Abstract
Computational modeling methods combined with non-invasive imaging technologies have exhibited great potential and unique opportunities to model new bone formation in scaffold tissue engineering, offering an effective alternate and viable complement to laborious and time-consuming in vivo studies. However, existing numerical approaches are still highly demanding computationally in such multiscale problems. To tackle this challenge, we propose a machine learning (ML)-based approach to predict bone ingrowth outcomes in bulk tissue scaffolds. The proposed in silico procedure is developed by correlating with a dedicated longitudinal (12-month) animal study on scaffold treatment of a major segmental defect in sheep tibia. Comparison of the ML-based time-dependent prediction of bone ingrowth with the conventional multilevel finite element (FE2) model demonstrates satisfactory accuracy and efficiency. The ML-based modeling approach provides an effective means for predicting in vivo bone tissue regeneration in a subject-specific scaffolding system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data are provided with this paper. Source data for Figs. 1 and 2, training data and raw/processed data required to reproduce these findings are available in a data repository in Zenodo (https://zenodo.org/record/5017032)65.
Code availability
The related code, neural networks and examples are available to academic researchers at public institutions from Zenodo (https://zenodo.org/record/5017032)65.
References
Hollister, S. J. Porous scaffold design for tissue engineering. Nat. Mater. 4, 518–524 (2005).
Petite, H. et al. Tissue-engineered bone regeneration. Nat. Biotechnol. 18, 959–963 (2000).
Choi, N. W. et al. Microfluidic scaffolds for tissue engineering. Nat. Mater. 6, 908–915 (2007).
Ringe, J. & Sittinger, M. Regenerative medicine: selecting the right biological scaffold for tissue engineering. Nat. Rev. Rheumatol. 10, 388–389 (2014).
Moutos, F. T., Freed, L. E. & Guilak, F. A biomimetic three-dimensional woven composite scaffold for functional tissue engineering of cartilage. Nat. Mater. 6, 162–167 (2007).
Schouman, T., Schmitt, M., Adam, C., Dubois, G. & Rouch, P. Influence of the overall stiffness of a load-bearing porous titanium implant on bone ingrowth in critical-size mandibular bone defects in sheep. J. Mech. Behav. Biomed. Mater. 59, 484–496 (2016).
Pobloth, A. M. et al. Mechanobiologically optimized 3D titanium-mesh scaffolds enhance bone regeneration in critical segmental defects in sheep. Sci. Transl. Med. 10, 8828 (2018).
Li, J. J. et al. A novel bone substitute with high bioactivity, strength, and porosity for repairing large and load‐bearing bone defects. Adv. Healthc. Mater. 8, 1801298 (2019).
Sharma, U. et al. The development of bioresorbable composite polymeric implants with high mechanical strength. Nat. Mater. 17, 96–102 (2018).
Entezari, A. et al. Architectural design of 3D printed scaffolds controls the volume and functionality of newly formed bone. Adv. Healthc. Mater. 8, 1801353 (2019).
Chen, Y., Zhou, S. & Li, Q. Microstructure design of biodegradable scaffold and its effect on tissue regeneration. Biomaterials 32, 5003–5014 (2011).
Chen, Y., Zhou, S. & Li, Q. Mathematical modeling of degradation for bulk-erosive polymers: applications in tissue engineering scaffolds and drug delivery systems. Acta Biomater. 7, 1140–1149 (2011).
Sturm, S., Zhou, S., Mai, Y. W. & Li, Q. On stiffness of scaffolds for bone tissue engineering—a numerical study. J. Biomech. 43, 1738–1744 (2010).
Adachi, T., Osako, Y., Tanaka, M., Hojo, M. & Hollister, S. J. Framework for optimal design of porous scaffold microstructure by computational simulation of bone regeneration. Biomaterials 27, 3964–3972 (2006).
Sanz-Herrera, J. A., García-Aznar, J. M. & Doblaré, M. On scaffold designing for bone regeneration: a computational multiscale approach. Acta Biomater. 5, 219–229 (2009).
Zhao, F., Melke, J., Ito, K., van Rietbergen, B. & Hofmann, S. A multiscale computational fluid dynamics approach to simulate the micro-fluidic environment within a tissue engineering scaffold with highly irregular pore geometry. Biomech. Model. Mechanobiol. 18, 1965–1977 (2019).
Marin, A. C., Grossi, T., Bianchi, E., Dubini, G. & Lacroix, D. µ-Particle tracking velocimetry and computational fluid dynamics study of cell seeding within a 3D porous scaffold. J. Mech. Behav. Biomed. Mater. 75, 463–469 (2017).
Kelly, D. J. & Prendergast, P. J. Mechano-regulation of stem cell differentiation and tissue regeneration in osteochondral defects. J. Biomech. 38, 1413–1422 (2005).
Huiskes, R., Van Driel, W. D., Prendergast, P. J. & Soballe, K. A biomechanical regulatory model for periprosthetic fibrous-tissue differentiation. J. Mater. Sci. Mater. Med. 8, 785–788 (1997).
Prendergast, P. J., Huiskes, R. & Søballe, K. Biophysical stimuli on cells during tissue differentiation at implant interfaces. J. Biomech. 30, 539–548 (1997).
Maslov, L. B. Mathematical model of bone regeneration in a porous implant. Mech. Compos. Mater. 53, 399–414 (2017).
Shi, Q., Shui, H., Chen, Q. & Li, Z. Y. How does mechanical stimulus affect the coupling process of the scaffold degradation and bone formation: an in silico approach. Comput. Biol. Med. 117, 103588 (2020).
Beaupré, G. S., Orr, T. E. & Carter, D. R. An approach for time‐dependent bone modeling and remodeling—theoretical development. J. Orthop. Res. 8, 651–661 (1990).
Sanz-Herrera, J. A., García-Aznar, J. M. & Doblaré, M. Micro-macro numerical modelling of bone regeneration in tissue engineering. Comput. Methods Appl. Mech. Eng. 197, 3092–3107 (2008).
Cheong, V. S., Fromme, P., Mumith, A., Coathup, M. J. & Blunn, G. W. Novel adaptive finite element algorithms to predict bone ingrowth in additive manufactured porous implants. J. Mech. Behav. Biomed. Mater. 87, 230–239 (2018).
Cheong, V. S., Fromme, P., Coathup, M. J., Mumith, A. & Blunn, G. W. Partial bone formation in additive manufactured porous implants reduces predicted stress and danger of fatigue failure. Ann. Biomed. Eng. 48, 502–514 (2020).
Taylor, M. & Prendergast, P. J. Four decades of finite element analysis of orthopaedic devices: where are we now and what are the opportunities? J. Biomech. 48, 767–778 (2015).
Checa, S. & Prendergast, P. J. A mechanobiological model for tissue differentiation that includes angiogenesis: a lattice-based modeling approach. Ann. Biomed. Eng. 37, 129–145 (2009).
Lecun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Nguyen, A. H. et al. Cardiac tissue engineering: state-of-the-art methods and outlook. J. Biol. Eng. 13, 57 (2019).
Kavakiotis, I. et al. Machine learning and data mining methods in diabetes research. Comput. Struct. Biotechnol. J. 15, 104–116 (2017).
Zhang, Y. & Ye, W. Deep learning–based inverse method for layout design. Struct. Multidiscip. Optim. 60, 527–536 (2019).
Alber, M. et al. Integrating machine learning and multiscale modeling—perspectives, challenges, and opportunities in the biological, biomedical, and behavioral sciences. npj Digit. Med. 2, 115 (2019).
Huang, D., Fuhg, J. N., Weißenfels, C. & Wriggers, P. A machine learning based plasticity model using proper orthogonal decomposition. Comput. Methods Appl. Mech. Eng. 365, 1–33 (2020).
Mozaffar, M. et al. Deep learning predicts path-dependent plasticity. Proc. Natl Acad. Sci. USA 116, 26414–26420 (2019).
Freiberg, A. H. Wolff’s law and the functional pathogenesis of deformity. Am. J. Med. Sci. 124, 956–971 (1902).
Lin, D., Li, Q., Li, W., Duckmanton, N. & Swain, M. Mandibular bone remodeling induced by dental implant. J. Biomech. 43, 287–293 (2010).
Lin, D., Li, Q., Li, W. & Swain, M. Dental implant induced bone remodeling and associated algorithms. J. Mech. Behav. Biomed. Mater. 2, 410–432 (2009).
Rungsiyakull, C. et al. Bone’s responses to different designs of implant-supported fixed partial dentures. Biomech. Model. Mechanobiol. 14, 403–411 (2015).
Weinans, H., Huiskes, R. & Grootenboer, H. J. Effects of material properties of femoral hip components on bone remodeling. J. Orthop. Res. 10, 845–853 (1992).
Liu, L., Shi, Q., Chen, Q. & Li, Z. Mathematical modeling of bone in-growth into undegradable porous periodic scaffolds under mechanical stimulus. J. Tissue Eng. 10, 204173141982716 (2019).
Feurer, M. et al. Efficient and robust automated machine learning. in Advances in Neural Information Processing Systems 28 (eds Ghahramani, Z. et al.) 2962–2970 (NIPS, 2015).
Snoek, J., Larochelle, H. & Adams, R. P. Practical Bayesian optimization of machine learning algorithms. Adv. Neural Inf. Process. Syst. 4, 2951–2959 (2012).
Perier-Metz, C., Duda, G. N. & Checa, S. Initial mechanical conditions within an optimized bone scaffold do not ensure bone regeneration – an in silico analysis. Biomech. Model. Mechanobiol. https://doi.org/10.1007/s10237-021-01472-2 (2021).
Cohen, D. O., Aboutaleb, S. M. G., Johnson, A. W. & Norato, J. A. Bone adaptation-driven design of periodic scaffolds. J. Mech. Des. Trans. ASME 143, 121701 (2021).
Göpferich, A. Polymer bulk erosion. Macromolecules 30, 2598–2604 (1997).
Shi, Q., Chen, Q., Pugno, N. & Li, Z. Y. Effect of rehabilitation exercise durations on the dynamic bone repair process by coupling polymer scaffold degradation and bone formation. Biomech. Model. Mechanobiol. 17, 763–775 (2018).
Wang, L. et al. Mechanical–chemical coupled modeling of bone regeneration within a biodegradable polymer scaffold loaded with VEGF. Biomech. Model. Mechanobiol. 19, 2285–2306 (2020).
Roohani-Esfahani, S.-I. I., Newman, P. & Zreiqat, H. Design and fabrication of 3D printed scaffolds with a mechanical strength comparable to cortical bone to repair large bone defects. Sci. Rep. 6, 19468 (2016).
Duda, G. N. et al. Analysis of inter-fragmentary movement as a function of musculoskeletal loading conditions in sheep. J. Biomech. 31, 201–210 (1997).
Guedes, J. & Kikuchi, N. Preprocessing and postprocessing for materials based on the homogenization method with adaptive finite element methods. Comput. Methods Appl. Mech. Eng. 83, 143–198 (1990).
Numerical experiments of the homogenization method. in Computing methods in applied sciences and engineering, 1977, I 330–356 (Springer, 1979).
Bendsøe, M. P. & Kikuchi, N. Generating optimal topologies in structural design using a homogenization method. Comput. Methods Appl. Mech. Eng. 71, 197–224 (1988).
Wu, C., Zheng, K., Fang, J., Steven, G. P. & Li, Q. Time-dependent topology optimization of bone plates considering bone remodeling. Comput. Methods Appl. Mech. Eng. 359, 112702 (2020).
Turner, C. H., Anne, V. & Pidaparti, R. M. V. A uniform strain criterion for trabecular bone adaptation: do continuum-level strain gradients drive adaptation? J. Biomech. 30, 555–563 (1997).
Checa, S., Prendergast, P. J. & Duda, G. N. Inter-species investigation of the mechano-regulation of bone healing: comparison of secondary bone healing in sheep and rat. J. Biomech. 44, 1237–1245 (2011).
Perier-Metz, C., Duda, G. N. & Checa, S. Mechano-Biological Computer Model of Scaffold-Supported Bone Regeneration: Effect of Bone Graft and Scaffold Structure on Large Bone Defect Tissue Patterning. Front. Bioeng. Biotechnol. 8, 585799 (2020).
Chen, G. et al. A new approach for assigning bone material properties from CT images into finite element models. J. Biomech. 43, 1011–1015 (2010).
Suquet, P. M. Elements of homogenization for inelastic solid mechanics, homogenization techniques for composite media. Lect. Notes Phys. 272, 193 (1985).
White, D. A., Arrighi, W. J., Kudo, J. & Watts, S. E. Multiscale topology optimization using neural network surrogate models. Comput. Methods Appl. Mech. Eng. 346, 1118–1135 (2019).
Funahashi, K. I. On the approximate realization of continuous mappings by neural networks. Neural Netw. 2, 183–192 (1989).
Hassoun, M. H. Fundamentals of artificial neural networks (MIT press, 1995).
Hogg, M. mhogg/pyvxray: an ABAQUS plug-in for the creation of virtual X-rays from 3D finite element bone/implant models (GitHub, 2013); https://github.com/mhogg/pyvxray
Pearson, K. Notes on regression and inheritance in the case of two parents. Proc. R. Soc. Lond. 58, 240–42 (1895).
Wu, C. Machine learning based multi-scale remodelling code (Zenodo, 2021); https://doi.org/10.5281/ZENODO.5017032
Acknowledgements
We acknowledge financial support from the Australian Research Council (ARC) through the Discovery (DP180104200, Q.L. and M.V.S.) and ARC Industrial Transformation Training Centre (IC170100022, Q.L. and H.Z.). The Artemis HPC provided by the Sydney Informatics Hub, a Core Research Facility of the University of Sydney, is acknowledged.
Author information
Authors and Affiliations
Contributions
C.W. and Q.L designed the research plan. C.W. performed the simulations. C.W., A.E., K.Z. and J.F analyzed the data. C.W., A.E. and Q.L. wrote the manuscript. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Computational Science thanks Jose A. Sanz-Herrera, Sara Esteban and Zhiyong Li for their contribution to the peer review of this work. Handling editor: Ananya Rastogi, in collaboration with the Nature Computational Science team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Figs. 1–3.
Source data
Source Data Fig. 4
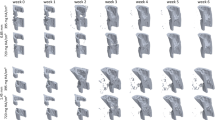
Original images for virtual X-ray images at months 3, 6, 9 and 12 and histological image at month 12.
Source Data Fig. 5
In vivo X-ray images taken at months 3, 6, 9 and 12.
Rights and permissions
About this article
Cite this article
Wu, C., Entezari, A., Zheng, K. et al. A machine learning-based multiscale model to predict bone formation in scaffolds. Nat Comput Sci 1, 532–541 (2021). https://doi.org/10.1038/s43588-021-00115-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-021-00115-x
This article is cited by
-
Machine Learning Approaches to 3D Models for Drug Screening
Biomedical Materials & Devices (2023)
-
Identification and ranking biomaterials for bone scaffolds using machine learning and PROMETHEE
Research on Biomedical Engineering (2023)
-
Predicting bone regeneration from machine learning
Nature Computational Science (2021)