Abstract
The cranial diversity of sharks reflects disparate biomechanical adaptations to feeding. In order to be able to investigate and better understand the ecomorphology of extant shark feeding systems, we created a x-ray computed tomography (CT) library of shark cranial anatomy with three-dimensional (3D) lower jaw reconstructions. This is used to examine and quantify lower jaw disparity in extant shark species in a separate study. The library is divided in a dataset comprised of medical CT scans of 122 sharks (Selachimorpha, Chondrichthyes) representing 73 extant species, including digitized morphology of entire shark specimens. This CT dataset and additional data provided by other researchers was used to reconstruct a second dataset containing 3D models of the left lower jaw for 153 individuals representing 94 extant shark species. These datasets form an extensive anatomical record of shark skeletal anatomy, necessary for comparative morphological, biomechanical, ecological and phylogenetic studies.
Design Type(s) | data integration objective • species comparison design |
Measurement Type(s) | morphology |
Technology Type(s) | computed tomography |
Factor Type(s) | animal body part • organism |
Sample Characteristic(s) | Brachelurus waddi • Echinorhinus brucus • Centrophorus uyato • Chlamydoselachus anguineus • Pristiophorus japonicus • Centrophorus seychellorum • Cetorhinus maximus • Orectolobus japonicus • Apristurus laurussonii • Centroselachus crepidater • Sphyrna tudes • Euprotomicrus bispinatus • Etmopterus spinax • Mustelus higmani • Galeus melastomus • Galeorhinus galeus • Scyliorhinus stellaris • Squatina squatina • Carcharhinus leucas • Scyliorhinus canicula • Chiloscyllium punctatum • Halaelurus boesemani • Atelomycterus marmoratus • Hemiscyllium strahani • Prionace glauca • Squalus megalops • Atelomycterus macleayi • Carcharhinus hemiodon • Hemiscyllium trispeculare • Chiloscyllium arabicum • Sphyrna tiburo • Sphyrna corona • Mustelus mustelus • Isistius brasiliensis • Scymnodalatias albicauda • Deania calcea • Chiloscyllium griseum • Heterodontus japonicus • Eusphyra blochii • Sphyrna zygaena • Carcharhinus amboinensis • Triakis semifasciata • Squalus suckleyi • Carcharhinus macloti • Carcharhinus dussumieri • Chaenogaleus macrostoma • Hemigaleus microstoma • Sphyrna lewini • Halaelurus buergeri • Stegostoma fasciatum • Chiloscyllium indicum • Chiloscyllium hasseltii • Eucrossorhinus dasypogon • Scoliodon laticaudus • Oxynotus centrina • Carcharhinus melanopterus • Carcharhinus falciformis • Heterodontus francisci • Squatina africana • Triaenodon obesus • Ginglymostoma cirratum • Alopias vulpinus • Squalus cubensis • Isogomphodon oxyrhynchus • Heptranchias perlo • Rhizoprionodon terraenovae • Dalatias licha • Squatina japonica • Nebrius ferrugineus • Squalus acanthias • Lamna nasus • Mustelus asterias • Scyliorhinus boa • Lamna ditropis • Carcharodon carcharias • Isurus oxyrinchus • Isurus paucus • Centroscymnus coelolepis • Cephaloscyllium ventriosum • Galeocerdo cuvier • Pseudocarcharias kamoharai • Mitsukurina owstoni • Carcharias taurus • Carcharhinus acronotus • Sphyrna mokarran • Hexanchus griseus • Negaprion acutidens • Alopias superciliosus • Orectolobus maculatus • Carcharhinus brachyurus • Carcharhinus galapagensis • Mustelus henlei • Orectolobus ornatus • Proscyllium habereri • lower jaw region • whole body • head • pectoral fin • tail |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Computed tomography (CT) scanning has opened new ways for studying various parts of an organism’s biology. This technique has been used in sharks to elucidate the development1, function2–5 and morphology of their feeding mechanics6–8. The majority of these studies have focused on small taxonomic subsets to gain detailed anatomical knowledge. However, phylogenetically broad statistical analyses of shark cranial mechanics are still lacking. The lead author of this work is currently undertaking such studies. This contribution provides a descriptor of a large CT dataset of shark anatomy. We used whole specimens from museum collections (a few specimens only comprise the head due to their conservation) to create a dataset of medical CT scans which covers approximately 75% of all extant shark families9.
The x-ray computed tomography library presented here was created for investigating the ecomorphological diversity of shark feeding systems. The data from the CT scans were used to create a second dataset comprising three-dimensional (3D) models of the lower jaw. These models were used to examine lower jaw disparity in extant sharks species in a separate study by quantifying jaw shape using landmark based geometric morphometrics. The lower jaw was selected because it displays a diversity of jaw morphologies between shark species10–15. Lower jaws present biomechanically important features related to jaw closing mechanics and are therefore expected to function as a predictor of ecological specialisation16–21.
This tomography and virtual 3D dataset can be applied to or used to supplement further comparative and functional analyses of shark morphology. These may include investigations of (a)symmetry, development, integration and modularity or shape change through evolutionary time22. Furthermore, 3D anatomical models provide an excellent visual resource for outreach and education. For example, 3D models can be integrated into 3D PDF documents as interactive figures or can be physically reproduced using rapid prototyping (also referred to as stereolithography or 3D printing)23.
Methods
Specimens
122 individuals representing 73 extant species of 25 out of 34 families from all 9 orders of extant sharks9 (Selachimorpha, Chondrichthyes) are used in the CT scan dataset (Data Citation 1). The specimens are, at the time of CT scanning, formalin preserved and stored in 70% alcohol except for 4 specimens which were stored frozen (RMNH.PISC.36345, RMNH.PISC.verznr.2, 3 and 7). All CT scanned specimens are curated in the spirit collections of the British Museum of Natural History (BMNH) and Naturalis Biodiversity Center (NBC) (see the accompanying metadata for a complete list of specimens). The NBC collections comprise material from the Rijksmuseum van Natuurlijke Historie (RMNH) and the Zoölogisch Museum Amsterdam (ZMA); those institutional identifiers continue to be used here.
CT scanning
CT scans of the specimens housed in the NBC collections were made at the Leiden University Medical Center, the Netherlands (LUMC) with a Toshiba Aquilion 64 medical scanner (Toshiba Medical Systems, Otawara, Japan) using a for the sharks customized scanning and reconstruction protocol (100 kV tube voltage, 150 mAs tube charge per rotation, 64 active channels, acquisition and reconstructed slice thickness 0.5 mm, reconstructed slice increment 0.5 mm, FC03 reconstruction filter, pitch factor 0.83). Specimens from the BMNH collection were scanned at the CT scanning facility of the Royal Brompton and Harefield NHS Trust (RBH), London, United Kingdom, with a Siemens Somatom Sensation 64 medical scanner using a customized scanning protocol with a variable reconstructed slice increment (100 kV tube voltage, 210 mAs tube charge per rotation, acquisition and reconstructed slice thickness 1.0 mm, B30f reconstruction filter).
3D segmentations
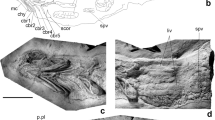
The image series from the CT scans were imported into Mimics, v 15.01, (Materialise Software) for segmentation and 3D modelling. We used manual segmentation with a threshold edit of Hounsfield units (i.e., grey values) that are associated to the calcified cartilage of the lower jaw (Fig. 1). The Hounsfield units produced by the medical CT scanners are calibrated according to standard procedures of the manufacturer. The structures of calcified cartilage are not very dense in the scans (meaning relatively low hounsfield units, ranging from 400 to 1,500), and at the border between calcified cartilage and soft tissue, there will always be a partial volume effect that give a 'smoothed' transition from calcified cartilage to soft tissue. Structures with high density, such as the lower jaws, are depicted bright, while structures with low or intermediate density are depicted as dark using optimized greyscales (Fig. 1a). A mask based on measured threshold values of the Hounsfield units, which aligned very closely with the high relative density of the calcified cartilages of the lower jaw is set to produce an exact overlay on the slice images (Fig. 1b). We used these masks to create 3D models of the left lower jaws (Fig. 1c). Threshold values couldn’t be standardised across different scans due to variable tissue densities across specimens. Most 3D models could be modeled without ambiguity. Where manual thresholds were required, ambiguity was checked against left-right symmetry of the skeleton.
(a) Each CT scan is build up of multiple slices (tomograms) showing structures with high attenuation of X-rays bright and structures with low attenuation dark, using a grey scale. Each pixel in the tomogram is associated with a Hounsfield unit. (b) A mask (green) is superimposed on the left lower jaw. This mask is based on predetermined threshold values of the Hounsfield unit of to produce an exact overlay on each tomogram. (c) When in each slice a mask is superimposed on the left lower jaw a 3D model is calculated based on the mask. The black line through the 3D model indicates the location of the tomogram in (a,b), mc l=left lower jaw.
Subsequently, each 3D model of the left lower jaw was exported as a *.PLY file from MIMICS (Data Citation 1).
Thirty-one additional 3D models of the left lower jaw were segmented using CT scans provided by other researchers (Table 1), resulting in a total of 153 3D models of 94 extant shark species.
Data Records
Data record 1—The data for this manuscript have been deposited in a Figshare repository (Data Citation 1). It comprises a dataset of 121 of volumes that consist of tomography images (‘slices’) in DICOM format reconstructed from medical computed tomography (CT) scans of sharks. The matrices of voxels in the dataset represent the Hounsfield units of the corresponding materials and tissues. The Hounsfield unit is associated with a well-defined physics quantity, being the linear attenuation coefficient of these materials and tissues. The majority of the data are whole body CT scans, while a few are CT scans of the head. From the CT scans we generated 153 3D models of the left lower jaws. The accompanying metadata describes the list of CT scanned specimens, their scan parameters and the derived 3D models included in Data record 1. Table 1 describes the specimen list and scan parameters of the additional 3D models of the left lower jaws that are created from CT scans provided by other researchers.
Technical Validation
CT scanner
Medical CT scanners are subject to a regular program for quality control and maintenance under the responsibility of a qualified medical physicist. However, discrepancies between the reconstructed values in an image and the true attenuation coefficients of the scanned object (i.e., image artifacts) can occur. Three common categories of CT image artefact appearances can be distinguished: streaking, shading, and rings and bands. Streaking artifacts appear as straight lines (bright and/or dark) across the image and are the result of the nature of the filtered backprojection reconstruction process. Shading artifacts can occur near objects of high contrast and usually appear in the soft tissue region near bony structures or near air pockets. This type of artifact is hard to identify since it shows a similar shape as the structure creating the shading. Ring and band artifacts can be visible as rings or bands overlaying the original image structure. The occurrence of artifacts can originate from the system design, x-ray tubes, detector, the specimen or operator24. In our dataset only two CT scans show significant artifacts. BMNH 1978.6.22.1 Cetorhinus maximus show ring artifacts in the dense vertebrae and BMNH 1978.6.22.1 Cetorhinus maximus shows ring and streaking artifacts in the dense vertebrae and posterior region of the head. Despite these artifacts skeletal structures were identifiable and useable for 3D segmentation. All other scans are free of artifacts or show only negligible artifacts.
The X-rays, generated by the CT scanner, are used to measure the transmission of X-ray through the specimens under hundreds of different angles. All these measurements are referred to as the raw data. This data is processed with a filtered backprojection, which generates a series of cross-sectional images25. Internal structures are visualized by their ability to attenuate the X-ray beams based on the linear attenuation coefficient. The parameters of the CT scanner were set to optimally visualize the jaws of the sharks. The jaws are well calcified compared to other skeletal structures in the head, therefore structures such as the neurocranium, basihyal and branchial chamber appear less clear in the scans.
Usage Notes
The CT scan data in DICOM format can be loaded into 3D analysis software such as the free software package SPIERS26 or in license based software packages such as MIMICS (http://biomedical.materialise.com/mimics), AVIZO (http://www.fei.com/software/avizo3d/), or VG StudioMax (http://www.volumegraphics.com/en/products/vgstudio-max/basic-functionality/). 3D models of the structures of interest can be produced and exported in various formats using one of these software packages. The exported models can be used in landmark-based geometric morphometric methods27,28 to quantitatively test hypotheses of morphological diversity. The IDAV Landmark Editor29 is a free software package well suited for placing landmarks on the 3D models and exporting the landmark coordinates. Note that Landmark Editor only accepts *.PLY files without binary encoding. The software package Meshab (http://meshlab.sourceforge.net/) can be used to analyse, view and convert the *.PLY files to a usable format for Landmark Editor.
The landmark coordinates generated in Landmark Editor can be used in statistical software designed for geometric morphometric approaches, such as MorphoJ30, Morphologika31, PAST32 or the geomorph package33 in R34.
Additional Information
How to cite this article: Kamminga, P. et al. X-ray computed tomography library of shark anatomy and lower jaw surface models. Sci. Data 4:170047 doi: 10.1038/sdata.2017.47 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Summers, A. P., Ketcham, R. A. & Rowe, T. Structure and function of the horn shark (Heterodontus francisci) cranium through ontogeny: development of a hard prey specialist. J. Morphol. 260, 1–12 (2004).
Wroe, S. et al. Three-dimensional computer analysis of white shark jaw mechanics: how hard can a great white bite? J. Zool. 276, 336–342 (2008).
Tomita, T., Sato, K., Suda, K., Kawauchi, J. & Nakaya, K. Feeding of the megamouth shark (Pisces: Lamniformes: Megachasmidae) predicted by its hyoid arch: a biomechanical approach. J. Morphol. 272, 513–524 (2011).
Ramsay, J. B. et al. Eating with a saw for a jaw: Functional morphology of the jaws and tooth-whorl in Helicoprion davisii. J. Morphol. 18, 1–18 (2014).
Ferrara, T. L. et al. Mechanics of biting in great white and sandtiger sharks. J. Biomech. 44, 430–435 (2011).
Lane, J. & Maisey, J. G. The Visceral Skeleton and Jaw Suspension In the Durophagous Hybodontid Shark Tribodus limae from the Lower Cretaceous of Brazil. J. Paleontol. 86, 886–905 (2012).
Motta, P. J. et al. Functional morphology of the feeding apparatus, feeding constraints, and suction performance in the nurse shark Ginglymostoma cirratum. J. Morphol. 269, 1041–1055 (2008).
Mara, K. R., Motta, P. J., Martin, A. P. & Hueter, R. E. Constructional morphology within the head of hammerhead sharks (sphyrnidae). J. Morphol. 276, 526–539 (2015).
Ebert, D. A., Fowler, S. & Compagno, L. J. V. Sharks of the world: a fully illustrated guide (Plymouth: Wild Nature Press, 2013).
Allis, E. P. Jr The cranial anatomy of Chlamydoselachus anguineus. Acta Zool 4, 123–221 (1923).
Ramsay, J. B. & Wilga, C. D. Morphology and mechanics of the teeth and jaws of white-spotted bamboo sharks (Chiloscyllium plagiosum). J. Morphol. 682, 664–682 (2007).
Wilga, C. D. Morphology and evolution of the jaw suspension in lamniform sharks. J. Morphol. 265, 102–119 (2005).
Motta, P. J. & Wilga, C. D. Anatomy of the feeding apparatus of the nurse shark, Ginglymostoma cirratum. J. Morphol. 241, 33–60 (1999).
Compagno, L. J. V. in Elasmobranchs as living resources: Advances in the Biology, Ecology, Systematics and the Status of the Fisheries (eds Pratt H. L., Gruber S. H. & Taniuchi T. ) 357–379 (NOAA Technical Report 90, 1990).
Denison, R. H. Anatomy of the head and pelvic fin of the whale shark, Rhineodon. Bull. Am. Museum Nat. Hist 523, 477–515 (1937).
Wainwright, P. C. Biomechanical limits to ecological performance: mollusk-crushing by the Caribbean hogfish, Lachnolaimus maximus (Labridae). J. Zool. 213, 283–297 (1987).
Wainwright, P. C. in Ecological Morphology: Integrative Organismal Biology (eds Wainwright, P. C. & Reilly, S. M. 42–59 (University of Chicago Press, 1994).
Wainwright, P. C. Ecological Explanation through Functional Morphology: The Feeding Biology of Sunfishes. Ecology 77, 1336–1343 (1996).
Ricklefs, R. E., Miles, D. B. in Ecological Morphology: Integrative Organismal Biology (eds Wainwright, P. C. & Reilly, S. M. 13–41 (University of Chicago Press, 1994).
Norton, S. F. A functional approach to ecomorphological patterns of feeding in cottid fishes. Environ. Biol. Fishes 44, 61–78 (1995).
Anderson, P. S. L., Friedman, M. & Ruta, M. Late to the table: diversification of tetrapod mandibular biomechanics lagged behind the evolution of terrestriality. Integr. Comp. Biol. 53, 197–208 (2013).
Adams, D. C., Rohlf, F. J. & Slice, D. A field comes of age: geometric morphometrics in the 21st century. Hystrix, Ital. J. Mammology 24, 7–14 (2013).
Lautenschlager, S. & Rücklin, M. Beyond the print—virtual paleontology in science publishing, outreach, and education. J. Paleontol. 88, 727–734 (2014).
Hsieh, J. Computed tomography: principles, design, artifacts, and recent advances. Second Edition (SPIE and John Wiley & Sons, Inc., 2009).
Herman, G. T. Fundamentals of Computerized Tomography-Image Reconstruction from Projections. Second Edition (Springer-Verlag, 2009).
Sutton, M. D., Garwood, R. J., Siveter, D. J. & Siveter, D. J. SPIERS and VAXML; a software toolkit for tomographic visualisation and a format for virtual specimen interchange. Paleontol. Electron 15, 1–15 (2012).
Rohlf, F. J. & Marcus, L. F. A revolution morphometrics. Trends Ecol. Evol. 8, 129–132 (1993).
Bookstein, F. Morphometric tools for landmark data: geometry and biology (Cambridge University Press, 1991).
Wiley, D. F. et al. Evolutionary morphing. in Proceedings of IEEE Visualization 2005 (VIZ’05) 431–438 (2005).
Klingenberg, C. P. MorphoJ: an integrated software package for geometric morphometrics. Mol. Ecol. Resour. 11, 353–357 (2011).
O’Higgins, P. & Jones, N. Tools for statistical shape analysis. Hull York Medical School. http://sites.google.com/site/hymsfme/resources (2006).
Hammer, Ø., Harper, D. A. T. & Ryan, P. D. PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol. Electron 4, 9 (2001).
Adams, D. C. & Otárola-Castillo, E. Geomorph: an R Package for the Collection and Analysis of Geometric Morphometric Shape Data. Methods Ecol. Evol. 4, 393–399 (2013).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, 2013).
Data Citations
Kamminga, P., De Bruin, P. W., Geleijns, K., & Brazeau, M. D. Figshare https://dx.doi.org/10.6084/m9.figshare.c.3662366 (2017)
Acknowledgements
We thank R. de Ruiter and J. Maclaine for granting access to the museum collections of Naturalis Biodiversity Center and the British Museum of Natural History respectively. For the reconstruction additional 3D models of the lower jaws of species that were not present in the museum collection we received scan from several scientists. Therefore, we thank F. Mollen, M. Dean, K. Mara, P. Motta, D. Huber, R. Robins, C. Crawford, D. Walpole, C. Perry, A. Frew and R. Berquist for sharing their data with us. Finally, we thank T. Patel of the RBH for granting access to the CT scan facility and her technical assistance.
Author information
Authors and Affiliations
Contributions
P.K. and M.D.B. conceived and designed the project, P.K. reconstructed the 3D models and wrote the paper. P.K., P.W.D.B. and J.G. collected and reconstructed the CT scan data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Kamminga, P., De Bruin, P., Geleijns, J. et al. X-ray computed tomography library of shark anatomy and lower jaw surface models. Sci Data 4, 170047 (2017). https://doi.org/10.1038/sdata.2017.47
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2017.47
This article is cited by
-
Evolutionary trends in the elasmobranch neurocranium
Scientific Reports (2024)
-
Shark mandible evolution reveals patterns of trophic and habitat-mediated diversification
Communications Biology (2023)
-
Skeletal convergence in thunniform sharks, ichthyosaurs, whales, and tunas, and its possible ecological links through the marine ecosystem evolution
Scientific Reports (2023)
-
Cross-sectional anatomy, computed tomography, and magnetic resonance imaging of the banded houndshark (Triakis scyllium)
Scientific Reports (2021)
-
Facial model collection for medical augmented reality in oncologic cranio-maxillofacial surgery
Scientific Data (2019)