Abstract
Extensive and expanding oxygen minimum zones (OMZs) exist at variable depths in coastal and open ocean waters. As oxygen levels decline, nutrients and energy are increasingly diverted away from higher trophic levels into microbial community metabolism, resulting in fixed nitrogen loss and production of climate active trace gases including nitrous oxide and methane. While ocean deoxygenation has been reported on a global scale, our understanding of OMZ biology and geochemistry is limited by a lack of time-resolved data sets. Here, we present a historical dataset of oxygen concentrations spanning fifty years and nine years of monthly geochemical time series observations in Saanich Inlet, a seasonally anoxic fjord on the coast of Vancouver Island, British Columbia, Canada that undergoes recurring changes in water column oxygenation status. This compendium provides a unique geochemical framework for evaluating long-term trends in biogeochemical cycling in OMZ waters.
Design Type(s) | time series design • data integration objective • observation design |
Measurement Type(s) | hydrographic profiling • nutrient level • mass of biological material • dissolved gases • water oxygen concentration |
Technology Type(s) | data acquisition system |
Factor Type(s) | sampling depth • temporal_interval |
Sample Characteristic(s) | Saanich Inlet • coastal sea water |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Marine oxygen minimum zones (OMZs) are widespread, naturally occurring water column features that arise from respiration of organic matter in subsurface waters with restricted circulation. Operationally defined by oxygen (O2) concentrations between 0 to 20 μM, and differential accumulation of nitrite (NO2−) and reduced sulphur compounds, OMZs currently constitute 1–7% of global ocean volume1–7. As oxygen levels decline, nutrients and energy are increasingly diverted away from higher trophic levels into microbial community metabolism2,8. As a result, OMZs are hotspots for the biogeochemical cycling of carbon, nitrogen and sulphur with resulting feedback on nitrogen loss processes and climate active trace gas production including nitrous oxide (N2O) and methane (CH4)9–13. The effects of climate change, including increased stratification and reduced O2-solubility in warming waters are resulting in OMZ expansion and intensification1,8,14–18 reinforcing the need to monitor changes in water column geochemistry in oxygen-deficient waters.
Oceanographic surveys in OMZ waters rely on a standard suite of measurements including temperature, salinity, density and conductivity. Additional parameters including irradiance, used to measure water column light penetration, fluorescence used to monitor chlorophyll concentrations and dissolved gases including O2 and carbon dioxide (CO2) provide information on primary production2,19,20. Chemical measurements of phosphate (PO43−), silicic acid (SiO2), and nitrate (NO3−) are measured as essential nutrients supporting growth and cell division15. Nitrite (NO2−) and ammonium (NH4+) are also measured to better constrain nitrogen cycling processes2,10,11,13. Because some OMZs can become completely anoxic, hydrogen sulfide (H2S) concentrations can be used as an indicator for sulphate reduction driving chemoautotrophic metabolism3,9. Measurements of N2O and CH4 can also be used to monitor potential climatological impacts of OMZ expansion9–13,21. Collectively, these measurements define geochemical gradients in OMZ water columns that shape the conditions for coupled biogeochemical cycling.
Saanich Inlet is a seasonally anoxic fjord on the coast of Vancouver Island, British Columbia, Canada22–25. Saanich Inlet is an inverse estuary where a glacial sill at the mouth restricts exchange between deep basin and external waters for most of the year. Freshwater is supplied at the inlet mouth predominantly by the Cowichan and Fraser Rivers, producing horizontal density differences that result in an inward flow into the inlet in the surface layer and outward flow at depth25,26. During spring and summer months, high levels of primary productivity in surface waters and limited vertical mixing of basin waters below the sill result in anoxia and the accumulation of CH4, NH4+ and H2S27–29. In late summer and fall, neap tidal flows produce an influx of denser water from the Northeastern subarctic Pacific (NESAP) Ocean that cascade over the sill, resulting in vertical mixing and the re-supplying of deep basin waters with O2 and nutrients25,26. The recurring seasonal development of water column anoxia followed by deep water renewal makes Saanich Inlet a model ecosystem for monitoring biogeochemical responses to changing levels of water column O2-deficiency2,30–32.
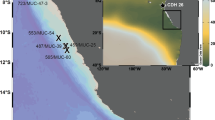
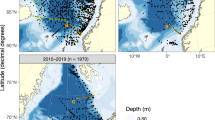
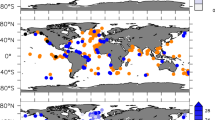
Here we present a compendium of time-series observations encompassing historical O2 measurements25,33 (Fig. 1a) and more recent monthly monitoring efforts in Saanich Inlet from 2006 through 2014, representing over 100 independent sampling expeditions (Fig. 1b). This compendium contains physical (temperature, salinity, density, irradiance, and fluorescence), chemical (PO43−, SiO2, NO3−, NO2−, NH4+, and H2S), dissolved gas (O2, CO2, N2, N2O, CH4), and biological (cell counts) parameter data (Fig. 1b,c) useful in comparing to other oceanographic time-series from the Northwest Atlantic to Eastern Tropical Pacific through the Global Ocean Sampling expeditions34, the Hawaii and Tara Oceans34–36 and Bermuda Atlantic Time-series37 and in the development of biogeochemical models. In addition, this geochemical compendium is paired with a cognate compendium of multi-omic sequence information (DNA, RNA, protein) focused on microbial diversity, abundance and function.38 Combined, these compendiums provide a community-driven framework for observing and predicting microbial community repsonses to changing levels of oxygen deficiency extensible to open ocean OMZs.
(a) Sampling station S3 location in the Saanich Inlet. (b) Historical sampling effort in Saanich Inlet depicted as O2 sampling points from 1953 to 2014. (c) Oxygen concentration contour for CTD data (February 2008 onward), and points for 16 sampling depths for nutrients and gases. (d) Sample inventory from February 2006 to October 2014 showing historical, CTD and nutrient datasets included in this manuscript (solid black), in previous publications (gray).
Methods
Time-series monitoring in Saanich Inlet was conducted on a monthly basis aboard the MSV John Strickland at station S3 (48°35.500 N, 123°30.300 W) as previously described32. Water samples from 16 high-resolution (HR) depths at station S3 (10, 20, 40, 60, 75, 80, 90, 97, 100, 110, 120, 135, 150, 165, 185 and 200 meters) spanning oxic (>90 μmol O2 kg−1), dysoxic (90–20 μmol O2 kg−1), suboxic (20–1 μmol O2 kg−1) anoxic (<1 μmol O2 kg−1) and sulfidic water column compartments2 were collected using Niskin or Go-Flow bottles for dissolved gasses: O2, CO2, CH4, Nitrogen gas (N2), N2O; nutrients: NO3−, NO2−, NH4+, SiO2, PO43−, H2S; and cell counts. Sampling methods for HR samples and additional six large-volume depths (10, 100, 120, 135, 150 and 200 meters) collected for time-series multi-omic sequence information analyses are published in an accompanying compendium38.
Environmental sampling
Historical dissolved O2 concentrations were obtained from station S3 by sampling with Niskin bottles at discrete depths and subsequently analyzing water samples using various modifications of the Winkler method25,33,39 (Data Citation 1). Historical water column profiles can also be accessed at the Ocean Sciences Data Inventory website hosted by the Institute of Ocean Sciences and Fisheries and Oceans Canada (http://www.pac.dfo-mpo.gc.ca/science/oceans/data-donnees/search-recherche/profiles-eng.asp). Samples collected from February 2006 to February 2008 were processed and analysed for dissolved gases and nutrients as first reported in Zaikova et al.32 (Fig. 2). Beginning on February 2008, a Sea-Bird SBE 25 CTD (conductivity, temperature and depth), with Sea-Bird SBE 43 dissolved O2 and Biospherical Instruments PAR sensors attached was used to measure conductivity, temperature, dissolved O2, PAR/Irradiance and fluorescence (Data Citation 1). To minimize the effects of off-gassing, waters were collected in the following order; dissolved O2 for Winkler titration (from select depths for CTD calibration), dissolved gases (N2O and CH4), NH4+, H2S, nutrients, cell counts (Data Citation 1) and salinity (from selected depths for CTD calibration). A detailed seawater sampling video protocol can be found online (http://www.jove.com/video/1159/seawater-sampling-and-collection).
Chemical data
CTD data analysis
CTD data were downloaded, converted and pre-processed in the laboratory using the SeaBirdSeasoft software. Downcast data of the deepest cast (200 m) was extracted and converted from ASCII format into a.cnv file for manual curation. Salinity and density were calculated using the Derive module with the corrected conductivity measurements. Temperature and salinity were exported using an ITS-90 scale. Oxygen sensor measurements collected in millilitre per litre (mll−1) were converted to micromolar (μM) units (Data Citation 1). Discrete winkler analyses from water samples spanning LV depths were used to calibrate the CTD O2 measurements (Data Citation 1).
Nitrate, phosphate and silicic acid
For each depth, sample water was filtered through a 0.2 μm acrodisc (Millipore) and used to rinse a 15 ml tube three times before filling with 14 ml. Samples were stored on ice and later in the lab at −20 °C for up to four months prior to analysis. A Bran Luebbe AutoAnalyser 3 using air-segmented continuous-flow and standard colorimetric methods was used for analysis. In brief, nitrate (NO3−) was reduced to nitrite by a copper-cadmium reduction column. Nitrite was then quantified by a modified colorimetric assay40, reading sample absorbance at 550 nm. Orthophosphate (PO43−) was quantified based on the colorimetric method for reduced phospho-molybdenum complex, reading samples absorbance at 880 nm41. Silicic acid (H4SiO4) was quantified by reduction to a molybdenum blue complex, reading sample absorbance at 820 nm. Oxalic acid was added to remove phosphate interference40 (Data Citation 1).
Ammonium
A fluorometric measurement protocol for NH4+ analysis was carried out as previously described in Holmes et al. for marine samples42. For each depth, glass amber scintillation bottles were rinsed three times, then filled to overflowing and capped immediately to minimize off-gassing of NH4+ and stored on ice for 1–3 h before processing. A total of 5 ml of sample water was transferred to vials with 7.5 ml o-phthaldialdehyde (OPA; Sigma) in triplicate. Simultaneously, 7.5 ml of OPA was added to prepared NH4+ standard curve (0.025–10.0 μM NH4Cl) and stored at room temperature for up to 4 h. Fluorescence at 380ex/420emm was read using a Turner Designs TD-700 fluorometer (2006–2009) or Varioskan plate reader (2009–2014) in triplicate with 300 μl of sample or standard in a 96-well round bottom plate (Corning) (Fig. 3) (Data Citation 1).
Nitrite
The protocol for NO2− analysis was carried out as previously described in Armstrong et al. modified for marine samples40. For each depth, sample water was filtered through a 0.2 μm acrodisc (Millipore) and used to rinse a 15 ml tube three times before filling with 14 ml filtered sample water and stored on ice for 1–3 h before processing. A total of 2 ml of sample water was transferred to 4 ml plastic cuvettes in triplicate, and 100 μl sulphanilamide and 100 μl nicotinamide adenine dinucleotide (NAD) were added. Simultaneously, reagents were added to prepared standards (0.025–5.0 μM NaNO2). Cuvettes were inverted and stored on ice for up to 4 h. Absorbance at 542 nm was read using a Cary60 spectrometer (Fig. 3) (Data Citation 1).
Hydrogen sulfide
The protocol for H2S was carried out as previously described in Cline43 modified for marine samples. For each depth, 10 ml sample water was collected directly into a 15 ml tube containing 200 μl 20% Zinc Acetate and stored on ice for 4–24 h before processing. A total of 300 μl of sample was transferred into triplicate wells of a 96-well round or flat-bottom plate (Corning), and 6 μl Hach Reagent (Hach) 1 and 2 for sulphide assay were added to each well. After 5 min incubation, absorbance at 670 nm was read using a spectrophotometer (2008–2009) or Varioskan plate reader (2009–2014) (Data Citation 1).
Cell counts
For each depth, 10 ml sample water was collected directly into a sterile 15 ml tube containing 1.1 ml of 37% formaldehyde and stored on ice. Back at the lab, samples were stored at 4 °C for up to two days prior to cell counting using a BD LSR II flow cytometer (2008–2012) or MACS Quant Analyzer (2012–2014) based on the following protocols. For BD LSR II, a dye mixture was prepared by diluting 3 μl of the SYBR Green I (Invitrogen) dye in 1,830 μl of sterile water. Six drops (Alignflow) alignment beads were then added to this mixture. In a round-bottom polystyrene tubes, 25 μl of the dye mix was added to 475 μl of the water sample (in triplicates). The cells and beads were then counted using BD LSR II flow cytometer. For MAXSQuant, a dye mixture was prepared by diluting 240 μl of seawater sample with 10 μl of SYBR Green I (Invitrogen) dye mix which contains 6 μm flow cytometry blue laser alignment beads (Alignflow), for calibration purposes. SYBR Green mix was prepared by diluting 4 μl of the dye in 1,570 μl of sterile water following an addition of 30 μl beads. Samples are prepared in triplicates in a 96-well flat bottom black plate (Corning) and run on MACSQuant Analyzer (MiltenyiBiotec) (Data Citation 1).
Dissolved gases
For each depth, sample water was collected through silicon tubing (~15 cm long and 1/4″ thick, pre-flushed for a few seconds with sample water) into a 30 or 60 ml borosilicate glass serum vial, overflowing three times the volume and taking care to remove air bubbles from the tubing and vial during filling. The vials were spiked with 50 μl saturated mercuric-chloride solution, then crimp-sealed with a butyl-rubber stopper and aluminium cap. Samples were stored in the dark at 4 °C until processing. Dissolved gases were analysed using either headspace for CO2, CH4, N2 and N2O (2006–2009, samples stored for up to 2 years) or automated purge-and-trap for CH4 and N2O only (2009–2014, samples stored for <3 months) coupled with gas chromatography-mass spectrometry (GC-MS)44 (Data Citation 1). Samples with >20% s.d. between replicates were excluded from our study to discard any long storage effects.
Data Records
Data record 1
The Saanich Inlet O2 historical data (1953–2000) is accessible in comma-separated-value format file ‘Historical_O2_DATA.csv’ on the Dryad Digital Repository [Data Citation 1] containing the data fields outlined in Table 1.
Data record 2
The Saanich Inlet time-series CTD data is accessible in comma-separated-value format file ‘Saanich_TimeSeries_CTD_DATA.csv’ on the Dryad Digital Repository [Data Citation 1] domain containing the data fields outlined in Table 2.
Data record 3
The Saanich Inlet time-series chemical data is accessible in comma-separated-value format file ‘Saanich_TimeSeries_Chemical_DATA.csv’ on the Dryad Digital Repository domain containing the data fields outlined in Table 3 [Data Citation 1].
Data record 4
The Saanich Inlet time-series Winkler O2 data is accessible in comma-separated-value format file ‘Saanich_TimeSeries_Winkler_DATA.csv’ on the Dryad Digital Repository [Data Citation 1] domain containing the data fields outlined in Table 4.
Technical Validation
Data quality control
Data in the Saanich Inlet time series was collected and processed by experienced scientists with extensive training in the sampling methods and data processing steps described above. People interested in becoming part of the scientific crew were invited to participate in training sessions with experienced scientists in the field and laboratory to gain practical experience. Once in the field, trainees were carefully supervised during sample collection for a minimum of 3 months for quality assurance. Following each cruise, the acting chief scientist compiled all chemical and physical data collected and conducted initial quality controls, checking for outliers and verifying standard curves. Data were then entered into an in-house database along with field notes and precise records of volumes of water filtered informing downstream analyses.
CTD and chemical data validation
The SeaBird 43 dissolved O2 sensor was calibrated by Winkler O2 measurements45. Samples from selected depths were collected into Winkler glass Erlenmeyer flasks using latex tubing, overflowing three times to ensure no air contamination. Oxygen concentration was determined using a Brinkman autotitrator, routinely calibrated with a potassium iodide standard. Stability of CTD O2 measurements was determined by comparing the high values with Winkler measurements, and low values with sulfidic profiles where the sensors levels off. Where H2S is detected we consider O2 measurements to be 0 μM based on spontaneous auto-oxidation reaction of H2S with O2. We have estimated our limit of detection for the automated Winkler method at ~0.007 mll−1 or ~0.3 μM.
The SeaBird conductivity sensor was calibrated using salinity samples collected at selected depths. Salinity glass bottles were rinsed 4 times and filled with water sample, stored at room temperature and analyzed within 4 months on a Guildline Portasal salinometer.
For each cruise, standard curves for NH4+ and NO2− were prepared. Stock solutions and reagents for both assays were freshly made every three months and stored in the dark at 4 °C and were tested prior to being used for analysis. Stock solution quality and assay validation was carried out using linear regression and calculating the r squared value (r2≥0.90) on the absorbance data (Fig. 3). Standard curve stock solutions and reagents for H2S assay were evaluated every three months based on manufacturer’s instructions. We have estimated our limit of detection for these assays to be 0.001 μM NH4+, 0.0006 μM NO2−, and 1.7 μM H2S.
Samples for NO3−, PO43− and H4SiO4 were run in single measurements. Autoanalyzer estimated limit of detection for these measurements are 0.020 μM NO3−, 0.012 μM PO43− and 0.100 μM H4SiO4.
Flow cytometry validation
Concentration of flow cytometry (FL) alignment beads was determined by microscopy using a hemocytometer. Bead counts for each FL run were then used to calculate the volume of sample measured. Two blanks were included in each FL run, and consisted of sterile water bead/dye solution with sterile water in place of sample water, to ensure instrument cleanliness and optics function. Size gates were set to include beads and bacterial and archaeal cell sizes and to reduce noise of any small particulate debris.
Gas analysis validation
A thorough review of the Purge and Trap GCMS (PT-GCMS) method validation has been previously described44. Standard curves were run at the start of each batch of 25 samples by injecting precisely measured quantities of a standard gas mixture (CH4, N2O, CO2 and N2) calibrated against National Ocean and Atmospheric Administration (NOAA) certified reference gas mixture. Single standards were also measured every ~2-hours (5–6 sample per run) to monitor instrument drift. The precision of CH4 and N2O measurements based on replicate measurements of air-equilibrated water samples was <4%. Accuracy was confirmed by measuring dissolved N2O and CH4 in carefully prepared air-equilibrated, temperature-controlled Milli-Q water and comparing this to expected concentrations based on gas-solubility equations46,47. Detection limits depend on the volume of sample being purged, and were 0.8 nM for CH4 and 0.5 nM for N2O for the samples analyzed in this time-series (2009–2014) (Fig. 3). Samples were run in duplicate or triplicate to ensure reproducible readings. The relative s.d. between replicate samples was calculated and included in the output data. The output data are also carefully inspected to ensure optimal instrument performance during sample analysis before being submitted to the database.
Usage Notes
Oxygen considerations
Based on the amount of dissolved O2 in the water column and the biogeochemical processes associated with it, thresholds for O2-defined water column conditions were determined as previously described2. As the range of O2 concentrations is wide and has great impact on biological processes in the lower concentrations, we suggest using the O2 thresholds described in Wright et al.2 for analysis.
Additional information
How to cite this article: Torres-Beltrán, M. et al. A compendium of geochemical information from the Saanich Inlet water column. Sci. Data 4:170159 doi: 10.1038/sdata.2017.159 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Paulmier, A. & Ruiz-Pino, D. Oxygen minimum zones (OMZs) in the modern ocean. Prog. Oceanogr. 80, 113–128 (2009).
Wright, J. J., Konwar, K. M. & Hallam, S. J. Microbial ecology of expanding oxygen minimum zones. Nat. Rev. Microbiol. 10, 381–394 (2012).
Ulloa, O., Canfield, D. E., DeLong, E. F., Letelier, R. M. & Stewart, F. J. Microbial oceanography of anoxic oxygen minimum zones. Proc. Natl Acad. Sci. USA 109, 15996–16003 (2012).
Codispoti, L. A. Interesting Times for Marine N2O. Science 327, 1339 (2010).
Fuenzalida, R., Schneider, W., Garcés-Vargas, J., Bravo, L. & Lange, C. Vertical and horizontal extension of the oxygen minimum zone in the eastern South Pacific Ocean. Deep-Sea Res. Pt. II 56, 992–1003 (2009).
Karstensen, J., Stramma, L. & Visbeck, M. Oxygen minimum zones in the eastern tropical Atlantic and Pacific oceans. Prog. Oceanogr. 77, 331–350 (2008).
Ulloa, O. & Pantoja, S. The oxygen minimum zone of the eastern South Pacific. Deep-Sea Res. Pt. II 56, 987–991 (2009).
Diaz, R. J. & Rosenberg, R. Spreading dead zones and consequences for marine ecosystems. Science 321, 926–929 (2008).
Canfield, D. E. et al. A cryptic sulfur cycle in oxygen-minimum-zone waters off the Chilean coast. Science 330, 1375–1378 (2010).
Lam, P. et al. Revising the nitrogen cycle in the Peruvian oxygen minimum zone. Natl Acad. Sci. USA 106, 4752–4757 (2009).
Ward, B. B. et al. Denitrification as the dominant nitrogen loss process in the Arabian Sea. Nature 461, 78–81 (2009).
Naqvi, S. W. A. et al. Marine hypoxia/anoxia as a source of CH4 and N2O. Biogeosciences 7, 2159–2190 (2010).
Lam, P. & Kuypers, M. M. M. Microbial Nitrogen Cycling Processes in Oxygen Minimum Zones. Ann. Rev. Mar. Sci. 3, 317–345 (2010).
Keeling, R. E., Kortzinger, A. & Gruber, N. Ocean deoxygenation in a warming world. Ann. Rev. Mar. Sci. 2, 199–229 (2010).
Arrigo, K. R. Marine microorganisms and global nutrient cycles. Nature 437, 349–355 (2005).
Stramma, L., Johnson, G. C., Sprintall, J. & Mohrholz, V. Expanding Oxygen-Minimum Zones in the Tropical Oceans. Science 320, 655 (2008).
Whitney, F. A., Freeland, H. J. & Robert, M. Persistently declining oxygen levels in the interior waters of the eastern subarctic Pacific. Prog. Oceanogr. 75, 179–199 (2007).
Schmidtko, S., Stramma, L. & Visbeck, M. Decline in global oceanic oxygen content during the past five decades. Nature 542, 335–339 (2017).
Lewis, M. R., Warnock, R. E. & Platt, T. Absorption and photosynthetic action spectra for natural phytoplankton populations: Implications for production in the open ocean1. Limnol. Oceanogr. 30, 794–806 (1985).
Kolber, Z. & Falkowski, P. G. Use of active fluorescence to estimate phytoplankton photosynthesis in situ. Limnol. Oceanogr. 38, 1646–1665 (1993).
Monteiro, P. M. S. et al. Variability of natural hypoxia and methane in a coastal upwelling system: Oceanic physics or shelf biology? Geophys. Res. Lett. 33, L16614 (2006).
Anderson, J. J. & Devol, A. H. Deep water renewal in Saanich Inlet, an intermittently anoxic basin. Estuar. Coast. Mar. Sci. 1, 1–10 (1973).
Carter, N. M. The oceanography of the fjords of southern British Columbia. Fish. Res. Bd. Canada Prog. Rept. Pacific Coast Sta. 12, 7–11 (1932).
Carter, N. M. Physiography and oceanography of some British Columbia fjords. Proc. Fifth. Pacific Sci. Cong. 1, 721–733 (1934).
Herlinveaux, R. H. Oceanography of Saanich Inlet in Vancouver Island, British Columbia. J. Fish. Res. Board Can. 19, 1–37 (1962).
Gargett, A. E., Stucchi, D. & Whitney, F. Physical processes associated with high primary production in Saanich Inlet, British Columbia. Estuar. Coast. Mar. Sci. 56, 1141–1156 (2003).
Lilley, M. D., Baross, J. A. & Gordon, L. I. Dissolved hydrogen and methane in Saanich Inlet, British Columbia. Deep-Sea Res. 29, 1471–1484 (1982).
Ward, B. B. & Kilpatrick, K. A. Relationship between substrate concentration and oxidation of ammonium and methane in a stratified water column. Cont. Shelf Res. 10, 1193–1208 (1990).
Ward, B. B., Kilpatrick, K. A., Wopat, A. E., Minnich, E. C. & Lidstrom, M. E. Methane oxidation in Saanich inlet during summer stratification. Cont. Shelf Res. 9, 65–75 (1989).
Walsh, D. A. & Hallam, S. J. in Handbook of Molecular Microbial Ecology II: Metagenomics in Different Habitats Vol. 2, (ed. de Brujin, F. J. (Ch. 25 253–267) (Wiley-Blackwell, 2011).
Walsh, D. A. et al. Metagenome of a versatile chemolithoautotroph from expanding oceanic dead zones. Science 326, 578–582 (2009).
Zaikova, E. et al. Microbial community dynamics in a seasonally anoxic fjord: Saanich Inlet, British Columbia. Environ. Microbiol. 12, 172–191 (2010).
Lee, B.-S., Bullister, J. L. & Whitney, F. A. Chlorofluorocarbon CFC-11 and carbon tetrachloride removal in Saanich Inlet, an intermittently anoxic basin. Mar. Chem. 66, 171–185 (1999).
Sunagawa, S. et al. Structure and function of the global ocean microbiome. Science 348, 1261359 (2015).
Karl, D. M. & Church, M. J. Microbial oceanography and the Hawaii Ocean Time-series programme. Nat. Rev. Microbiol. 12, 699–713 (2014).
Pesant, S. et al. Open science resources for the discovery and analysis of Tara Oceans data. Sci. Data 2, 150023 (2015).
Steinberg, D. K. et al. Overview of the US JGOFS Bermuda Atlantic Time-series Study (BATS): a decade-scale look at ocean biology and biogeochemistry. Deep-Sea Res. Pt. II 48, 1405–1447 (2001).
Hawley, A. K. et al. A compendium of multi-omic sequence information from the Saanich Inlet water column. Sci. Data 4, 170160 doi:10.1038/sdata.2017.160 (2017).
Carpenter, J. H. The Chesapeake Bay Institute Technique for the Winkler Dissolved Oxygen Method. Limnol. Oceanogr. 10, 141–143 (1965).
Armstrong, F. A. J., Stearns, C. R. & Strickland, J. D. H. The measurement of upwelling and subsequent biological process by means of the Technicon Autoanalyzer® and associated equipment. Deep-Sea Res. Oceanogr. Abstr. 14, 381–389 (1967).
Murphy, J. & Riley, J. P. A modified single solution method for the determination of phosphate in natural waters. Anal. Chim. Acta 27, 31–36 (1962).
Holmes, R. M., Aminot, A., Kérouel, R., Hooker, B. A. & Peterson, B. J. A simple and precise method for measuring ammonium in marine and freshwater ecosystems. Can. J. Fish Aquat. Sci. 56, 1801–1808 (1999).
Cline, J. D. Spectrophotometric determination of hydrogen sulfide in natural waters. Limnol. Oceanogr. 14, 454–458 (1969).
Capelle, D. W., Dacey, J. & Tortell, P. D. An automated, high-throughput method for accurate and precise measurements of dissolved nitrous-oxide and methane concentrations in natural seawaters. Limnol Oceanogr Methods 13, 345–355 (2015).
Winkler, L. W. Die Bestimmung des im Wasser gelösten Sauerstoffes. Ber. Dtsch. Chem. Ges. 21, 2843–2854 (1888).
Weiss, R. F. & Price, B. A. Nitrous oxide solubility in water and seawater. Mar. Chem. 8, 347–359 (1980).
Wiesenburg, D. A. & Guinasso, N. L. Equilibrium solubilities of methane, carbon monoxide, and hydrogen in water and sea water. J. Chem. Eng. Data 24, 356–360 (1979).
Data Citations
Torres-Beltran, M. Dryad Digital Repository http://dx.doi.org/10.5061/dryad.nh035 (2017)
Acknowledgements
We thank Captain Ken Brown and his crew for their engaged effort on every cruise aboard the RSV Strickland. We thank Rich Pawlowicz and the UBC Department of Earth, Ocean & Atmospheric Sciences (EOAS) for facilitating the use of the EOAS CTD. We also thank past and present members of the Hallam lab and the many undergraduate trainees, aka minions, for contributions to cruise preparation and clean up, water filtration and analytical chemistry. We also thank members of the Tortell, Crowe and Suttle Laboratories at UBC and members of the Varela Lab at UVic for logistical support, and Stephen Romaine for his efforts to host the metadata database in the LineP DFO domain. We thank Luis Malpica Cruz for his help with R data wrangling. This work was performed under the auspices of the US Department of Energy (DOE) Joint Genome Institute an Office of Science User Facility, supported by the Office of Science of the US Department of Energy under Contract DE-AC02- 05CH11231, the G. Unger Vetlesen and Ambrose Monell Foundations, the Tula Foundation-funded Centre for Microbial Diversity and Evolution, The Natural Sciences and Engineering Research Council of Canada, Genome British Columbia, the Canada Foundation for Innovation, and the Canadian Institute for Advanced Research through grants awarded to S.J.H. Ship time support was provided by NSERC between 2007–2014 through grants awarded to S.J.H. and P.D.T. M.T.-B. was funded by Consejo Nacional de Ciencia y Tecnología (CONACyT) and the Tula Foundation. A.K.H. was supported by the Tula Foundation.
Author information
Authors and Affiliations
Contributions
A.K.H. and M.T.-B. provided extensive logistical support and planning for the time series and curated the datasets. D.C. carried out all trap and purge dissolved gases analysis. E.Z. was a field technician over multiple years and wrote several protocols. D.A.W. and S.J.H. designed many methods at the initiation of the time series. A.M., M.S., S.K., J.F., M.B., O.S., E.A.G., D.F., C.M. were sea-going lab technicians, and graduate and post-doctoral fellows who contributed with data collection and curation over multiple years. C.P. and L.P. were sea-going technicians and carried out CTD data processing and winkler and salinity measurements. S.A.C and C.A.S. contributed with material and data collection, and edited manuscript. F.W. compiled and curated historical O2 and CTD measurements. P.D.T. oversaw time series planning and dissolved gas measurements for both head-space and trap-and-purge methods. S.J.H. designed and initiated the time series with P.D.T. S.J.H. managed and directed the project. A.K.H., M.T.-B. and S.J.H. wrote the manuscript with editorial input from remaining co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Torres-Beltrán, M., Hawley, A., Capelle, D. et al. A compendium of geochemical information from the Saanich Inlet water column. Sci Data 4, 170159 (2017). https://doi.org/10.1038/sdata.2017.159
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2017.159
This article is cited by
-
A compendium of bacterial and archaeal single-cell amplified genomes from oxygen deficient marine waters
Scientific Data (2023)
-
An 8-year record of phytoplankton productivity and nutrient distributions from surface waters of Saanich Inlet
Scientific Data (2022)
-
Pangenomics reveals alternative environmental lifestyles among chlamydiae
Nature Communications (2021)
-
Mercury methylation by metabolically versatile and cosmopolitan marine bacteria
The ISME Journal (2021)
-
Monitoring microbial responses to ocean deoxygenation in a model oxygen minimum zone
Scientific Data (2017)