Abstract
We have interrogated a 12-chemokine gene expression signature (GES) on genomic arrays of 14,492 distinct solid tumors and show broad distribution across different histologies. We hypothesized that this 12-chemokine GES might accurately predict a unique intratumoral immune reaction in stage IV (non-locoregional) melanoma metastases. The 12-chemokine GES predicted the presence of unique, lymph node-like structures, containing CD20+ B cell follicles with prominent areas of CD3+ T cells (both CD4+ and CD8+ subsets). CD86+, but not FoxP3+, cells were present within these unique structures as well. The direct correlation between the 12-chemokine GES score and the presence of unique, lymph nodal structures was also associated with better overall survival of the subset of melanoma patients. The use of this novel 12-chemokine GES may reveal basic information on in situ mechanisms of the anti-tumor immune response, potentially leading to improvements in the identification and selection of melanoma patients most suitable for immunotherapy.
Similar content being viewed by others
Introduction
Skin cancers are the most common of all human malignancies. Although melanoma constitutes a minority of skin cancers, it is considered to be the most deadly. The treatment of primary melanoma is predominantly surgical. Once melanoma has metastasized to lymph nodes, radical surgery to remove the nodes in the affected basin is performed, often followed by immunotherapy and sometimes by radiation. Melanoma metastatic to organ sites beyond the regional lymph nodes requires, in most cases, systemic therapy. Melanoma is notoriously insensitive to standard chemotherapy drugs that are widely used for other forms of cancer. However, melanoma is occasionally and sometimes dramatically, responsive to immunotherapy.
Immunotherapy is fast becoming an important part of the treatment armamentarium for advanced melanoma, but the degree of its clinical effectiveness varies among patients. In spite of the clinical success observed with antibodies against CTLA-4 and PD-11,2,3,4, cytokines (e.g., high-dose IL-24), as well as the adoptive transfer of tumor infiltrating lymphocytes4,5, dramatic responses are observed in a minority of patients, while the majority of patients treated with those agents do not show significant clinical benefit. This current limitation underscores the need for the discovery of immune-related biomarkers and gene expression signatures that can identify/predict the likelihood of melanoma patients achieving a robust response of prolonged duration to immunotherapy.
It is becoming widely accepted that immune cell infiltrates in human melanoma and other solid tumors have prognostic value6,7. Many of these studies described the subset composition of the lymphocytic infiltrates that were either distributed diffusely within tumor parenchyma or localized diffusely to the peripheral rim of the mass and the tumor/stroma interface. In addition, recent attention has been focused on defining and potentially utilizing a so-called “immune score” based on the types and degrees of immune cell infiltrates for classification of human cancer8.
We have been studying the intriguing observation that ectopic lymph node-like structures reside within the parenchyma of some solid tumor masses9,10. Our initial work focused on invasive colorectal carcinoma (CRC)11. We found that a metagene (i.e. several tightly correlated sets of genes) score for inflammation and immunity accurately predicted host immune reaction when applied to this tumor type. Indeed, we identified, through the use of a 12-chemokine GES, a subset of CRCs that contained unique, tumor-localized ectopic lymph node-like structures (TL-ELNs). This molecular classifier and the presence of these lymphoid structures were associated with better patient outcome, irrespective of other defined clinical and biologic variables. Based on their known selective capacity to potently recruit and concentrate immune cell subsets in vivo and also their known association with peripheral lymph node formation and function12,13,14, it is possible that chemokines locally produced within the microenvironment of the tumor mass are responsible for the formation of these ectopic lymph node-like structures. It is also conceivable that these lymph nodal structures are active and are playing an important, positive role in eliciting an endogenous anti-tumor immune response (initially local but becoming systemic) that results in improved patient survival.
In the current study, the 12-chemokine GES has been interrogated on genomic arrays of 14,492 distinct solid tumors (primaries and metastases) and is shown to have broad distribution across different histologies. Further, we show that this signature can, in fact, identify the presence of unique, TL-ELNs in melanoma metastases, which also appears to associate with better patient outcome. The implication of these findings for current and future immunotherapies for melanoma is discussed.
Results
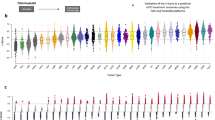
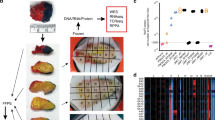
We had earlier identified in colorectal cancer a unique 12-chemokine (CCL2, CCL3, CCL4, CCL5, CCL8, CCL18, CCL19, CCL21, CXCL9, CXCL10, CXCL11 and CXCL13) GES from a metagene grouping with overwhelming enrichment for immune-related and inflammation-related genes11. In the current work, we interrogated this signature on 14,492 distinct solid tumors (primaries and metastases) with at least 30 per tumor type using gene chip technology. For this study, we employed principal component analysis (PCA), calculating a PCA score based on the first principal component - PC1 - that approximates an average Z-score based on the log2 intensities for each of the 12 genes in the chemokine GES. Figure 1 shows the distribution of the 12-chemokine GES across different solid tumor histologies.
Interrogation of a 12-chemokine gene expression signature across solid tumors of differing histology: Principal component analysis (PCA).
Gene expression microarrays on 14,492 samples (primaries and metastases) were examined and total PCA score was derived as a measure of chemokine signal. The top 10% of samples were selected as high. For this study, we calculated a PCA score based on PC1 that approximates an average Z-score based on the log2 intensities for each of the 12 chemokine genes in the gene expression signature. RNA was extracted from tumors frozen within 15 minutes of surgical removal (according to strict standard operating procedures). *The 569 skin tumor samples include non-melanomas (i.e. basal cell, squamous cell and Merkel cell carcinomas), primary melanomas, as well as 120 non-locoregional metastases from stage IV patients. **Compared to all other samples, RNA was extracted from paraffin of suboptimal amounts of tumor tissue.
Gene expression for the 12 chemokine genes was combined using PCA to generate a chemokine signature score for 120 individual, stage IV patient non-locoregional metastasis samples (Figure 2a.). The first principal component (~57% variance) is used to represent the chemokine signature. Melanomas with a high chemokine signature showed increased expression values in all 12 chemokine genes when compared to melanomas with a low chemokine signature. Statistical tests of differences among the 12 genes confirmed the PCA results, indicating up-regulation of gene expression (p value ranges, 3.25e-05 to 2.22e-16, Figure 2b.). Melanomas that had the highest (n = 9; red) or lowest (n = 12; green) chemokine signatures were identified (Figure 2c.).
Melanoma metastasis gene expression of 12 chemokines.
(a.) Gene expression for the 12 chemokine genes (13 Affymetrix probe sets) was combined using PCA. The scatter plot consists of melanoma samples plotted by the first two principal components. The first component, representing 57% of the variability within samples, was used as the chemokine signature score (GSE) for 120 non-locoregional melanoma metastases samples. (b.) The melanoma samples with high chemokine signature scores (n = 9) showed significantly increased expression values in all 12 chemokine genes (13 probe sets) when compared to melanomas with low chemokine signature (n = 12), as shown by mean and standard deviation of each gene, together with p value for differences. (c.) Patients whose melanoma samples had the very highest (n = 9, red) or lowest (n = 12, green) chemokine signatures are shown in the heatmap with the expression of the 12 chemokine genes (13 probe sets). Samples are labeled by rank order; samples 1–12 have low chemokine GSE and samples 112–120 have high GSE.
Microscopically, representative tissue sections of resected metastatic melanoma specimens from patients with the lowest 12-chemokine gene scores revealed, in all cases, a minimally to absent lymphocytic peritumoral host response, with low to no appreciable expression of lymphocytic markers (not shown). Conversely, the highest 12-chemokine gene-scoring melanoma samples revealed, in all cases, a marked peritumoral lymphocytic host response consistent with ectopic lymph node-like structures, evident by H&E staining (Figure 3a.,b.,c.) and by IHC (Figure 4 a.,b.,c.,d.), particularly at the interface between tumor and stroma. Ectopic lymph node-like structures were observed intratumorally as well, occasionally accompanied by a low number of tumor infiltrating lymphocytes. The tumor-localized ectopic lymph node-like structures (TL-ELNs) were found to contain prominent lymphoid follicles. As shown in Figure 4, CD20+ B cells were present almost exclusively within the follicular structures. The majority of the CD3+ T cells (both CD4+ and CD8+ subsets) were concentrated in parafollicular cortex-like zones; scattered T cells were also seen within these tumors. CD86+ antigen presenting/dendritic cells were present within the structures as well; however, FoxP3+ cells (putative T regulatory cells) were found to be very few in numbers.
Presence of ectopic lymph node-like structures in melanoma.
The nine highest 12-chemokine gene expression signature scored melanomas revealed a marked peritumoral lymphocytic host response, organized as ectopic lymph node-like structures by H&E staining (a.,b.,c.). Arrows (a. and b.) depict individual ectopic lymph node-like structures; Circle (b.) depicts an area of numerous ectopic lymph node-like structures.
Representative ectopic lymph node-like structures in melanoma examined by immunohistochemistry.
CD20+ B cells are concentrated as a follicle, with CD3+, CD4+ and CD8+ T cells appearing in the parafollicular cortex or marginal zones and with some dispersion into the follicle. CD86+ antigen presenting cells appear to be dispersed throughout the ectopic lymph node-like structure whereas Fox P3+ staining cells are few in numbers (a.,b.,c.). A melanoma metastasis displaying numerous CD20+ B cell, CD3+ T cell-containing, ectopic lymph node-like structures is also shown (d.). Magnification: x200.
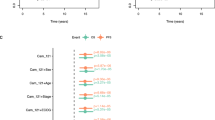
Clinical information was available on 10 of the 21 selected melanoma patients. Table 1 shows the overall survival of these patients and the 12-chemokine GES that identifies the presence or absence of TL-ELNs for these 10 cases. There was a highly significant (p = 0.008) association between increased survival and the value of the mean score of the 12-chemokine GES. We also obtained descriptive information between the score of the 12-chemokine GES, site of the metastasis and treatment received, which did not appear to show associations. The patients were those who predominantly had their metastatic lesions resected to render them free of disease and for whom long survival can often be seen, especially from the time of the primary diagnosis and death. Of interest, one patient with a long survival duration had received immunotherapy with an anti-CTLA4 antibody (ipilimumab) and has had a partial response of over 30 months duration.
Discussion
We have hypothesized that immune gene-related expression signatures can predict the presence of unique histological features of lymphoid cell infiltrates in solid tumor masses that correlate with better overall patient survival. In the current pilot study, we have shown that a 12-chemokine GES can accurately identify the presence of unique, TL-ELNs in metastatic melanoma, which also appear to be associated with better patient outcome. We focused on these melanoma cases in particular because such a patient population are considered as a candidate to receive immune-based therapies. Moreover, this signature has now been interrogated on genomic arrays of 14,492 distinct primary and metastatic solid tumors (Figure 1), raising the intriguing possibility that our findings, earlier in CRC and now in melanoma, may extend to other histologies as well. The use of the 12-chemokine GES was logistically easier, more cost-effective and far less time-consuming to predict the presence of TL-ELNs than histologic analysis based on our experience with CRC. In the earlier study, the presence of TL-ELNs often required analysis of numerous stained tumor sections, sometimes requiring multiple cuts through the paraffin block for positive identification of lymphoid infiltrates consistent with these structures. Remarkably, this was not the case for the melanoma metastases in the current study, which required only a single cut section.
Tumor infiltrating lymphocytes can have profound anti-tumor activity in melanoma4,5, so we are in the process of determining the functional nature of the TL-ELNs in the laboratory. These structures have the general appearance of “typical” peripheral lymph nodes, albeit much smaller in size and are constructed of the necessary immune components, including lymphatics (not shown), CD86+ antigen presenting cells and B cell follicles with adjacent T cell (CD4+ and CD8+) zones. Of interest, FoxP3+ cells (putative T regulatory cells) were found to be very few in numbers. In this regard, notably absent from the 12-chemokine GES are CCL1, CCL20 and CCL22, which have been shown to selectively recruit and/or maintain T regulatory cells15,16,17,18. The functional immune nature of the TL-ELNs with respect to local (and perhaps also systemic) tumor response and antigen(s) reactivity remains unknown. We plan to employ laser capture microdissection with DNA and RNA isolation and sequencing methodologies to address questions of clonality and functional relationships of the resident B cells and T cells within the tumor microenvironment19,20, including in depth T cell receptor and immunoglobulin transcript repertoire analyses. The presence of these unique lymphoid structures also appears to be associated with better survival, but this observation will require corroboration in larger, prospectively-defined sets of patients.
In spite of considerable progress made to date, it is well known that the majority of melanoma patients treated with immune-based therapies do not achieve a response or show clinical benefit. We believe that our findings raise the intriguing possibility of eventually selecting patients for immunotherapy interventions based on immune-related GES from their tumor21. We plan to evaluate the capacity of the 12-chemokine GES to predict the likelihood of patients achieving a response of prolonged duration to immune checkpoint inhibitory antibodies (e.g., anti-PD-1 and anti-CTLA-4) and/or cytokines (e.g., high-dose IL-2) – first, retrospectively and, later if appropriate, prospectively. We will also attempt to extend our findings to other solid tumor types (e.g., colon, lung and ovarian carcinomas), with the intent of possibly broadening the use of immunotherapies beyond melanoma. Lastly, the use of novel immune-related GES may reveal basic information about the mechanism of the anti-tumor effects underlying successful immune-based therapies, leading to improvements in current immune checkpoint antibody therapy and in the identification and ex vivo expansion of tumor infiltrating lymphocytes for broader adoptive immunotherapeutic application.
Methods
mRNA microarray analysis
Tumors from patients treated at the Moffitt Cancer Center under a University of South Florida Institutional Review Board-approved protocol were arrayed on Affymetrix HuRSTA-2a520709 GeneChips (Affymetrix, Santa Clara, CA). The chips contain ~60,000 probe sets representing ~25,037 unique genes (Affymetrix HuRSTA-2a520709, GEO: http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GPL10379). The expression data for the 14,492 distinct solid tumors were normalized using MAS5 and expression data for the 12 chemokine genes (13 probe sets) were extracted. The first principal component, PC1, (explaining ~53%) was calculated using the EvinceTM software (www.umbio.com). The tumor samples were grouped by tissue type and the distribution of the PC1 score was calculated. The expression data for the 120 individual patients, stage IV non-locoregional metastasis samples were normalized using RMA and expression data for the 12 chemokine genes (13 probe sets) were extracted. A principal component analysis (PCA) was performed from these 13 probe sets. The PC1, representing the most variability (~57%) within the metastatic melanoma samples, was used to represent the chemokine signature. Based on PC1, samples with values >+4 (9 patient samples) or <−4 (12 patient samples) were identified as the high and low expressers of the chemokine signature, respectively. A heat map was generated using Cluster 3.0 with unsupervised, hierarchical method and then visualized using Java Treeview on expression data of 12 chemokine genes (13 probe sets) in 21 selected samples. The two-sided Student's t test and Bonferroni correction were used to test for differences in high (n = 9) and low (n = 12) gene expresseion across the genes in the chemokine signature. PCA was performed and visualized using EvinceTM.
Selection of human tissues
The histological sections corresponding to 21 cases (prepared from the mirror-image of the portion of tumor submitted for the mRNA microarray analysis) were retrieved from the Moffitt Cancer Center Anatomic Pathology Division's repository. All of the specimens were preserved in 10% buffered formalin before assessing the presence of microscopically evident host immune response. The final pathology report for each case was also reviewed and the pathological data were collected. Linked, annotated but deidentified clinical data (i.e. survival) and treatments received were also available in the database. A single, random representative section through all selected tissue blocks was evaluated for the presence or absence of dense peritumoral infiltrates consistent with TL-ELNs. The tissue sections of melanoma metastases were stained using the avidin-biotin complex method with retrieval under high pH. Prediluted monoclonal antibodies (mAb) to CD3, CD4, CD8, CD20, CD86 and FoxP3 (Ventana Medical Systems, Tucson, AZ) were used for the analysis of lymphoid infiltrates. The slides were scanned using the Aperio™ ScanScope XT (Vista, CA) with a 200x/0.8NA objective lens at a rate of 2 minutes per slide via Basler tri-linear-array. These digital slides are housed in the Aperio server located in the Moffitt Network Operations Center and are available via the password protected web-based Spectrum database package. This ensures rapid quality control of images and analysis by all investigators. Clinical information is accessible only to the PI and authorized collaborators and all samples are anonymously coded prior to analysis.
Statistical analysis
Significance of linkages of the 12-chemokine GES profile to patient survival was analyzed statistically by the exact Wilcoxon rank sum test. For data presented in Figure 1, significance was analyzed by Fisher's exact test.
References
Sarnaik, A. A. & Weber, J. S. Recent advances using anti-CTLA-4 for the treatment of melanoma. Cancer J. 15, 169–173 (2009).
Weber, J. S. Immune checkpoint proteins: A new therapeutic paradigm for cancer-Preclinical background: CTLA-4 and PD-1 blockade. Semin. Oncol. 37, 430–439 (2010).
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nature Rev. Cancer 12, 252–264 (2012)
DeVita, Jr, V. T. Rosenberg SA: Two hundred years of cancer research. N. Engl. J. Med. 366, 2207–2214 (2012).
Rosenberg, S. A. et al. Durable complete responses in heavily pretreated patients with metastatic melanoma using T-cell transfer immunotherapy. Clin. Cancer Res. 17, 4550–4557 (2011).
Jochems, C. & Schlom, J. Tumor-infiltrating immune cells and prognosis: the potential link between conventional cancer therapy and immunity. Exp. Biol. Med. 236, 567–579 (2011).
Fridman, W. H. et al. Immune infiltration in human cancer: prognostic significance and disease control. Curr. Top. Microbiol. Immunol. 344, 1–24 (2011).
Galon, J. et al. Editorial: The immune score as a new possible approach for the classification of cancer. J. Transl. Med. 10, 1–4 (2012).
Coppola, D. & Mulé, J. J. Ectopic lymph nodes within human solid tumors. J. Clin. Oncol. 26, 4369–4370 (2008).
Dubinett, S. M., Lee, J. M., Sharma, S. & Mulé, J. J. Chemokines: can effector cells be re-directed to the site of tumor? Cancer J. 167, 325–335 (2010).
Coppola, D. et al. Unique ectopic lymph node-like structures present in human colorectal carcinoma are predicted by immune gene array profiling. Amer. J. Pathol. 179, 37–45 (2011).
Kirk, C. J., Hartigan-O'Connor, D. & Mulé, J. J. The dynamics of the T-cell antitumor response: chemokine-secreting dendritic cells can prime tumor-reactive T cells extranodally. Cancer Res. 61, 8794–8802 (2001).
Balkwill, F. Cancer and the chemokine network. Nat. Rev. Cancer 4, 540–550 (2004).
Charo, I. F. & Ransohoff, R. M. The many roles of chemokines and chemokine receptors in inflammation. N. Engl. J. Med. 354, 610–621 (2006).
Hoelzinger, D. B., Smith, S. E., Mirza, N., Dominguez, A. L., Manrique, S. Z. & Lustgarten, J. Blockade of CCL1 inhibits T regulatory cell suppressive function enhancing tumor immunity without affecting T effector responses. J. Immunol. 184, 6833–6842 (2010).
Mira, E. et al. Statins induce regulatory T cell recruitment via a CCL1 dependent pathway. J. Immunol. 181, 3524–3534 (2008).
Yamazaki, T. et al. CCR6 regulates the migration of inflammatory and regulatory T cells. J. Immunol. 181, 8391–8401 (2008).
Toulza, F. et al. Human T-lymphotropic virus type 1-induced CC chemokine ligand 22 maintains a high frequency of functional FoxP3+ regulatory T cells. J. Immunol. 185, 183–189 (2010).
Cheng, J. et al. Ectopic B-cell clusters that infiltrate transplanted human kidneys are clonal. Proc. Natl. Acad. Sci. USA 108, 5560–5565 (2011).
Yakirevich, E. et al. Analysis of T-cell clonality using laser capture microdissection and high-resolution microcapillary electrophoresis. J. Mol. Diagn. 9, 490–497 (2007).
Yeatman, T., Mulé, J. J., Dalton, W. S. & Sullivan, D. On the eve of personalized medicine in oncology. Cancer Res. 68, 7250–7252 (2008).
Acknowledgements
This work was supported in part by the NCI-NIH (1 R01 CA148995-01 to JJM), the V Foundation and the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation. We thank the Moffitt Total Cancer CareTM (TCC) team for their efforts in creating and updating the tumor genomic and clinical databases, as well as Dr. William Dalton for his vision of TCC.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: JLM, JJM. Performed the experiments: JLM, MCL. Analyzed the data, including biostatistics: MJS, DAM, SE, XQ, AEB, JSW. Contributed materials/specimens: VKS. Wrote the paper: JLM, JSW, SE, VKS, JJM.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Messina, J., Fenstermacher, D., Eschrich, S. et al. 12-Chemokine Gene Signature Identifies Lymph Node-like Structures in Melanoma: Potential for Patient Selection for Immunotherapy?. Sci Rep 2, 765 (2012). https://doi.org/10.1038/srep00765
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00765
This article is cited by
-
How chemokines organize the tumour microenvironment
Nature Reviews Cancer (2024)
-
Biomarkers for immune checkpoint inhibition in sarcomas – are we close to clinical implementation?
Biomarker Research (2023)
-
Tertiary Lymphoid Structures Are Associated with a Favorable Prognosis in High-Grade Serous Ovarian Cancer Patients
Reproductive Sciences (2023)
-
Machine learning and bioinformatics analysis revealed classification and potential treatment strategy in stage 3–4 NSCLC patients
BMC Medical Genomics (2022)
-
Avelumab maintenance in advanced urothelial carcinoma: biomarker analysis of the phase 3 JAVELIN Bladder 100 trial
Nature Medicine (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.