Abstract
The study of non-model organisms stands to benefit greatly from genetic and genomic data. For a better understanding of the molecular mechanisms driving neuronal development, and to characterize the entire leech Hirudo medicinalis central nervous system (CNS) transcriptome we combined Trinity for de-novo assembly and Illumina HiSeq2000 for RNA-Seq. We present a set of 73,493 de-novo assembled transcripts for the leech, reconstructed from RNA collected, at a single ganglion resolution, from the CNS. This set of transcripts greatly enriches the available data for the leech. Here, we share two databases, such that each dataset allows a different type of search for candidate homologues. The first is the raw set of assembled transcripts. This set allows a sequence-based search. A comprehensive analysis of which revealed 22,604 contigs with high e-values, aligned versus the Swiss-Prot database. This analysis enabled the production of the second database, which includes correlated sequences to annotated transcript names, with the confidence of BLAST best hit.
Design Type(s) | replicate design • transcription profiling by high throughput sequencing design |
Measurement Type(s) | transcription profiling assay |
Technology Type(s) | RNA sequencing |
Factor Type(s) | replicate analysis |
Sample Characteristic(s) | Hirudo medicinalis • ganglion |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
For historic reasons, certain organisms, such as Escherichia coli, Saccharomyces cerevisiae, Arabidopsis thaliana, Caenorhabditis elegans, Drosophila melanogaster, Danio rerio or Mus musculus have gained the status of model organisms, due to their biological attributes and to technical advantages1. The study of these organisms has benefited from the concerted effort of a large research community allocating vast resources, resulting in a significantly larger body of genetic and genomic information concerning these organisms than any other. However, many other organisms have unique traits that make them valuable models for the study of specific biological processes. Until recently, genetic and genomic information for this latter group was scarce due to limitations of cost and labour intensity.
Next Generation Sequencing (NGS) tools have made the production of genetic and genomic information more accessible. NGS has made such research feasible for the study of all organisms, including those for whom no reference genetic data is available, the so-called non-model organisms1. Moreover, the innovative approaches to the analysis of these new sequencing data turned previously perceived obstacles into potentially surmountable challenges.
In the study presented here, we chose to focus on the transcriptome of the medicinal leech, Hirudo medicinalis CNS, which is a non-model organism, but serves as a well-studied model in neurobiology, specifically in neuronal development, regeneration and repair2,3. The leech CNS is composed of 6 fused ganglia at the head, 21 highly similar body ganglia and 7 fused tail ganglia4. Each ganglion contains approximately 200 pairs of neurons and is linked to its neighbours by thousands of axons5. This valuable model offers an interesting platform for use of molecular and cellular scientific methods for the evaluation of the involvement of specific cells in the regenerative processes6. Previous work has yielded characterization of specific genes in leech CNS7,8, and genes of interest were studied using the candidate gene approach9–12. Recently, an expression sequence tag (EST) database was constructed and is now available to the scientific community13. Yet, functional genomic studies in the Hirudo medicinalis are in their infancy13,14.
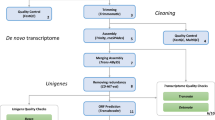
While several previous studies have shed some light on particular genes and gene expression patterns13–15, the full transcriptomic data of the Hirudo medicinalis CNS is still limited. In our related work at Bioinformatics we provided an in-depth spatial regulation analysis of the CNS transcriptome data and showed the potential of combining expression distribution patterns to produce a spatio-transcripto map along the ganglia chain15. As illustrated in Fig. 1, we collected RNA from three distinct locations along the leech CNS (ganglia 2, 10 and 19). To achieve the single ganglion resolution, we collected the RNA content of these organs and had to amplify the RNAs, using the NuGEN amplification kit which known to be beneficial in low amounts of RNA16, before sequencing. In total, we sequenced 221.1 million high-quality short reads 50 bp single-end from the leech CNS. Then, using the de-novo assembly program, Trinity, we reconstructed these reads to produce the first model of the leech CNS transcriptome. By combining those three distinct points along the leech CNS we assumed that our data reveals most of the transcripts that are expressed in steady state of neuronal cells in the leech CNS.
We collected 12 ganglia in total from the Hirudo medicinalis. Three biological replicates were harvested from ganglia 2, 10 and 19 and 3 technical replicates harvested from ganglion 10. Due to the low initial yield of total RNA we first amplified the RNA using NuGEN kit and then used TruSeq kit to prepare the cDNAs. Next, we sequenced the cDNAs on Illumina HISeq2000 in 50 bp single end (SE) reads. The analysis started with assembling the short reads, using the de-novo assembly program Trinity, and continued with functional analysis and evolutionary relationships analysis using BLASTX.
In this Data Descriptor, we provide the full assembly and annotation datasets, aimed at making our data accessible to others for use in their research and for expanding the community understanding of this data. This study complements previous approaches to address similar questions in the leech CNS9,10,17. Sequencing the transcriptome is a prerequisite to the expansion of our knowledge on the nervous systems (physiological and pathological conditions). Utilizing these assembly data through an annotation dataset for these new transcripts may help in the accessibility and understanding of this data. The use of a simple model, the leech CNS, together with a novel assembly and analysis approaches, combine the transcriptome with a spatial configuration, thus producing a novel transcript database of the leech CNS. Furthermore, these leech databases can be used to define the underlying conserved genetic modules controlling the equivalent patterning processes along the CNS as well as serving to cross-validate each other. Similarity, these data will offer insights into the molecular mechanism that underpin the fundamental patterning differences between leech and related organisms.
Methods
These methods have been presented in an abbreviated form in the journal Bioinformatics15.
Animal conditions
The experiments were performed on the Hirudo medicinalis leech. All leeches were obtained from an adult Hirudo medicinalis colony grown in France at Ricarimpex Farm. Further to the transportation from the farm, leeches were maintained in our animal facility in tanks populated with about 20 leeches in a controlled environment, at 16 °C and 12 h/12 h day/night cycle.
Experimental design
Twelve samples with a focus on the Hirudo medicinalis CNS were taken from six different leeches for this experiment. Before use, leeches were placed on ice for 30 min and then dissected dorsally. Three ganglia (2, 10 and 19) were harvested from three leeches (Fig. 1). For technical replicas, ganglion number 10 was harvested from three additional leeches, pooled together for RNA isolation and separated into three samples for RNA-seq.
CNS/Ganglia collection
Total RNA was extracted from each ganglion using RNeasy Lipid Tissue (Qiagen). The quality and quantity of each RNA sample was assessed by Agilent’s 100 Bioanalyzer pico chip (Fig. 2a,b).
Representative electropherogram of Bioanalyzer analysis of total RNA integrity from the 2 samples. In (a) and (b) are the pico chip analysis for 2 representative RNA samples (ganglion 10). The x axis outlines the time in seconds and the y axis provides the fluorescence. The first peak, in the ~22 s represents the ladder. The second peak in ~25 s represents the RNA. The third peak in the ~41 s represents the ribosomal RNA. For our study we selected only RNA samples that were found to have RNA concentration >45 pg/μl. In (c) and (d) are the DNA 1000 chip for 2 representative amplified cDNAs samples (ganglia 2 and 10, respectively). The first peak, in ~42 is the ladder and from ~60–110 is the cDNA. The distribution of the diagrams is due to the different sizes of the cDNAs after the amplification. All cDNAs showed concentration >72 ng/ul.
RNA amplification
The initial RNA yield was low, requiring amplification of RNAs using a specific kit prior to the use of the mRNA-TruSeq preparation kit. RNA was amplified using Ovation Kit v2.0 (NuGEN) (Fig. 2c,d). Before amplification, all samples were lyophilized using a SpeedVac instrument and then suspended in 5 μl of nuclease-free water. This was the starting volume of NuGEN kit. Then, (as suggested by NuGEN), the 2 μg (in 100 μl) of cDNAs were fragmented by a Bioruptor instrument 3 cycles of 10 s of sonication and 90 s of pause. Library preparation proceeded with an ‘END-REPAIR’ reaction of NEB kit, then with TruSeq DNA/RNA libraries preparation according to Illumuna protocol. The 12 stock libraries were loaded on a High Sensitivity Chip and quantified on a QuBIT instrument, in order to prepare the two 6-plex pools were separated into two pools. The two pools were quantified (molarity) on Bioanalyzer HighSensitivity as stocks. In order to better balance the single libraries inside the pools, we decide to quantify libraries with qPCR, following the NuGEN suggestion for pool preparation.
Illumina sequencing
The cDNA libraries were generated using messenger RNA-seq (mRNA-seq) assay for transcriptome sequencing on Illumina Hiseq2000 (Data Citation 1 and Table 1). Three cDNA libraries were generated from the total RNA of ganglion number 19 and three cDNA libraries were generated from the pooled total RNA of ganglion 10 in equal amounts, and sequencing was performed in one lane to generate 50 bp single end (SE) reads. A similar procedure was carried out for ganglia numbers 2 and 10. Library construction and sequencing was performed by a commercial service provider (IGA, Applied Genomics Institute).
De novo assembly
Due to the fact that the genome/transcriptome of the Hirudo medicinalis is not available yet, we used de-novo tools for the reconstruction the transcriptome. The tools we used are Trinity18 (version trinityrnaseq_r2012-03-17) and Trans-ABySS19 (version 1.3.2). Here we report only the procedure and the results from Trinity, the full process and considerations can be seen in our analysis paper15. Trinity has been developed for assembly of short reads using de Bruijn graph algorithm by single k-mer. Trinity was executed in the inchworm method and the minimum contig length set to 200 nucleotides. The other parameters we used are default for Trinity single-end assembly (Trinity.pl—seqType fq—kmer_method inchworm—single seq.input—output seq.output—min_contig_length 200) (Data Citation 2 and Table 2).
Annotation dataset
In general, the creation of an annotation dataset is derived from a draft genome or transcriptome20,21. For the Hirudo medicinalis none of these are available. Therefore, we used BLASTX (version 2.2.23) to identify sequence conservation and to create an annotated dataset for the 73,493 contigs, generated by Trinity, against Swiss-Prot database downloaded from the National Center for Biotechnology Information (NCBI). We used blast commands setting the minimum e-value to be e−10 for maximal confidence of the contigs (blastall -p blastx -i Trinity.fasta -d Swiss-prot e−10 −o Trinity.fasta.out). Next, to improve the readability of blast output we kept only the first result for each contig by using Linux common command (grep -E ‘Query\=|No\ hits\ found|^gi\|’ Trinity.fasta.blastx.out|grep -A1 ‘Query\=\contig_name’| sed ‘/^--/d’| sed ‘s/path.*//’ | paste - - -d’;’Trinity.fasta.blasx.out.best-hit). Applying the described pipeline has led to a set of reliable annotated contigs (Data Citation 3).
Data Records
In this study we deposited four datasets. The first dataset is the RNA-Seq raw reads (Data Citation 1 and Table 1). This dataset contains 12 samples in total. Three biological replicates from each ganglia (numbers 2, 10 and 19), and three more technical replicates from ganglion number 10. The second dataset is the expression value of each of the Trinity assembly contigs (Data Citation 2 and Table 2). The third dataset contains the actual contigs (Data Citation 2 and Table 3). The fourth dataset is the annotation file (Data Citation 3). The annotation file is a comma separated value (CSV) format file with all of the annotated contigs generated by Trinity. The annotation file deposited in figshare depository. In general, there are 7 columns. Column #1 is the name of the contig, #2 is the length, #3 is the gi accession number, #4 is the sp accession number, #5 is the entry name, #6 is the BLASTX score and #7 is the e-value. The first two datasets described above (Data Citation 1 and Table 1, and Data Citation 2 and Table 2) were previously published in our related work in the journal Bioinformatics15, and the third dataset in Data Citation 2 (Table 3) and the forth dataset in Data Citation 3 are the core of this work and have not been published before.
Technical Validation
CNS collection quality control
Prior to processing, after samples were harvested, we assessed the quality and quantity of each sample using Agilent’s 100 Bioanalyzer pico chip. Only RNA samples which contained a concentration of 45–170 pg/μl were used for this experiment (Fig. 2a,b). Since the harvesting was only from single ganglions (~400 cells), we expected low concentrations of total RNA.
RNA amplification quality control
Following amplification, cDNAs were quantified using Nanodrop and Bioanalyzer DNA 1000 Chip (Fig. 2c,d). After balancing the two pools for a similar total pool volume, we proceeded with the procedure. The first pool volume was 47.2 μl and the second pool was 45.9 μl.
Sequencing quality control
We used multiple steps for testing sequencing quality. The first step included a count of total reads and total bases for each of the samples to ensure that the amounts are approximately of the same order of magnitude. These amounts were 10–26 million reads. As a second step, we tested samples to pass FastQC22 for basic statistics as quality estimation, per base sequence quality, per sequence quality score, length distribution and raw reads quality control (Table 4). As a third step, we estimated the sequencing depth of coverage by estimating the size of the transcriptome. We used the know information on a related organism, Helobdella robusta, which its transcriptome size is estimated to be 29032248 bp (http://metazoa.ensembl.org/Helobdella_robusta/Info/Annotation/#assembly). Thus, we estimated the sequencing depth of coverage to be (the number of sequenced bases divided by the predicted transcriptome size).
Assembly quality control
Following the use of Trinity, to make sure that the produced contigs are correct, we compared our model of transcriptome also to Helobdella robusta published transcriptome. We start with comparing basic statistics such as average length of contigs (Table 4). We calculated the average contig length for the Hirudo medicinalis and we found it to be 1124 (Table 5), which is very similar to the average transcript size for the Helobdella robusta which is 1239 according to the Joint Genome Institute (http://genome.jgi-psf.org/Helro1/Helro1.home.html). Next, we used Bowtie23 to map reads back to the contigs to test the mapping rate for each sample (Fig. 3a). Finally, we correlated the technical replicates between themselves, which showed a strong correlation (Fig. 3b) (Table 3). Moreover, for examining the completeness of the data we tested full-length over the innexin genes family to see if the transcripts are really full without any bias to particular side, and we found that 11 out of the 21 innexin reconstructed perfectly, and the other 10 almost in fully15. Moreover, for strengthening the completeness evaluation of our data we estimated the transcriptome size according to the EnsemblMetazoa known information of the Helobdella robusta (see above), and found that the number of coding genes for the Helobdella robusta is 23432 which is very similar to the number of contigs with high e-value that when aligning to the Swiss-Prot (Fig. 3c).
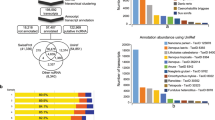
In (a) is the samples’ contribution to the assembly. The percentage in the pie chart outlines the percentage that each sample contributed to the reconstruction of the transcriptome dataset. The numbers in the pie chart outline the sample number. Out of all the 73,493 contigs, sample number 5 contributed to reconstruct 5% and sample number 9 contributed 15%. As can be seen in the chart, the percentiles vary from 5 to 15% and the dataset is not biased towards any specific sample. In (b) is a scatter plot of the technical replicates expression values for ganglion number 10 to verify that the samples sequenced without bias. The x axis outlines sample number 11 expression values, which represent technical replica number 2. The y axis provides sample number 12 expression values, which represent technical replica number 3. In the top right is the R2 correlation. We also tested the correlation between technical replica 1 to 2 and 1 to 3, and they have also showed strong R2, 0.789 and 0.7845 respectively. In (c) is the hits Scores Distribution. The x axis outlines the upper limit of the score. The left y axis provides the frequency, and the right y axis provides the cumulative. Out of the 73,493 transcripts reconstructed by Trinity 22,604 had high e-value hits in the Swiss-Prot database. The blast score gives an indication of the quality of the alignment. As can be seen, most transcripts aligned with a score higher than 50 and only 22 had a score under/equal 50.
Annotation quality control
To ensure the quality of our annotation we set the blast e-value threshold to e−10. We found that the blast scores were high (Fig. 3c). By counting the number of transcripts that passed the threshold, we saw that the approximate predicted size of the Hirudo medicinalis CNS transcriptome is 22,604. This result is very similar to size of the Helobdella robusta and the Capitella teleta transcriptomes (19,487 and 37,908, respectively).
Usage Notes
The data provided in this experimental set can be used for several purposes. First, it is possible to use the raw reads for executing a new experiment, with different analysis approaches. Second, each analysis step can be performed differently as all the technical experimental information is publicly available.
De novo assembly
By using our non-redundant set of 73,493 transcripts we generated using Trinity, the search in the dataset for genes of interest can be easily done by homologs using blast, or text based search if using the annotation table. Moreover, the raw reads provided here make it possible to use alternative methods for de novo assembly that may assemble the reads into a different set of transcripts than the ones we constructed. The de novo assembly can be performed using SOAPdenovo-Trans24 or Velvet/Oases25,26.
Differential expression
The combination of raw reads and transcripts allows the re-calculation of expression values for each transcript for each sample. There are different approaches for discovering differentially expressed genes. This can be further refined into new groups of genes expressed in each individual ganglion, or combinations of ganglia. This can be performed by using, for example, free Bioconductor packages such as DESeq27 or edgeR28 (http://www.bioconductor.org/).
Downstream analysis
The datasets provided here represented by 3 biological replicates for each condition and technical replicates for one condition. Comparing the ganglia sequenced here to other ganglia to determine the set of genes statistically significantly differentially expressed along the whole CNS, and not only three points.
Additional information
How to cite this article: Hibsh, D. et al. De novo transcriptome assembly databases for the central nervous system of the medicinal leech. Sci. Data 2:150015 doi: 10.1038/sdata.2015.15 (2015).
References
References
Tagu, D., Colbourne, J. K. & Nègre, N. Genomic data integration for ecological and evolutionary traits in non-model organisms. BMC Genomics 15, 490 (2014).
Coggeshall, R. E. & Fawcett, D. W. The fine structure of the central nervous system of the leech, Hirudo Medicinalis. J. Neurophysiol. 27, 229–289 (1964).
Meriaux, C. et al. Multiple changes in peptide and lipid expression associated with regeneration in the nervous system of the medicinal leech. PLoS One 6, 1–20 (2011).
Muller K. J., Nicholls J. G., Stent G. S. (Eds) Neurobiology of the Leech. (Cold Spring Harbor, 1981).
Macagno, E. R. Number and distribution of neurons in leech segmental ganglia. J. Comp. Neurol. 190, 283–302 (1980).
Wang, W.-Z., Emes, R. D., Christoffers, K., Verrall, J. & Blackshaw, S. E. Hirudo medicinalis: A platform for investigating genes in neural repair. Cell. Mol. Neurobiol. 25, 427–440 (2005).
Harvey, R. P. et al. Cloning and expression of a cDNA coding for the anticoagulant hirudin from the bloodsucking leech, Hirudo medicinalis. Proc. Natl Acad. Sci. USA 83, 1084–1088 (1986).
Wysocka-Diller, J. W., Aisemberg, G. O., Baumgarten, M., Levine, M. & Macagno, E. R. Characterization of a homologue of bithorax-complex genes in the leech Hirudo medicinalis. Nature 341, 760–763 (1989).
Blackshaw, S. E., Babington, E. J., Emes, R. D., Malek, J. & Wang, W. Z. Identifying genes for neuron survival and axon outgrowth in Hirudo medicinalis. J. Anat. 204, 13–24 (2004).
Vergote, D. et al. Up-regulation of neurohemerythrin expression in the central nervous system of the medicinal leech, Hirudo medicinalis, following septic injury. J. Biol. Chem. 279, 43828–43837 (2004).
Vergote, D., Macagno, E. R., Salzet, M. & Sautière, P. E. Proteome modifications of the medicinal leech nervous system under bacterial challenge. Proteomics 6, 4817–4825 (2006).
Aisemberg, G., Kuhn, J. & Macagno, E. Netrin signal is produced in leech embryos by segmentally iterated sets of central neurons and longitudinal muscle cells. Dev. Genes Evol. 211, 589–596 (2001).
Macagno, E. R. et al. Construction of a medicinal leech transcriptome database and its application to the identification of leech homologs of neural and innate immune genes. BMC Genomics 11, 407 (2010).
Kandarian, B. et al. The medicinal leech genome encodes 21 innexin genes: Different combinations are expressed by identified central neurons. Dev. Genes Evol. 222, 29–44 (2012).
Hibsh, D., Schori, H., Efroni, S. & Shefi, O. Spatial regulation dominates gene function in the ganglia chain. Bioinformatics 30, 310–316 (2014).
Shanker, S. et al. Evaluation of commercially available RNA amplification kits for RNA sequencing using very low input amounts of total RNA. J. Biomol. Tech. 26, 1–15 (2015).
Salzet, M., Chopin, V., Baert, J. L., Matias, I. & Malecha, J. Theromin, a novel leech thrombin inhibitor. J. Biol. Chem. 275, 30774–30780 (2000).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Robertson, G. et al. De novo assembly and analysis of RNA-seq data. Nat. Methods 7, 909–912 (2010).
Kenny, N. J., Quah, S., Holland, P. W. H., Tobe, S. S. & Hui, J. H. L. How are comparative genomics and the study of microRNAs changing our views on arthropod endocrinology and adaptations to the environment? Gen. Comp. Endocrinol. 188, 16–22 (2013).
Dos Santos, G. et al. FlyBase: introduction of the Drosophila melanogaster Release 6 reference genome assembly and large-scale migration of genome annotations. Nucleic Acids Res. 43, D690–D697 (2015).
Andrews, S. FastQC: A quality control tool for high throughput sequence data. Babraham Bioinforma 1, http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Xie, Y. et al. SOAPdenovo-Trans: De novo transcriptome assembly with short RNA-Seq reads. Bioinformatics 30, 1660–1666 (2014).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Schulz, M. H., Zerbino, D. R., Vingron, M. & Birney, E. Oases: Robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28, 1086–1092 (2012).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Data Citations
Hibsh, D., Schori, H., Efroni, S., & Shefi, O. NCBI Sequence Read Archive SRP020225 (2013)
Hibsh, D., Schori, H., Efroni, S., & Shefi, O. GEO GSE45569 (2013)
Hibsh, D., Schori, H., Efroni, S., & Shefi, O. Figshare http://dx.doi.org/10.6084/m9.figshare.1289242 (2015)
Acknowledgements
The authors wish to thank Shahar Alon for fruitful discussions and invaluable advice, Helit Cohen for her help with experimentation. The Helobdella robusta and Capitella teleta sequence data were produced by the US Department of Energy Joint Genome Institute http://www.jgi.doe.gov/in collaboration with the user community. This work was supported (in part) by the EU-FP7 People IRG Marie Curie Grants (239482) (to O.S.) and by the Israel Science Foundation for Individual Research Grants (1403/11) (to O.S.).
Author information
Authors and Affiliations
Contributions
D.H., O.S. and S.E. are responsible for the concept of the research and writing the manuscript. H.S has performed the RNA extraction. D.H. responsible for database and data analyses, deposit the data; all authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Hibsh, D., Schori, H., Efroni, S. et al. De novo transcriptome assembly databases for the central nervous system of the medicinal leech. Sci Data 2, 150015 (2015). https://doi.org/10.1038/sdata.2015.15
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2015.15
This article is cited by
-
The genome of medicinal leech (Whitmania pigra) and comparative genomic study for exploration of bioactive ingredients
BMC Genomics (2022)
-
Draft genome sequences of Hirudo medicinalis and salivary transcriptome of three closely related medicinal leeches
BMC Genomics (2020)
-
Hirudins of the Asian medicinal leech, Hirudinaria manillensis: same same, but different
Parasitology Research (2019)
-
9-Phenanthrol modulates postinhibitory rebound and afterhyperpolarizing potentials in an excitatory motor neuron of the medicinal leech
Journal of Comparative Physiology A (2017)
-
More than just one: multiplicity of Hirudins and Hirudin-like Factors in the Medicinal Leech, Hirudo medicinalis
Molecular Genetics and Genomics (2016)