Abstract
We provide a detailed description of a gadoteridol-derivatized lysozyme (gadolinium lysozyme) two-colour serial femtosecond crystallography (SFX) dataset for multiple wavelength anomalous dispersion (MAD) structure determination. The data was collected at the Spring-8 Angstrom Compact free-electron LAser (SACLA) facility using a two-colour double-pulse beam to record two diffraction patterns simultaneously in one diffraction image. Gadolinium lysozyme was chosen as a well-established model system that has a very strong anomalous signal. Diffraction patterns from gadolinium lysozyme microcrystals were recorded to a resolution of 1.9 Å in both colours. This dataset is publicly available through the Coherent X-ray Imaging Data Bank (CXIDB) as a resource for algorithm development.
Design Type(s) | protocol testing objective |
Measurement Type(s) | X-ray diffraction data |
Technology Type(s) | X-ray free electron laser |
Factor Type(s) | |
Sample Characteristic(s) | Gallus gallus |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
SACLA is one of the two currently operating X-ray free-electron laser (XFEL) facilities in the world (with several XFELs coming online this year) producing a hard X-ray beam for the investigation of matter, with various applications in biology1,2, chemistry, physics and material science. In general, XFELs have the capacity to deliver a pulsed beam that is ten billion times brighter than synchrotron X-ray beams, with pulses ten thousand times shorter. So far, single wavelength anomalous dispersion (SAD)3–10 and single or multiple isomorphous replacement (SIR10,11, MIR12) methods have been used for de novo structure determination with XFELs to solve the phase problem. However, the stochastic nature of the experiment (randomly oriented crystals, strongly varying pulse intensity and wavelength distribution) demands large amounts of sample and beam time to average out the stochastic effects. Since 2013, two distinct X-ray pulses13 that have an unprecedentedly large energy separation can be created simultaneously by the split undulator operation of the SACLA XFEL. Thus, a large and flexible wavelength separation of the double-pulse of more than 30% with a precisely controlled time interval down to the attosecond regime can be realized. This allows targeting of multiple absorption edges of a suitable anomalous scatterer simultaneously. Using this operation mode of the SACLA XFEL, two distinct datasets can be recorded from the same sample thereby reducing the costs in both beam time and sample for structure determination significantly.
Here we describe the deposition of a two-colour serial femtosecond crystallography (SFX) dataset acquired at SACLA as reported in Gorel et al.14. The two diffraction patterns were first identified and then processed individually for successful MAD phasing. Analysis of this dataset showed that 5,000 images were required for MAD-based de novo structure determination. Furthermore, it was shown that phases retrieved by MAD phasing are more accurate than phases retrieved by SAD phasing using the same number of diffraction images.
Methods
Data acquisition
The two-colour experiment (proposal number 2015B8045) was performed in January 2016 at the SACLA XFEL in Hyogo, Japan. The photon energies for the two colours were chosen to be above the M-edges (7 keV) and L-edges (9 keV) of gadolinium, respectively. A High Viscosity Extrusion injector15 (HVE) mounted in the DAPHNIS16 chamber was used to introduce gadolinium lysozyme microcrystals14 suspended in grease medium17 into the two-colour double-pulsed X-ray beam. X-ray diffraction data was recorded at beamline 3 at the SACLA facility using the multiport CCD (MPCCD18) detector. Two diffraction patterns, one for each colour, were acquired simultaneously in one diffraction image.
Data analysis
Data analysis was performed on the SACLA High Performance Computing Cluster consisting of several steps of parameter optimization and special data processing for the two-colour data. In the following we provide a more detailed description of the work published in Gorel et al.14.
Raw data hit files
During a period of 12 h, a dataset of 570,000 diffraction images with consistent experimental parameters (attenuation, transmission, detector distance etc.) was collected. From this set 208,373 hits were identified with the programme Cheetah19 (40.6% of the total dataset). The raw data images were obtained using the Cheetah Dispatcher20 graphical user interface (GUI) from the set of all recorded images as a subset of images with more than 20 peaks. By default, this data contains only the wavelength of the 7 keV colour pulse saved in the HDF5 dataset 'photon_energy_ev' and 'photon_wavelength_A' in the diffraction image. The raw images were written in the multi-event data format and thus had to be split into individual diffraction images using the split.py module before further data processing was performed.
Silicon powder files
Silicon powder patterns were recorded and used for detector distance parameter determination. To this end, grease was mixed with silicon nanocrystals and attached to the injector capillary tip. To calculate the detector distance, the radii of the Debye Scherrer diffraction rings had to be retrieved. However, due to the low concentration of silicon nanocrystals the high resolution powder rings were faint and could not be distinguished from the background by a threshold approach. This made another pattern recognition technique necessary. Thus, 300 interest points14 (i.e., points with certain characteristics like signal over background and signal divided by background exceeding a dynamically calculated value) per CCD (2,400 in total) were calculated for each diffraction image of the silicon powder data and written into the HDF5 dataset 'poi' in the diffraction image container. Using these interest points the detector distance was calculated to be 51.03 mm as described in the supplementary information of Gorel et al.14.
Spectra files
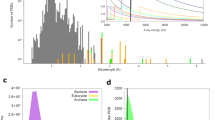
By default, the raw-images in HDF5 file format contain only the wavelength value for one particular colour because the energy separation of the two colours exceeds the spectral range of the narrow range inline spectrometer. Thus, only the 7 keV colour or the 9 keV colour wavelength is available for each shot but not both at the same time. Since knowledge of the wavelength values of both colours is required for the two-colour data processing, the missing wavelength had to be retrieved from the recordings of a wide range inline spectrometer21. To this end, the measured spectral profiles were recorded into spectra.h5 files for later use by the write_calib_color.py module to obtain accurate wavelength values and to add these to the diffraction images. The wide range inline spectrometer images were obtained with the application programming interface to SACLA metadata database. Spectral profiles were obtained from these 1,024×512 pixel images by collapsing them into a 1,024 pixel sized one-dimensional image. These profiles were recorded into a HDF5 dataset named 'spectrum' within the spectra.h5 files by the write_spectra.py module. A double Lorentzian model was fitted to these profiles resulting in values for the amplitudes of both peaks, a constant offset, the positions of both Lorentzians and their widths. These parameters were written into the HDF5 datasets 'Amp1', 'Amp2', 'Const', 'Peak1', 'Peak2', 'Width1' and 'Width2', respectively, in the spectra.h5 file by the write_spectra.py module. The HDF5 dataset 'tags' in the spectra.h5 file contains the tag name of the respective diffraction image and thus identifies the set of parameters of the two-colour double-pulse that belongs to the respective diffraction event. Two calibration runs are available (run 392,732 for 7 keV and run 392,738 for 9 keV colour) which were used to find the photon energy calibration functions for the respective energy ranges of the wide range inline spectrometer.
Since the resolving power of the narrow range spectrometer is smaller than that of the wide range spectrometer, the same energy value is retrieved for different readings of the wide range spectrometer. By calculating the median values from the wide range spectrometer readings that have the same narrow range spectrometer reading, calibration points were obtained, which were then used for the photon energy calibration function estimation as described in Gorel et al.14 supplement.
7 keV and 9 keV indexable files
The respective photon energy calibration function was applied by the write_calib_color.py module to the double-pulse energy profile fit parameters. This way the photon energy was obtained and written into the HDF5 datasets 'photon_energy_ev_color1', 'photon_energy_ev_color2', 'photon_wavelength_A_color1' and 'photon_wavelength_A_color2', where color1 corresponds to the 7 keV and color2 to 9 keV colour. The raw data was processed by CrystFEL's22 indexamajig with the colour information added, against given cell parameters (a=b=78.3 Å, c=39.1 Å α=β=γ=90°). All correctly indexed images are contained in the hits7kev (21,830) and hits9kev (33,297) datasets.
Two-colour indexable files
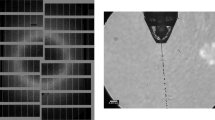
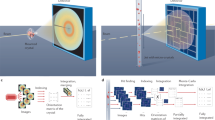
Due to an anti-correlation of the double-pulse intensities and thus of the respective diffraction pattern intensities, the two-colour images typically contain one strong and one weak diffraction pattern. Because two completely different diffraction patterns are present in the diffraction image (see Fig. 1) and since the processing programme can deal with only one pattern at a time, processing of the two-colour diffraction data is not straightforward and was achieved by the following three steps: Firstly, the diffraction images were processed with CrystFEL's indexamajig module for the brighter colour identifying the stronger diffraction pattern in the diffraction image (threshold 200, signal-to-noise ratio 5). Secondly, before the other diffraction pattern of the weaker colour could be processed, the peak search parameters, i.e., threshold and signal-to-noise ratio, were lowered to select a broader set of diffraction peaks by thresholding (threshold 150, signal-to-noise ratio 3). Then, all peaks of the stronger diffraction pattern were removed with the write_subtract.py module from this set such that the residual peaks possibly constituted the weaker diffraction pattern. Subsequently, in the third step these peaks were processed with indexamajig. To compensate for the residual error in peak position prediction, large integration radii were applied with indexamajig (--int-radius=6,6,8).
(a) The two similar diffraction patterns in this image likely belong to a twinned crystal. Indexing in a second colour was not possible. (b) The diffraction patterns from 7 keV (red) and 9 keV (blue) strongly differ since very different areas in reciprocal space are probed by the two-colour double-pulse.
For two-colour indexing all 9 keV indexable images from the hits9kev dataset were reprocessed to identify the second (weak) diffraction pattern. These points were saved in the HDF5 dataset 'residual_points_7keV' in the diffraction images and were processed with CrystFEL’s indexamajig module. Thus, 14,782 two-colour indexable images were found from the 33,297 images of the hits9kev data subset (44.4%). An overview of all HDF5 data fields used in data processing is provided in Table 1. A summary of all parameters used in data processing is given in Table 2 as well as an overview of the indexing rates at the various stages of data analysis in Table 3.
Mean phase error
A reference structure was built using data from all 14,782 9 keV indexable images. The cosine difference defined as Cos[phase(model_obtained_with_all_images)]-Cos[phase(model_obtained_with_subset_of_images)] was calculated to assess the quality of the phases. This is a comparison between a well-defined reference structure and the structure obtained with fewer images. By contrast, the figure of merit is an intrinsic measure without reference. The calculated figure of merit and the cosine differences are tabulated in Table 4 along with the number of correctly built residues by ARP/wARP.
Code availability
For data processing CrystFEL 0.6.2 and newly implemented python modules were employed. CrystFEL 0.6.2 is a free open source software under the GNU Public License version 3 and can be downloaded from http://www.desy.de/~twhite/crystfel/. stream2h5.py, write_calib_color.py, write_pca_peaks.py, write_spectra.py, split.py and write_subtract_peaks.py are free open source software under the GNU Public License version 3 and can be downloaded from https://github.com/AlexanderGorel/crystallography.
Data Records
Due to the large size of the raw data we only deposited the hit images (hitsrawdata.tar) in the Coherent X-ray Imaging Data Bank website (CXIDB) with the CXIDB ID 66 (Data Citation 1). Furthermore, we deposited the spectra.h5 files (spectra.tar.gz) with spectrum information, the silicon powder files (silicon-powder-poi.tar.gz) with the calculated interest points, the 7 and 9 keV indexable data (hits7kev.tar and hits9kev.tar) and the two-colour indexable data (two-color.tar) with residual points from the weak diffraction pattern in HDF5 file format as well as the geometry file and the cell file (supplement-files.tar.gz) at CXIDB (Data Citation 1).
Technical Validation
We have successfully phased the deposited data by MAD and solved the structure of the lysozyme gadoteridol complex using AutoSHARP23 with data to 1.9 Å resolution. The final structure built by ARP/wARP24 was refined against 5,000 9 keV diffraction images. It is available from the Protein Data Bank (Data Citation 2).
Usage Notes
Our complete data processing scheme is shown in the data flow diagram published in Gorel et al.14 supplement. Each data processing step can be repeated with the provided command line tools. It would be interesting to see a comparison between our processing strategy and others, such as the recently published FELIX25 algorithm which is capable of indexing more than ten different diffraction patterns per diffraction image. Moreover, the two-colour data set can be used for further software development. Indexing the reflections belonging to one colour yields the orientation matrix of the unit cell relative to the laboratory system. Future software may then use this matrix as a starting point for the initial indexing of the Bragg reflections of the second colour. Since they provide a different set of diffraction conditions, the matrix can be optimized for the second colour and through iterative refinement using the two sets of reflections, an extremely accurate orientation matrix can be obtained, in particular for the weak high resolution reflections. Ideally, a global refinement including both colours should be performed. We expect that such new analysis algorithms will greatly improve serial femtosecond crystallography (SFX) data processing in general and facilitate MAD phasing at XFELs in particular.
Additional information
How to cite this article: Gorel, A. et al. Two-colour serial femtosecond crystallography dataset from gadoteridol-derivatized lysozyme for MAD phasing. Sci. Data 4:170188 doi: 10.1038/sdata.2017.188 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Schlichting, I. Serial femtosecond crystallography: the first five years. IUCrJ 2, 246–255 (2015).
Spence, J. XFELs for structure and dynamics in biology. IUCrJ 4, 322–339 (2017).
Barends, T. R. et al. De novo protein crystal structure determination from X-ray free-electron laser data. Nature 505, 244–247 (2014).
Nakane, T. et al. Native sulfur/chlorine SAD phasing for serial femtosecond crystallography. Acta Crystallogr. Section D Biol. Crystallogr 71, 2519–2525 (2015).
Nakane, T. et al. Membrane protein structure determination by SAD, SIR, or SIRAS phasing in serial femtosecond crystallography using an iododetergent. Proc. Nat. Acad. of Sci. USA 113, 13039–13044 (2016).
Nass, K. et al. Protein structure determination by single-wavelength anomalous diffraction phasing of X-ray free-electron laser data. IUCrJ 3, 180–191 (2016).
Batyuk, A. et al. Native phasing of x-ray free-electron laser data for a G protein-coupled receptor. Sci Adv 2, e1600292 (2016).
Fukuda, Y. et al. Redox-coupled proton transfer mechanism in nitrite reductase revealed by femtosecond crystallography. Proc. Nat. Acad. Sci. USA 113, 2928–2933 (2016).
Hunter, M. S. et al. Selenium single-wavelength anomalous diffraction de novo phasing using an X-ray-free electron laser. Nat. Commun. 7, 13388 (2016).
Yamashita, K. et al. Experimental phase determination with selenomethionine or mercury-derivatization in serial femtosecond crystallography. IUCrJ 4 (2017).
Yamashita, K. et al. An isomorphous replacement method for efficient de novo phasing for serial femtosecond crystallography. Nat. Sci. Rep 5, 14017 (2015).
Colletier, J. P. et al. De novo phasing with X-ray laser reveals mosquito larvicide BinAB structure. Nature 539, 43–47 (2016).
Hara, T. et al. Two-colour hard X-ray free-electron laser with wide tunability. Nat. Commun. 4, 2919 (2013).
Gorel, A. et al. Multi-wavelength anomalous diffraction de-novo phasing using a two-colour X-ray free-electron laser with wide tunability. Nat. Commun. 4, 1170 (2017).
Botha, S. et al. Room-temperature serial crystallography at synchrotron X-ray sources using slowly flowing free-standing high-viscosity microstreams. Acta Crystallogr. Section D Biol. Crystallogr 71, 387–397 (2015).
Tono, K. et al. Diverse application platform for hard X-ray diffraction in SACLA (DAPHNIS): application to serial protein crystallography using an X-ray free-electron laser. J. Synch. Rad 22, 532–537 (2015).
Sugahara, M. et al. Grease matrix as a versatile carrier of proteins for serial crystallography. Nat. Meth 12, 61–63 (2015).
Kameshima, T. et al. Development of an X-ray pixel detector with multi-port charge-coupled device for X-ray free-electron laser experiments. Review of Scientific Instruments 85, 033110 (2014).
Barty, A. et al. Cheetah: software for high-throughput reduction and analysis of serial femtosecond X-ray diffraction data. J. Appl. Cryst. 47, 1118–1131 (2014).
Nakane, T. et al. Data processing pipeline for serial femtosecond crystallography at SACLA. J. Appl. Cryst 49, 1035–1041 (2016).
Tamasaku, K. et al. Inline spectrometer for shot-by-shot determination of pulse energies of a two-color X-ray free-electron laser. J. Synch. Rad 23, 331–333 (2016).
White, T. A. et al. CrystFEL: a software suite for snapshot serial crystallography. J. Appl. Cryst. 45, 335–341 (2012).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007).
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protocols 3, 1171–1179 (2008).
Beyerlein, K. R. et al. FELIX: an algorithm for indexing multiple crystallites in X-ray free-electron laser snapshot diffraction images. J. Appl. Cryst. 50, 1075–1083 (2017).
Data Citations
Gorel, A., & Foucar, L. Coherent X-ray Imaging Data Bank http://dx.doi.org/10.11577/1369634 (2017)
Gorel, A. Protein Data Bank 5OER (2017)
Acknowledgements
This work was supported by the X-ray Free-Electron Laser Priority Strategy Program (Ministry of Education, Culture, Sports, Science and Technology of Japan) and partially by the Strategic Basic Research Program (JST) and RIKEN Pioneering Project Dynamic Structural Biology. We acknowledge computational support from the SACLA High Performance Computing system. The research was supported by the Max Planck Society and Dynamic Alliance for Open Innovation Bridging Human, Environment and Materials and TAGEN project of Tohoku University. We thank Dr Roland van Gessel, Bracco Imaging Deutschland, Konstanz, Germany, for the very generous sample of gadoteridol.
Author information
Authors and Affiliations
Contributions
I.S., M.K., G.N.K. prepared and characterized samples, R.B.D., R.L.S., G.N.K., M.L.G., M.K. designed and operated sample injection hardware, M.Y., Y.J, S.I. were involved in preparations for the experiment, R.B.D., R.L.S, G.N.K., M.L.G., M.K., I.S., M.H., C.M.R., K.N., T.R.M.B., K.M., H.F., K.U., I.I., K.T., E.N., R.T, performed the experiment, C.M.R., M.H., K.N., T.R.M.B, and L.F. performed online processing, A.G. performed off-line processing, T.R.M.B., A.G. phased the data, I.S. designed and coordinated the project, A.G. and I.S. wrote the manuscript with input from all the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Gorel, A., Motomura, K., Fukuzawa, H. et al. Two-colour serial femtosecond crystallography dataset from gadoteridol-derivatized lysozyme for MAD phasing. Sci Data 4, 170188 (2017). https://doi.org/10.1038/sdata.2017.188
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2017.188