changing in expectation. For full details of the model, please refer to the SI.
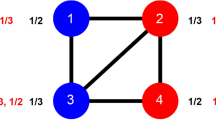
Before discussing the equilibria of the model, we use a simple example to provide an intuition for why there is an advantage to homophily. Suppose there are N1 individuals in group 1 with phenotype 1 and N2 in group 2 with phenotype 2. Further suppose that all individuals within each group have the same preference (p1, p2, respectively). The payoff to an individual in group 1 is then  , where N = N1 + N2. Taking two extreme cases, perfect homophilic preferences (p1 = 1) and perfect heterophilic preferences (p1 = 0), we can see that homophilic preferences yield a payoff advantage to individuals in group 1 if a > N2(1 − p2)b/(N1 − 1). Remarkably, this equation shows that homophily pays even if the benefit, a, to same type interactions is lower than the benefit, b, to opposite type interactions. This happens when group 1 is in the majority (it is easier to connect to a similar phenotype individual in a larger group) and/or when group 2 is also homophilic (which reduces the payoff to heterophilic interactions).
, where N = N1 + N2. Taking two extreme cases, perfect homophilic preferences (p1 = 1) and perfect heterophilic preferences (p1 = 0), we can see that homophilic preferences yield a payoff advantage to individuals in group 1 if a > N2(1 − p2)b/(N1 − 1). Remarkably, this equation shows that homophily pays even if the benefit, a, to same type interactions is lower than the benefit, b, to opposite type interactions. This happens when group 1 is in the majority (it is easier to connect to a similar phenotype individual in a larger group) and/or when group 2 is also homophilic (which reduces the payoff to heterophilic interactions).
Using coalescent theory30, we derive a closed-form solution for the limiting distribution of the preferences that survive in the population at equilibrium31,32 and we replicate all predictions with computational simulations. Figure 1 shows an example of three such equilibria.
Equilibrium distribution of homophilic preferences, p and the networks that result.
Solid blue lines in (a), (b) and (c) represent the theoretical distributions. Arrows show the theoretical critical point pc where the distribution crosses the uniform distribution (i.e., becomes more common or less common than expected due to chance). (d), (e), display networks of homophilic and heterophilic structures of social relationships that emerge from computer simulations of (a) and (c), respectively. Parameters: N = 30, β = 0.005, M = 3, u = 0.04, v = 0.06, (a, d) a = 1, b = 0.1, (b) a = 5/9, b = 1, (c, e) a = 0.1, b = 1. Results are averaged over T = 109 time steps.
When the payoff to homophilic interactions is high, those with a preference for homophily (high p) are more common than those with a preference for heterophily. Compared to a population with preferences distributed uniformly at random, the theory establishes a specific critical point pc above which individuals are favoured and below which they are disfavoured by natural selection. When the benefit to interacting with similar and dissimilar others is more in balance, a bimodal distribution of preferences emerges with most individuals strongly preferring either homophilic or heterophilic interactions. Finally, when the payoff to specialisation is high, those with a preference for heterophily (low p) emerge and are favoured when they have a preference below a critical point pc.
While these results may seem unsurprising, Figure 2 shows that the theory generates an elegant critical threshold a > Kb that determines whether the average individual in the population will evolve to become homophilic. The slope of this threshold is:

where M is the number of possible phenotypes and µ and ν are mutation rates that are rescaled by the population size N, µ = uN and ν = vN.
Evolutionary determinants of homophily.
Shown are the population average 〈p〉 as a function of the payoff values a and b. Compared with the base case in (a), decreases in (b) the number of phenotypes, M, (c) the strategy mutation rate, u and (d) the phenotypic mutation rate, v, always make it easier for homophily to evolve. For each b value, squares denote the critical a values determined by simulations. The straight lines are the theoretical predictions. We find good agreement between simulation results and our analytical theory. Parameters: N = 50, β = 0.005, (a) M = 4, u = 0.06, v = 0.06, (b) M = 2, u = 0.06, v = 0.06, (c) M = 4, u = 0.01, v = 0.06, (d) M = 4, u = 0.06, v = 0.02. Results are averaged over T = 108 time steps.
An important implication of this condition is that homophily can evolve even when the benefit b to heterophily (specialisation) exceeds the benefit a to homophily (synergy). This happens when K < 1. For example, Figure 2 shows that decreasing the number of phenotypes in the population (M), the phenotypic mutation rate (ν), or the preference mutation rate (µ), all decrease K and in turn increase the range of values under which homophily can emerge. Figure 3 shows that the average individual in the population becomes less homophilic as we increase each of these three parameters. And, in Figure 4, we show the full set of conditions under which homophily evolves for a given population size. We emphasise that homophily evolves even if a ≪ b as long as the (phenotypic) mutation rates are sufficiently low.
Population average of homophilic preferences, 〈p〉.
Shown are the population average 〈p〉 as a function of (a) the number of phenotypes, M, (b) the strategy mutation rate, u and (c) the phenotypic mutation rate, v. The filled circles are simulation results, which agree well with theoretical predictions (solid lines). As M, u, v increase, the population becomes less homophilic. Parameters: N = 50, β = 0.002, a = 3/4, b = 1, (a) u = 0.04, v = 0.02, (b) M = 5, v = 0.02, (c) M = 5, u = 0.04. Results are averaged over T = 5 × 108 time steps.
The full set of conditions under which homophily evolves.
The coloured 3D regions in (a), (c) show the combinations of parameters (µ, ν, M) that allow natural selection to favour homophily for given payoff values: (a) a = 3/4, b = 1 and (c) a = 1/10, b = 1. Corresponding to (a) and (c) respectively, the shaded areas in (b), (d) denote the set of parameters (µ, ν) favouring homophily with the number of phenotypes, M = 2. The boundaries of these shaded areas shrink to the dashed red lines, as M increases to ∞. Homophily evolves even if a ≪ b, as long as the mutation rates are low.
We studied the behaviour of the critical point pc (see SI) and found a surprisingly strong tendency toward homophily when mutation rates are low. In fact, when we let both ν and µ approach 0, the model shows that natural selection favours all individuals with  . In other words, all homophilic individuals and even those individuals that weakly prefer heterophily tend to do better than those that strongly prefer heterophily. And this result is independent of the payoff to heterophilic interactions, b and phenotypic diversity, M. The only requirement is that the payoff to homophilic interactions, a, be positive.
. In other words, all homophilic individuals and even those individuals that weakly prefer heterophily tend to do better than those that strongly prefer heterophily. And this result is independent of the payoff to heterophilic interactions, b and phenotypic diversity, M. The only requirement is that the payoff to homophilic interactions, a, be positive.
Given constraints on mobility, it may be unrealistic to assume that individuals are equally likely to meet with all other members of the population7,33. In many species (including humans) interactions tend to be more likely between individuals of the same type because they are drawn to and/or drawn from similar environments. We therefore extended the basic model by introducing an additional parameter to allow for such assortativity. With probability 0 < φ < 1, individuals interact with others of the same phenotype and with probability 1 − φ they interact as before (see SI).
As it turns out, this extension generates an almost identical critical threshold, with homophily emerging when a > (1 − φ)Kb. Since φ < 1, this result shows that any degree of assortative matching that brings similar types into contact with one another more often makes it even more likely that homophily will evolve.
Another strong assumption of the basic model is that increasing a preference to interact with individuals of the same phenotype (p) yields an equal and opposite decrease in the preference to interact with individuals of different phenotypes (1 − p). Yet some individuals might want to interact with both similar and different phenotypes. We relaxed this assumption by allowing a separate preference q (independent of the preference p) to evolve that indicates the probability that an individual interacts with individuals of different phenotypes. Thus, each individual was characterised by a triplet {Gi, pi, qi} denoting phenotype, homophilic preference and heterophilic preference, respectively.
In this extended version of the model (see SI), we found that the average homophilic preference 〈p〉 was greater than the average heterophilic preference 〈q〉 in equilibrium if and only if a > Kb. Note that this is exactly the same condition that results from the basic model. Moreover, under the same condition (a > Kb), the population consists of more homophilic individuals (p > q) than the counterpart, heterophilic ones (p < q). In other words, the population tends to show more homophily than heterophily.
Finally, in all models, we assume natural selection is weak, which means that fitness differences are very small. Increasing the strength of natural selection relaxes this assumption and magnifies the fitness difference between traits. As a result, the evolutionary dynamics become increasingly deterministic and the critical point pc and the distribution of preferences in equilibrium become more skewed towards the extremes (p = 0 and p = 1).
Discussion
Our model differs from these prior studies that have taken into account assortativity in mating choice or ecological competition interactions. Among them, one study34 concerns whether a particular assortative mating choice of females is favoured for a fixed composition of phenotypes in the population. In this model, the mating choice is exclusively unilateral and up to females and the derived condition only gives the direction of the evolution at a given population composition. In contrast, interactions in our model are based on bilateral agreement, individuals' preferences and phenotypes are allowed to co-evolve and most importantly, the mutation-selection equilibrium is analytically derived that explicitly accounts for the abundance of any homophilic trait in the long run evolution. Some other previous studies, for example, consider the role of pre-existing assortative competition, but rather than the emergence of assortativity per se, in asexual35 or sexual36 selection contexts. Complementing and strengthening these prior works34,35,36, our model here explicitly addresses how homophily evolves in the first place.
Furthermore, our evolutionary model shows how homophily can emerge under a wide variety of conditions, particularly when mutation rates are low. It is not surprising that the payoff to homophilic interactions must be high relative to the payoff to heterophilic interactions in order for homophily to evolve. However, the analysis also suggests that this relationship is only relative, since the threshold for the ratio between these benefits can be less than one. This means that synergy may have a powerful effect on evolution, even when there are substantial benefits to specialization, helping to explain the ubiquity of homophily in nature. In higher-order species, the emergence of language or other forms of communication, or of certain cognitive capacities, might serve such a function and may help to promote a general tendency to seek out similar individuals with whom to cooperate or interact.
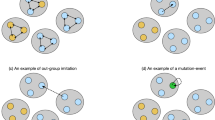
The model also shows that even small advantages to synergy can significantly reduce phenotypic diversity. Heterophilic populations maintain diversity by privileging rare phenotypes, generally causing their distribution to become uniform. Homophilic populations, on the other hand, privilege common phenotypes, helping to drive alternative phenotypes to extinction. Even if all phenotypes are themselves fitness neutral (as they are in our model), advantages to synergy will tend to yield populations dominated by a single phenotype and in the long run the population will tend to oscillate from one dominant phenotype to another, with rapid phase transitions in between.
Our results may also shed light on the observation that evolution in humans is accelerating37. The human capacity to collaborate not only with kin but also with unrelated members of our species may have dramatically increased the potential gains from synergy and this shift in payoffs would not only favour interactions with similar partners, but would also affect the overall desire to search out such partners. A wide variety of studies suggest that humans particularly seek out similar others6,38, even when there are no obvious benefits to these interactions39 (on the contrary, there may be more benefits to specialisation). Hence, it is possible that we evolved a strong predilection for homophily once we started to interact frequently with unrelated individuals. Such an effect would especially accelerate the evolution of phenotypes that are intrinsically synergistic, such as those related to communication or other collaborative activities.
Here, we focus on only one phenotypic dimension, but it is possible to extend our model to multiple dimensions of phenotypes (see SI). Payoffs to synergy and specialisation may vary for different sets of phenotypes, as each set serves a different function in social interactions. Therefore, multi-layer networks superimposed on various types of social interactions could result and each layer could play a different role. It is possible that in one layer ‘similarity attracts’, leading to stable, long-term relationships6, while in another layer ‘opposite attracts’, which often results in short-lived task-related ties8. In this way, natural selection could lead to the delicate, if very lopsided, balance between homophily and heterophily that we observe today in the real world.
Methods
In the Supplementary Information, we elaborate on the model presented above and analyse it, characterising equilibria under mutation-selection, explaining selection criteria, deriving a formula for continuous preferences and showing long form equations for the triplet correlations at neutrality that allow us to use coalescent theory to derive conditions for the evolution of homophily. We then describe how we derived the critical slopes and critical preferences shown in the main text and characterise equilibria for the whole parameter space. Finally, we turn our attention to several extensions of the model, analytically characterising models with biased matching, local mutation, strong selection, full strategy space and multiple sets of phenotypes.
We also describe the application of the model to derive estimates of homophily from empirical data, as shown in Table 1.
References
Sundaresan, S. R., Fischhoff, I. R., Dushoff, J. & Rubenstein, D. I. Network metrics reveal differences in social organization between two fission-fusion species, Grevys zebra and onager. Oecologia 151, 140–149 (2007).
Lusseau, D. & Newman, M. E. J. Identifying the role that animals play in their social networks. Proc. R. Soc. Lond. B 271, S477–S481 (2004).
Kutsukake, N., Suetsugu, N. & Hasegawa, T. Pattern, distribution and function of greeting behavior among black-and-white colobus. Int. J. Primatol. 27, 1271–1291 (2006).
Brent, L. J. H., Lehmann, J. & Ramos-Fernández, G. Social network analysis in the study of nonhuman primates: A historical perspective. Am. J. Primatol. 73, 720–730 (2011).
Madden, J. R., Drewe, J. A., Pearce, G. P. & Clutton-Brock, T. H. The social network structure of a wild meerkat population: 3. Position of individuals within networks. Behav. Ecol. Sociobiol. 65, 1857–1871 (2011).
McPherson, M., Smith-Lovin, L. & Cook, J. M. Birds of a feather: Homophily in social networks. Annu. Rev. Sociol. 27, 415–444 (2001).
Apicella, C. L., Marlowe, F. W., Fowler, J. H. & Christakis, N. A. Social networks and cooperation in hunter-gatherers. Nature 481, 497–501 (2012).
Moody, J. The structure of a social science collaboration network: Disciplinary cohesion from 1963 to 1999. Am. Sociol. Rev. 69, 213–238 (2004).
Fowler, J. H., Settle, J. E. & Christakis, N. A. Correlated genotypes in friendship networks. Proc. Natl. Acad. Sci. U.S.A. 108, 1993–1997 (2011).
Wakayama, T. et al. Heterophilic binding of the adhesion molecules poliovirus receptor and immunoglobulin superfamily 4A in the interaction between mouse spermatogenic and Sertoli cells. Biol. Reprod. 76, 1081–1090 (2007).
Kleene, R. & Schachner, M. Glycans and neural cell interactions. Nat. Rev. Neurosci. 5, 195–208 (2004).
Edelman, G. M. Cell adhesion and the molecular processes of morphogenesis. Ann. Rev. Biochem. 54, 135–169 (1985).
Kaplan, H. & Gurven, M. in Moral Sentiments and Material Interests: The Foundations of Cooperation in Economic Life. (eds Gintis, H., Bowles, S., Boyd, R. & Fehr, E.) (MIT Press, 2005) 75–113.
Schelling, T. Dynamic models of segregation. J. Math. Sociol. 1, 143–186 (1971).
Axelrod, R. The dissemination of culture: A model with local convergence and global polarization. J. Conflict Resol. 41, 203–226 (1997).
Currarini, S., Jackson, M. O. & Pin, P. An economic model of friendship: Homophily, minorities and segregation. Econometrica 77, 1003–1045 (2009).
Centola, D. An experimental study of homophily in the adoption of health behavior. Science 334, 1269–1272 (2011).
Bergstrom, T. C. The algebra of assortative encounters and the evolution of cooperation. Int. Game Theory Rev. 5, 1–18 (2003).
Rand, D. G., Arbesman, S. & Christakis, N. A. Dynamic social networks promote cooperation in experiments with humans. Proc. Natl. Acad. Sci. U.S.A. 108, 19193–19198 (2011).
Smith, A. An Inquiry into the Nature and Causes of the Wealth of Nations (University Of Chicago Press, 1977).
Wuchty, S., Jones, B. F. & Uzzi, B. The increasing dominance of teams in production of knowledge. Science 316, 1036–1039 (2007).
Fischer, C. S. & Oliker, S. J. A research note on friendship, gender and the life cycle. Soc. Forces 62, 124–133 (1983).
Traulsen, A., Shoresh, N. & Nowak, M. A. Analytical results for individual and group selection of any intensity. Bull. Math. Biol. 70, 1410–1424 (2008).
Nowak, M. A., Sasaki, A., Taylor, C. & Fudenberg, D. Emergence of cooperation and evolutionary stability in finite populations. Nature 428, 646–650 (2004).
Doebeli, M. & Hauert, C. Models of cooperation based on the Prisoners Dilemma and the Snowdrift game. Ecol. Lett. 8, 748–766 (2005).
Roca, C. P., Cuesta, J. A. & Sánchez, A. Time scales in evolutionary dynamics. Phys. Rev. Lett. 97, 158701 (2006).
Helbing, D. & Yu, W. The outbreak of cooperation among success-driven individuals under noisy conditions. Proc. Natl. Acad. Sci. U.S.A. 106, 3680–3685 (2009).
Gracia-Lazaro, C., Cuesta, J. A., Snchez, A. & Moreno, Y. Human behavior in Prisoners Dilemma experiments suppressesnetwork reciprocity. Sci. Rep. 2, 325 (2012).
Szolnoki, A., Perc, M. & Szabó, G. Defense mechanisms of empathetic players in spatial ultimatum game. Phys. Rev. Lett 109, 078701 (2012).
Wakeley, J. Coalescent Theory: An Introduction (Roberts & Company Publishers, 2008).
Antal, T., Ohtsuki, H., Wakeley, J., Taylor, P. D. & Nowak, M. A. Evolution of cooperation by phenotypic similarity. Proc. Natl. Acad. Sci. U.S.A. 106, 8597–8600 (2009).
Tarnita, C. E., Wage, N. & Nowak, M. A. Multiple strategies in structured populations. Proc. Natl. Acad. Sci. U.S.A. 108, 2334–2337 (2011).
Onnela, J. P., Arbesman, S., Gonzalez, M. C., Barabasi, A. L. & Christakis, N. A. Geographic constraints on social network groups. PLoS ONE 6, e16939 (2011).
Otto, S. P., Servedio, M. R. & Nuismer, S. L. Frequency-dependent selection and the evolution of assortative mating. Genetics 179, 2091–2112 (2008).
Dieckmann, U. & Doebeli, M. On the origin of species by sympatric speciation. Nature 400, 354–357 (1999).
M'Gonigle, L. K., Mazzucco, R., Otto, S. P. & Dieckmann, U. Sexual selection enables longterm coexistence despite ecological equivalence. Nature 484, 506–509 (2012).
Hawks, J., Wang, E. T., Cochran, G. M., Harpending, H. C. & Moyzis, R. K. Recent acceleration of human adaptive evolution. Proc. Natl. Acad. Sci. U.S.A. 104, 20753–20758 (2007).
Rivera, M. T., Soderstrom, S. B. & Uzzi, B. Dynamics of dyads in social networks: Assortative, relational and proximity mechanisms. Annu. Rev. Sociol. 36, 91–115 (2010).
Tajfels, H. Social psychology of intergroup relations. Annu. Rev. Psychol. 33, 1–39 (1982).
Acknowledgements
We are grateful for support from the National Institute on Aging (P-01 AG031093 and P30 AG034420), the John Templeton Foundation, the National Science Foundation/National Institute of Health joint program in mathematical biology (NIH grant no. R01GM078986) and the Bill and Melinda Gates Foundation (Grand Challenges grant 37874).
Author information
Authors and Affiliations
Contributions
F.F., M.A.N., N.A.C. and J.H.F. performed analyses, discussed the results and contributed to the text of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information for the evolution of homophily
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Fu, F., Nowak, M., Christakis, N. et al. The Evolution of Homophily. Sci Rep 2, 845 (2012). https://doi.org/10.1038/srep00845
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00845
This article is cited by
-
Assortativity in cognition
Scientific Reports (2023)
-
A new similarity measure to increase coverage of rating predictions for collaborative filtering
Applied Intelligence (2023)
-
Association patterns and community structure among female bottlenose dolphins: environmental, genetic and cultural factors
Mammalian Biology (2022)
-
Research collaboration for a knowledge-based economy: towards a conceptual framework
Triple Helix (2018)
-
Similar neural responses predict friendship
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.