Abstract
An evolutionary tree of key enzymes from the Complex-Iron-Sulfur-Molybdoenzyme (CISM) superfamily distinguishes “ancient” members, i.e. enzymes present already in the last universal common ancestor (LUCA) of prokaryotes, from more recently evolved subfamilies. The majority of the presented subfamilies and, as a consequence, the Molybdo-enzyme superfamily as a whole, appear to have existed in LUCA. The results are discussed with respect to the nature of bioenergetic substrates available to early life and to problems arising from the low solubility of molybdenum under conditions of the primordial Earth.
Similar content being viewed by others
Introduction
A vast number of enzymes rely on metal cofactors for catalysis and/or redox conversions. Different types of such “metalloenzymes” appear to have evolutionary roots which reach more or less deeply into the distant past of life on Earth. The varying depths of evolutionary pedigrees likely reflect constraints imposed by paleogeochemistry such as abundances and solubilities of metals in specific environments and eons of our planet's history1. Iron-sulphur cluster-containing ferredoxins, for example, probably go back to the very origin of life2. Copper enzymes, by contrast, are argued to have emerged only after the oxygenation of the environment some 2.5 billion years ago due to its insolubility under anaerobic conditions3.
Whereas the details of iron's and copper's involvement in numerous biological reactions have been studied for more than a century, the precise role of other metals, although recognized as vital trace elements in enzyme catalysis, became elucidated only recently. Molybdenum (Mo) has during the last 2 decades been shown to constitute an essential cofactor in at least 3 distinct enzyme superfamilies4, the most widespread of which is the so-called Complex Iron-Sulfur Molybdoenzyme (CISM) superfamily of molybdo-pterin containing enzymes. Incidentally, this denomination ignores the fact that a few members of the family use tungsten (W) instead of molybdenum in their active sites. In the periodic table of elements, tungsten lies directly below molybdenum in the d-block and is thus expected to feature chemical properties related to those of Mo.
As for the case of copper, chemists and geochemists have argued that molybdenum is unlikely to have played a biological role prior to the advent of oxygenic photosynthesis5. This is in conflict with the molecular phylogeny of the CISM-superfamily which we present below.
Results
Individual enzymes from the CISM superfamily catalyze an astonishing variety of reactions with diverse substrates, some of which will be discussed in more detail below. Furthermore, many individual subfamilies comprise representatives from both Bacteria and Archaea. Setting out to reconstruct the composite phylogeny of a large part of the superfamily therefore implies dealing with strongly diverging sequences, as well as a high chance for the presence of several paralogs in the same species. Obtaining inferences from reconstructed phylogenies of enzyme superfamilies presents two major obstacles6,7,8,9. (1) The reliability of conventional multiple alignment algorithms frequently breaks down when strongly divergent sequences are analyzed. (2) The presence of several paralogs had led to erroneous annotations in the databases blurring the evolutionary message suggested by the trees.
We have in the past turned to assisting multiple alignments by 3D structure comparisons6 which in many cases substantially improved alignment quality. The annotation problem was addressed by taking into account all available phylogenetic markers such as operon organization or functionally important amino acid residues7 to arrive at as reliable as possible enzyme identifications. The case of the CISM enzymes exacerbates the mentioned adversities for phylogenetic reconstruction due to their extraordinary variability in enzyme function and wide species distribution. Extensive resorting to methods remedying these problems therefore proved crucial for arriving at sufficiently reliable phylogenetic tree topologies.
Redressing annotations and multiple alignments
Amino acid sequences of a representative member from each enzyme family for which a 3D structure is available were used as queries for BLAST searches on completed and draft genomes in the NCBI-database. The cases of ethylbenzene dehydrogenase and acetylene hydratase were, despite the presence of 3D structures, not included in this analysis since only very few members of the respective families have been identified so far. For the sake of balanced weights of individual enzyme subtrees, the Nap and Nas families of nitrate reductases were omitted. The composite tree of the nitrate reductase enzymes was published previously9 and inclusion of Nap and Nas does not alter the topology of the Nar subtree9. In the absence of a 3D structure for a given family, i.e. arsenate reductase (Arr) for the ensemble of considered subfamilies, the member best characterized with respect to biochemistry and enzymatics was chosen as query.
Obtained hits were analyzed with respect to their genomic context, i.e. the ordering of flanking genes (coding for further subunits of the enzyme complexes) in corresponding gene clusters (as previously described7), the nature of residues known to play critical roles in the respective catalytic reactions of the various enzyme families and structural idiosyncrasies of individual families6. In all retrieved cases, phylogenetic clustering coincided with these three additional phylogenetic markers. Many retrieved sequences turned out to be incorrectly annotated (see Table_sequences.xls in Supplementary Information).
When using automated multiple alignment algorithms, positions of roots within subtrees varied greatly with the set of sequences considered due to sequence-sample-dependent fluctuations in alignments. A close inspection of these alignments revealed a significant lack of robustness for nearly half of the total sequence length. The presence of extensive insertions/deletions (indels) characterizing individual families obviously hampers reliable alignments between subfamilies. To overcome this problem, we have resorted to structural alignments of the catalytic Molybdenum-subunit based on available 3D structures. X-ray structures have in fact been reported for members of 5 out of 6 of the enzyme families dealt with in this work, i.e. nitrate reductase (pdb entry 1Q16), arsenite oxidase (1G8J), polysulfide reductase (2VPZ), dimethylsulfoxide (DMSO) reductase (4DMR) and formate dehydrogenase (1KQF). As described previously for the case of the Rieske protein superfamily6, Mo-subunits were structurally superimposed using DeepView.
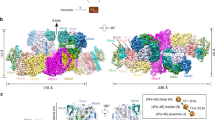
The severity of discrepancies between the structural approach and conventional alignment methods will be exemplified by a pair of enzymes, nitrate reductase (Nar) and polysulfide reductase (Psr). Figure 1A shows the 3D structural alignment of these two enzymes with Nar in purple and Psr in blue and green. The green structural elements in Psr denote the parts of the sequence where automated and 3D alignments yield equivalent results. Fig. 1B schematically indicates the deviations between sequence-only (Clustal and T-Coffee) and the structure-based alignments for the Nar/Psr pair. Among the automated alignment algorithms tested (Clustal, Muscle, T-Coffee and Mafft), T-Coffee scored slightly superior to the other methods (see Fig. 1B) while still aligning more than half of the sequence incorrectly. The structural alignment approach thus indeed yields results substantially different from those suggested by the automated alignment algorithms in sequence areas which do not feature very high sequence homology, that is, in the major part of the protein.
(A) Structural superposition of the catalytic Molybdenum subunits from respiratory nitrate reductase (Nar, light and dark purple) and polysulfide reductase (Psr, blue and green). (B) Schematic sequence alignments between Nar and Psr. The second and third lines represent the alignment obtained fully automatically by ClustalX and T-Coffee whereas the bottom-line alignment is based on the structural information gathered from (A) and further processed as described above. In both (A) and (B), light purple stretches indicate parts of the Nar polypeptide chain for which no structural equivalents exist in Psr. The same applies for the light blue sequences stretches in Psr. The green stretches denote the part of the sequences for which Clustal- or T-Coffee- and structure-based alignments coincide.
The multiple sequence alignment of stretches for which the structural approach has shown structural equivalence was subsequently refined using ClustalX and Seaview. The family for which no X-ray structure has so far been reported, i.e. Arr, was first individually submitted to a multiple alignment procedure and the obtained alignment result subsequently profile-aligned to the bulk of structurally aligned sequences.
In summary, if remedying erroneous annotations was omitted, “messy” trees deviating from species phylogenies, very much like the one we obtain for the Arr enzyme, were obtained for the whole superfamily misleadingly indicating extensive lateral gene transfer. Exclusive reliance on sequence-based multiple alignments, on the other hand, resulted in roots sliding into either the archaeal or the bacterial domains in several subfamilies (e.g. in Nar), which would pretend a post-LUCA emergence of the corresponding enzyme followed by inter-domain transfer.
These two observations may rationalize the deviance of conclusions presented in a recent purely bioinformatic study of evolutionary histories of almost 4000 gene families10 from our results to be presented below. This analysis10 suggested a late appearance of Mo-based enzymes and in particular Nar (and consequently the denitrifying pathway). This latter conclusion is in conflict with molecular phylogenies of other enzymes involved in this pathway8,9 and appears unlikely in the light of several paleogeochemical data11,12,13 which suggest large scale abiotic generation and hence abundance of nitrogen oxides, the substrates of the denitrifying pathway, in the Hadean and early Archaean. Moreover, another bioinformatic survey of enzyme superfamilies also indicated the presence of the CISM-superfamily significantly before the Archaean/Proterozoic transitions14. However, this latter study does not allow conclusions to be drawn as to the absence or presence of these enzymes in LUCA.
In this context, we furthermore need to note that the vast majority of previously published phylogenies on the CISM superfamily suffers from the experimental flaw that in calculating trees the possibility of multiple substitutions was not taken into account. This approach would be valuable for relatively closely related sequences but is certainly erroneous when members from different subfamilies in both prokaryotic domains are compared to each other. The resulting trees suffer from weak resolution in deep branching and therefore preclude the kind of analysis we present in this work.
Composite phylogenetic tree of CISM superfamily enzymes
The phylogenetic tree encompassing several CISM subfamilies based on the 3D-structure assisted multiple alignment and taking multiple substitutions into account is shown in Fig. 2. Whereas specific subfamilies (see below) indeed appear to have originated only at a later stage, the majority of subfamilies show distinctive features indicating their presence in the last universal common ancestor (LUCA), that is, well before the rise of O2 in the environment1. Figure 2A shows a schematic representation of the composite phylogeny of several CISM-enzymes and the detailed tree is available as Supplementary Information.
(A) 3D structure-based NJ-phylogenetic tree (see Supplementary Information) of the CISM-enzymes nitrate reductase (Nar), DMSO/TMAO reductase (Dms/Dor/Tor), arsenate reductase (Arr), formate dehydrogenase (Fdh), arsenite oxidase (Aro) and polysulfide reductase (Psr).Violet and orange denote eury- and crenarchaeal branches, dark green, cyan and light green stand for Proteobacteria, Firmicutes and other Bacteria, respectively. Open and closed dots indicate bootstrap values for the deep branchings exceeding 70 and 90%, respectively. In 3D structures, the Mo-subunit is in violet. (B) Redox potential range of relevant substrates and corresponding enzymes.
It is noteworthy that a given clade is tacitly assumed to correspond to an enzyme subfamily with a unique function. This inference is supported for the Nar and the Fdh clades by experimental evidence showing that archaeal and bacterial representatives perform the same chemical reaction. The presence of clade-specific particularities of substrate binding-site residues in all clades as mentioned above further corroborates the one-clade/one-enzyme notion.
Discussion
The clades representing DMSO/TMAO reductase (Dms/Dor/Tor) and arsenate reductase (Arr) contain only Bacteria and furthermore strongly diverge from 16S rRNA-based species trees. They therefore likely are late-emerging enzymes distributed predominantly via horizontal gene transfer. The clades corresponding to formate dehydrogenase (Fdh), polysulfide reductase (Psr), arsenite oxidase (Aro) and nitrate reductase (Nar) by and large resemble species trees, feature a prominent Archaea/Bacteria cleavage and their roots fall in between the archaeal and bacterial subtrees. The combined occurrence of these features strongly suggests the enzymes making up these clades to have been present in LUCA. The structural unit of the CISM protein thus appears to have served multiple purposes for life, especially in energy harvesting, right from its very beginnings. These results allow inferences on the available geochemical energy substrates. CISM enzymes in LUCA likely performed energy conversion through the reduction of carbon dioxide, polysulfide or nitrate as well as from the oxidation of arsenite. Reduction of CO2 and sulfur with H2 as electron donor would be viable bioenergetic pathways in the geochemical setting of the early Archaean and have indeed been put forward as ancestral bioenergetic mechanisms15,16. Bioenergetic oxidation of arsenite in the early Archaean has been proposed previously. The latter process requires sufficiently oxidizing e− acceptors which may have been nitrogen oxides (ref. 8 and references cited therein), a scenario supported by the pre-LUCA character of Nar (Fig. 2), a key enzyme of the denitrification pathway. The ensemble of pre-LUCA bioenergetic CISM enzymes in fact suggests that LUCA's energy conserving processes tapped into electrochemical disequilibria approaching 1 V (Fig. 2B) and furthermore bears witness to the astonishing redox versatility of the Mo-cofactor.
This redox versatility in part certainly arises from the fact that molybdenum and tungsten are 2-electron redox compounds, that is, they can shuttle between the +4/+5 and the +5/+6 redox couples. However, it is precisely the property of Mo and W to feature 2-electron transitions which allows these two elements to perform energetically challenging redox conversions. As detailed in references 17 and 18, several 2-electron compounds under certain circumstances feature so-called crossed-over individual redox transitions which allows them to redox bifurcate electrons with one of the two reducing equivalents going seemingly uphill towards very low potential electron acceptors18. Specific members of the CISM family have been shown to feature crossed-over redox potentials19. It is tempting to hypothesize that the possibility to redox bifurcate electrons may play a crucial role in energetically challenging redox reactions such as the reduction of CO2 to formate in aceto- and methanogens. Both these reactions indeed rely on members of the CISM superfamily.
Apart from informing on the nature of primordial energy sources, the presence of the CISM superfamily in LUCA implies a vital role of its metal cofactors in early life. As mentioned, a few members of the superfamily use W rather than Mo and a few “Mo-enzymes” have been shown to insert W under Mo-depletion conditions or at high temperatures20. However, many other members have turned out to be specific for Mo.
What then of the availability of these two transition metals? W occurs in both acid and alkaline solutions and was thus available to emerging life21, whereas Mo is relatively insoluble in reduced and neutral waters5, but does occur in mixed valence sulfide and selenide and/or oxide complexes in alkaline solutions. Mo's insolubility at neutral pH values, exacerbated by an anoxic atmosphere1, suggested a low bioavailability of this element for early life1,3. Mo-isotope analyses on samples from the Archaean era indeed show substantially lower levels than during Phanerozoic times22. Two scenarios can reconcile the results of molecular phylogeny and paleogeochemistry. (i) The ancestral CISM enzyme exclusively used W which was later replaced by Mo. (ii) CISM-catalyzed reactions in early life used Mo supplied by alkaline hydrothermal vents, proposed as cradles for life23. The exclusiveness for Mo of many CISM-members as well as findings that primary productivity involving Mo has been comparable to the present since the geological record began at 3.8 Ga24 lead us to favor the second scenario.
The results described above thus require that certainly tungsten and most likely molybdenum ought to be added to the list of metals vital already to earliest life on Earth24. Both Mo and W are heavier than Fe and thus aren't bred in standard stellar fusion reactions. Their nucleosynthesis requires the much higher energies only attained in rapidly spinning gas giants and during supernova explosions. Intriguingly, astrophysical data indicate a crucial role of a nearby supernova in the birth of our solar system25. The origin of life on our and any other wet rocky world may thus have been assisted by (and potentially depended on) supernova-generated elements.
Methods
Sequence retrieval and multiple alignment
Amino acid sequences of CISM superfamily members were retrieved via BLAST searches on the NCBI-database (http://www.ncbi.nlm.nih.gov) using biochemically or structurally characterized representatives of the subfamilies nitrate reductase (Nar), polysulfide reductase (Psr), formate dehydrogenase-N (Fdh), DMSO reductase (Dms/Dor/Tor), arsenite oxidase (Aro) and arsenate reductase (Arr) as queries. The number of hits used for each major species phylum was limited in order to avoid unreadable trees induced by the strong oversampling of specific classes of prokaryotes in the databases (e.g. proteobacteria or pathogenic actinobacteria). Gene cluster context was assayed by analyzing the genomic environment of each sequence on the respective genome web-pages of the NCBI server.
Multiple alignments of recognized subfamilies were automatically produced using Clustal26, MEGA27, Mafft (http://mafft.cbrc.jp/alignment/software/about.html), Muscle28 and T-Coffee29 and in cases subsequently refined with respect to functionally conserved residues using Seaview30. The second and third line sequence alignments in Fig. 1B were performed using Clustal and T-Coffee, respectively.
The ensemble of enzymes for which 3D structures are available were aligned via DeepView31 (www.expasy.org/spdbv/; see Figure and alignment file in Supplementary Information)
Phylogenetic tree reconstruction
NJ-trees were reconstructed using ClustalX26 and MEGA-427. These two programs yielded tree topologies differing only in a few very late branching orders. The tree shown in Fig. 2A and available as phb-file in Supplementary Information has been obtained using ClustalX. Gap positions were taken into account. Multiple substitutions were allowed for using Kimura's correction algorithm26. Calculated bootstrap values correspond to the frequency of occurrence of nodes in 1,000 bootstrap replicates.
References
Anbar, A. D. Elements and evolution. Science 322, 1481–1482 (2008).
Eck, R. V. & Dayhoff, M. O. Evolution of the structure of ferredoxin based on living relics of primitive amino acid sequences. Science 152, 363–366 (1966).
Williams, R. J. P. & Frausto da Silva, J. J. R. Evolution was chemically constrained. J. Theor. Biol. 220, 323–343 (2003).
Rothery, R. A., Workun, G. J. & Weiner, J. H. The prokaryotic complex iron-sulfur molybdoenzyme family. Biochim. Biophys. Acta 1778, 1897–1929 (2008).
Williams, R. J. P. & Frausto da Silva, J. J. R. The involvement of molybdenum in life. Biochem. Biophys. Res. Commun. 292, 293–299 (2002).
Lebrun, E. et al. The Rieske protein: a case study on the pitfalls of multiple sequence alignments and phylogenetic reconstruction. Mol. Biol. Evol. 23, 1180–1191 (2006).
Duval, S., Ducluzeau, A.-L., Nitschke, W. & Schoepp-Cothenet, B. Enzyme phylogenies as markers for the oxidation state of the environment: the case of the respiratory arsenate reductase and related enzymes. BMC Evol. Biol. 8, 206–219 (2008).
Ducluzeau, A.-L. et al. Was nitric oxide the first deep electron sink? Trends Biochem. Sci. 34, 9–15 (2009).
van Lis, R., Ducluzeau, A.-L., Nitschke, W. & Schoepp-Cothenet, B. in Nitrogen Cycling in Bacteria: Molecular Analysis (ed. Moir, J. W. B.) 1–21 (Caister Academic, Norwich/UK., 2011).
David, L. A. & Alm, E. J. Rapid evolutionary innovation during an Archaean genetic expansion. Nature 469, 93–96 (2011).
Martin, R. S., Mather, T. A. & Pyle, D. M. Volcanic emissions and the early Earth atmosphere. Geochim. Cosmochim. Acta 71, 3673–3685 (2007).
Summers, D. P. & Khare, B. Nitrogen fixation on Mars and other terrestrial planets: experimental demonstration of abiotic fixation reactions to nitrite and nitrate. Astrobiology. 7, 333–341 (2007).
Kasting, J. F. Bolide impacts and the oxidation state of carbon in the Earth's early atmosphere. Orig. Life Evol. Biosph.20, 199–231 (1990).
Dupont, C. L., Butcher, A., Valas, R. E., Bourne, P. E. & Caetano-Anollés, G. History of biological metal utilisation inferred through phylogenomic analysis of protein structures. Proc. Natl. Acad. Sci. USA 107, 10567–10572 (2010).
Fuchs, G. in Autotrophic Bacteria (eds. Schlegel H. G., & Bowen B.) 365–382 (Science Technology, Madison, WI, 1989).
Martin, W. </author> & Russell, M. J. On the origin of biochemistry at an alkaline hydrothermal vent. Phil. Trans. R. Soc. London B 362, 1887–1925 (2007).
Nitschke, W. & Russell, M. J. Hydrothermal focusing of chemical and chemiosmotic energy, supported by delivery of catalytic Fe, Ni, Mo/W, Co, S and Se, forced life to emerge. J. Mol. Evol. 69, 481–496 (2009).
Nitschke, W. & Russell, M. J. Redox Bifurcations; how they work and what they mean to extant life and (potentially) to its inorganic roots. BioEssays 34, 106–109 (2012).
Hoke, K. R., Cobb, N., Armstrong, F. A. & Hille, R. Electrochemical studies of arsenite oxidase: an unusual example of a highly cooperative two-electron molybdenum center. Biochemistry 43, 1667–1674 (2004).
Adams, M. W. W. in Thermophiles (eds. Wiegel, J. & ., Adams, M.W.M.) 325–338 (Taylor & Francis, London, 1998).
Kishida, K., Sohrin, Y., Okamura, K. & Ishibashi, J. Tungsten enriched in submarine hydrothermal fluids. Earth Planet. Sci. Lett. 222, 819–827 (2004).
Scott, C. et al. Tracing the stepwise oxygenation of the Proterozoic ocean. Nature 452, 456–460 (2008).
Russell, M. J. & Hall, A. J. The emergence of life from iron monosulphide bubbles at a submarine hydrothermal redox and pH front. J. Geol. Soc. Lond. 154, 377–402 (1997).
Zerkle, A. L., House, C. H. & Brantley, S. L. Biogeochemical signatures through time as inferred from whole microbial genomes .Am. J. Sci. 305, 467–502 (2005).
Bizzarro, M. J. et al. Evidence for a late supernova injection of 60Fe into the protoplanetary disk. Science 316, 1178–1181 (2007).
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research 25, 4876–4882 (1997).
Tamura, K., Dudley, J., Nei, V. & Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Di Tommaso, P. et al. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39,W13–7 (2011).
Galtier, N., Gouy, M. & Gautier, C. Seaview and Phylo_win: two graphic tools for sequence alignment and molecular phylogeny. Comput. Appl. Biosci. 12, 543–548 (1996).
Guex, N. & Peitsch, M. C. SWISS-MODEL and the Swiss-PdbViewer: An environment for comparative protein modeling. Electrophoresis 18, 2714–2723 (1997).
Acknowledgements
MJR's contribution was carried out at the Jet Propulsion Laboratory, California Institute of Technology, under a contract with the National Aeronautics and Space Administration.
Author information
Authors and Affiliations
Contributions
BS-C, RvL and WN performed sequence analyses, structural alignments and phylogenetic reconstruction. BS-C, PP, AM, RvL, MJR and WN wrote and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
3D_alignment in pdb-format
Supplementary Information
table of sequence entries
Supplementary Information
phylogenetic tree in phb format
Supplementary Information
picture of the total multiple structural alignment
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Schoepp-Cothenet, B., van Lis, R., Philippot, P. et al. The ineluctable requirement for the trans-iron elements molybdenum and/or tungsten in the origin of life. Sci Rep 2, 263 (2012). https://doi.org/10.1038/srep00263
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00263
This article is cited by
-
Kinetic consequences of the endogenous ligand to molybdenum in the DMSO reductase family: a case study with periplasmic nitrate reductase
JBIC Journal of Biological Inorganic Chemistry (2021)
-
Iron is not everything: unexpected complex metabolic responses between iron-cycling microorganisms
The ISME Journal (2020)
-
Metatranscriptomic analysis of prokaryotic communities active in sulfur and arsenic cycling in Mono Lake, California, USA
The ISME Journal (2017)
-
Thermodynamics, Disequilibrium, Evolution: Far-From-Equilibrium Geological and Chemical Considerations for Origin-Of-Life Research
Origins of Life and Evolution of Biospheres (2017)
-
Redox cofactors insertion in prokaryotic molybdoenzymes occurs via a conserved folding mechanism
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.