Abstract
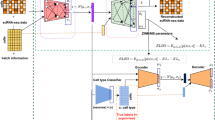
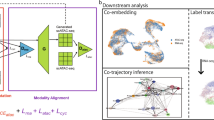
The rapid emergence of large-scale atlas-level single-cell RNA-seq datasets presents remarkable opportunities for broad and deep biological investigations through integrative analyses. However, harmonizing such datasets requires integration approaches to be not only computationally scalable, but also capable of preserving a wide range of fine-grained cell populations. We have created Portal, a unified framework of adversarial domain translation to learn harmonized representations of datasets. When compared to other state-of-the-art methods, Portal achieves better performance for preserving biological variation during integration, while achieving the integration of millions of cells, in minutes, with low memory consumption. We show that Portal is widely applicable to integrating datasets across different samples, platforms and data types. We also apply Portal to the integration of cross-species datasets with limited shared information among them, elucidating biological insights into the similarities and divergences in the spermatogenesis process among mouse, macaque and human.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data used in this work are publicly available from online sources as follows: for mouse brain cells from ref. 8 (http://dropviz.org), ref. 9 (http://mousebrain.org/downloads.html) and ref. 34 (GSE110823), the mouse cell atlas from the Tabula Muris Consortium7 (https://figshare.com/projects/Tabula_Muris_Transcriptomic_characterization_of_20_organs_and_tissues_from_Mus_musculus_at_single_cell_resolution/27733) and the mouse lemur cell atlas from the Tabula Microcebus Consortium31 (https://figshare.com/projects/Tabula_Microcebus/112227); for human PBMCs from ref. 39 (GSE156478), ref. 53 (GSE132044) and 10X Genomics35 (https://support.10xgenomics.com/single-cell-gene-expression/datasets/1.1.0/pbmc3k); for mouse spermatogenesis cells from ref. 45 (https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-6946/); for human spermatogenesis cells from ref. 42 (GSE142585); for macaque spermatogenesis cells from ref. 42 (GSE142585); for hematopoietic stem cells from ref. 54 (GSE72857) and ref. 55 (GSE81682); for reprogramming of induced pluripotent stem cells from ref. 56 (GSE122662); for human brain cells from ref. 36 (GSE164485) and ref. 37 (https://github.com/LieberInstitute/10xPilot_snRNAseq-human#work-with-the-data). Source data are provided with this paper.
Code availability
Portal software is available at https://github.com/YangLabHKUST/Portal. The codes for reproducing the results are available at https://github.com/jiazhao97/Portal-reproducibility. All codes are deposited in Zenodo repositories57,58.
References
Villani, A.-C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes and progenitors. Science 356, eaah4573 (2017).
Brbić, M. et al. MARS: discovering novel cell types across heterogeneous single-cell experiments. Nat. Methods 17, 1200–1206 (2020).
Iacono, G., Massoni-Badosa, R. & Heyn, H. Single-cell transcriptomics unveils gene regulatory network plasticity. Genome Biol. 20, 110 (2019).
Cuomo, A. S. E. et al. Single-cell RNA-sequencing of differentiating iPS cells reveals dynamic genetic effects on gene expression. Nat. Commun. 11, 810 (2020).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
The Tabula Muris Consortium. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030 (2018).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014 (2018).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Stegle, O., Teichmann, S. A. & Marioni, J. C. Computational and analytical challenges in single-cell transcriptomics. Nat. Rev. Genet. 16, 133–145 (2015).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Gao, C. et al. Iterative single-cell multi-omic integration using online learning. Nat. Biotechnol. 39, 1000–1007 (2021).
Hu, J., Chen, M. & Zhou, X. Effective and scalable single-cell data alignment with non-linear canonical correlation analysis. Nucleic Acids Res. 50, e21 (2022).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Polański, K. et al. BBKNN: fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2020).
Chazarra-Gil, R., van Dongen, S., Kiselev, V. & Hemberg, M. Flexible comparison of batch correction methods for single-cell RNA-seq using BatchBench. Nucleic Acids Res. 49, e42 (2021).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Goodfellow, I. et al. Generative adversarial nets. In Proc. Advances in Neural Information Processing Systems (eds. Ghahramani, Z. et al.) 2672–2680 (NIPS, 2014).
Zhu, J.-Y., Park, T., Isola, P. & Efros, A. A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proc. IEEE International Conference on Computer Vision 2223–2232 (IEEE, 2017).
Liu, M.-Y., Breuel, T. & Kautz, J. Unsupervised image-to-image translation networks. In Proc. Advances in Neural Information Processing Systems (eds. Guyon, I. et al.) 700–708 (NIPS, 2017).
Choi, Y. et al. StarGAN: unified generative adversarial networks for multi-domain image-to-image translation. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 8789–8797 (IEEE, 2018).
Büttner, M., Miao, Z., Wolf, F. A., Teichmann, S. A. & Theis, F. J. A test metric for assessing single-cell RNA-seq batch correction. Nat. Methods 16, 43–49 (2019).
Hubert, L. & Arabie, P. Comparing partitions. J. Classification 2, 193–218 (1985).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. Theory Exp. 2008, P10008 (2008).
Ezran, C. et al. Tabula Microcebus: a transcriptomic cell atlas of mouse lemur, an emerging primate model organism. Preprint at https://www.biorxiv.org/content/10.1101/2021.12.12.469460v1 (2021).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Selewa, A. et al. Systematic comparison of high-throughput single-cell and single-nucleus transcriptomes during cardiomyocyte differentiation. Sci. Rep. 10, 1535 (2020).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
3k peripheral blood mononuclear cells (PBMCs) from a healthy donor from 10X Genomics (10X Genomics); https://support.10xgenomics.com/single-cell-gene-expression/datasets/1.1.0/pbmc3k
Fullard, J. F. et al. Single-nucleus transcriptome analysis of human brain immune response in patients with severe COVID-19. Genome Med. 13, 118 (2021).
Tran, M. N. et al. Single-nucleus transcriptome analysis reveals cell-type-specific molecular signatures across reward circuitry in the human brain. Neuron 109, 3088–3103 (2021).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Mimitou, E. P. et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat. Biotechnol. 39, 1246–1258 (2021).
Lin, Y. et al. scJoint integrates atlas-scale single-cell RNA-seq and ATAC-seq data with transfer learning. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01161-6 (2022).
Cui, C., Zhou, Y. & Cui, Q. Defining the functional divergence of orthologous genes between human and mouse in the context of miRNA regulation. Brief. Bioinform. 22, bbab253 (2021).
Shami, A. N. et al. Single-cell RNA sequencing of human, macaque, and mouse testes uncovers conserved and divergent features of mammalian spermatogenesis. Dev. Cell 54, 529–547 (2020).
Green, C. D. et al. A comprehensive roadmap of murine spermatogenesis defined by single-cell RNA-seq. Dev. Cell 46, 651–667 (2018).
Hermann, B. P. et al. The mammalian spermatogenesis single-cell transcriptome, from spermatogonial stem cells to spermatids. Cell Rep. 25, 1650–1667 (2018).
Ernst, C., Eling, N., Martinez-Jimenez, C. P., Marioni, J. C. & Odom, D. T. Staged developmental mapping and X chromosome transcriptional dynamics during mouse spermatogenesis. Nat. Commun. 10, 1251 (2019).
Lau, X., Munusamy, P., Ng, M. J. & Sangrithi, M. Single-cell RNA sequencing of the cynomolgus macaque testis reveals conserved transcriptional profiles during mammalian spermatogenesis. Dev. Cell 54, 548–566 (2020).
Arjovsky, M. & Bottou, L. Towards principled methods for training generative adversarial networks. Preprint at https://arxiv.org/abs/1701.04862 (2017).
Bendall, S. C. et al. Single-cell trajectory detection uncovers progression and regulatory coordination in human B cell development. Cell 157, 714–725 (2014).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Powers, D. Evaluation: from precision, recall and F-measure to ROC, informedness, markedness and correlation. J. Mach. Learn. Technol. 2, 37–63 (2011).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
McInnes, L., Healy, J., Saul, N. & Grossberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Ding, J. et al. Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat. Biotechnol. 38, 737–746 (2020).
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015).
Nestorowa, S. et al. A single-cell resolution map of mouse hematopoietic stem and progenitor cell differentiation. Blood 128, e20–e31 (2016).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943 (2019).
Zhao, J. et al. Portal (Zenodo); https://doi.org/10.5281/zenodo.6467690
Zhao, J. et al. Portal-reproducibility (Zenodo); https://doi.org/10.5281/zenodo.6467711
Acknowledgements
We acknowledge grants as follows: Hong Kong Research Grant Council grants nos. 16307818, 16301419 and 16308120, Hong Kong University of Science and Technology’s startup grant no. R9405, Guangdong-Hong Kong-Macao Joint Laboratory grant no. 2020B1212030001 and the RGC Collaborative Research Fund grant no. C6021-19EF to C.Y.; Hong Kong Research Grant Council grant no. 16101118, Hong Kong University of Science and Technology’s startup grant no. R9364 and the Lo Ka Chung Foundation through the Hong Kong Epigenomics Project and the Chau Hoi Shuen Foundation to A.R.W.; the Hong Kong University of Science and Technology Big Data for Bio Intelligence Laboratory (BDBI), the Hong Kong University of Science and Technology Center for Aging Science Research Program to C.Y. and A.R.W.; Hong Kong Research Grant Council grants nos. 24301419 and 14301120, the Chinese University of Hong Kong’s startup grant no.4930181 to Z.L.; the Shanghai Sailing Program grant no. 21YF140600 to J.M.
Author information
Authors and Affiliations
Consortia
Contributions
J.Z. and G.W. conceived and developed the method. A.R.W. and C.Y. supervised the project. J.Z., G.W., Z.L., A.R.W. and C.Y. designed the experiments, performed the analyses and wrote the manuscript. J.M., Y.W. and T.M.C. provided critical feedback during the study and helped revise the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Mengjie Chen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Fernando Chirigati, in collaboration with the Nature Computational Science team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Source Data Fig. 2
Source data for Fig. 2.
Source Data Fig. 3
Source data for Fig. 3.
Source Data Fig. 4
Source data for Fig. 4.
Source Data Fig. 5
Source data for Fig. 5.
Source Data Fig. 6
Source data for Fig. 6.
Rights and permissions
About this article
Cite this article
Zhao, J., Wang, G., Ming, J. et al. Adversarial domain translation networks for integrating large-scale atlas-level single-cell datasets. Nat Comput Sci 2, 317–330 (2022). https://doi.org/10.1038/s43588-022-00251-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-022-00251-y
This article is cited by
-
An organism-wide atlas of hormonal signaling based on the mouse lemur single-cell transcriptome
Nature Communications (2024)
-
Cellcano: supervised cell type identification for single cell ATAC-seq data
Nature Communications (2023)
-
scBridge embraces cell heterogeneity in single-cell RNA-seq and ATAC-seq data integration
Nature Communications (2023)
-
Construction of a 3D whole organism spatial atlas by joint modelling of multiple slices with deep neural networks
Nature Machine Intelligence (2023)