Abstract
Human APOBEC3 proteins play pivotal roles in intracellular defense against viral infection by catalyzing deamination of cytidine residues, leading to base substitutions in viral DNA. Activation-induced cytidine deaminase (AID), another member of the APOBEC family, is capable of editing immunoglobulin (Ig) and non-Ig genes and aberrant expression of AID leads to tumorigenesis. However, it remains unclear whether APOBEC3 (A3) proteins affect stability of human genome. Here we demonstrate that both A3A and A3B can induce base substitutions into human genome as AID can. A3B is highly expressed in several lymphoma cells and somatic mutations occur in some oncogenes of the cells highly expressing A3B. Furthermore, transfection of A3B gene into lymphoma cells induces base substitutions in cMYC gene. These data suggest that aberrant expression of A3B can evoke genomic instability by inducing base substitutions into human genome, which might lead to tumorigenesis in human cells.
Similar content being viewed by others
Introduction
It is widely recognized that the accumulation of genetic changes in tumor-related genes is essential for cancer development1. With the innovation of high-throughput sequencing technology, genome-wide analyses on various types of cancer cells have revealed numerous somatic mutations in tumor-related genes2. Some of these mutations are caused by defects in DNA repair systems (e.g., DNA mismatch repair deficiencies give rise to hereditary non-polyposis colon cancer3), whereas mechanisms that account for the majority of genetic changes in cancer cells are poorly understood. Referring to somatic base substitution spectra in cancer cells, C/G to T/A transitions are most prevalent, especially in gastric cancer, colorectal cancer, glioma and melanoma2,4,5. This strong bias in somatic mutations suggests the existence of active mechanisms that induce C/G to T/A transitions into genomic DNA. It is obviously attributable to ultraviolet irradiation and following repair process against pyrimidine dimer in case of melanoma, but not in others.
The human APOBEC family proteins can induce C to T (G to A, in complementary sequences) transitions into target DNA through cytidine deamination. The APOBEC family is comprised of a series of molecules with conserved cytidine deaminase domains (CDAs), including AID, APOBEC1, APOBEC2, APOBEC3A to H and APOBEC46,7. Among them, AID plays a crucial role in somatic hypermutation and class switch recombination of Ig genes, which enables diversification of immune system8. AID has been considered the only molecule that can induce C/G to T/A transitions into genomic DNA. The expression of AID is highly regulated and restricted in germinal center B-cells under physiological conditions, but with inflammatory stimulations, AID can be overexpressed in not only B-cells but also other types of cells (e.g., epithelial cells) via activation of NF-κB9. Aberrant expression of AID results in the accumulation of mutations in non-Ig genes10, which leads to development of various cancers such as gastric and hepatic cancers as well as lymphomas9,11,12,13.
A series of seven A3 genes are tandemly arrayed on human chromosome 22 and the main function of the resulting gene products is to protect the cells from retroviruses and endogenous mobile retroelements14,15. A3B, A3D, A3F and A3G contain two CDAs, instead of one in A3A, A3C and A3H. A3G is a powerful anti-retroviral molecule that induces cytidine deamination in viral genome and acts as a host defensive factor against viruses such as HIV-116. A3A and A3B have been reported as potent inhibitors of retrotransposons17. Thus, A3 proteins act as sentinels in innate immunity against mobile DNA/RNA including viruses, while little is known about the effect of these proteins on nuclear DNA, in other words, host human genome. Recent studies have demonstrated that A3A impairs nuclear DNA under the condition of suppressing uracil DNA-glycosylase (UNG) which prevents base alterations by eliminating uracil from DNA and initiating the base-excision repair pathway18,19. However, it is still unclear whether A3 proteins can induce somatic mutations into human genome with intact DNA repair systems. Here we first demonstrate that expression of A3B and A3A as well as AID can induce somatic mutations in genomic DNA in human cells even in the presence of UNG. We also find that high expression of A3B leads to somatic mutations in tumor-related genes. These data suggest that aberrant expression of A3B might be one of the active mechanisms that induce somatic mutations in cancer cells.
Results
A3 and AID induce hypermutations into foreign DNA
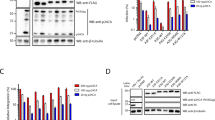
Besides A3A, we focused on A3B because it is localized predominantly in the nucleus20,21 and highly expressed in many types of cancer cells14 referring to microarray database (e.g., NextBio: http://www.nextbio.com). Previous studies have shown that A3B contains two enzymatically active CDAs in restricting HIV-122, whereas only carboxyl-terminal CDA is responsible for inhibiting HBV replication23,24 and editing bacterial DNA22. A3B is also shown to restrict foreign DNA in mammalian cells25, but it has not been tested which CDA is active in this context. First, to examine whether A3 and AID induce mutations in foreign DNA in human cells and which CDA is responsible for this DNA editing, we constructed amino- and/or carboxyl-terminal CDA mutants (H66R, H253R and H66/253R) by site directed mutagenesis (Fig. 1a) and confirmed their expression in HEK293 cells by immunoblotting (Fig 1b). We transfected expression vectors for these together with EGFP expression vector into HEK293 cells and examined base substitutions in EGFP sequences. The expression vector for UNG inhibitor (UGI) was also co-transfected to avoid UNG-triggered degradation of uracil-containing foreign DNA as described previously25. We recovered total DNA from the cells 2 days after transfection and performed differential DNA denaturation PCR (3D-PCR) to efficiently recover edited DNA sequences26. 3D-PCR is based on the principle that DNA sequences with fewer interstrand hydrogen bonds dissociates easier. If cytidine deamination takes place frequently, resulting AT-rich EGFP gene can be amplified at lower denaturation temperatures. Although PCR products were obtained from all samples at 92°C of denaturation temperature (Td), we obtained robust PCR products at 83.8°C of Td only from A3A-, A3B wild-type (WT)- and AID-expressing cells (Fig 2a). Amplification of EGFP at the lowest Td was impaired in H66R-expressing cells compared to A3B WT-expressing cells and undetectable in H253R- or H66/253R-expressing cells (Fig 2a, bottom). To ascertain whether EGFP gene was actually hyperedited, we cloned and sequenced the amplicons at 83.8°C of Td. As can be seen from the mutation matrices, high levels of C/G to T/A transitions were introduced into EGFP sequences (Fig 2b). To compare the extent of baseline mutations and that of A3B-induced mutations, we also cloned and sequenced the amplicons at 94.0°C of Td. Mutation frequency of A3B-expressing cells were about 6 times higher than that of mock-transfected cells (Supplementary Fig. S1 online). The mutation frequency in H66R-expressing cells was approximately a half compared to that in A3B WT-expressing cells in the amplicons at the lowest Td (Fig 2c). These data suggest that carboxyl-terminal CDA of A3B is mainly responsible for foreign DNA editing, but both domains are requisite for full editing activity. It is worth noting that AID is also capable of inducing cytidine deamination into foreign DNA.
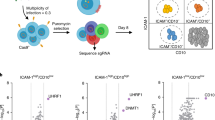
Expression of A3A, A3B wild-type and mutants and AID.
(A) Schematic of expression vectors. The consensus amino acid residues for zinc-coordinating motifs are shown. Substituted residues are shown in white. (B) Expression of HA-tagged proteins. Expression vectors were transfected into HEK293 cells and cell lysates were analyzed by immunoblotting with anti-HA antibody (top panel) and anti-ß-actin antibody (bottom panel) for loading control.
Foreign DNA editing by A3A, A3B and AID.
(A) Agarose gel analyses of 3D-PCR products from HEK293 cells. Cells were transfected with expression vector for A3A, A3B wild-type or mutant, or AID together with pEGFP-N3 and pEF-UGI. Total DNA was recovered 2 days after transfection and EGFP gene was amplified by 3D-PCR at the indicated denaturation temperatures (Td). (B) Mutation matrices of hyperedited EGFP sequences derived from cloned amplicons at 83.8°C of Td. “n” indicates the number of bases sequenced. We sequenced 5 clones (2,225 base pairs in total) in each group. (C) Frequencies of C/G to T/A transitions in hyperedited EGFP genes. C/G to T/A transitions per 1,000 sequenced base pairs are shown. (D) Dinucleotide contexts in foreign DNA editing. The rates of indicated dinucleotide sequence at the C to T transitions are shown. Asterisks indicate statistical significance in a χ2 test (p < 0.01).
Human A3 proteins have preferred target dinucleotide sequences in the substrate DNA; A3A and A3B prefer to deaminate cytosine residues flanked by 5′ thymine residue, 5′-TC, whereas A3G prefers to deaminate cytosine residues flanked by 5′ cytosine residue, 5′-CC25,27,28,29. We analyzed the context of C/G to T/A transitions in hyperedited EGFP sequences. We observed a strong bias toward deamination at 5′-TC dinucleotides in A3A-, A3B WT- and H66R-expressing cells, but not in AID-expressing cells (Fig 2d). 5′-TC dinucleotide preference of A3B was also confirmed by sequencing amplicon at 94.0°C of Td which is supposed to be unbiased (Supplementary Fig. S1 online). These data suggest that the preference of editing sites in foreign DNA by A3s coincides with that seen in viral DNA.
A3A and A3B can edit genomic DNA in human cells
We next investigated whether A3 proteins induce C/G to T/A transitions into not only foreign DNA but also nuclear DNA in human cells. We first established a HEK293 cell line stably expressing EGFP (HEK293/EGFP) using retrovirus vector that carries EGFP. We transfected HEK293/EGFP cells with expression vectors for A3A, A3B WT or mutant (H66R, H253R, or H66/253R), or AID by lipofection and then recovered total DNA from these cells after 7-day culture. We performed 3D-PCR of EGFP gene and obtained amplicons from A3A-, A3B WT-, H66R- and AID-expressing cells at lower Td (Fig 3a). EGFP gene was recovered at Td as low as 86.3°C from A3B WT-expressing cells, while as low as 86.5°C from A3A-, H66R- and AID-expressing cells. By contrast, EGFP gene was not amplified below Td of 87°C from cells transfected with mock, H253R or H66/253R. We repeated this procedure consisting of transfection, DNA extraction and 3D-PCR three times and obtained similar results (Fig 3b). To unambiguously confirm the presence of C/G to T/A transitions, we cloned and sequenced amplicons obtained at the lowest Td in A3A-, A3B WT-, H66R- and AID-expressing cells (Fig 3c). These analyses revealed approximately 2 to 5 C/G to T/A transitions per EGFP sequence from each sample (Fig 3d). The transitions were detected most frequently in A3A-expressing cells and deaminase activity of H66R mutant was approximately a half compared to that of A3B WT as seen in foreign DNA assays. The contexts of C/G to T/A transitions detected from the lowest Td amplicons in genomic DNA editing in A3-expressing cells were distinct from those in foreign DNA editing (Fig 3e). A preference for 5′-TC dinucleotide was not apparently observed, alternatively, 5′-GC dinucleotides were preferred in all samples. However, this bias fails to reach statistical significance (p < 0.01) in a χ2 test. The preferred target sequences of AID editing were 5′-GC and 5′-AC dinucleotides as described by many prior studies27,30. Mutation frequencies and preferred target sequence of A3B was also analyzed by using amplicons at 94.0°C of Td. Mutation frequency of A3B-expressing cells were about 3 times higher than that of mock-transfected cells (Supplementary Fig. S2 online). A preference for 5′-TC dinucleotide was impaired, compared to that in foreign DNA editing assays (Supplementary Fig. S2 online). Our results reveal that in addition to AID, A3A and A3B can induce C/G to T/A transitions into human nuclear DNA without repressing proofreading enzymes (e.g., UNG). Mutation frequencies were 6 to 9 per 1000 base pairs in A3A-, A3B WT- and AID-expressing cells and much less frequent compared to those in foreign DNA editing. As seen with foreign DNA editing, carboxyl-terminal CDA is mainly responsible for catalytic activity but not sufficient for full editing activity. The preference context of genomic DNA editing by A3A and A3B is different from that of viral or foreign DNA editing.
Hypermutations in EGFP genes integrated in genomic DNA of HEK293 cells.
(A) Agarose gel analyses of 3D-PCR products of EGFP genes extracted from HEK293/EGFP cells. Cells were transfected with expression vector for A3A, A3B wild-type or mutant, or AID. Total DNA was recovered 7 days after transfection and EGFP genes were amplified by 3D-PCR at the indicated denaturation temperatures (Td). (B) Distributions of the lowest denaturation temperatures for positive PCR amplification in each sample. Each circle represents independent experiment consisting of transfection, DNA extraction and 3D-PCR. (C) Mutation matrices of hyperedited EGFP sequences derived from cloned PCR products at Td lower than 87°C. “n” indicates the number of bases sequenced. We sequenced 25 clones (11,125 base pairs in total) in each group. (D) Frequencies of C/G to T/A transitions in hyperedited EGFP genes. C/G to T/A transitions per 1,000 sequenced base pairs are shown. (E) Dinucleotide contexts in genomic DNA editing. The rates of indicated dinucleotide sequence at the C to T transitions are shown. Deviations in the editing contexts do not reach statistical significance (p < 0.01) in a χ2 test.
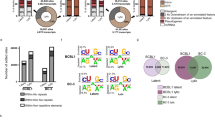
Deep sequencing reveals hyperediting of human genomic DNA by A3 proteins
Amplicon sequencing by next-generation sequencer has enabled to detect extremely low levels of mutations of targeted regions in genomic DNA. To verify more certainly that A3 proteins edit human nuclear DNA, we performed deep sequencing of A3-expressing cells. HEK293/EGFP cells were transfected with an empty vector or expression vectors for A3A, A3B WT, H66/253R, or AID by lipofection and total DNA were extracted after 7-day culture. We amplified a portion of EGFP gene with 443 base pair length (from thymine 56 to cytosine 498) by conventional PCR protocol, not by 3D-PCR and performed amplicon sequencing with the coverage of 1337 to 2654 reads per sample. This analysis revealed that extremely large numbers of nucleotides were substituted over the full length of amplicons in A3A-, A3B WT- and AID-expressing cells, whereas very few mutations were detected in mock and H66/253R-expressing cells (Supplementary Table 1 online). C/G to T/A transitions were observed most frequently in A3A-expressing cells as variation rates reach approximately 7% at the maximum, while below 3% at most in A3B- and AID-expressing cells (Fig 4a and Supplementary Table 1 online). The mutation frequency analysis revealed that large numbers of C/G to T/A substitutions were induced in A3A-, A3B- and AID-expressing cells, whereas other types of base substitutions were very few (Fig 4b). These results are similar to the data obtained by 3D-PCR and clonal sequencing of A3- and AID-expressing cells and further demonstrated that A3A and A3B as well as AID can induce C/G to T/A transitions into genomic DNA in human cells with intact DNA repair systems. Dinucleotide preference of target sequence for deamination by A3A, A3B and AID was also analyzed, however, we did not found any preference in this experiment (Fig. 4C), suggesting the difference between foreign DNA editing and genomic DNA editing.
Deep sequencing of EGFP genes in genomic DNA.
(A) The distributions of C/G to T/A substitutions in the EGFP sequences. Total DNA was recovered form HEK293/EGFP cells 7 days after transfection with expression vector for A3A, A3B wild type or H66/253R or AID. We amplified a portion of EGFP sequence from thymine 47 to cytidine 504 (top schematic) by PCR with high-fidelity polymerase and sequenced the amplicons by GS-junior bench top system (Roche). Sequence data were analyzed with equipped software. “Coverage” indicates the total numbers of sequenced reads. (B) Frequencies of base substitutions in hyperedited EGFP genes. Base substitutions were classified to 6 groups and substituted base number of each group per 1,000 sequenced base pairs are show. (C) Dinucleotide contexts in genomic DNA editing. The rates of indicated dinucleotide sequence at the C to T transitions are shown. Deviations in the editing contexts do not reach statistical significance (p < 0.01) in a χ2 test.
Expression of A3B and somatic mutations in lymphoma cells
Although AID has been reported to play important roles in lymphomagenesis by inducing mutations in both Ig and non-Ig genes11,12,31,32,33,34, AID-independent mechanisms are also suggested, because AID is not expressed in all types of B-cell lymphomas31,35. We hypothesized that A3 may contribute to somatic mutations in some lymphoma cells. To examine this hypothesis, we first determined expression levels of A3A, A3B and AID by quantitative RT-PCR in several B-cell lymphoma cell lines using peripheral blood lymphocytes (PBL) as control (Fig 5a). Our analysis revealed that A3B was highly expressed in 3 of 4 cell lines, particularly, markedly high in KIS1 cells, whereas expression of A3A transcripts was not detected in any lymphoma cell lines consistent with prior work suggesting myeloid specificity25,36. AID transcripts were detected in 2 of 4 cell lines, which is consistent with previous studies31,32,37. We also examined expression of A3B in two lymph node samples of diffuse large B-cell lymphoma and found that A3B is actually expressed (supplementary Fig. 3 online).
Expression of A3B and somatic mutations in oncogenes in human lymphoma cell lines.
(A) Quantitative RT-PCR for A3A, A3B and AID in lymphoma cell lines. The levels of target cDNA were normalized to the endogenous hypoxanthine phosphoribosyl transferase 1 (HPRT1) and then compared to those in peripheral blood lymphocytes. (B) Mutational analyses of C-myc, Pax5 and A20 in SUDH6 and KIS1 cells. We recovered total DNA from the cells and amplified the sequence between exon1 and intron1 of C-myc, Pax5 and A20 by PCR and performed direct sequencing of the amplicons. Locations of somatic mutations are shown below the loci with their positions. (C) The expression levels of transcripts of C-myc, Pax5 and A20 in KIS1 and SUDHL6 cells. Quantitative RT-PCR was similarly performed with (a).
To investigate the correlation between A3B expression and frequency of somatic mutations, we next performed direct sequencing of cMYC, PAX5 and A20 genes which are exemplary genes mutated frequently in B-cell lymphoma33,38. We compared mutation frequencies of these genes in SUDHL-6 and KIS-1, because the expression of A3B was the lowest in the former and the highest in the latter, while AID was not expressed in either cell line. DNA sequences between exon 1 and intron 1 of these three genes were analyzed (899 base pairs of cMYC, 1550 base pairs of Pax5 and 1088 base pairs of A20), since it have been reported that somatic mutations induced by cytidine deaminases were concentrated within 2 kb downstream from transcription initiation sites33,34. We found nine mutations within investigated sequences of cMYC and PAX5 in KIS-1, but not in SUDHL-6, in which five of nine mutations detected were C/G to T/A transitions. On the other hand, no mutation was detected within sequenced region of A20 in either cells (Fig 5b). To analyze ongoing mutations in the genome in individual cells, we next sequenced the same region of cMYC sub-cloned from KIS-1 and SUDHL-6 and found several more C to T mutations in KIS-1 cells, but not in SUDHL-6 cells (Supplementary Fig. S4 online). We next determined expression of these tumor-related genes by quantitative RT-PCR and found that the transcripts of cMYC and PAX5 were highly expressed in both SUDHL6 and KIS1 cells as compared to PBL, whereas A20 was less transcribed in these lymphoma cells. These results suggest that high expression of A3B resulted in accumulation of base alterations, especially C/G to T/A transitions, in actively transcribed tumor-related genes in lymphoma cells.
To ascertain more definitely that A3B can edit tumor-related genes in lymphoma cells, we introduced A3B into a lymphoma cell line and analyzed somatic mutations in cMYC. SUDHL-6 cells were transfected with expression vector for A3B WT, H66/253R, or mock by electroporation and total DNA was extracted after 7-day culture. With 3D-PCR analysis of cMYC, we obtained the amplicon from only A3B WT-expressing cells at the lower Td (Fig 6a). Clonal sequencing of amplicons at 85.9°C revealed 2 to 7 nucleotide substitutions per strand and more than 80% of these mutations were C/G to T/A transitions, with a preference for 5′-GC dinucleotide sites (Fig 6b and c). We also sequenced the amplicons at 94.0°C of Td and found A3B-induced C/G to T/A transitions without 5′-TC dinucleotide preference (supplementary Fig. 5 online). These data demonstrate that expression of A3B can induce somatic mutations into actively transcribed tumor-related genes in lymphoma cells.
A3B induced somatic mutations into c-myc gene in human lymphoma cells.
(A) Agarose gel analyses of 3D-PCR products of c-Myc genes in SUDHL6. We transfected expression vector for A3B wild-type or H66/253R or empty vector and recovered total DNA 7 days after transfection. C-myc genes were amplified by 3D-PCR at the indicated denaturation temperatures (Td). (B) Clonal sequencing of amplicons from A3B-WT expressing SUDHL6 cells. We sequenced 11 clones (5104 base pairs in total). Seventy six bases from thymine 310 to adenine 385 in which mutations are concentrated among sequenced 464 base pairs are shown. The numbers of C/G to T/A substitutions in sequenced 464 base pair length are shown at the right end. (C) Dinucleotide contexts of somatic mutations in c-Myc gene by A3B. The rates of indicated dinucleotide sequence at the C to T transitions are shown. Asterisks indicate statistical significance in a χ2 test (p < 0.01).
Discussion
To date, most studies on A3 proteins have focused on their abilities as antiviral or antitransposon factors, whereas the capability of A3 proteins to induce mutations into genomic DNA in host cells has been scarcely verified. In contrast, many studies have elucidated that AID induces somatic mutations into not only Ig genes, but also tumor-related genes in human cells and that ubiquitous expression of AID in mice leads to cancers of various organs as well as lymphomas, with the accumulation of nucleotides alterations9,10,11,12,13. Thus, AID has been considered as the only DNA cytosine deaminase that can induce somatic mutations into human genome and has potential to cause cancers or hematologic malignancies.
Suspène et al. have recently reported that hyperediting of both mitochondrial and nuclear DNA was detected in human cells defective for UNG derived from hyper IgM syndrome patients and demonstrated that A3A-induced mutations in nuclear DNA are detectable under the condition of suppressing UNG in human cells19. In their report, deamination of nuclear DNA was not observed in cells expressing other A3 proteins or in cells expressing A3A without UNG suppression. Furthermore, Landry et al. have reported that expression of A3A together with UGI in mammalian cell lines resulted in breaking of DNA and activation of DNA damage response in a deaminase-dependent manner18. In these two reports, the effect on genomic integrity by A3A was dependent on the presence of UGI. Thus, there has been no direct evidence that A3 proteins induce mutations in genomic DNA in the cells with intact DNA repair systems.
In this study, we demonstrate that A3A and A3B as well as AID can induce C/G to T/A transitions into nuclear DNA without suppressing UNG by two different assays, 3D-PCR and deep sequencing. We assume that increased number of cytidine deamination catalyzed by highly expressed A3A or A3B exceeded the processivity of DNA repair enzymes such as UNG and resulted in leaving C/G to T/A transitions in nuclear DNA. Mutation frequencies were considerably lower compared to Suspène's report. However, Yoshikawa et al. reported that AID induced hypermutations into an actively transcribed gene in fibroblasts and that the mutation frequency was approximately 4 to 6 per 1000 base pairs10, almost to the same extent as our results. Hence the frequency of mutations induced by cytidine deaminases into nuclear DNA is probably this extent or less in the cells with intact DNA repair systems.
We also find that A3B is highly expressed in several lymphoma cell lines and that the cells expressing high levels of A3B actually possesses somatic mutations, especially C/G to T/A transitions, in actively transcribed tumor-related genes. Furthermore, we reveal that introduction of A3B into lymphoma cells induces the accumulation of C/G to T/A transitions in cMYC gene. This is the first report that suggests the involvement of A3B in inducing somatic mutations of oncogenes in tumor cells. Together with the microarray database of A3B expression in miscellaneous cancer cell lines14 (NextBio: http://www.nextbio.com), it is possible that A3B may induce somatic mutations into tumor related genes in various types of cancers.
Several questions remain open. First, it remains unclear what is preferred target sequences of A3 proteins in genomic DNA. Because in several cancers such as breast cancer and melanoma, 5′-TC is the most prevalent target in C to T base substitutions, A3 is the most potential candidate to induce these mutations25,28. However, in our results, neither A3A nor A3B had a definite preference of editing site in nuclear DNA editing, whereas a preference for 5′-TC dinucleotide was observed in foreign DNA editing as previously reported. It is possible that A3A and A3B have no distinct favorite context in nuclear DNA editing, unlike viral and bacterial DNA editing, because human genomic DNA is more profoundly protected in transcription than viral or bacterial DNA and is under survey of DNA repair systems. However, Suspene et al. reported that target contexts of cytidine deamination in A3A+UGI-expressing cells were 5′-TC and 5′-CC dinucleotides, which were identical to the contexts of viral or bacterial DNA editing19. We assume that this discrepancy might be attributable to cell types or expression levels in cells. Hence, further analyses should be required to clarify the favorite target contexts of A3 proteins in nuclear DNA editing. The second question is how transcriptional control and post-translational modification of A3 proteins regulate A3 activity. Because the molecules that possess a capability of editing nuclear DNA threaten cell homeostasis, expression and activity of A3A and A3B must be strictly controlled. AID is known to be regulated at multiple steps39, for example, transcriptional regulation40,41,42, post-transcriptional regulation by micro-RNA43,44, regulation of intracellular localization45,46 and phosphorylation by PKA47,48. In contrast to AID, little is known about how A3 proteins are regulated. It has been reported that A3A is abundantly expressed in CD14+ monocytes and upregulated by interferon-α stimulation25,49,50. Meanwhile, it is not clear where A3B is expressed normally25,42,49,50 and how it is regulated. As for post-translational modification, we previously reported that PKA-mediated phosphorylation of A3G regulates the interaction between A3G and HIV Vif51. To better understand the physiological roles of A3 proteins, it is important to elucidate how their expression and activity are regulated. The last question is whether A3 proteins can serve as an “initiator” of tumorigenesis. Our results suggest A3B indeed induces somatic mutations into genomic DNA in various human tumor cells, however, it is unclear whether A3B proteins impair genomic DNA from the early stage of oncogenesis. To address this question, hereafter, histopathological and genetic analyses of transgenic mouse constitutively expressing A3 proteins are necessary.
In conclusion, our findings provides the first evidence that A3A and A3B can induce C/G to T/G transitions into genomic DNA without suppressing DNA repair system. Our data also show that high expression of A3B is related to mutation frequencies of oncogenes in lymphoma cells. Our results suggest that A3B is an oncogene, like AID, which may have the capacity to evoke genomic instability through base substitutions in human cells. Further studies will be required to test whether endogenous A3B is capable of impairing genomic integrity as a DNA mutator and contributing to the development of human cancers and hematologic malignancies.
Methods
DNA constructs and cell lines
Plasmids containing coding sequence of human A3A and A3B were kindly provided by Dr. Kenzo Tokunaga21. Expression vectors for HA-tagged A3A, A3B and AID were generated by sub-cloning of coding sequences into pCAG-GS vector. A3B catalytic domain mutants (H66R, H253R and H66/253R) were generated by KOD-plus mutagenesis Kit (Toyobo). Expression vector for Uracil-DNA glycosylase inhibitor, pEF-UGI was kindly provided by Dr. Ruben S Harris25. HEK293 and HEK293T cells were maintained with Dulbecco's modified Eagle's medium containing 10% of fetal bovine serum (FBS) and penicillin, streptomycin and glutamine (PSG). All B-cell lymphoma cell lines were maintained with RPMI1640 containing 10% FBS and PSG. Retrovirus containing EGFP sequence was produced by co-transfection of pMLV gag-pol, VSV-G and pDON-EGFP into HEK293T cells. HEK293/EGFP cells were generated by retroviral transduction of EGFP and selection of 1 mg/ml G418 for two months.
Immunoblotting
HEK293 cells were transfected with expression vector for A3A, A3B wild-type or mutant (H66R, H253R or H66/253R) or AID and lysed with RIPA buffer (50 mM Tris-HCl pH7.5, 150 mM NaCl, 1 mM EDTA, 1% Triton-X, 0.1% SDS, 0.1% DOC) after 2-day culture. After centrifugation at 20,000 x g for 15 min, supernatant was mixed with sample buffer (Biorad), boiled for 5 minutes, resolved on 12% (w/v) polyacrylamide gel, transferred to PVDF membrane (Immobilon, Millipore) and analyzed by standard immunoblotting procedure with anti-HA monoclonal antibody (12CA5, Roche) or anti-β-actin monoclonal antibody (AC-15, Sigma).
3D-PCR and clonal sequencing
For foreign DNA editing assay, HEK293 cells were transfected with pEGFP-N3, pEF-UGI and expression vector for A3A, A3B WT or mutant, or AID by using Fugene HD (Roche). After two-day culture, total DNA was extracted by using Quick Gene DNA whole blood kit (Fuji Film). First round PCR was performed with primers listed in Supplementary Table S2 using rTaq DNA polymerase (Takara), with the following reaction profile; 30 s at 94°C, 25cycles of 30 s at 94°C, 40 s at 62°C and 90 s at 72°C followed by 10 min at 72°C. The amplicons were separated by electrophoresis on 1% (w/v) agarose gel and extracted from the gel using Qiaquick Gel Extraction kit (Qiagen). We used 25 ng of first-round PCR products as template for nested PCR using Hotstar Hifidelity DNA polymerase (Qiagen), with the following reaction profile; 5 min at 95°C, 35 cycles of 15 s at 83–92°C, 60 s at 62°C, 80 s at 72°C, followed by 10 min at 72°C. The amplicons derived at 83.8°C were cloned into pT7-blue vector (Novagen). For nuclear DNA editing assay, HEK293/EGFP cells were transfected with expression vector for A3A, A3B WT or mutants, or AID using Fugene HD (Roche). Seven days after transfection, we extracted total DNA from these cells with the same method of foreign DNA editing assay. First round PCR was performed using Advantage HF2 polymerase kit (Clontech), with the following reaction profile;1 min at 94°C, 30 cycles of 30 s at 94°C followed by 2 min at 68°C, followed by 3 min at 68°C. We used 25 ng of first-round PCR products for nested PCR using Hotstar Hifidelity DNA polymerase (Qiagen) with the following reaction profile; 5 min at 95°C, 35 cycles of 15 s at 86–89°C, 60 s at 62°C and 80 s at 72°C, followed by 10 min at 72°C. The amplicons derived at 86.5°C and 83.8°C were cloned into pT7-blue vector (Novagen). For c-myc gene editing assay in lymphoma cells, we transfected SUDHL6 cells with expression vectors for A3B WT or H66/253R by electroporation using Nucleofector (Amaxa) and extracted total DNA from the cells 7 days after transfection. First round PCR and gel extractions of amplicons were performed with the same methods of nuclear DNA editing assay. We used 25 ng of first-round PCR products for nested PCR using Hotstar Hifidelity DNA polymerase (Qiagen), with the following reaction profile; 5 min at 95°C, 35 cycles of 15 s at 85–88°C, 60 s at 62°C, 80 s at 72°C, followed by 10 min at 72°C. The amplicons derived at 85.3°C were cloned into pT7-blue vector (Novagen) and sequenced using 3130xl Genetic Analyzer (Applied Biosystems).
Deep sequencing
Total DNA was extracted from HEK293/EGFP transfected with expression vectors for A3A, A3B WT or H66/253R or AID 7 days after transfection. A portion of EGFP gene with 443 base pair length, from thymine 56 to cytosine 498, was amplified with the primers listed in Supplementary Table 2 using Advantage HF2 polymerase kit (Clontech), with the following reaction profile; 1 min at 94°C, 30 cycles of 30 s at 94°C followed by 2 min at 68°C and followed by 3 min at 68°C. The amplicons were separated by electrophoresis on 1% (w/v) agarose gel and extracted from the gel using Quiaquick Gel Extraction Kit (Qiagen). Purified amplicons were sequenced using GS junior bench top system (Roche) according to the manufacturer's protocol and analyzed with equipped software, GS Amplicon Variant Analyzer.
Lymphoma cell lines and patient samples
Four B-cell lymphoma cell lines (SUDHL6, KIS1, KM-H2 and Granta519) were cultured in RPMI1640 containing 10% of fetal bovine serum (FBS) and penicillin, streptomycin and glutamine (PSG). We extracted total DNA from these cells by using Quick Gene DNA whole blood kit (Fuji Film) and total RNA by using mir Vana miRNA isolation kit (Ambion). Tumor biopsy specimens prior to treatment were obtained from two patients with diffuse large B-cell lymphoma. The study was approved by the Kyoto University Institutional Review Board and written informed consent was obtained from each patient. Total RNA were extracted similarly to lymphoma cell lines. Naïve B-cells were isolated from healthy donor's peripheral blood by using MACS® naive B cell isolation kit (Miltenyi Biotec).
Quantitative RT-PCR
Complementary DNA was synthesized from 200 ng of total RNA using Revertra Ace qPCR RT Master Mix (Toyobo). Real-time PCR were performed with Thunderbird SYBR qPCR Mix (Toyobo) according to manufacturer's protocol. Target cDNAs were normalized to the endogenous expression level of the house keeping reference gene for hypoxanthine-guanine phosphoribosyl transferase 1 (HPRT1) or glyceraldehyde 3-phosphate dehydrogenase (GAPDH). All primers for real-time PCR are listed in Supplementary Table 2.
Sequencing of oncogenes from lymphoma cell lines
We amplified portions of C-myc, Pax5 and A20 from with the primers listed in Supplementary Table 2 using Advantage HF2 polymerase kit (Clontech), with the following reaction profile; 1 min at 94°C, 30 cycles of 30 s at 94°C followed by 4 min at 68°C and followed by 3 min at 68°C. The amplicons were separated by electrophoresis on 1% (w/v) agarose gel, extracted from the gel using Qiaquick Gel Extraction kit (Qiagen) and sequenced using 3130xl Genetic Analyzer (Applied Biosystems). In c-Myc clonal sequencing, the amplicon was subcloned into pTA2-vector (TOYOBO) and subsequently sequenced.
References
Hahn, W. C. & Weinberg, R. A. Rules for making human tumor cells. N Engl J Med 347, 1593–603 (2002).
Pleasance, E. D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, 191–6 (2010).
Bronner, C. E. et al. Mutation in the DNA mismatch repair gene homologue hMLH1 is associated with hereditary non-polyposis colon cancer. Nature 368, 258–61 (1994).
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–8 (2007).
Prickett, T. D. et al. Analysis of the tyrosine kinome in melanoma reveals recurrent mutations in ERBB4. Nat Genet 41, 1127–32 (2009).
Macduff, D. A. & Harris, R. S. Directed DNA deamination by AID/APOBEC3 in immunity. Curr Biol 16, R186–9 (2006).
Conticello, S. G. The AID/APOBEC family of nucleic acid mutators. Genome Biol 9, 229 (2008).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–63 (2000).
Matsumoto, Y. et al. Helicobacter pylori infection triggers aberrant expression of activation-induced cytidine deaminase in gastric epithelium. Nat Med 13, 470–6 (2007).
Yoshikawa, K. et al. AID enzyme-induced hypermutation in an actively transcribed gene in fibroblasts. Science 296, 2033–6 (2002).
Okazaki, I. M. et al. Constitutive expression of AID leads to tumorigenesis. J Exp Med 197, 1173–81 (2003).
Pasqualucci, L. et al. AID is required for germinal center-derived lymphomagenesis. Nat Genet 40, 108–12 (2008).
Endo, Y. et al. Activation-induced cytidine deaminase links between inflammation and the development of colitis-associated colorectal cancers. Gastroenterology 135, 889–98, 898 e1-3 (2008).
Jarmuz, A. et al. An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics 79, 285–96 (2002).
Goila-Gaur, R. & Strebel, K. HIV-1 Vif, APOBEC and intrinsic immunity. Retrovirology 5, 51 (2008).
Mangeat, B. et al. Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature 424, 99–103 (2003).
Bogerd, H. P. et al. Cellular inhibitors of long interspersed element 1 and Alu retrotransposition. Proc Natl Acad Sci U S A 103, 8780–5 (2006).
Landry, S., Narvaiza, I., Linfesty, D. C. & Weitzman, M. D. APOBEC3A can activate the DNA damage response and cause cell-cycle arrest. EMBO Rep 12, 444–50 (2011).
Suspène, R. et al. Somatic hypermutation of human mitochondrial and nuclear DNA by APOBEC3 cytidine deaminases, a pathway for DNA catabolism. Proc Natl Acad Sci U S A 108, 4858–63 (2011).
Lackey, L. et al. APOBEC3B and AID have similar nuclear import mechanisms. J Mol Biol 419, 301–14 (2012).
Kinomoto, M. et al. All APOBEC3 family proteins differentially inhibit LINE-1 retrotransposition. Nucleic Acids Res 35, 2955–64 (2007).
Bogerd, H. P., Wiegand, H. L., Doehle, B. P. & Cullen, B. R. The intrinsic antiretroviral factor APOBEC3B contains two enzymatically active cytidine deaminase domains. Virology 364, 486–93 (2007).
Suspène, R. et al. Extensive editing of both hepatitis B virus DNA strands by APOBEC3 cytidine deaminases in vitro and in vivo. Proc Natl Acad Sci U S A 102, 8321–6 (2005).
Bonvin, M. & Greeve, J. Effects of point mutations in the cytidine deaminase domains of APOBEC3B on replication and hypermutation of hepatitis B virus in vitro. J Gen Virol 88, 3270–4 (2007).
Stenglein, M. D., Burns, M. B., Li, M., Lengyel, J. & Harris, R. S. APOBEC3 proteins mediate the clearance of foreign DNA from human cells. Nat Struct Mol Biol 17, 222–9 (2010).
Suspène, R., Henry, M., Guillot, S., Wain-Hobson, S. & Vartanian, J. P. Recovery of APOBEC3-edited human immunodeficiency virus G->A hypermutants by differential DNA denaturation PCR. J Gen Virol 86, 125–9 (2005).
Vartanian, J. P. et al. Massive APOBEC3 editing of hepatitis B viral DNA in cirrhosis. PLoS Pathog 6, e1000928 (2010).
Bishop, K. N. et al. Cytidine deamination of retroviral DNA by diverse APOBEC proteins. Curr Biol 14, 1392–6 (2004).
Harris, R. S. et al. DNA deamination mediates innate immunity to retroviral infection. Cell 113, 803–9 (2003).
Beale, R. C. et al. Comparison of the differential context-dependence of DNA deamination by APOBEC enzymes: correlation with mutation spectra in vivo. J Mol Biol 337, 585–96 (2004).
Greeve, J. et al. Expression of activation-induced cytidine deaminase in human B-cell non-Hodgkin lymphomas. Blood 101, 3574–80 (2003).
Deutsch, A. J. et al. MALT lymphoma and extranodal diffuse large B-cell lymphoma are targeted by aberrant somatic hypermutation. Blood 109, 3500–4 (2007).
Pasqualucci, L. et al. Hypermutation of multiple proto-oncogenes in B-cell diffuse large-cell lymphomas. Nature 412, 341–6 (2001).
Kotani, A. et al. A target selection of somatic hypermutations is regulated similarly between T and B cells upon activation-induced cytidine deaminase expression. Proc Natl Acad Sci U S A 102, 4506–11 (2005).
Smit, L. A. et al. Expression of activation-induced cytidine deaminase is confined to B-cell non-Hodgkin's lymphomas of germinal-center phenotype. Cancer Res 63, 3894–8 (2003).
Chen, H. et al. APOBEC3A is a potent inhibitor of adeno-associated virus and retrotransposons. Curr Biol 16, 480–5 (2006).
Mottok, A., Hansmann, M. L. & Bräuninger, A. Activation induced cytidine deaminase expression in lymphocyte predominant Hodgkin lymphoma. J Clin Pathol 58, 1002–4 (2005).
Kato, M. et al. Frequent inactivation of A20 in B-cell lymphomas. Nature 459, 712–6 (2009).
Delker, R. K., Fugmann, S. D. & Papavasiliou, F. N. A coming-of-age story: activation-induced cytidine deaminase turns 10. Nat Immunol 10, 1147–53 (2009).
Crouch, E. E. et al. Regulation of AID expression in the immune response. J Exp Med 204, 1145–56 (2007).
Gonda, H. et al. The balance between Pax5 and Id2 activities is the key to AID gene expression. J Exp Med 198, 1427–37 (2003).
Pauklin, S., Sernández, I. V., Bachmann, G., Ramiro, A. R. & Petersen-Mahrt, S. K. Estrogen directly activates AID transcription and function. J Exp Med 206, 99–111 (2009).
Teng, G. et al. MicroRNA-155 is a negative regulator of activation-induced cytidine deaminase. Immunity 28, 621–9 (2008).
de Yébenes, V. G. et al. miR-181b negatively regulates activation-induced cytidine deaminase in B cells. J Exp Med 205, 2199–206 (2008).
Ito, S. et al. Activation-induced cytidine deaminase shuttles between nucleus and cytoplasm like apolipoprotein B mRNA editing catalytic polypeptide 1. Proc Natl Acad Sci U S A 101, 1975–80 (2004).
Patenaude, A. M. et al. Active nuclear import and cytoplasmic retention of activation-induced deaminase. Nat Struct Mol Biol 16, 517–27 (2009).
Basu, U. et al. The AID antibody diversification enzyme is regulated by protein kinase A phosphorylation. Nature 438, 508–11 (2005).
McBride, K. M. et al. Regulation of class switch recombination and somatic mutation by AID phosphorylation. J Exp Med 205, 2585–94 (2008).
Koning, F. A. et al. Defining APOBEC3 expression patterns in human tissues and hematopoietic cell subsets. J Virol 83, 9474–85 (2009).
Berger, G. et al. APOBEC3A is a specific inhibitor of the early phases of HIV-1 infection in myeloid cells. PLoS Pathog 7, e1002221 (2011).
Shirakawa, K. et al. Phosphorylation of APOBEC3G by protein kinase A regulates its interaction with HIV-1 Vif. Nat Struct Mol Biol 15, 1184–91 (2008).
Acknowledgements
We thank R. Harris for thoughtful feedback and for UGI vector and K. Tokunaga for A3A and A3B vectors.
Author information
Authors and Affiliations
Contributions
M.S. performed most of the experiments, analyzed the data and wrote the manuscript; I.K., M.M., T.S. and K.T. performed sequencing of lymphoma cell lines; K.S. and N.K. wrote the manuscript; M.K. established q-PCR; A.T.K. designed the experiments and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Materials
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Shinohara, M., Io, K., Shindo, K. et al. APOBEC3B can impair genomic stability by inducing base substitutions in genomic DNA in human cells. Sci Rep 2, 806 (2012). https://doi.org/10.1038/srep00806
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00806
This article is cited by
-
Research on the influence of APOBEC family on the occurrence, diagnosis, and treatment of various tumors
Journal of Cancer Research and Clinical Oncology (2023)
-
Multiple roles of apolipoprotein B mRNA editing enzyme catalytic subunit 3B (APOBEC3B) in human tumors: a pan-cancer analysis
BMC Bioinformatics (2022)
-
Impact of the APOBEC3A/B deletion polymorphism on risk of ovarian cancer
Scientific Reports (2021)
-
Alternative splicing of APOBEC3D generates functional diversity and its role as a DNA mutator
International Journal of Hematology (2020)
-
Endogenous APOBEC3B Overexpression Constitutively Generates DNA Substitutions and Deletions in Myeloma Cells
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.