Abstract
Camera traps can be used to address large-scale questions in community ecology by providing systematic data on an array of wide-ranging species. We deployed 225 camera traps across 1,125 km2 in Serengeti National Park, Tanzania, to evaluate spatial and temporal inter-species dynamics. The cameras have operated continuously since 2010 and had accumulated 99,241 camera-trap days and produced 1.2 million sets of pictures by 2013. Members of the general public classified the images via the citizen-science website www.snapshotserengeti.org. Multiple users viewed each image and recorded the species, number of individuals, associated behaviours, and presence of young. Over 28,000 registered users contributed 10.8 million classifications. We applied a simple algorithm to aggregate these individual classifications into a final ‘consensus’ dataset, yielding a final classification for each image and a measure of agreement among individual answers. The consensus classifications and raw imagery provide an unparalleled opportunity to investigate multi-species dynamics in an intact ecosystem and a valuable resource for machine-learning and computer-vision research.
Design Type(s) | observation design • time series design • species comparison design |
Measurement Type(s) | mammalian species abundance |
Technology Type(s) | Camera-trap |
Factor Type(s) | |
Sample Characteristic(s) | Serengeti National Park • savanna |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Over the last 20 years, camera traps—remote, automatic cameras—have revolutionized wildlife ecology and conservation and are now emerging as a key tool in the broader disciplines of behavioural, population, and community ecology1. Historically, cameras have been used to document the presence of rare species in understudied protected areas2,3 or estimate densities of individually identifiable animals4. But advances in digital technology have increased capacity while lowering prices, resulting in a dramatic increase in the number and diversity of camera trap studies5. While traditional analytical approaches for camera trapping data require individually identifiable animals1,4,6, recent developments have allowed the expansion of camera trap inference to multiple ‘unmarked’ species1,7–9. Although camera-trap surveys are increasing in popularity and scope, they can produce overwhelming amounts of data10–12, highlighting the need for efficient image processing techniques. Here we describe the datasets generated by Snapshot Serengeti, a large-scale survey that (1) deployed 225 camera traps across a 1,125 km2 area in the Serengeti National Park, Tanzania from 2010–2013, (2) used a citizen science website (www.snapshotserengeti.org) to process millions of images, and (3) used a simple algorithm to ensure high reliability of the resultant species classifications.
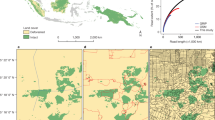
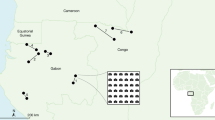
We established the Snapshot Serengeti camera survey to evaluate spatial and temporal dynamics of large predators and their prey. Serengeti National Park (SNP) is the core of a 25,000 km2 savannah ecosystem that straddles the Kenya-Tanzania border in East Africa. The Serengeti is dominated by the annual migration of the combined 1.6 million wildebeest and zebra that follow the seasonal rainfall onto the nutrient rich plains13. Since the 1960s, the Serengeti Lion Project has monitored lion population numbers and ranging patterns14, and the Tanzania Wildlife Research Institute (TAWIRI) has surveyed major herbivore numbers via flight counts and aerial photography15–19. Our camera survey expands upon historical monitoring by providing the first continuous systematic data on all of the larger predator and prey species, day and night, across several years. We set out 225 cameras within a 1,125 km2 grid inside the long-term lion study area that covers the intersection of open plains and savannah woodlands (Fig. 1 and Fig. 2), and spans a 1.67-fold rainfall gradient and 1.44-fold productivity gradient (T.M. Anderson, unpublished data.). The camera-trap grid offers systematic coverage of the entire study area (as per O’Brien et al.20) and ensures at least two cameras per home range for each medium to large mammalian species.
(a) Serengeti National Park. Long-term lion project study area is indicated by dotted line; camera-trap study area is indicated by dashed line. (b) Camera trap layout within the long-term Lion Project Study Area. Camera locations are plotted over tree cover (extracted from Landsat imagery), with darker green indicating increased tree cover per 30 m2-grid cell.
Between June 2010 and May 2013, the survey operated for a total of 99,241 camera-trap days and produced 1.2 million image sets (each image set contains 1–3 photographs taken in a single burst of ~1 s). In collaboration with The Zooniverse (www.zooniverse.org), the world’s most popular citizen science platform, we developed the website www.snapshotserengeti.org that allowed members of the general public to view and classify each image, identifying species, counting the number of individuals, and characterizing behaviours (Fig. 3).
Every image set was circulated to multiple users to improve data accuracy. More than 28,000 registered users and ~40,000 unregistered users contributed 10.8 million classifications for the 1.2 million image sets. We applied a simple plurality algorithm to produce a ‘consensus dataset’ of final classifications for each image set. The consensus classifications were validated against 4,149 ‘gold-standard’ image-sets that had been classified by experts, revealing 96.6% accuracy for species identifications and 90% accuracy for species counts..
Of the 1.2 million image sets, Snapshot Serengeti volunteers indicated that 322,653 contained animals; the remainder were misfires that had been triggered by heat or vegetation. The volunteers identified 48 different species and species groups, including rare and elusive animals such as aardwolf and zorilla (see Table 1).
In this report we describe the field methods, citizen science interface, and consensus algorithm used to produce the following datasets:
(1) Images: Full-resolution images produced by the survey.
(2) Raw classification data: All individual classifications made by all users on all image sets.
(3) Consensus data: Single classification per image set produced by applying the consensus algorithm to raw classifications, along with image metadata (date, time, location).
(4) Operation Dates: Metadata of when each camera was operational
(5) Gold-standard data: Expert classifications for a subset of 4,149 image sets.
We anticipate broad interdisciplinary re-use of these datasets with applications that span basic and applied ecology, citizen-science research, machine learning, and computer vision. The consensus dataset provides species-specific capture histories that can be analysed in a number of ways to evaluate population and community dynamics, either within Serengeti or as part of a larger cross-reserve analysis (see Usage Notes for details). The applications for this dataset extend beyond ecological research. For example, increasing the quality of citizen-science data is an area of active research21–24. Computer science and informatics researchers can use the raw (un-aggregated) citizen-science answers to develop more complex aggregation algorithms and to test their performance against the gold-standard dataset25. Additionally, computer-vision researchers need large human-annotated sets of imagery as training sets in machine-learning algorithms26. Our collaborators are currently using this dataset to automate species detection, classification and similar-species differentiation, as well as to develop combined human-machine learning systems and imaging systems for searchable colour. Subsets of the consensus dataset have also been used in classrooms to engage students in authentic research that spans ecology, animal behaviour, and computer science (see Usage Notes for examples).
Methods
Field methods
We set up an initial camera survey at 200 sites within the long-term Serengeti Lion Project study area from June to November 2010 (Fig. 1a). Cameras were re-installed in February 2011 and have operated continuously thereafter. We expanded the survey from 200 to 225 traps in February 2012 and it is currently on-going. This paper describes data collected until May 2013.
Layout
The camera-trap layout placed each camera at the centre of a 5 km2 grid cell (Fig. 1b), so as to offer systematic coverage of the entire study area and allow simultaneous monitoring of multiple species1,20,27–29. The precise location of each camera was selected as the nearest suitable tree to the pre-determined centre point of each grid cell, which was typically within 250 m of the centre. We selected sites to minimize camera misfires by prioritizing trees that offered shade and by avoiding trees surrounded by tall grass. Where no trees were available within 1 km of the grid cell centre point, we placed cameras on metal poles (Fig. 2).
Sites
We set cameras ~50 cm above ground level to capture medium to large vertebrates, housed in steel cases that were attached to trees with 10 cm hardened-steel lag bolts. We trimmed tall grass to <30 cm, and removed low-hanging branches to minimize risk of camera misfires and improve the unobstructed view from the camera. Cameras were pointed to minimise obstructions or risk of misfires rather than with respect to compass direction.
Cameras
We primarily used Scoutguard (SG565) incandescent cameras. We initiated the survey using DLC Covert II cameras with an infrared flash, but poor night-image quality prompted the transition to incandescent cameras. The cameras deployed in 2011 included a mixture of DLC Covert Reveal and SG565. Since 2012, all cameras and replacements have been SG565. Animals and weather damaged approximately 15% of cameras annually, requiring repeated replacement.
All survey cameras used passive infrared sensors that were triggered by a combination of heat and motion. Although standard camera-trapping protocols recommend setting sensitivity to ‘high’ for warm climates, this produced unacceptable levels of misfires by the movements of tall vegetation or shadows, thus we set sensor sensitivity to ‘low’ to minimize misfires. The detection radius and field of view were approximately 14 m and 45° for all cameras.
We set all cameras to take 3 photos per trigger in the daytime. At night, infrared-flash cameras took 3 photos per trigger, but incandescent-flash cameras could only take 1 image per trigger due to flash limitations (and occasional camera malfunction created a small number of image sets with varying numbers of images). We refer to each trigger as a ‘capture event’ and the resulting 1–3 images as an ‘image set’; capture events are the units of analysis for ecological studies and comprise the results presented here. We set cameras to ensure at least 1-minute delay between capture events to prevent the memory card being filled to capacity by a single individual or herd.
Maintenance
We checked each camera every 6–8 weeks. Except in cases of camera malfunction or damage, this schedule was sufficient to replace batteries and SD cards and ensure continuous operation. We labelled SD cards with the Site ID and the date retrieved and reviewed images in the field to ensure that the camera had functioned properly. We then installed new SD cards and triggered cameras to photograph placards that indicated Site ID, date, and time.
Data management
We wrote Python scripts to extract date/time from the image files and season, site, and card information from the directory structure. Common errors that arose from camera malfunction (typically due to animal or weather damage) included: the recording of videos instead of still images, incorrect time-stamps for a portion of images, and only 1–2 photos per capture event instead of three. We wrote code in Python, MySQL, and R to flag and correct these errors in the metadata.
Data processing
Platform
We partnered with the online citizen science platform The Zooniverse (www.zooniverse.org) to develop the Snapshot Serengeti website (www.snapshotserengeti.org), an online interface where the general public helps process camera trap data. The Snapshot Serengeti website utilizes the Zooniverse’s platform Ouroboros, written in Ruby on Rails (https://github.com/zooniverse/serengeti). Volunteer classifiers interact with a custom-built JavaScript front-end to classify image sets and results are saved in a MongoDB datastore. Each classification is recorded alongside the time of classification and the identity of the classifier in the form of either a unique identifier assigned by the Zooniverse (for logged in users) or an IP address (for users who have not logged in). Ouroboros also allows for custom rules for image-set retirement, as discussed below, and the system can scale rapidly to cope with the demands of a popular site. The interface and images are hosted on Amazon Web Services via Amazon’s Simple Storage Service (S3).
Task flow
On the Snapshot Serengeti interface (Fig. 3), volunteers identify species in each image set, count the number of individuals, classify behaviour, and indicate the presence/absence of young. For image sets that contain more than one image, volunteers initially see the second image in the set and can toggle between images or use the ‘play’ feature to animate the images. We designed the task flow to help guide people with no background knowledge through the process of identifying the animal(s) in question from 48 possible species and species groups while still providing a rapid route to classification for more knowledgeable participants. Users filter potential species matches by morphological characteristics such as horn shape, body shape, colour, pattern, and tail shape or jump straight to selecting from a list of all species. A ‘nothing here’ button allows users to classify image sets without any animals present. We do not offer an ‘impossible’ or ‘I don’t know’ option because previous testing on a small-scale prototype indicated that such answers were overused and provided no information on the actual species classification, thus wasting volunteer effort. Image difficulty (and probability of being correct) can instead be assessed by measuring variance across individual volunteer answers (see Technical Validation)
Circulation and retirement
We circulate each image set to multiple users and retire image sets from circulation when they have met one of the following criteria (see Table 2 and 3 & Fig. 4 for examples):
Sample images from image sets retired from Snapshot Serengeti as (a) blank: receiving five consecutive ‘nothing here’ classifications, (b-c) consensus: receiving 10 matching species classifications, and (d) complete: receiving 25 classifications regardless of agreement. Note that the plurality algorithm correctly arrived at ‘giraffe,’ ‘spotted hyena,’ and ‘impala’ for images b-d, respectively (see Tables 2 and 3 for individual classifications).
-
Blank: the first 5 classifications are ‘nothing here’.
-
Blank_Consensus: 10 ‘nothing here’ classifications, not necessarily consecutive.
-
Consensus: 10 matching classifications of species or species combination (e.g., 10 identifications of ‘lion’ or 10 identifications of ‘lion-zebra’); these classifications do not have to be consecutive.
-
Complete: 25 total non-‘nothing here’ classifications (does not require consensus for any single species).
Note that volunteers classified Snapshot Serengeti data faster than images were produced, and images were re-circulated for classroom use and testing the value of additional classifications. As a result, the number of classifications (11–57 for images containing animals) generally exceeded the number needed for retirement under the above rules.
Data aggregation
We implemented a simple plurality algorithm to transform the volunteer classifications for each image set into a single aggregated species identification. First, we calculated the number of different species present in an image set as the median number of different species identified across all users for that image set. For all image sets, we assigned the one (or more) species with the most ‘votes’ as the aggregated answer.
We calculated the number of individuals present for each identified species as the median number reported for that image set for that species by all volunteers. We also calculated the proportion of users who chose each behavioural activity or presence of young.
To assess the accuracy of aggregated classifications, we calculated an evenness index, using all non-blank classifications for each image set. When all classifications were in agreement, we assigned the value zero, indicating high accuracy. Otherwise, we used Pielou’s evenness index (Pielou 1966), calculated as , where S is the number of different species chosen among all volunteers, and pi is the proportion of ‘votes’ that species i received. The Pielou evenness index ranges from 0 to 1, with 0 indicating low evenness and high accuracy and 1 indicating high evenness and low accuracy. Note that the Pielou evenness index is expected to be high for image sets with multiple species and therefore is not a useful gauge of accuracy in these cases.
Code availability
Classification Interface
The code used to create the Snapshot Serengeti web interface is publicly available at https://github.com/zooniverse/serengeti (current) and archived on figshare30.
Data processing and consensus calculation
The scripts used to process the data and calculate the consensus classifications are publicly available at https://github.com/mkosmala/SnapshotSerengetiScripts (current) and archived on figshare31.
Data Records
All classification data and metadata are publicly available at Dryad (Data Citation 1).
Images: (all_images.csv; 3,198,737 data rows) URL information for retrieving each image; 1 record per image. All images in this data descriptor can be accessed at https://snapshotserengeti.s3.msi.umn.edu by appending the URL_Info field. For example, appending the value ‘S1/B04/B04_R1/S1_B04_R1_PICT0012.JPG’ yields the full URL: https://snapshotserengeti.s3.msi.umn.edu/S1/B04/B04_R1/S1_B04_R1_PICT0012.JPG. Pasting this value into a browser will display the image in the browser. Note that while we provide all images via the University of Minnesota Supercomputing Institute, this is not a proper archive site. Currently, there are no archiving systems or organizations available for storing the terabytes of images from our study. We hope that image archiving options will become available in the near future.
-
CaptureEventID: A unique identifier for each capture event and resultant image set.
-
URL_Info: A URL suffix to be appended to ‘https://snapshotserengeti.s3.msi.umn.edu/’ to yield the full URL of the image.
Raw classification data: (raw_data.csv; 10,530,564 data rows) Raw classification dataset; 1 record per unique user, capture event, and species. Includes images retired as ‘Blank’ and ‘Blank_consensus.’
-
CaptureEventID: A unique identifier for each capture event and resultant image set.
-
ClassificationID: A unique identifier for each classification event (one user classifying a single capture event). If a single user identifies multiple species within a capture event, they share the same classification ID.
-
UserID: Unique user ID for logged-in users; sessionID (unique computer & browser information) for non-logged-in users.
-
Species: Species selected from a list of 48 options or ‘blank’ for ‘nothing here’
-
Count: Number of individuals, estimated as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11–50 or 51+.
-
Standing: Binary indicator of whether any individuals are standing.
-
Resting: Binary indicator of whether any individuals are resting.
-
Moving: Binary indicator of whether any individuals are moving.
-
Eating: Binary indicator of whether any individuals are eating.
-
Interacting: Binary indicator of whether any individuals are interacting (including both intra- and inter-specific interactions).
-
Babies: binary indicator of whether young were present.
Consensus classification data and metadata: (consensus_data.csv; 334,671 data rows) Applying the plurality algorithm to the raw classification data yielded a single classification per capture event, accompanied by measures of uncertainty and difficulty. Each species classified in a single capture event receives its own record and species-specific measures of uncertainty. Metadata (data/time & location) are provided to facilitate ecological analyses. This dataset excludes all images retired as ‘Blank’ or ‘Blank_consensus.’
-
CaptureEventID: A unique identifier for each capture event and resultant image set. Links to CaptureEventID from raw classification data.
-
NumImages: The number of images in the image set/capture event.
-
DateTime: The date-time stamp is reported in yyyy-mm-dd hh:mm:ss. Time zone is UTC + 3:00. Note that daylight savings time is not observed in Tanzania.
-
SiteID: The alpha-numeric site code.
-
LocationX, LocationY: UTM X and Y coordinates of the site (datum Arc1960, zone 36S)
-
NumSpecies: The number of species in this capture event.
-
Species: Species of animal in the capture event (one of 48 possibilities).
-
Count: Median number of individuals, estimated as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11–50 or 51+.
-
Standing: The proportion of users who selected this behaviour for this species.
-
Resting: The proportion of users who selected this behaviour for this species.
-
Moving: The proportion of users who selected this behaviour for this species.
-
Eating: The proportion of users who selected this behaviour for this species.
-
Interacting: The proportion of users who selected this behaviour for this species.
-
Babies: The proportion of users who selected ‘young present’ for this species.
-
NumClassifications: The total number of users who classified this capture event.
-
NumVotes: The total number of users who selected this species for this capture event.
-
NumBlanks: The number of users who selected ‘nothing here’ for this capture event.
-
Evenness: The Pielou evenness index of species classifications for the capture event.
Operation dates: (search_effort.csv; 1,128 data rows) The dates that each camera was active and functioning properly, extracted from the image EXIF data as the first and last dates of valid photographs on a given SD card. Valid photographs are defined as those taken while the camera was secured on the tree pointing outwards (as opposed to photos taken after a camera was torn down and facing the ground).
-
Site ID: The alpha-numeric site code.
-
Start date: Date of first valid image on a given SD card.
-
End date: Date of last valid image on the SD card.
Gold standard data: (gold_standard_data.csv; 4,432 data rows) Expert classifications for 4,149 capture events. Note that gold-standard answers are more accurate than answers provided by a single expert because multiple experts reviewed all images for which any single expert expressed uncertainty. In 0.2% of images, the panel of experts agreed that no authoritative species identification could be made; those images are marked as ‘impossible.’
-
CaptureEventID: Same as in the raw and reduced classification data.
-
NumSpecies: The number of species in this capture event.
-
Species: One of the 48 possibilities or ‘impossible.’
-
Count: Number of individuals, estimated as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11–50 or 51+.
Technical Validation
We asked five researchers with extensive wildlife identification experience to classify 4,149 randomly selected image sets containing animals using the Snapshot Serengeti interface; 263 image sets received two expert classifications and 8 image sets received three, for a total of 4,428 classifications. The experts noted whether any image sets were especially difficult or whether they thought the image was identifiable at all. In cases where experts disagreed with the results of the plurality algorithm or had marked an image set as particularly difficult or impossible, AS and CP made the final authoritative identification. Thus, the gold standard dataset included a small number of images that were agreed by multiple experts to be ‘impossible’ to identify. Because the Snapshot Serengeti interface does not allow ‘impossible’ as an option, the consensus answers for these images are incorrect by definition. We compared citizen-science classifications derived from the plurality algorithm with the expert-classified ‘gold standard’ dataset to assess accuracy of species identifications and counts of individuals.
Of the 4,149 image sets viewed by experts, 96.6% of algorithm-derived answers agreed with the expert species classification, though the accuracy rate varied by species (Table 4). Of the 142 image sets in which the algorithm did not agree with the experts, 21% (n=30) were marked as ‘impossible’ by experts, 29.5% (n=42) reflected cases where the algorithm only identified one of two species identified by experts (for example, only zebra in an image set where both wildebeest and zebra had been present), 3.5% involved cases where the algorithm indicated two species whereas the experts only reported one, and 45.8% (n=65) reflected true errors in which the algorithm reported the wrong species. The most common mistakes included misidentification of birds (n=11) and incorrectly identifying Grant’s gazelles as Thomson’s gazelles (n=11).
Variance in raw classifications strongly predicted whether image sets were classified correctly. Image sets for which the algorithm differed from expert IDs had higher levels of disagreement among raw classifications: the mean evenness score (+/−standard error) was 0.451 (+/−0.004) for correct answers versus 0.725 (+/−0.014) for incorrect answers. Classifications of images that experts identified as ‘impossible’ are considered to be incorrect. We provide guidelines in the Usage Notes for using measures of disagreement to measure certainty that a consensus classification is correct and to target image sets for review or exclusion in any given analysis.
For image sets where the plurality algorithm accurately captured all or a subset of species present, we compared the species-specific counts reported by the algorithm to expert classifications (n=4,269 species counts). 76.4% of algorithm-derived counts matched expert counts exactly, and 92.98% of algorithm-derived counts were within +/−1 bin of the expert classification (Table 5). Accuracy varied by the number of individuals present: users were > 97% accurate when counting a single individual and least accurate distinguishing between 5–10 individuals.
Usage Notes
We envision broad applications for these datasets in ecology, informatics, computer vision, and education. Here we provide additional details and guidelines.
Ecological analyses
The consensus classifications are equivalent to raw data produced by standard camera trapping surveys and include all metadata necessary for applying robust analytical procedures that explicitly consider variations in detection probability. We provide dates of activity for every camera trap, as well as dates, times, and locations for every image. Researchers can thus aggregate camera activity (‘search effort’) and species-specific capture histories into time spans suitable for relative abundance indices32–34, single and multi-season occupancy modelling8,35–38 across multiple scales39, dynamic and multi-species occupancy modelling1,40–42, hierarchical binomial or n-mixture models43,44. All images are downloadable and identified to species, so capture histories of individually recognized animals can be constructed for species with distinct pelage patterns (e.g., cheetahs, leopards, hyenas, civets, genets, etc.). In such cases, sophisticated mark-recapture analyses can permit spatially-explicit inference45,46.
Note that ecological analyses sometimes require higher species-identification accuracy than the 97% returned by the consensus algorithm. Our three ‘certainty’ metrics for consensus answers reflect image difficulty and likelihood of being correct and thus provide guidelines for targeting ‘uncertain’ images for expert review or exclusion from the analysis (see certainty and difficulty section below).
Citizen science and informatics analyses
Crowdsourcing and citizen science are being used increasingly often to produce science datasets22–24, but they require robust methods to measure and validate data quality. While our consensus dataset derives from a simple plurality algorithm, more complex algorithms can improve upon these results. For example, Hines et al.25 weighted raw classifications by individual accuracy, raising overall accuracy to 98%. Our raw classification dataset could be used to develop and test algorithms that employ user-weighting or even apply a Bayesian framework to incorporate information about species likelihood based on previous or subsequent images.
Computer vision
Object search-and-recognition research requires large data sets of labelled imagery. Reliable data sets of wild animals are rare, due to the enormous task of hand-annotating large numbers of images. By using the raw images together with the consensus dataset, machine-learning algorithms could be developed to automatically detect and identify species, using part of the dataset for training the image-recognition algorithm and the rest for testing the algorithm. Raw images could be used separately, or in conjunction with the consensus data set, to research automatic detection of textures, patterns, and other characteristics of outdoor scenes.
Education
Higher education teachers can use the consensus dataset and metadata to develop curricula around the scientific method, using charismatic fauna to engage students. As examples, students can ask questions about species abundances and distributions, daily activity patterns and seasonal movements. The dataset can be tailored by the instructor for use by undergraduates for authentic research experiences. (See http://www.cbs.umn.edu/explore/education-outreach/hhmi/projects/nonmajor/serengeti as an example.) This dataset is also suitable for graduate level coursework in ecology, informatics, and computer vision. (See http://rogerioferis.com/VisualRecognitionAndSearch2014/Projects.html as an example.)
Dataset and calculation details
The 48 possible ‘species’ options include four ‘group’ categories: human, bird (other), reptiles, and rodents. ‘Human’ reflects any human activity, including vehicles and hot-air balloons. We made no effort to distinguish among species of hare, jackal, and mongoose. Additionally, lions were split into two categories: Lion (male) and Lion (female & cubs).
Users typically selected as many behaviours as applicable for a given species in each image but sometimes classified two individuals as displaying two different behaviours by listing the same species twice. For example, an image with one standing zebra and one moving zebra might receive one classification of ‘1 zebra, standing’ and ‘1 zebra, moving’—resulting in multiple classifications of the same species with the same ClassificationID. We standardized classifications by combining multiple classifications of the same species within a single ClassificationID before applying the consensus algorithm. So ‘1 zebra, standing’ and ‘1 zebra, moving’ were combined to form ‘2 zebras, standing and moving,’ which was the most common classification for such images. Note that the raw classification data set contains separate assessments made by each volunteer and thus does not combine multiple records within ClassificationID for any single image.
Certainty & difficulty measures
The plurality algorithm produces classifications that are97% correct on average. However, the accuracy varies by species and difficulty and certain analyses may require greater accuracy than obtained from the plurality algorithm. ‘Percent support’ for each species in each image set can be calculated as NumVotes/NumClassifications and reflects the proportion of users who identified that species as present in the image set. High values indicate high certainty. High values for NumBlanks and Evenness (for single species image sets) tend to reflect more difficult image sets for which the consensus vote is more likely to be incorrect. We found that when some users were unsure of their classification, they used the ‘nothing here’ option instead of guessing a species. As a result, difficult images are more likely to have more blank classifications. As described in the Technical Validation, higher evenness score reflects lower agreement among classifications and were more likely to be incorrect for single species captures. To increase certainty of datasets for analyses, analyses can be restricted to data that meet any threshold for percent support, Evenness, or NumBlanks.
Additional Information
How to cite this article: Swanson, A. et al. Snapshot Serengeti, high-frequency annotated camera trap images of 40 mammalian species in an African savanna. Sci. Data 2:150026 doi: 10.1038/sdata.2015.26 (2015).
References
References
O’Connell, A. F., Nichols, J. D. & Karanth, K. U. Camera Traps in Animal Ecology: Methods and Analyses. (Springer, 2011).
Surridge, A., Timmins, R., Hewitt, G. & Bell, D. Striped rabbits in Southeast Asia. Nature 400 (1999).
Holden, J., Yanuar, A. & Martyr, D. J. The Asian Tapir in Kerinci Seblat National Park, Sumatra: evidence collected through photo-trapping. Oryx 37, 34–40 (2003).
Karanth, K. U. Estimating tiger Panthera tigris populations from camera-trap data using capture—recapture models. Biol. Conserv. 71, 333–338 (1995).
Rowcliffe, J. M., Field, J., Turvey, S. T. & Carbone, C. Estimating animal density using camera traps without the need for individual recognition. J. Appl. Ecol. 45, 1228–1236 (2008).
O’Brien, T. G. & Kinnaird, M. F. Density estimation of sympatric carnivores using spatially explicit capture-recapture methods and standard trapping grid. Ecol. Appl. 21, 2908–2916 (2011).
Royle, J. A. & Dorazio, R. M . Hierarchical Modeling and Inference in Ecology: The Analysis of Data from Populations, Metapopulations and Communities. (Academic, 2008).
MacKenzie, D. I., Nichols, J. D., Hines, J. E., Knutson, M. G. & Franklin, A. B. Estimating site occupancy, colonization, and local extinction when a species is detected imperfectly. Ecology 84, 2200–2207 (2003).
Kéry, M. Introduction to WinBUGS for Ecologists – A Bayesian Approach to Regression, ANOVA, Mixed Models and Related Analyses. (Academic Press, 2010).
Fegraus, E. H. et al. Data acquisition and management software for camera trap data: A case study from the TEAM Network. Ecol. Inform. 6, 345–353 (2011).
Krishnappa, Y. S. & Turner, W. C. Software for minimalistic data management in large camera trap studies. Ecol. Inform. 24, 11–16 (2014).
Swinnen, K. R. R., Reijniers, J., Breno, M. & Leirs, H. A novel method to reduce time investment when processing videos from camera trap studies. PLoS ONE 9, e98881 2014).
Holdo, R. M., Holt, R. D. & Fryxell, J. M. Opposing rainfall and plant nutritional gradients best explain the wildebeest migration in the Serengeti. Am. Nat. 173, 431–445 (2009).
Packer, C., Swanson, A., Ikanda, D. & Kushnir, H. Fear of darkness, the full moon and the nocturnal ecology of African lions. PLoS ONE 6, e22285 2012).
Sinclair, A. R. E. & Norton-Griffiths, M. . Serengeti: Dynamics of an Ecosystem. (University of Chicago Press, 1979).
Sinclair, A. R. E., Packer, C., Mduma, S. A. R. & Fryxell, J. M. Serengeti III: Human Impacts on Ecosystem Dynamics. (University of Chicago Press, 2008).
TAWIRI. Aerial Census in the Serengeti Ecosystem. (Tanzania Wildlife Research Institute, 2008).
TAWIRI. Aerial Census in the Serengeti Ecosystem. (Tanzania Wildlife Research Institute, 2010).
Strauss, M. K. L. & Packer, C. Using claw marks to study lion predation on giraffes of the Serengeti. J. Zool. 289, 134–142 (2013).
O’Brien, T. G., Baillie, J. E. M., Krueger, L. & Cuke, M. The Wildlife Picture Index: monitoring top trophic levels: The Wildlife Picture Index. Anim. Conserv. 13, 335–343 (2010).
Tulloch, A. I. T., Possingham, H. P., Joseph, L. N., Szabo, J. & Martin, T. G. Realising the full potential of citizen science monitoring programs. Biol. Conserv. 165, 128–138 (2013).
Bonney, R. et al. Citizen Science: A developing tool for expanding science knowledge and scientific literacy. BioScience 59, 977–984 (2009).
Dickinson, J. L., Zuckerberg, B. & Bonter, D. N. Citizen science as an ecological research tool: challenges and benefits. Annu. Rev. Ecol. Evol. Syst. 41, 149–172 (2010).
Sauermann, H. & Franzoni, C. Crowd science user contribution patterns and their implications. Proc. Natl Acad. Sci 112, 679–684 (2015).
Hines, G., Swanson, A., Kosmala, M. & Lintott, C. J. in Proceedings of the Twenty-Seventh Annual Conference on Innovative Applications of Artificial Intelligence (IAAI-15) (2015).
Russell, B. C., Torralba, A., Murphy, K. P. & Freeman, W. T. LabelMe: a database and web-based tool for image annotation. Int. J. Comput. Vis 77, 157–173 (2008).
Tobler, M. W., Carrillo-Percastegui, S. E., Leite Pitman, R., Mares, R. & Powell, G. An evaluation of camera traps for inventorying large- and medium-sized terrestrial rainforest mammals. Anim. Conserv. 11, 169–178 (2008).
Rovero, F. & Marshall, A. R. Camera trapping photographic rate as an index of density in forest ungulates. J. Appl. Ecol. 46, 1011–1017 (2009).
O’Brien, T. G. On the use of automated cameras to estimate species richness for large- and medium-sized rainforest mammals. Anim. Conserv. 11, 179–181 (2008).
Kosmala, M. et al. SnapshotSerengetiScripts. figsharehttps://doi.org/10.6084/m9.figshare.1397507 (2015).
Lintott, C. et al. Snapshot Serengeti. figsharehttps://doi.org/10.6084/m9.figshare.1397518 (2015).
Carbone, C. et al. The use of photographic rates to estimate densities of tigers and other cryptic mammals. Anim. Conserv. 4, 75–79 (2001).
O’Brien, T. in Camera Traps in Animal Ecology. Methods and Analyses. (eds O’Connell, A. F., Nichols, J. D. & Karanth, U. D. 71–96 (Springer, 2011).
O’Brien, T. G., Kinnaird, M. F. & Wibisono, H. T. Crouching tigers, hidden prey: Sumatran tiger and prey populations in a tropical forest landscape. Anim. Conserv. 6, 131–139 (2003).
MacKenzie, D. I. et al. Estimating site occupancy rates when detection probabilities are less than one. Ecology 83, 2248–2255 (2002).
MacKenzie, D. I. et al. Occupancy Estimation and Modeling: Inferring Patterns and Dynamics of Species. (Elsevier, 2006).
Mackenzie, D. I. & Royle, J. A. Designing occupancy studies: general advice and allocating survey effort. J. Appl. Ecol. 42, 1105–1114 (2005).
Bailey, L. L., Hines, J. E., Nichols, J. D. & MacKenzie, D. I. Sampling design trade-offs in occupancy studies with imperfect detection: examples and software. Ecol. Appl. 17, 281–290 (2007).
Nichols, J. D., Hines, J. E., Mackenzie, D. I., Seamans, M. E. & Gutiérrez, R. J. Occupancy estimation and modeling with multiple states and state uncertainty. Ecology 88, 1395–1400 (2007).
MacKenzie, D. I., Bailey, L. L. & Nichols, J. D. Investigating species co-occurrence patterns when species are detected imperfectly. J. Anim. Ecol. 73, 546–555 (2004).
Kéry, M. & Andrew Royle, J. Hierarchical modelling and estimation of abundance and population trends in metapopulation designs. J. Anim. Ecol. 79, 453–461 (2010).
Royle, J. A. & Kéry, M. A Bayesian state-space formulation of dynamic occupancy models. Ecology 88, 1813–1823 (2007).
Kéry, M., Royle, J. A. & Schmid, H. Modeling avian abundance from replicated counts using binomial mixture models. Ecol. Appl. 15, 1450–1461 (2005).
Royle, J. A. N-Mixture models for estimating population size from spatially replicated counts. Biometrics 60, 108–115 (2004).
Chandler, R. B. & Royle, J. A. Spatially explicit models for inference about density in unmarked or partially marked populations. Ann. Appl. Stat 7, 936–954 (2013).
Sollmann, R. et al. A spatial mark–resight model augmented with telemetry data. Ecology 94, 553–559 (2013).
Data Citations
Swanson, A. Dryad (2015) https://doi.org/10.5061/dryad.5pt92
Acknowledgements
Research clearance was provided by the Tanzania Wildlife Research Institute and Tanzania National Parks. We thank members of the Serengeti Lion Project, particularly Daniel Rosengren, George Gwaltu Lohay, and Stanslaus Mwampeta, the Zooniverse staff, and the 28,040 volunteers who contributed to Snapshot Serengeti classifications (complete list at www.snapshotserengeti.org/#/authors). This work was supported by NSF grant DEB-1020479 to CP for maintenance of the long-term Lion Project, the University of Minnesota Supercomputing Institute, private donations raised during the Serengeti Live and Save Snapshot Serengeti crowd-funding campaigns, and by grants to AS from Explorer’s Club, UMN Thesis Research Grants, UMN Office of International Programs, American Society of Mammalogists, and Minnesota Zoo Ulysses S. Seal Conservation fund. Snapshot Serengeti website development was funded by awards to the Zooniverse from the Alfred P. Sloan Foundation.
Author information
Authors and Affiliations
Contributions
A Swanson designed and implemented the camera survey, guided development of the Snapshot Serengeti interface, contributed to data management, and wrote the paper. MK guided development of the Snapshot Serengeti interface, designed and implemented data aggregation approaches and data quality control, and contributed to data management and to paper writing. CL oversaw the development of the Snapshot Serengeti interface as the PI for www.zooniverse.org. RS led the development of the Snapshot Serengeti interface and managed the volunteer community. ASmith led the development of the Snapshot Serengeti interface. CP provided funding and guidance for the implementation of the camera survey and contributed to paper writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Swanson, A., Kosmala, M., Lintott, C. et al. Snapshot Serengeti, high-frequency annotated camera trap images of 40 mammalian species in an African savanna. Sci Data 2, 150026 (2015). https://doi.org/10.1038/sdata.2015.26
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2015.26
This article is cited by
-
Fusion of visible and thermal images improves automated detection and classification of animals for drone surveys
Scientific Reports (2023)
-
Wildlife Population Assessment: Changing Priorities Driven by Technological Advances
Journal of Statistical Theory and Practice (2023)
-
Neural architecture search based on packed samples for identifying animals in camera trap images
Neural Computing and Applications (2023)
-
Dynamic Curriculum Learning for Great Ape Detection in the Wild
International Journal of Computer Vision (2023)
-
Towards Automated Ethogramming: Cognitively-Inspired Event Segmentation for Streaming Wildlife Video Monitoring
International Journal of Computer Vision (2023)