Abstract
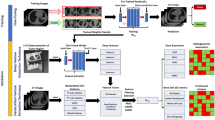
Medical image biomarkers of cancer promise improvements in patient care through advances in precision medicine. Compared to genomic biomarkers, image biomarkers provide the advantages of being non-invasive, and characterizing a heterogeneous tumor in its entirety, as opposed to limited tissue available via biopsy. We developed a unique radiogenomic dataset from a Non-Small Cell Lung Cancer (NSCLC) cohort of 211 subjects. The dataset comprises Computed Tomography (CT), Positron Emission Tomography (PET)/CT images, semantic annotations of the tumors as observed on the medical images using a controlled vocabulary, and segmentation maps of tumors in the CT scans. Imaging data are also paired with results of gene mutation analyses, gene expression microarrays and RNA sequencing data from samples of surgically excised tumor tissue, and clinical data, including survival outcomes. This dataset was created to facilitate the discovery of the underlying relationship between tumor molecular and medical image features, as well as the development and evaluation of prognostic medical image biomarkers.
Design Type(s) | database creation objective • data integration objective • disease state design • image analysis objective |
Measurement Type(s) | non-small cell lung carcinoma • transcription profiling assay |
Technology Type(s) | computed tomography scanner • microarray • RNA sequencing |
Factor Type(s) | ethnic group • histology • tumor grading • age at diagnosis • smoking status measurement |
Sample Characteristic(s) | Homo sapiens • lung |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background and Summary
Advances in high-throughput molecular technologies hold great promise for the development of genomic biomarkers that enable precision medicine tailored to specific patients. These molecular biomarkers deliver powerful diagnostic information, as well as high prognostic significance. Similarly, medical imaging technologies provide tools for measuring the structural, functional and physiologic properties of tissue. Identifying image-based properties of tumors through medical images is a standard part of diagnosis, clinical staging, and treatment planning. Because image interpretation can be subjective, for medical imaging to have a role in personalized medicine, the development of robust, standardized image features that can be used to predict molecular properties, prognosis and/or treatment response, is required. These standardized features can be in the form of semantic annotations acquired from human observers, or radiomic features, i.e. quantitative image features computed from the image pixels. Quantitative image features include tumor size and shape, image intensity distributions, and image texture. While the adoption of molecular technologies can be limited by cost and the invasiveness of the procedure, medical imaging is, more commonly, part of the standard of care1. Moreover, in comparison to molecular profiling, radiomic characterization provides a more comprehensive representation of the tumor. Since molecular profiling is restricted to the region of the biopsy, it results in an incomplete representation of the heterogeneous tissue of the tumor. On the other hand, molecular technologies allow profiling of genes expressed in the tissue sample. This complementary relationship suggests that combining the use of molecular and imaging biomarkers has the potential to improve patient care and to provide insight into how molecular mechanisms give rise to imaging phenotypes.
The prognostic power of medical image features and their link to molecular properties has only been recently investigated for certain cancer types2–20. An important challenge in such radiogenomic studies is the scarcity of large data sets containing medical images, extracted image features, gene expression profiles, and clinical data with survival outcomes. Specifically, for NSCLC, which is the leading cause of cancer death21, there is a dearth of available datasets that contain medical images, molecular features, and associated clinical data. In NSCLC, CT and PET/CT are the investigation tools of choice for diagnosis, staging and monitoring of response to treatment. From these scans, one can compute a large number of quantitative image features for associations with tumor molecular features and clinical outcomes. Molecular profiles of tumors can be obtained through needle biopsies or samples of surgically-excised tumors. Clinical data and outcomes can be obtained from standard medical follow-up. While large molecular datasets with clinical data are readily available22–25, there are fewer public medical imaging datasets combined with clinical and molecular data. For example, while five independent NSCLC datasets containing collectively 788 subjects were used in a radiogenomics study7, only 89 subjects had imaging, molecular and clinical data. Moreover, that dataset included CT scans but did not contain PET/CT data. It is important to continue to create large integrated databases available for discovery and validation of biomarkers, and so we created this dataset to allow researchers to investigate the relationships between image features, tumor molecular phenotype, and survival outcomes.
Between 2008 and 2012, we collected clinical and imaging data for 211 subjects referred for surgical treatment and obtained tissue samples from the excised tumors, where available. Tissue samples were analyzed to produce molecular phenotypes using gene microarrays, RNA sequencing technology, or both, in addition to standard-of-care NSCLC mutational testing. We also collected clinical data, such as: age, gender, weight, ethnicity, smoking status, TNM stage, histopathological grade. In addition, we included 3D tumor segmentations of the CT studies that were used for computation of 3D quantitative image features. Not all data are available for all subjects due to limitations in resources; out of the 211 subjects, 116 have all data types expect for micro-array (the data type with the smallest number of subjects), 130 have clinical, imaging (CT and PET/CT), and molecular (RNA-Seq) as detailed in Tables 1 and 2.
Methods
Subject Demographics and Clinical Data
With approval of our respective Institutional Review Boards (IRB), we recruited a total of 211 subjects for the following two cohorts: (1) The R01 cohort consisted of 162 NSCLC subjects (38 females, 124 males, age at scan: mean 68, range 42–86) from Stanford University School of Medicine (69) and Palo Alto Veterans Affairs Healthcare System (93). Subjects were recruited between April 7th, 2008 and September 15th, 2012. Subjects signed written consent forms according to the guidelines of institutions’ IRBs. The subjects were selected from a pool of early stage NSCLC patients, referred for surgical treatment with preoperative CT and PET/CT performed prior to surgical procedures. Samples of excised tissues were later used to obtain mutation data and gene expression data using gene expression microarrays, or RNA sequencing, or both. Identifiers for this set of 162 subjects are in the format R01-XXXXXX. (2) The AMC cohort, consisting of 49 additional subjects (33 females, 16 males, age at scan: mean 67, range 24–80), was retrospectively collected from Stanford University School of Medicine based on the same criteria in addition to the availability of the following clinical mutational test results: Epidermal Growth Factor Receptor (EGFR), Kirsten Rat Sarcoma viral oncogene homolog (KRAS), and Anaplastic Lymphoma Kinase (ALK). Identifiers for this set of 49 subjects are in the format AMC-XXXXXX. For both cohorts, clinical data included, where available, smoking history (211), survival (211), recurrence status (210), histology (211), histopathological grading (162) and Pathological TNM staging (161). There were 172 adenocarcinomas and 35 squamous cell carcinomas and 4 not otherwise specified with grades ranging from poorly to well-differentiated. Clinical date features (e.g. recurrence date and scan dates) are shifted for anonymization purposes and are chronologically ordered relative to each other. Table 3 summarizes clinical data of the cohorts, and Table 4 lists all clinical features.
Imaging Data
Subjects received preoperative CT and PET/CT scans at Stanford University Medical Center and Palo Alto Veterans Affairs Healthcare System prior to surgical treatment as part of their care. Different scanners were used depending on the institution and physician choice and scanning protocols also varied.
De-Identification of Imaging Data
All imaging data were de-identified prior to analysis at Stanford. For subjects from Stanford, we de-identified the imaging data using the Medical Imaging Resource Center (MIRC) Clinical Trial Processor (CTP) (RSNA, Oakbrook, IL). The MIRC CTP is a software tool designed to Anonymize DICOM objects to remove protected health information. Medical image data from VA subjects were de-identified using PACSGEAR (Perceptive Software, Pleasanton, CA).
Prior to making the data available on The Cancer Imaging Archive (TCIA)26, we performed a second round of de-identification using CTP, further assuring complete removal of any identifying information. TCIA complies with HIPAA de-identification standards using the Safe Harbor Method as defined in section 164.514(b)(2) of the HIPPA Privacy Rule. This is done by incorporating the “Basic Application Confidentiality Profile” which is amended by inclusion of the following profile options: Clean Pixel Data Option, Clean Descriptors Option, Retain Longitudinal with Modified Dates Option, Retain Patient Characteristics Option, Retain Device Identity Option, and Retain Safe Private Option. The de-identification rules applied to each object are recorded by TCIA in the DICOM sequence Method Code Sequence [0012,0063] by entering the Code Value, Coding Scheme Designator, and Code Meaning for each profile and option that were applied to the DICOM object during de-identification27.
CT Data
CT images in DICOM format28 are available from 211 subjects. Since this is a retrospectively collected dataset, different subjects were scanned using different scanners, scanning protocols and scanning parameters: slice thickness of 0.625–3 mm (median: 1.5 mm) and an X-ray tube current of 124–699 mA (mean 220 mA) at 80–140 kVp (mean 120 kVp). Detailed scanning parameters, including scanner make and model are specified in the DICOM headers. Scans were acquired with subjects in supine position with arms at sides, from the apex of the lung to the adrenal gland within a single breath-hold. Table 5 summarizes the ranges of CT parameters used for our cohort.
PET/CT Data
Fasting Fluorodeoxyglucose 18F-FDG PET/CT data are available for 201 subjects. A GE Discovery D690 PET/CT was used for PET/CT scanning at Stanford University Medical Center, while the Palo Alto VA employed a GE Discovery PET/CT scanner. (The exact model of PET/CT scanners are specified DICOM image headers.) FDG Dose and uptake time were 138.90–572.25 MBq (mean 309.26 MBq) and 23.08–128.90 min (mean 66.58 minutes), respectively. PET images were generated at both sites using a similar protocol. Specifically, CT-based attenuation correction was utilized with iterative Ordered Subset Expectation Maximization (OSEM) reconstruction. Image acquisition included routine coverage of base-of-skull to mid-thigh with additional spot views where necessary. Each bed position was 1–5-minute acquisition, dependent on su weight. Table 6 summarizes ranges of scan parameters used to obtain PET/CT images. This PET/CT data set was used to identify tumor PET-FDG uptake features associated with gene expression signatures and survival29.
CT and PET/CT acquisition protocols
It has been recognized that the results of quantitative analyses (including e.g., radiomics) of images will vary as a function of image acquisition and reconstruction protocol30–38. However, we note that the imaging datasets reported here were acquired over several years and from several institutions, and not as part of a prospective trial. For these reasons there was no attempt to harmonize the acquisition and reconstruction protocols.
Semantic Annotations
Semantic annotations are available for axial CT series of 190 subjects. The template of semantic terms was developed in consensus by two academic thoracic radiologists (A.N.C.L. and D.A.) with expertise and interest in lung cancer imaging. The template was developed for nodules as they are the most common manifestation of lung cancer. As a result, we provide no semantic annotations for cancers of other manifestations, e.g., central obstructive tumors or "pneumonic tumors”. The template contains 28 nodule analysis features and parenchymal features comprising conventional and newly developed features used for diagnosis and staging using the CT images. Nodule features describe anatomy location, geometry, internal features and other associated findings of the nodules.
Parenchymal features characterize lung emphysema, bronchi and lumen. The selected terms are in common usage in radiology clinical practice and are derived from descriptions in the radiology literature; definitions of some of these, such as “nodule” are found in the Fleischner Society: Glossary of Terms for Thoracic Imaging39. Table 7 (available online only) describes the semantic features included in the template. The ePAD template that we developed forces complete annotation for each nodule, resulting in all applicable features being collected. There are some features whose presence are conditioned upon other features being present. For example, the primary emphysema pattern feature is not collected when emphysema is not present in the lung. ePAD creates annotations in the Annotation and Image Mark-up (AIM) file format using a controlled vocabulary. The AIM information model is designed to be semantically inoperable. Information such as annotator identity, annotation date, and a reference to the annotated image, complement information on anatomic entities and imaging observation characteristics of the referenced image. AIM files supplement DICOM and other image formats which do not contain information on the meaning of the pixels in the image40,41. One radiologist (A.N.L.) with more than 20 years of experience ascribed the semantic annotations for all subjects’ CT scans using ePAD, an open-source and freely available web-based quantitative imaging informatics platform41. While we acknowledge that semantic annotations are subjective and subject to intra-and inter-reader variability, these were used in several studies, e.g., to predict EGFR and KRAS mutation status42, and to create a radiogenomic map linking semantic features to gene expression profiles generated by RNA sequencing13.
Segmentations
Initial segmentations for 144 subjects were obtained from an axial CT image series using an unpublished automatic segmentation algorithm. All of these segmentations were viewed by a thoracic radiologist (M.K.) with more than 5 years of experience and edited as necessary using ePAD. Final segmentations were reviewed by an additional thoracic radiologist (A.N.L.); disagreements in tumor boundaries were discussed and edited as appropriate, with final approval by A.N.L. All segmentations are stored as DICOM Segmentation Objects28.
Molecular Data
Tumor Preparation
All tumor samples were collected from treatment-naïve subjects during surgical procedure. Following excision, the surgeon cut a 3–5-mm-thick slice along the longest axis of the excised tissue, which was frozen within 30 minutes of excision. It was later retrieved for RNA extraction. Molecular data are available from EGFR, KRAS, ALK mutational testing, gene expression microarrays, and RNA sequencing. Tumors from 17 subjects were analyzed using both gene expression microarrays and RNA sequencing.
Mutational testing
EGFR, KRAS and ALK mutation status are available from clinical records in 206, 205, and 196 subjects, respectively. Single nucleotide mutation detection was performed using SNaPshot technology based on dideoxy single-base extension of oligonucleotide primers after multiplex polymerase chain reaction (PCR). Exons 18, 19, 20 and 21 were tested for EGFR mutations. Exon 2 Positions 12 and 13 were tested for missense KRAS mutations with amino acid substitution. Mutation results were a combination of mutation at any location of the tested exons. For ALK, EML4-ALK translocation detection test was performed using fluorescence in situ hybridization (FISH).
Gene Expression Microarray Data
Gene expression microarray data was collected for the subset of 26 subjects, who underwent surgical treatment between April 7, 2008 and May 21, 2010. RNA was processed at the Stanford Functional Genomics Facility using Illumina Whole Genome Bead Chips (Human HT-12; Illumina, San Diego, CA). These data were preprocessed as follows: First, we filtered the microarray probes on the basis of a significant detection call in at least 60% of the samples. Next, we log transformed the microarray data and used quantile normalization to normalize between arrays. These data, along with the corresponding CT images, were used to describe associations between image features, gene expression, and survival10,29.
RNA Sequencing Data
Based on availability and quality of available tissue, RNA sequencing was performed on samples from 130 subjects (17 of which intersect with the gene expression microarray dataset described in the previous section). We excluded RNASeq for tissue samples with RNA integrity number (RIN) below 2.5. Total RNA was extracted from the tissue samples and converted into a library for paired-end sequencing on Illumina Hiseq according to the protocol for the Illumina TruSeq Sample preparation kit (Centrillion Biosciences, Palo Alto, CA). Briefly, total RNA quality and quantity were measured by BioAnalyzer (Agilent). For library preparation, the TruSeq Total Stranded RNA with Ribo-Zero Reduction (Illumina) was used following manufacturer’s instructions. This method includes a Ribo-Zero rRNA depletion step, followed by fragmentation and cDNA synthesis using SuperScript II (Life Technologies). The cDNA was A-tailed, ligated and amplified using the materials in the TruSeq Total Stranded RNA with Ribo-Zero Reduction kit. Quality was confirmed using the BioAnalyzer and finally the concentration evaluated by KAPA qPCR (KAPA Biosystems). Prior to sequencing, samples were diluted to 4 nmol and pooled. Pooled libraries were clustered via the cBOT and sequenced on the HiSeq 2500 (illumina) following manufacturer’s instructions. The set of 130 tissue samples was sequenced in three batches of sizes 16, 66, 48.
Data processing was performed by Centrillion Biosciences as follows: reads were aligned to the human genome (hg19) using the alignment algorithm STAR43 version 2.3 with 91 bases of splice junction overhangs. Next, Cufflinks version 2.0.244 was used to determine the expression calls in each sample using Fragments Per Kilobase of transcript per Million mapped reads (FPKM).
Data Records
Subject Identifiers
A unique identifier for each subject is identical in all four public data records in this dataset. Subject ID’s are 6-digit numbers in the form of R01-XXXXXX or AMC-XXXXXX.
Data Record 1
Clinical, image, semantic data for all subjects are stored in The Cancer Imaging Archive (TCIA) (Data Citation 1). One comma-delimited file contains clinical data for all subjects with unique subject identifiers. Semantic features for each subject are stored in Annotation and Image Markup (AIM) files45. CT and PET/CT Images are in DICOM format. Where available, segmentations are provided as DICOM Segmentation Objects.
Data Record 2
Image data of 26 subjects had been previously deposited in the TCIA repository (Data Citation 2). These images were given new subject names in the form R01-XXXXXX as part of the new dataset described in this work.
Data Record 3
Gene expression microarray data, available for 26 subjects, were deposited in National Center for Biotechnology Information Gene Expression Omnibus (NCBI GEO)46 (Data Citation 3). The subject identifiers are identical to subject names in Data Record 2. Processed gene clusters were deposited in tab-delimited files with column values corresponding to microarray ID, log2 transformed quantile normalized and probe selection detection-p-value, respectively. This data record also contains raw expression data, as well as matrix data obtained prior to normalization.
Data Record 4
Raw and processed sequencing data obtained from RNASeq for 130 subjects are available at NCBI GEO (Data Citation 4). The subject IDs are identical to subject names in Data Record 1.
Technical Validation
All CT and PET/CT data were collected as part of patient care and therefore all quality assurance was performed by the institution that collected the data.
Usage Notes
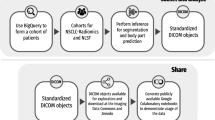
All data are freely available to browse, download, and use for commercial, scientific and educational purposes as outlined in the Creative Commons Attribution 3.0 Unported License. Users should properly cite this source for any work based on this dataset.
Additional information
How to cite this article: Bakr, S. et al. A radiogenomic dataset of non-small cell lung cancer. Sci. Data. 5:180202 doi: 10.1038/sdata.2018.202 (2018).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Lambin, P. et al. Predicting outcomes in radiation oncology–multifactorial decision support systems. Nat Rev Clin Oncol 10, 27–40, 10.1038/nrclinonc.2012.196 (2013).
Segal, E. Decoding global gene expression programs in liver cancer by noninvasive imaging. Nat Biotechnol 25, 675–680 (2007).
Diehn, M. Identification of noninvasive imaging surrogates for brain tumor gene-expression modules. Proc Natl Acad Sci USA 105, 5213–5218 (2008).
Tixier, F. et al. Intratumor heterogeneity characterized by textural features on baseline 18F-FDG PET images predicts response to concomitant radiochemotherapy in esophageal cancer. J Nucl Med 52, 369–378, 10.2967/jnumed.110.082404 (2011).
El Naqa, I. et al. Exploring feature-based approaches in PET images for predicting cancer treatment outcomes. Pattern Recognit 42, 1162–1171, 10.1016/j.patcog.2008.08.011 (2009).
Nair, V. S. & Prognostic, P. E. T. 18F-FDG uptake imaging features are associated with major oncogenomic alterations in patients with resected non-small cell lung cancer. Cancer Res 72, 3725–3734 (2012).
Aerts, H. J. et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5, 4006, 10.1038/ncomms5006 (2014).
Coroller, T. P. CT-based radiomic signature predicts distant metastasis in lung adenocarcinoma. Radiother Oncol 114, 345–350 (2015).
Ganeshan, B., Skogen, K., Pressney, I., Coutroubis, D. & Miles, K. Tumour heterogeneity in oesophageal cancer assessed by CT texture analysis: preliminary evidence of an association with tumour metabolism, stage, and survival. Clin Radiol 67, 157–164 (2012).
Gevaert, O. Non-Small Cell Lung Cancer: Identifying Prognostic Imaging Biomarkers by Leveraging Public Gene Expression Microarray Data—Methods and Preliminary Results. Radiology 264, 387–396 (2012).
Ganeshan, B., Panayiotou, E., Burnand, K., Dizdarevic, S. & Miles, K. Tumour heterogeneity in non-small cell lung carcinoma assessed by CT texture analysis: a potential marker of survival. Eur Radiol 22, 796–802 (2012).
Itakura, H. et al. Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities. Sci Transl Med 7, 303ra138, doi:10.1126/scitranslmed.aaa7582 (2015).
Zhou, M. et al. Non-Small Cell Lung Cancer Radiogenomics Map Identifies Relationships between Molecular and Imaging Phenotypes with Prognostic Implications. Radiology 161845, 10.1148/radiol.2017161845 (2017).
Bakr, S. et al. Noninvasive radiomics signature based on quantitative analysis of computed tomography images as a surrogate for microvascular invasion in hepatocellular carcinoma: a pilot study. J Med Imaging (Bellingham) 4, 041303, 10.1117/1.JMI.4.4.041303 (2017).
Liu, Y. et al. Radiologic Features of Small Pulmonary Nodules and Lung Cancer Risk in the National Lung Screening Trial: A Nested Case-Control Study. Radiology 161458, 10.1148/radiol.2017161458 (2017).
Li, Q. et al. Imaging features from pretreatment CT scans are associated with clinical outcomes in nonsmall-cell lung cancer patients treated with stereotactic body radiotherapy. Med Phys 44, 4341–4349, 10.1002/mp.12309 (2017).
Rios Velazquez, E. et al. Somatic Mutations Drive Distinct Imaging Phenotypes in Lung Cancer. Cancer Res 77, 3922–3930, 10.1158/0008-5472.CAN-17-0122 (2017).
Wu, J. et al. Heterogeneous Enhancement Patterns of Tumor-adjacent Parenchyma at MR Imaging Are Associated with Dysregulated Signaling Pathways and Poor Survival in Breast Cancer. Radiology 162823, 10.1148/radiol.2017162823 (2017).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images Are More than Pictures, They Are Data. Radiology 278, 563–577, 10.1148/radiol.2015151169 (2016).
O'Connor, J. P. et al. Imaging biomarker roadmap for cancer studies. Nat Rev Clin Oncol 14, 169–186, 10.1038/nrclinonc.2016.162 (2017).
Jemal, A., Siegel, R., Xu, J. & Ward, E. Cancer statistics, 2010. CA Cancer J Clin 60, 277–300, 10.3322/caac.20073 (2010).
Lee, E. S. et al. Prediction of recurrence-free survival in postoperative non-small cell lung cancer patients by using an integrated model of clinical information and gene expression. Clin Cancer Res 14, 7397–7404, 10.1158/1078-0432.CCR-07-4937 (2008).
Parkinson, H. et al. ArrayExpress update--an archive of microarray and high-throughput sequencing-based functional genomics experiments. Nucleic Acids Res 39, D1002–D1004, 10.1093/nar/gkq1040 (2011).
Cancer Genome Atlas Research, N. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525, 10.1038/nature11404 (2012).
Cancer Genome Atlas Research, N. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550, 10.1038/nature13385 (2014).
Clark, K. The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository. J Digit Imaging 26, 1045–1057 (2013).
Digital imaging and communication in medicine (DICOM) (1997).
Kahn, C. E Jr., Carrino, J. A, Flynn, M. J, Peck, D. J . & Horii, S. C. DICOM and radiology: past, present, and future. J Am Coll Radiol 4, 652–657, 10.1016/j.jacr.2007.06.004 (2007).
Nair, V. S. et al. Prognostic PET 18F-FDG uptake imaging features are associated with major oncogenomic alterations in patients with resected non-small cell lung cancer. Cancer Res 72, 3725–3734, 10.1158/0008-5472.CAN-11-3943 (2012).
Lambin, P. et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14, 749–762, 10.1038/nrclinonc.2017.141 (2017).
Nyflot, M. J. et al. Quantitative radiomics: impact of stochastic effects on textural feature analysis implies the need for standards. J Med Imaging (Bellingham) 2, 041002, 10.1117/1.JMI.2.4.041002 (2015).
Lo, P., Young, S., Kim, H. J., Brown, M. S. & McNitt-Gray, M. F. Variability in CT lung-nodule quantification: Effects of dose reduction and reconstruction methods on density and texture based features. Med Phys 43, 4854, 10.1118/1.4954845 (2016).
Solomon, J., Mileto, A., Nelson, R. C., Roy Choudhury, K. & Samei, E. Quantitative Features of Liver Lesions, Lung Nodules, and Renal Stones at Multi-Detector Row CT Examinations: Dependency on Radiation Dose and Reconstruction Algorithm. Radiology 279, 185–194, 10.1148/radiol.2015150892 (2016).
Zhao, B. et al. Reproducibility of radiomics for deciphering tumor phenotype with imaging. Sci Rep 6, 23428, 10.1038/srep23428 (2016).
Balagurunathan, Y. et al. Reproducibility and Prognosis of Quantitative Features Extracted from CT Images. Transl Oncol 7, 72–87 (2014).
Oxnard, G. R. et al. Variability of lung tumor measurements on repeat computed tomography scans taken within 15 minutes. J Clin Oncol 29, 3114–3119, 10.1200/JCO.2010.33.7071 (2011).
Zhao, B. et al. Evaluating variability in tumor measurements from same-day repeat CT scans of patients with non-small cell lung cancer. Radiology 252, 263–272, 10.1148/radiol.2522081593 (2009).
Fave, X. et al. Can radiomics features be reproducibly measured from CBCT images for patients with non-small cell lung cancer? Med Phys 42, 6784–6797, 10.1118/1.4934826 (2015).
Hansell, D. M. et al. Fleischner Society: glossary of terms for thoracic imaging. Radiology 246, 697–722, 10.1148/radiol.2462070712 (2008).
Channin, D. S., Mongkolwat, P., Kleper, V. & Rubin, D. L. The Annotation and Image Mark-up project. Radiology 253, 590–592, 10.1148/radiol.2533090135 (2009).
Rubin, D. L. et al. Automated tracking of quantitative assessments of tumor burden in clinical trials. Transl Oncol 7, 23–35 (2014).
Gevaert, O. et al. Predictive radiogenomics modeling of EGFR mutation status in lung cancer. Sci Rep 7, 41674, 10.1038/srep41674 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner Bioinformatics 29, 15–21, 10.1093/bioinformatics/bts635 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7, 562–578, 10.1038/nprot.2012.016 (2012).
Mongkolwat, P., Kleper, V., Talbot, S. & Rubin, D. The National Cancer Informatics Program (NCIP) Annotation and Image Markup (AIM) Foundation model. J Digit Imaging 27, 692–701, 10.1007/s10278-014-9710-3 (2014).
Barrett, T. NCBI GEO: archive for high-throughput functional genomic data. Nucleic Acids Res 37, D885–D890 (2009).
Data Citations
Bakr, S. et al. The Cancer Imaging Archive https://doi.org/10.7937/K9/TCIA.2017.7hs46erv (2017)
Napel, S., & Plevritis, S. K. The Cancer Imaging Archive https://doi.org/10.7937/K9/TCIA.2014.X7ONY6B1 (2014)
Gene Expression Omnibus GSE28827 (2012)
Gene Expression Omnibus GSE103584 (2018)
Acknowledgements
This work was funded by the National Cancer Institute and the National Institute of Biomedical Imaging and Bioengineering of the National Institutes of Health under Award Numbers: R01 CA160251, U01 CA187947, U01 CA142555, U01 CA190214, and R01 EB020527. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. We are grateful to Professor Denise Aberle for providing semantic annotations for the CT image dataset. Finally, we sincerely thank Justin Kirby, Kirk Smith, Tracy Nolan, and William Bennett for help in curating and incorporating the imaging and related data on The Cancer Imaging Archive (https://cancerimagingarchive.net).
Author information
Authors and Affiliations
Contributions
Data acquisition or data analysis/interpretation all authors; manuscript drafting: S.B., O.G.; manuscript revision and/or approval: all authors.
Corresponding author
Ethics declarations
Competing interests
S.N. is a consultant for Carestream Inc, and a member of the scientific advisory boards for EchoPixel, Inc.; Fovia, Inc. and Radlogics, Inc. All other authors declare no competing interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Bakr, S., Gevaert, O., Echegaray, S. et al. A radiogenomic dataset of non-small cell lung cancer. Sci Data 5, 180202 (2018). https://doi.org/10.1038/sdata.2018.202
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2018.202
This article is cited by
-
End-to-end reproducible AI pipelines in radiology using the cloud
Nature Communications (2024)
-
SAROS: A dataset for whole-body region and organ segmentation in CT imaging
Scientific Data (2024)
-
Evaluating ChatGPT-4V in chest CT diagnostics: a critical image interpretation assessment
Japanese Journal of Radiology (2024)
-
MultiRoiMix: A Data Augmentation Method for PET/CT Multimodal Medical Images
Journal of Medical and Biological Engineering (2024)
-
Enhancing NSCLC recurrence prediction with PET/CT habitat imaging, ctDNA, and integrative radiogenomics-blood insights
Nature Communications (2024)