Abstract
European common ash, Fraxinus excelsior, is currently threatened by Ash dieback (ADB) caused by the fungus, Hymenoscyphus fraxineus. To detect and identify metabolites that may be products of pathways important in contributing to resistance against H. fraxineus, we performed untargeted metabolomic profiling on leaves from five high-susceptibility and five low-susceptibility F. excelsior individuals identified during Danish field trials. We describe in this study, two datasets. The first is untargeted LC-MS metabolomics raw data from ash leaves with high-susceptibility and low-susceptibility to ADB in positive and negative mode. These data allow the application of peak picking, alignment, gap-filling and retention-time correlation analyses to be performed in alternative ways. The second, a processed dataset containing abundances of aligned features across all samples enables further mining of the data. Here we illustrate the utility of this dataset which has previously been used to identify putative iridoid glycosides, well known anti-herbivory terpenoid derivatives, and show differential abundance in tolerant and susceptible ash samples.
Design Type(s) | parallel group design • disease analysis objective |
Measurement Type(s) | metabolite profiling |
Technology Type(s) | quadrupole time-of-flight mass spectrometry |
Factor Type(s) | phenotype • experimental condition • age • geographic location • biological replicate |
Sample Characteristic(s) | Fraxinus excelsior • leaf • Kingdom of Denmark |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
The globalisation of trade and logistical biosecurity challenges at port of entry has led to an increasing number of alien species invading countries where they have often adapted to new environments and infected exotic flora. Furthermore, climate change is likely to modify plant pathogen profiles, further contributing to emerging pathogens1. Most visible and socially impactful are tree pathogens, which have the potential to dramatically modify the landscapes of countries or even continents. Recent examples include Dutch elm disease which began in the UK and chestnut blight in the USA2. Today, European common ash (Fraxinus excelsior) is currently threatened by Ash dieback (ADB) which was first reported in the early 1990’s in north-eastern Poland3 and from where it has rapidly spread across Europe4–8. ADB, caused by the fungus Hymenoscyphus fraxineus, is currently found in most European countries, and was confirmed in the UK and Ireland in 2012. In the UK alone, where in excess of 100 million trees are at threat, it is estimated that over 1,000 species rely on ash trees for all or part of their lifecycle, including wood mice, squirrels, bullfinches, wrens, bats and beetles. Forty five of these species are considered obligate9. The causal agent of ash dieback, H. fraxineus most probably originated from East Asia. Hosoya et al.10 reported the presence of the fungus10,11 in Japan on its native host Manchurian ash (Fraxinus mandshurica) where it exhibits a hemi-biotrophic lifestyle12,13. By contrast, on a range of exotic hosts, not just exclusively Fraxinus spp. but extending into other Oleaceae, H. fraxineus displays a necrotrophic lifestyle. Interestingly, emerald ash borer (Agrilus planipennis), a native Chinese buprestid beetle, has devastated tens of millions of Fraxinus americana (American ash) in the USA14. Yet, both A. planipennis and H. fraxineus appear to co-exist with Manchurian ash in their native habitats.
The recent discovery of a genetic basis to ADB susceptibility provides the opportunity for selective breeding approaches15–18 that will facilitate the identification and propagation of superior ash trees. Tree breeding programmes have been established to select tolerant/resistant germplasm. Nevertheless, such programmes take many years to establish, ideally require access to replicated field trials, and need large breeding population sizes to avoid loss of rare alleles and maximisation of long-term commercial traits and may not reveal the genetic background behind the resistance. An ideal solution would be to have robust markers to assist in selective breeding for ADB and recent progress on whole genome sequencing19 and association transcriptomics20 will facilitate identification of molecular markers for assisted ash breeding and help provide a molecular understanding of the tolerance identified by field testing. However, these studies are confounded by the high degree of genetic heterozygosity due to extensive outcrossing, making the identification of genetic markers challenging. Moreover, breeding value assessments of Danish tolerant ‘Tree 35’ concluded that resistance was quantitative and that there was no evidence to suggest that resistance to ADB operating in F. excelsior was a consequence of a single or a few resistance genes (qualitative resistance), thus race specificity is unlikely21.
Despite the extensive genetic heterogeneity in European F. excelsior, ADB tolerant trees are increasingly being reported across Europe, e.g., Havrdová et al.22 and Muñoz et al.23. Given (i) the evidence for quantitative resistance, (ii) that resistance mechanisms of deciduous trees are thought to include a combination of constitutive and induced chemical defences21 and (iii) as wind borne ascospores are present for a sustained period over the summer, the mechanism underpinning foliar infection tolerance to ADB is highly likely to have a major constitutive component. Based upon this knowledge we undertook an unbiased global metabolic profiling of ash. We used material from the highly advanced Danish study that identified ‘Tree 35’21. We first tested whether the methodological approach could discriminate tolerant and susceptible ash. Then, using a second independent set of individuals, we undertook a detailed metabolomics profiling of these Danish ash. Here we provide our untargeted LC-MS metabolomics raw data from ash trees with high-susceptibility and low-susceptibility to ADB, a processed dataset containing abundances of aligned features across all samples, and use this dataset to highlight a family of small molecules from the iridoid glycoside class, that discriminate tolerant and susceptible ash. We demonstrate that iridoid glycosides, well known antifeedant molecules, are markedly reduced in tolerant ash leaves. The ecological and breeding implications of this is intriguing, as it implies breeding for ADB tolerance may unintentionally confer enhanced susceptibility to emerald ash borer. Given its presence in Russia, and hence representing an emergent threat to Europe, these findings certainly warrant further investigation.
Our study provides a wealth of potential information for future investigation and are the only set of replicated, unbiased LC-MS metabolite data from verified tolerant and susceptible European ash and these are available with associated metadata in Metabolights24,25 [MTBLS372]. Thus different data processing algorithms can be applied to reuse these data. As a striking 25% of the ash genome encoded unique (orphan) genes19, we see genuine utility in using this dataset to facilitate metabolite predictions to support studies aimed at identifying gene function. While we have focussed on the iridoid glycosides, the processed dataset contains features that discriminate tolerant trees and may be developed into rapid screens for ADB tolerant ash, e.g., in breeding trials.
Methods
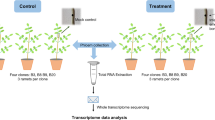
These methods are expanded from descriptions previously published in Nature19. A schematic overview of the methods are shown in Fig. 1.
(a) Sample processing and mass spectrometry. (b) Data extraction (MassHunter), peak identification, alignment (XCMS) and isotope/adduct annotation (CAMERA). (c) Statistics (JMP and MetaboAnalyst). (d) Fragment ion mass extraction and molecular formula prediction (MassHunter), and feature interpretation(Cytoscape & Molecular Structure Correlator). (e) Feature identification (Various databases and literature searches).
Collection of leaf material
Origin of genotypes used: Leaf material was sampled from trees with very different levels of susceptibility to ADB. Five trees were selected among the individuals exhibiting very low levels of symptoms when subject to natural infection of H. fraxineus: R14164C (HGH-A), R14184A (HGH-B), R14193A (HGH-C), R14198B (HGH-D), R14181 (HGH-E) and five trees were selected among trees exhibiting severe symptoms: R14127 (UGH-F), R14120 (UGH-G), R14169 (UGH-H), R14156 (UGH-I), and 25UTaps (UGH-J). All sampled trees were considered unrelated.
The R-trees were selected from a genetic field trial established in 2004 near Randers, Denmark (N56°50, E10°04). The trial comprised 2,448 trees derived from seed collected in 2001 from 101 mature Danish trees. The progenies were monitored annually for symptoms of natural infections from 2008–2011. Material used in this study was derived from selections made in 2012, by which time mortality had increased to 68%, based upon symptom presentation15.
The 25UTaps tree was selected among 39 clones based on the average level of crown damage assessed between 2007–2011 across the two Danish test sites at Tuse Næs (N55°45, E11°42) and Tapse (N55°24, E9°27). These represent mature trees selected in Denmark between 1934–1944 (except 25UTap and 27UTaps selected in 1994) and grafted on F. excelsior rootstock before establishment in a clonal trial16. The level of natural infections in both clonal trials was very high resulting in substantial mortality although all 39 genotypes survive as the clones were originally established with >50 replications (ramets) per clone.
ADB damage of the selected trees was re-assessed in trials in June 2013 and 2014, using the average score to characterise their level of susceptibility (Table 1). Three R-trees sampled as highly susceptible in 2012 were dead by 2013 and by 2016 all susceptible R trees were dead in Randers. 25UTaps is alive and still present in the clonal trials (Tuse Næs and Tapse). All healthy trees retained a 0–10% damage score in 2016, except for R14181 (HGH-E), which had some crown damage (25–50%).
Scions were collected from the trees in either Randers or Tapse during January and February 2013, grafted onto rootstocks of F. excelsior seedlings, potted and placed in a greenhouse at University of Copenhagen. Leaves were sampled in September 2014. All plants were symptom free at the time of sampling. Three leaves were sampled from each graft, flash frozen in liquid nitrogen and stored at −80 °C.
Sample processing
In order to understand whether ash trees with low and high susceptibility to ADB vary in their metabolite profiles as well as their transcriptomes, we undertook untargeted metabolite profiling on a subset of trees of Danish origin from a genetic field trial (R-trees)15 and a test panel (25UTaps)16. Untargeted metabolomics has not previously been applied to natural populations but has the potential to identify small molecules (or small molecule associations) that directly contribute to tolerance or resistance, particularly given limited evidence for qualitative resistance to ADB and that deciduous trees appear to deploy a combination of constitutive and induced chemical defences. We compared triplicate samples from five low-susceptibility Danish trees (R-14164C, R-14184A, R-14193A, R-14198B and R-14181) and five high-susceptibility trees (R-14169, R-14127, R-14156 R-14120 and 25UTaps). Three leaflets from each triplicate sample were freeze dried and gently crushed to mix tissue types. Approximately 100–150 mg of this material was ground to a fine powder in a 2 ml polypropylene microfuge tube using a TissueLyser (Qiagen; 2×1 min at 25 Hz). 10 mg was extracted in 400 μl 80% MeOH containing d5-IAA internal standard at 2.5 μg/ml ([2H5] indole-3-acetic acid; OlChemIm Ltd, Czech Republic), centrifuged (10,000 g, 4 °C, 10 min) and the pellet re-extracted in 80% MeOH. The pooled supernatants were filtered through a 0.2 μm PVDF syringe tip filter (Chromacol, Thermo Scientific, MA, USA).
Mass spectrometry
These leaf extracts were analysed using a Polaris C18 1.8 μm, 2.1×250 mm reverse phase analytical column (Agilent Technologies, Palo Alto, USA). 5 μl samples were resolved on an Agilent 1200 series Rapid Resolution HPLC system coupled to a quadrupole time-of-flight QToF 6520 mass spectrometer (Agilent Technologies, Palo Alto, USA). Mobile phases were as follows: positive ion mode; mobile phase A (5% acetonitrile, 0.1% formic acid), mobile phase B (95% acetonitrile with 0.1% formic acid). Negative ion mode; mobile phase A (5% acetonitrile with 1 mM ammonium fluoride), mobile phase B (95% acetonitrile). The following gradient was used: 0–10 min-0% B; 10–30 min-0–100% B; 30–40 min-100% B. The flow rate was 0.25 ml min−1 and the column temperature was held at 35 °C throughout. The source conditions for electrospray ionisation were as follows: gas temperature was 325 °C with a drying gas flow rate of 9 l min−1 and a nebuliser pressure of 35 psig. The capillary voltage was 3.5 kV in both positive and negative ion mode. The fragmentor voltage was 115 V and skimmer 70 V. Scanning was performed using the auto MS/MS function at 5 scans sec−1 for precursor ion surveying and 4 scans sec−1 for MS/MS with a sloped collision energy of 3.5 V/100 Da with an offset of 5 V. Scan speed varied based on precursor abundance with a maximum of 1.15 s between MS and final (5th) MS/MS (in practice this was never observed to exceed 0.8).
Data processing
Positive and negative ion data (centroid) were converted into mzData using the export option in Agilent MassHunter. Peak identification and alignment was performed using the Bioconductor R package XCMS26 and features were detected using the centWave method27 for high resolution LC/MS data in centroid mode at 30 ppm. The samples were grouped into ‘tolerant’ and ‘susceptible’ folders/groups before processing and all samples were aligned and peaks identified in a single batch. Changes to the default parameters were: mzdiff=0.01, peakwidth=10–80, noise=1,000, prefilter=3,500. Peaks were matched across samples using the density method with a bw=5 and mzwid=0.025 and retention time correlated using the obiwarp algorithm28 with profStep=0.5. Missing peak data was filled in the peaklists generated from the ADB low susceptibility ash leaf samples compared to the peaklists generated from the ADB susceptible leaves. The resulting peaklists were annotated using the Bioconductor R package, CAMERA29. The peaks were grouped using 0.05% of the width of the full width at half maximum (FWHM) and groups correlated using a P-value of 0.05 and calculating correlation inside and across samples. Isotopes and adducts were annotated using a 10 ppm error.
Statistics
Statistical analysis and modelling was performed using MetaboAnalyst v3.02830,31, with the following parameters. Missing values were replaced using a (K-nearest neighbour) KNN missing value estimation. Data was filtered (40%) to remove non-informative variables using the interquartile range (IQR). Samples were normalised using the internal standard d5-IAA (POS: M181T1448; NEG: M179T1382). Data was auto-scaled.
Peaks from the three replicates, run in positive and negative mode were aligned with XCMS and features tested for practical significance to determine the differences between the tolerant and susceptible genotypes. In addition, PLS-DA was performed using MetaboAnalyst 3.0 allowing clear discrimination of tolerant and susceptible genotypes based on their metabolic profiles (Fig. 2a).
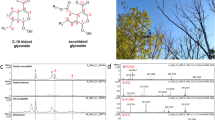
(a) Multivariate analysis PLS-DA score plot showing discrimination of five susceptible and five tolerant trees, each replicated three times, based on their leaf metabolite profiles. (b) MS/MS fragmentation network of LC-ESI-MS/MS fragmentation data for discriminatory features of high and low susceptibility genotypes. The size of each circle represents the fragment size. The intensity of blue increases as the number of times each fragment is present in all precursor ions increases. The edges are in shades of red based on retention time; the paler the colour the earlier the retention time. The black (POS) and grey (NEG) features are the discriminatory features. The outside ring shows fragments unique to that precursor ion (i.e., not in the other precursor ions). The inside circle is ‘shared’ fragments, present in more than one precursor ion. Those fragment masses shaded in green have been previously reported from fragmentation of iridoid glycosides41. Figure adapted from Sollars et al.19.
The individual features (putative metabolites) that contribute to the separation between the different classes were further characterised. We first applied a range of univariate and multivariate statistical tests to determine the importance of these features. This included variable influence on the projection (VIP) values derived from PLS-DA scores, practical significance, t-test, P-value, Benjamini and Hochberg FDR (False Discovery Rate) adjusted P-value, effect size, Random Forest analysis and MS/MS fragmentation network analysis. For example, using Random Forest, significant features were ranked by mean decrease in classification accuracy with 14/15 susceptible samples (OOB error: 0.033; class error 0.07) and 15/15 tolerant samples correctly classified. For all further analyses we chose to use statistical and practical significance (Response screening, JMP version 12) to identify features with a practical significance and validate these using a MS/MS fragmentation network (Fig. 2b). Features that were shown to be of interest using the above statistical tests were individually checked by returning to the raw data and determining if they had been properly identified and aligned by XCMS. We also checked that those features of interest were absent in the blanks and the extraction buffer only samples.
Putative feature identification
Putative identification of features was performed using several approaches. Each feature of interest was analysed using MassHunter molecular formula estimation. This information, along with accurate mass and MS/MS spectra were used to interrogate existing literature, and databases including, but not limited to, KNApSAcK (http://kanaya.naist.jp/KNApSAcK/), Metlin (https://metlin.scripps.edu), ReSpect (http://spectra.psc.riken.jp/), PubChem (https://pubchem.ncbi.nlm.nih.gov/), ChemSpider (http://www.chemspider.com/), mzCloud (https://www.mzcloud.org/) and Massbank (http://www.massbank.jp/). Additionally, features of interest identified in negative mode were extracted from MassHunter as CEF (compound exchange format) files. These were imported into MSC and searches against ChemSpider and PubChem databases and a custom database using mol files generated from papers. Searches were performed with default parameters apart from the minimum MSC score was set to 50.0. The following parameters were used: MS/MS isolation window was set to first isotope only with default ionization set to protons. Multiple C.E. treatment was set to simple average. Only the 30 most abundant ions were used and formulae contained in the data files was used (i.e., from MassHunter Molecular Formula Prediction). Elements used for MFG composition were H: 2–1,000, C: 1–1,000, N 0–1,000, O:0–1,000 and S: 0–1,000. Height uncertainty was set to 7.5%. Only a representative tautomer was set, the number of total rings and the number of aromatic rings were unconstrained.
Negative mode feature identification
N1
The fragmentation pattern of feature N1 shows a neutral loss of 162 Da (813→651) indicating a possible loss of hexose. Using MassHunter, we predicted the aglycone fragment had a molecular formula estimation of C30H35O16 (diff=C6H11O5) and therefore a potential formula of C36H46O21.
N2
N2 also shows a neutral loss of 162 Da consistent with loss of hexose (565→403). The exact mass is close to a secoiridoid glucoside identified in Ligustrum japonicum (Oleaceae)32 with a molecular formula of C23H34O16. It is also the predicted molecular formula based on isotopic distribution (MassHunter) with a score of 98.44. Although fraxiresinol hexoside has the same mass, the MS/MS fragment sizes are different with abundant peaks of 181 and 373. Another possible candidate is demethylated Excelside A.
N3
N3 is predicted to be (7 R/S)-10-Hydroxy-7-methoxyoleuropein (Ligustrum vulgare (Oleaceae))33, a formic acid [M+FA-H]− adduct of the bitter secoiridoid glucoside Oleuropein/Oleuroside or an acetate [M+CH3COO]− adduct of Demethyloleuropein.
N4
Using molecular formula prediction N4 was suggested to be C31H42O17 (score: 99.47). A neutral loss of 685→453 (232) indicates a loss of C10H16O6 followed by a neutral loss of 453→299 (154), which is likely to be C7H6O4. The mass of the molecule is similar to that of Excelside B/Nuezhenide which has previously been identified in F. excelsior seeds34. Additionally, Nuezhenide has been previously reported to have fragment masses of 453.1389 and 299.113035 which corresponds to m/z peaks of 453.1424 and 299.1108 in the ash leaf spectrum.
N5
N5 has a similar fragmentation pattern and retention time to N3 with a mass difference of 384.12 (i.e., C21H20O7). The abundant 403 fragment has the same mass as Oleoside 11-methyl ester/Oleoside 7-methyl ester and shares a fragmentation fragment (223) with these isomers suggests a glycosylated dimer of Oleoside 11-methyl ester or Oleoside 7-methyl ester (((404×2)−1)+162=969)35.
Positive mode feature identification
P1
P1a and its M+1 isotope, (predicted by CAMERA to be an ammonium adduct ([M+H+NH3]+ 686.568) was confirmed by MassHunter as being an ammonium adduct with a formula of C31H42O17 correspond to Excelside B/Nuezhenide. In corroboration, a sodium adduct identified in the practically significant dataset (P1b) and its M+1 isotope, predicted by MassHunter to be C31H42O17 with a sodium adduct. 709→547 represents the loss of hexose. Although the fragmentation patterns of adducts are different it was also detected in negative mode at the same retention time (N4).
P2
P2b 589→427, neutral loss of 162 indicates a hexose moiety. CAMERA suggests a sodium adduct, and MassHunter confirms this as a sodium adduct with a predicted formula of C23H34O16 which corresponds to a secoiridoid glucoside from L. japonicum32, most likely dimethyl-Excelside A, which was also putatively detected in negative mode (N2). There was also an ammonium adduct represented by P2a and its isotope. This is supported by MassHunter as an ammonium adduct with a predicted molecular formula of C23H34O16. As with the Excelside/Nuezhenide feature, adducts fragment differently.
P3/4
Both P3 and P4 have a similar mass (m/z: 225.0759, 225.0752 respectively) but elute at different retention times (1,236 and 1,399 s respectively). They have the same predicted formula of 12O5 and share some similar fragment ions (95, 151, 165). This could be sinapic acid although based on the spectrum of sinapic acid (http://www.massbank.jp, MCH00015, PR020014, PR101042), the base peak for this compound should be 207. It could be a fragment from an iridoid glucoside, which elutes at the same retention time, some of which have fragment ions with a mass of 225 Da. For example, the ammonium adduct of the Excelside A-like feature P2a which elutes at 1,236 s and the ammonium adduct of the Excelside B-like feature P1a which elutes at 1,398 s all contain fragment ions with a mass of 165 and 151 Da.
P5
The isomer of P1, predicted to be Excelside B/Nuezhenide is found as a sodium adduct, P5b, along with its isotope and as an ammonium adduct P5a, although the isomer of the ammonium adduct was not significant.
P6
P6b and P6a are sodium and ammonium adducts of the same compound. For the sodium adduct, P6b there is a loss of 162 (501→339) indicating a hexose moiety. The predicted molecular formula of both features is C20H30O13. This is the same molecular formula as the antibacterial phenolic apioglucoside Forsythoside D from Forsythia suspensa (Oleaceae)36 and kelampayoside A. P6 is possibly an isomer of kelampayoside A as the fragmentation pattern is different to previously reported in Trachelospermum jasminoides which had fragment ions of 411 and 369 for the sodium adduct37, whereas the F. excelsior compound has a base peak of 339.
P7
The sodium adduct, P7b and its isotope M929T1536 and the ammonium adduct, P7a and its isotopes M934T1536 ([M+1]) and M935T1536 ([M+2]) suggest a predicted compound formula of C42H54O22. This is the molecular formula of Jaspolyanoside identified in Jasminum polyanthum (Oleaceae)38 and Olea europaea (Oleaceae)39. This compound is an aromatic conjugated secoiridoid glucoside and the loss of 162 Da (933→771) is consistent with it being a glucoside.
Data Records
For this study, we submitted the raw untargeted LC-MS metabolomics data for three replicates of 5 susceptible and 5 tolerant ash dieback genotypes of Danish F. excelsior run in both positive and negative mode to MetaboLights24,25 (Data Citation 1). Additionally, the XCMS alignment files were deposited (Data Citation 1). The datasets described (Data Citation 1) were previously published in our related work in the journal Nature19.
Technical Validation
Experimental
Samples were randomised with blanks run after every 5 samples. An aliquot of extraction buffer including the internal standard were run at the beginning (RT=1443.48 s), middle (RT=1442.07 s) and end (RT=1443.78 s) of the run to assess retention time drift. There was a maximum retention time difference of 1.08 s. The maximum retention time difference based on the internal standard in the samples was 3.24 s in positive mode and 7.44 s in negative mode. Retention time values are shown in Table 2. Internal standards had relative standard deviations (RSD) values of 10.6% in negative mode 14.3% in positive mode and ppm differences of −1.3 ppm (negative mode) and −11.4 ppm (positive mode) from theoretical monoisotiopic mass for d5-IAA. Features described in this study had a ppm difference on average of −1.3 ppm difference from the theoretical value with a maximum difference of 5.4 ppm. To assess the experimental error, the mean coefficient of variation (CV) was calculated for each samples in positive and negative mode by using the peak areas of all features reported by XCMS after missing values were imputed by MetaboAnalyst (Table 3).
Statistics
Several methods were used to validate discriminatory features including equivalence testing, PLS-DA and random forest analysis as described in the methods section.
Putative feature identification
Several approaches were taken to validate the putative identification of discriminatory features. An MS/MS fragmentation network was generated after extracting the m/z of the MS/MS product ions (where abundance >5% and m/z >100) from the 13 discriminatory features identified in positive and negative mode using MassHunter Qualitative Analysis Version 4 and visualized using Cytoscape40 indicating product ion masses which have been previously reported from fragmentation of iridoid glycosides41. Further validation was performed through a mirror plot (Fig. 3) comparing the MS/MS spectra of four features (N2–5) detected in negative mode with an ESI-TOF/IT-MS spectra of elenolic acid glucoside taken from the literature42. Finally, the accurate mass of MS/MS product ions from four discriminatory features identified in negative mode (N2-N4) were correlated with the structure of the putatively identified feature using MassHunter Molecular Structure Correlator (Agilent) (Fig. 4).
The spectra share 4 peaks in common, m/z 179, 223, 371 and 403 (elenolic acid glucoside molecular ion). These fragments correspond to a loss of a methyl and hydroxyl group (403–371), loss of hexose (403–223) which is followed by a loss of CO2 (223–179). Elenolic acid corresponds to the secoiridoid part of oleuropein-related compounds suggesting that these four features are secoiridoids42. Figure from Sollars et al.19 extended data.
Predicted structures for three key m/z peaks using Molecular Structure Correlator (Agilent; http://www.agilent.com/cs/library/usermanuals/public/G3335-90176_MSC_QuickStart.pdf) and the structure of putative IDs. Features of interest were compared to the closest known compounds based on molecular formula and accurate mass. The Excelside A-like feature s demethylated Excelside A. The overall score (top right) and individual fragment scores (below molecular formula) obtained from MSC are shown. Bonds and atoms in black are present in that fragment, whereas grey indicates loss. The molecular formula and the MSC score are displayed next to each fragment structure. CID=collision induced dissociation. Figure adapted from Sollars et al.19 extended data.
Identification was not possible for those features with no fragmentation data, or lacking significant supporting adducts. Many additional features of interest were identified but require further validation to allow confident attributions, while some features did not provide fragmentation patterns. We thus restricted further identification and characterisation to discriminatory features from the iridoid glycoside class of compounds. These iridoid gylcosides, which have been previously documented in Oleaceae, are summarised in Supplementary Table S1 and Supplementary Fig. 1. Fraxinus excelsior is a member of the Olive family (Oleaceae), therefore data reported for this family were used to aid identification. To further validate our identifications we used an MS/MS fragmentation network approach and graphically illustrate the resulting network in Fig. 2b.
Based on ESI fragmentation patterns, the majority of features identified as having a practical significant difference between tolerant and susceptible genotypes of F. excelsior are likely to be from the same class of compounds. Common fragment ions are evident in many of these discriminating features, including m/z 179, 223 and 403, which are indicative of elenolic acid glucoside related molecules. Molecular networking confirms that these features are likely to be from the same family of compounds and we provide a detailed putative identification of each feature that is represented by box plots and spectra (Supplementary Fig. 1).
Usage Notes
The general approach for sample processing and data analysis is described in the schematic (Fig. 1). Links to the software, resources and repositories used are stated below.
-
XCMS26: https://bioconductor.org/packages/release/bioc/html/xcms.html
-
CAMERA29: https://bioconductor.org/packages/release/bioc/html/CAMERA.html
-
MetaboAnalyst 3.030,31: http://www.metaboanalyst.ca/
-
JMP®, Version 12: SAS Institute Inc., Cary, NC, 1989–2007: https://www.jmp.com
-
CytoScape40: http://www.cytoscape.org/
-
Molecular Structure Correlator (Agilent):] http://www.agilent.com/cs/library/usermanuals/public/G3335-90176_MSC_QuickStart.pdf
-
MetaboLights24,25: http://www.ebi.ac.uk/metabolights/
-
MassHunter Qualitative Analysis Version 4: http://www.agilent.com/cs/library/usermanuals/Public/G3336-90018_Qual_Familiarization.pdf
Additional information
How to cite this article: Sambles, C. M. et al. Ash leaf metabolomes reveal differences between trees tolerant and susceptible to ash dieback disease. Sci. Data 4:170190 doi: 10.1038/sdata.2017.190 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Fisher, M. C. et al. Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 186–194 (2012).
Woodward, S. & Boa, E. Ash dieback in the UK: a wake-up call. Mol. Plant Pathol. 14, 856–860 (2013).
Przybyl, K. Fungi associated with necrotic apical parts of Fraxinus excelsior shoots. Forest Pathol 32, 387–394 (2002).
Gross, A., Holdenrieder, O., Pautasso, M., Queloz, V. & Sieber, T. N. Hymenoscyphus pseudoalbidus, the causal agent of European ash dieback. Mol. Plant Pathol. 15, 5–21 (2014).
Luchi, N., Montecchio, L. & Santini, A. Situation with Ash in Italy: stand characteristics, health condition, ongoing work and research needs. COST Action FP1103 FRAXBACK 1st MC/WG Meeting, Vilnius, Lithuania (2012).
Solheim, H., Timmermann, V., Talgø, V. & Røsberg, I. Ash dieback in Norway. Forstschutz Aktuell 55, 49–51 (2011).
Halmschlager, E. & Kirisits, T. First report of the ash dieback pathogen Chalara fraxinea on Fraxinus excelsior in Austria. Plant Pathol 57, 1177–1177 (2008).
Bakys, R., Vasaitis, R., Barklund, P., Thomsen, I. M. & Stenlid, J. Occurrence and pathogenicity of fungi in necrotic and non-symptomatic shoots of declining common ash (Fraxinus excelsior) in Sweden. European Journal of Forest Research 128, 51–60 (2009).
Lawrence, R. & Cheffings, C. M. A summary of the impacts of ash dieback on UK biodiversity, including the potential for long-term monitoring and further research on management scenarios. Joint Nature Conservation Committee Report 501 (2014).
Hosoya, T., Otani, Y. & Furuya, K. Materials for the fungus flora of Japan (46). Transactions of the Mycological Society of Japan 34, 429–432 (1993).
Zhao, Y. J., Hosoya, T., Baral, H. O., Hosaka, K. & Kakishima, M. Hymenoscyphus pseudoalbidus the correct name for Lambertella albida reported from Japan. Mycotaxon 122, 25–41 (2012).
Cleary, M. et al. Friend or foe? Biological and ecological traits of the European ash dieback pathogen Hymenoscyphus fraxineus in its native environment. Sci. Rep. 6, 21895 (2016).
Cleary, M. R., Arhipova, N., Gaitnieks, T., Stenlid, J. & Vasaitis, R. Natural infection of Fraxinus excelsior seeds by Chalara fraxinea. Forest Pathol 43, 83–85 (2013).
MacQuarrie, C. J. K., Scarr, T. A. & Ryall, K. L. The science of the emerald ash borer (Coleoptera: Buprestidae): where are we after 10 years of research? Can Entomol 147, 249–251 (2015).
Kjaer, E. D., McKinney, L. V., Nielsen, L. R., Hansen, L. N. & Hansen, J. K. Adaptive potential of ash (Fraxinus excelsior) populations against the novel emerging pathogen Hymenoscyphus pseudoalbidus. Evolutionary applications 5, 219–228 (2012).
McKinney, L. V., Nielsen, L. R., Hansen, J. K. & Kjaer, E. D. Presence of natural genetic resistance in Fraxinus excelsior (Oleraceae) to Chalara fraxinea (Ascomycota): an emerging infectious disease. Heredity 106, 788–797 (2011).
Pliura, A., Lygis, V., Suchockas, V. & Bartkevicius, E. Performance of Twenty Four European Fraxinus excelsior Populations in Three Lithuanian Progeny Trials with a Special Emphasis on Resistance to Chalara fraxinea. Balt For 17, 17–34 (2011).
Stener, L. G. Clonal differences in susceptibility to the dieback of Fraxinus excelsior in southern Sweden. Scand J Forest Res 28, 205–216 (2013).
Sollars, E. S. et al. Genome sequence and and genetic diversity of European ash trees. Nature 541, 212–216 (2017).
Harper, A. L. et al. Molecular markers for tolerance of European ash (Fraxinus excelsior) to dieback disease identified using Associative Transcriptomics. Sci Rep 6, 19335 (2016).
McKinney, L. V. et al. The ash dieback crisis: genetic variation in resistance can prove a long-term solution. Plant Pathol 63, 485–499 (2014).
Havrdová, L. et al. Differences in susceptibility to ash dieback in Czech provenances of Fraxinus excelsior. Forest Pathol 46, 281–288 (2016).
Muñoz, F., Marçais, B., Dufour, J. & Dowkiw, A. Rising Out of the Ashes: Additive Genetic Variation for Crown and Collar Resistance to Hymenoscyphus fraxineus in Fraxinus excelsior. Phytopathology 106, 1535–1543 (2016).
Haug, K. et al. MetaboLights-an open-access general-purpose repository for metabolomics studies and associated meta-data. Nucleic Acids Res 41, 781–786 (2013).
Kale, N. S. et al. MetaboLights:An Open-Access Database Repository for Metabolomics Data. Curr Protoc Bioinformatics 53, 14.13.1–14.13.18 (2016).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. Xcms: Processing mass spectrometry data for metabolite profiling using Nonlinear peak alignment, matching, and identification. Anal Chem 78, 779–787 (2006).
Tautenhahn, R., Bottcher, C. & Neumann, S. Highly sensitive feature detection for high resolution LC/MS. BMC Bioinformatics 9, 504 (2008).
Prince, J. T. & Marcotte, E. M. Chromatographic alignment of ESI-LC-MS proteomics data sets by ordered bijective interpolated warping. Anal Chem 78, 6140–6152 (2006).
Kuhl, C., Tautenhahn, R., Bottcher, C., Larson, T. R. & Neumann, S. Camera: an integrated strategy for compound spectra extraction and annotation of liquid chromatography/mass. Anal Chem, spectrometry data sets 84, 283–289 (2012).
Xia, J., Sinelnikov, I. V., Han, B. & Wishart, D. S. MetaboAnalyst 3.0-making metabolomics more meaningful. Nucleic Acids Res 43, 251–257 (2015).
Xia, J. & Wishart, D. S. Using MetaboAnalyst 3.0 for Comprehensive Metabolomics Data Analysis. Curr Protoc Bioinformatics 55, 14.10.1–14.10.48 (2016).
Kuwajima, H. et al. A secoiridoid glucoside from Ligustrum japonicum. Phytochemistry 28, 1409–1411 (1989).
Tanahashi, T. et al. Secoiridoid glucosides and unusual recyclized secoiridoid aglycones from Ligustrum vulgare. Phytochemistry 70, 2072–2077 (2009).
Bai, N. et al. Iridoids from Fraxinus excelsior with adipocyte differentiation-inhibitory and PPARalpha activation activity. Journal of Natural Products 73, 2–6 (2010).
Kanakis, P. et al. From olive drupes to olive oil. An HPLC-orbitrap-based qualitative and quantitative exploration of olive key metabolites. Planta Med 79, 1576–1587 (2013).
Endo, K. & Hikino, H. Structures of Forsythoside C and D, antibacterial principles of Forsythia suspensa fruits. Heterocycles 19, 2033–2036 (1982).
Liu, X. T. et al. Qualitative and quantitative analysis of lignan constituents in Caulis trachelospermi by HPLC-QTOF-MS and HPLC-UV. Molecules 20, 8107–8124 (2015).
Tanahashi, T. et al. Six secoiridoid glucosides from Jasminum polyanthum. Chemical and pharmaceutical bulletin 45, 367–372 (1997).
Pérez-Bonilla, M. et al. Isolation of antioxidative secoiridoids from olive wood (Olea europaea L.) guided by on-line HPLC-DAD-radical scavenging detection. Food Chemistry 124, 36–41 (2011).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504 (2003).
Li, C. M. et al. Structural characterization of iridoid glucosides by ultra-performance liquid chromatography/electrospray ionization quadrupole time-of-flight tandem mass spectrometry. Rapid communications in mass spectrometry: RCM 22, 1941–1954 (2008).
Gupta, S. D. Reactive Oxygen Species and Antioxidants in Higher Plants p323 (CRC, 2010).
Data Citations
Sambles, C. M. MetaboLights MTBLS372 (2016)
Acknowledgements
CMS was funded by the ‘Nornex’ project jointly by UK BBSRC (BBS/E/J/000CA5323) and DEFRA and a BBSRC Tools and Resources grant (BB/N021452/1) awarded to M.G. and D.J.S.
Author information
Authors and Affiliations
Contributions
C.M.S. conceived and designed the experiments, developed methodology, analysed the data, interpreted and validated the data and wrote the manuscript. M.G. conceived and designed the experiments, interpreted and validated the data and wrote the manuscript D.J.S. conceived and designed the experiments. D.L.S. processed samples. HF ran the mass-spectrometer. T.P.H. assisted with statistics. N.S. assisted with methodology and validation R.J.A.B. assisted with interpretation and presentation. E.D.K. generated, selected and collected leaf samples. L.R.N. generated, selected and collected leaf samples. L.V.M. generated, selected and collected leaf samples. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Sambles, C., Salmon, D., Florance, H. et al. Ash leaf metabolomes reveal differences between trees tolerant and susceptible to ash dieback disease. Sci Data 4, 170190 (2017). https://doi.org/10.1038/sdata.2017.190
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2017.190
This article is cited by
-
Transcriptional responses in developing lesions of European common ash (Fraxinus excelsior) reveal genes responding to infection by Hymenoscyphus fraxineus
BMC Plant Biology (2020)
-
Diversity of secoiridoid glycosides in leaves of UK and Danish ash provide new insight for ash dieback management
Scientific Reports (2020)