Abstract
Coral reefs are severely threatened by global and local environmental changes. However, susceptibility to perturbations and subsequent mortality varies among coral species. In this study, we tested the contribution of genetic and environmental conditions to coral’s phenotypic response in Pocillopora spp. and Porites spp. sampled together at a large ecological and temporal scale throughout the Pacific Ocean. We assessed coral phenotype signatures using a multi-biomarker approach (animal and symbiont biomasses, protein carbonylation and ubiquitination and total antioxidant capacities). In both genera, we highlighted a strong anticorrelation between the redox state and the animal and symbiont biomasses. In addition, Pocillopora exhibited high phenotypic plasticity, responding to various environmental variables such as temperature, nutrients, phosphate, and carbonate chemistry. In contrast, Porites displayed more robust phenotypes influenced by both genetics and past climate events. In conclusion, co-located coral species display different phenotypic response strategies that are influenced by different environmental conditions.
Similar content being viewed by others
Introduction
Climate change is expected to significantly increase the occurrence of cold and warm ocean sea-surface temperature extremes1,2 that have already been well documented in the Pacific Ocean3,4. Accentuated by direct anthropogenic stressors due to local pollution and overfishing, climate change will have devastating consequences on marine organisms. Among them, reef-building corals are highly vulnerable to environmental changes, experiencing repeated mass coral bleaching (loss of the intracellular photosynthetic protist symbionts belonging to the Symbiodiniaceae family), which have already led to substantial coral mortalities over large spatial scales2.
However, susceptibility to environmental changes, and subsequent bleaching that may lead to mortality, vary among coral species5,6, but also within species7. Consequently, the surviving species will play a key role for coral reef resilience in a changing climate8. In these last decades, monitoring of reefs over the world has allowed to identify several species able to persist after perturbations and therefore considered today as winner species7,9,10. Among them, two groups of corals have been identified:11 (i) massive and long-living species (as Porites species) persisting during the environmental perturbations and considered highly stress-tolerant and (ii) ‘weedy’ species (as Pocillopora species), with fast growth and reproduction allowing a rapid recolonisation of the environment after stress. However, predictive models of global change forecast increasingly drastic and rapid climate changes that species have never experienced before and consequently make it difficult to predict who will be the future winners. It is therefore still imperative to know which coral species may have the adaptive potential and innate resilience to survive the disturbances of the Anthropocene for conservation and restoration purposes12.
The response of coral species to the environment has been attributed to different factors as, for example, the host genetic background13,14,15, specific holobiont composition (the combination of different host species with one or multiple Symbiodiniaceae and bacterial lineages)16,17,18,19, nutrient supply20,21,22, ocean deoxygenation23,24 and/or preconditioning to past repeated environmental changes25,26,27. However, the contribution of each of these factors to the coral susceptibility remains unclear, as the extent to which it is conserved within a coral genus, or within diverse Scleractinia corals living in sympatry in a broad range of environmental conditions.
To predict the ability of coral species to respond to stress, it is necessary to define the coral phenotype with pertinent traits. The search of relevant, sensitive and quantifiable biomarkers of coral traits remains a key goal28. The coral phenotypic response is an unconstrained notion mainly due to their intrinsically morphological plasticity linked to environment-induced skeleton changes29 and to their photoacclimation capacity that induced significant colour changes by modifying their cellular pigments and/or their symbiont population contents30,31. As corals can strongly depend on the high-energy products of their algal symbionts, those acclimation processes are essential for trophic equilibrium32. In addition, environmental perturbations (especially variations in temperature) induce in extreme cases bleached coral phenotypes which reflect symbiotic homeostasis breakdown33. To prevent bleaching, corals could then have (i) specific host-symbiont genotypes as specific adaptation to the local environment, or (ii) display a large spectrum of phenotypic plasticity. The latter allows the emergence of diverse physiological states from a unique genotype, in response to the environmental conditions12. Several studies performed under controlled laboratory or field conditions have identified biomarkers as indicators/proxy of coral physiological states. Among them, biomarkers of the trophic coral state, as polyp tissue thickness and/or the hosted symbiont density, have been linked to coral sensitivity to environmental perturbations and reflected the existing nutrient intake by the coral34,35,36,37. In addition indicators of intracellular redox state modification (as protein carbonylation and oxidative scavenging capacities) or more general damage signature as the protein ubiquitination, have been validated reflecting imbalances between reactive oxygen species overproduction and antioxidant defences during environmental stress38,39,40,41,42,43,44,45,46. Accordingly, one consequence of this imbalance was the disruption of the cnidarian–Symbiodiniaceae association. However, to what extent the diverse biomarker signatures reflect an inherited and genotypically fixed character or diverse coral response to specific environmental factors is not clear.
Hence, a large coral sampling was carried out, at an unprecedented ecological scale throughout the Pacific Ocean during the Tara Pacific expedition 2016–201847. In this study, we assessed the biological response to the environment from 321 colonies from 11 islands (across East-West and South-North gradients from the coasts of Panama to Micronesia) of two emblematic coral-species complexes with different morphologies and lifestyles, Pocillopora spp. and Porites spp.. Pocillopora species consist mostly of branched morphotypes, fast-growing and with a moderate lifespan of some decades11,48. Pocillopora species display flexibility towards Symbiodiniaceae, with the ability to harbour diverse species of Symbiodinium, Cladocopium or Durisdinium genera49,50,51,52,53,54. In contrast, even if they are found in sympatry with Pocillopora, Porites species display contrasted characteristics. Porites species consist mostly of massive corals, with slow growth and long lifespan (several centuries)48,55, with a strict symbiont fidelity to the genus Cladocopium, specifically with subclade C15 in the Indo-Pacific18,52.
In this study, we questioned whether visually healthy colonies, from two reef-building corals, co-located in the same wide range of habitats, showed common or divergent phenotypic signatures in response to identical environmental variations. Our first objective was to test the ability of quantitative and reliable biomarkers to define different physiological states for healthy corals, and subsequently establish interpretive coral phenotypic signatures. We used multiple physiological biomarkers at a macroscopic scale (quantification of animal and symbiont biomasses to assess polyp tissue thickness and symbiont density) and at a cellular scale (evaluation of the redox state by quantifying protein carbonylation, protein ubiquitination and total antioxidant capacities). Our second objective was to test, by a comparative approach between the two coral genera, to what extent these diverse physiological traits reflect genetic relatedness (in the coral host and its symbionts) and/or environmental correlations. For this purpose, a holistic integration of those signatures with putative explanatory parameters (as host, symbiont and microbiome diversities and environmental variables), collected during the Tara Pacific expedition, was performed.
Results
Distinct coral biomarker profiles across the Pacific Ocean
To obtain a physiological/phenotype profiling of Porites and Pocillopora colonies collected in reefs of different Pacific Ocean islands, we measured five different biomarkers revealing diverse trophic (animal and symbiont biomasses) and redox states (protein carbonylation, ubiquitination and Total Oxidant Scavenging Capacity—TOSC). The comparison of the different biomarkers across the 11 islands and within the two genera, revealed a heterogeneous pattern both among islands and among species (Figs. 1 and 2).
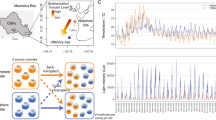
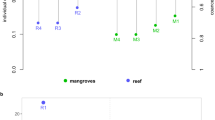
Variability of the biomarkers measured in Pocillopora (purple-pink) and Porites (cyan) genera and in function of sampled islands. a Animal biomass assay. b Symbiont biomass assay (normalised by mg of protein). c Protein carbonylation. d Protein ubiquitination. e Antioxidant capacity. The boxes represent the 1st and the 3rd quartiles around the median (horizontal black bar). Outlier points are plotted individually (dots). Statistically significant differences among the islands within the two genera have been tested by a Kruskall–Wallis’ test and a Dunn’s test as post hoc. The identical letter above boxplots indicates no statistically significant differences. NA: not applicable as no Pocillopora samples were available for Malpelo. Island coordinates and dates are indicated in Supplementary Table S8.
Variability of the biomarkers measured among species within Pocillopora (purple-pink) and within Porites (cyan) genera (a) Animal biomass assay. b Symbiont biomass assay (normalised by mg of protein). c Protein carbonylation. d Protein ubiquitination. e Antioxidant capacity. The coral-species attribution is detailed in Table 3. The boxes represent the 1st and the 3rd quartiles around the median (horizontal black bar). Outlier points are plotted individually (dots). Statistically significant differences among the species within the two genera have been tested by a Kruskall–Wallis’ test and a Dunn’s test as post hoc. Identical letters above boxplots indicate no statistically significant differences. Species distributions and lineages are indicated in Supplementary Fig. S6 and Table 3.
For Pocillopora, animal biomass was high in the first three islands and then declined to a minimal mean in Moorea Island which presented around three times less proteins compared to eastern colonies or to Guam (Fig. 1a). For Porites, we observed an eightfold reduction comparing colonies from Las Perlas to Guam colonies (Fig. 1a). Thanks to the identification of the coral species56,57 (five species within Pocillopora and three species in Porites), we could assess the species specificity of this phenotype within each genus. Figure 2a shows that despite some statistically significant differences among Pocillopora species, no species-specific animal biomass signature was identified. In contrast, all three Porites species identified in our sample showed a statistically significant species-specific pattern for animal biomass.
Symbiont biomass normalised per milligram of protein was highest in Eastern Pacific for Pocillopora colonies (specifically for colonies from Panama islands, Las Perlas and Coiba) and lowest in Niue and Samoa islands (10- to 20-fold lower, respectively; Fig. 1b). Porites colonies harboured a more homogenous symbiont biomass even if Niue showed a lower symbiont content (Fig. 1b). Very similar trends were observed following a tissue surface normalisation (Supplementary Fig. S1a). The comparison of symbiont biomass in respect to host lineages showed a higher symbiont content of Pocillopora species 4 (highly frequent in the Eastern Pacific) and in Porites species 3 (present in the Central Pacific) (Fig. 2b and Supplementary Fig. S1b). Due to the strong correlation measured between the two symbiont biomass datasets (Supplementary Fig. S2), we hence conserved only symbiont biomass normalised per milligram of protein for the following analyses.
For both genera, carbonyl content was higher in the three central islands of the transect, Moorea, Aitutaki and Niue (Fig. 1c). For example, in these three islands, the coral colonies of Pocillopora and Porites respectively presented carbonyl content means of two- to eightfold higher than the colonies from the East Pacific Ocean. The comparison of carbonyl content in respect to host lineages (Fig. 2c) showed that despite differences among the Pocillopora species, no species-specific response on protein carbonyl was measured. Contrastingly, the Porites species 2 (present in Central Pacific) had higher carbonyl content than the other two Porites species.
With regard to the protein ubiquitination biomarker, the analyses showed a high specific content of ubiquitinated proteins in Pocillopora from Gambier, with a median more than three-time higher than elsewhere and with a statistically significant variation among the sampled colonies (Fig. 1d). Although less pronounced, a low specific amount of ubiquitinated proteins was also observed in Porites colonies from Niue, Samoa and Guam (Fig. 1d). No species specificity was observed for this marker within either genera (Fig. 2d).
For Pocillopora, the colonies from the two Panama, Easter and Guam islands had the lowest TOSC values. Contrastingly, Aitutaki colonies harboured maximal TOSC values, with around 20 times more antioxidant capacities (Fig. 1e). For Porites, TOSC values were more homogenous with a slight increase along the transect line (Guam colonies presented a sevenfold increase of TOSC compared to Las Perlas). The comparison of TOSC in respect to host lineages revealed a species-specific reduced amount of TOSC on Pocillopora species 4 and Porites species 1 (Fig. 2e), although this species was only found in Eastern Pacific islands.
Island- and species-specific coral phenotypic signatures
We then used a multimarker approach using the five biomarkers to establish phenotype signatures for each colony and compared them in respect to the genus, species and islands of origin. A first PERMANOVA analysis testing the genus affiliation shows different phenotypic profiles (P value = 0.0001) with greater phenotypic dispersion of Pocillopora phenotype than Porites (P value = 0.001) visible in Supplementary Table S1 and Supplementary Fig. S3. We have therefore analysed more in detail the Pocillopora and Porites signatures independently, using principal component analysis (PCA) and PERMANOVA tests. The PCA of Pocillopora biomarker signatures revealed the gradual distribution of the phenotypes along the first axis (Fig. 3a), mostly driven by both biomasses and carbonyls (Fig. 3b and Supplementary Table S2). For Porites, the phenotypes were less spread into the PCA space but dispersed more homogeneously along the two first axes (Fig. 3c), due to a stronger independence between animal biomass and symbiont biomass (Fig. 3d and Supplementary Table S2) than in Pocillopora.
PCA on biomarkers data from Pocillopora spp. a, b and Porites spp. c, d The dispersion of the phenotypic signatures (defined by the five measured biomarkers) are shown in (a, c), where the colours represent the island origin, and the shapes represent the species. The importance of each biomarker explaining the PCA distribution of data is shown with correlation circles in (c, d), where the colour scale is the contribution of each biomarker to the plotted PCA dimensions.
The statistical partition of the phenotypic signatures by geographic origin or by coral species has been tested for both genera by PERMANOVA (Table 1 and Fig. 4). For Pocillopora, the colonies from the eastern and the western islands displayed statistically different phenotypes (Fig. 4 and Supplementary Table S3). Pocillopora colonies from Las Perlas and Coiba had a strong phenotypic differentiation with high biomasses and low redox state. In addition, the results show some phenotypic dissimilarity among Pocillopora species: with the often sympatric species 4 and 5 different from species 1 and from the other sympatric pair the species 2 and 3 (Fig. 4 and Supplementary Table S4). The Porites phenotypes were less structured by geographical regions, but few islands appeared as one-off pairwise differentiation (as Niue or Coiba; Fig. 4 and Supplementary Table S5). Only Porites species 3 had statistically different phenotypes from the two other species (Fig. 4 and Supplementary Table S6).
The phenotypic signature of Pocillopora (purple-pink) and Porites (cyan) colonies by biomarker is represented by scaled average values (from grey: the minimum value; to genus colour: the maximum value) used here as an illustrative barcode-like phenotypic signature. The letters associated to the barcodes are the groups of statistical significance based on pairwise PERMANOVAs (Supplementary Tables S3 and S5 for island comparisons; Supplementary Tables S4 and S6 for species comparisons within Pocillopora and Porites genera, respectively). The same letter indicates no statistically significant difference.
To test the influence, on the global phenotype signature, of the island or species origins but also the associated microorganism diversity (testing separately the Symbiodiniaceae and the microbiome diversities), we performed a variation partition analysis. To avoid any bias, we considered only coral colonies from islands containing more than one coral species per genus. The analysis confirmed a main effect of the geographical origin on the variance of the diverse phenotypic signature for Pocillopora (Fig. 5a) and Porites (Fig. 5b). For Porites, the coral species seemed to be an additional explanatory factor. A complementary Mantel’s test performed within each species did not reveal any correlation between the genetic distance and the coral phenotype (Supplementary Fig. S4). On the other hand, the associated microorganism diversity was a marginal driver of the coral phenotypes, compared to the variance explained by the partition into islands.
Proportion of the variance explained by the different partitions of the Pocillopora (a) and Porites (b) phenotypic signatures (defined by the five measured biomarkers): island of origin (black), host species (grey), Symbiodiniaceae composition (light grey) or the microbiome composition (white). The white boxes represent the 1st and the 3rd quartiles around the median (horizontal black bar). Outlier points are plotted individually (black dots). The shaded surface of each violin displays the distribution of the data.
Differing coral sensitivities to environmental conditions
Because the phenotypes of both genera were demonstrated to be influenced mainly by the geographical origin of the corals, we identified putative drivers, by exploring the correlations between biomarkers and several environmental variables, collected during the Tara Pacific expedition or recovered by historical temperature data extracted from satellite imagery (see details in material and methods). Using sPLS (sparse Partial Least Squares method), we identified clusters based on similar correlations between biomarker data and environmental variables, and found that the two coral genera did not show the same clustering of these data.
For Pocillopora, the 20 variables most correlated with the studied biomarkers were included in all environmental categories (for details, see 'Methods') and structured in two clusters (Fig. 6a and Table 2). Among the first cluster, characterised by strong correlation with antioxidant capacity and protein carbonylation, we identified three biological variables (C_Bio_086, C_Bio_092, C_Bio_094; corresponding respectively to high content on chlorophyll, nanoplankton and total Eukaryotes analysed in the seawater); five physico-chemical variables measured the sampling days (C_Phy_081, C_Phy_090, C_Phy_099, C_Phy_100 and C_Phy_101; corresponding respectively to high oxygen, turbidity, pH, silica content and sun exposure duration, Table 2); three climate variables (the sea surface temperature of the sampling day, recent cold temperature anomaly and past cold sea surface temperature anomalies, respectively C_Cli_084, R_Cli_045 and H_Cli_009). Among the second cluster, characterised by high biomasses and low redox state, we identified one dynamics variable (the sea surface height; C_Dyn_077), three contemporary physico-chemistry variables (C_Phy_083, C_Phy_096, C_Phy_098; corresponding respectively to high carbon dioxide content, total alkalinity, phosphate concentration) and three historical climate variables (H_Cli_013, H_Cli_015, H_Cli_029, H_Cli_071; corresponding to high intensity of sea surface temperature deviations, recent heatwaves episodes and short recovery time after recent cold waves). In Pocillopora, all those variables displayed strong correlation (between −0.66 and 0.57) with the biomarkers, except for protein ubiquitination, whose correlations never reached the 0.3 fixed threshold. For all correlated environmental variables, two main biomarker clusters were also observed showing anti-correlation between TOSC and carbonyl content biomarkers versus animal and symbiont biomass biomarkers.
Correlations among Pocillopora (a) or Porites (b) biomarkers and the environment data are represented with a classification of the environmental data by grey shades and with 3-letters code (from darker to lighter: Biological data (Bio); Hydro- and Aerodynamics data (Dyn); Physico-chemical data (Phy); and Climatic data (Cli)). In addition, environmental data are also identified as historical (H), recent (R) and contemporary to the sampling (C). Finally, an interpretation of these environmental variables is proposed in Table 2, and displayed by a symbol code present on the bottom of the figure. The corresponding original variable names are available in Supplementary Table S9. The correlation coefficient values are given by the central colour key, and only biomarkers or environmental variables with an absolute coefficient value higher than 0.3 are displayed (leaving the lower values blank). In the top, left and right are represented by pie charts of the proportion of environmental variables correlated with at least one biomarker for Pocillopora and Porites, respectively.
On the other hand for Porites, the 34 most correlated environmental variables with the biomarkers, belonged mainly on historical climatological variables and were more weakly correlated (around −0.55 and 0.5, Fig. 6b) than what we observed in Pocillopora. In Porites, the most correlated variables were clustered in three groups (Fig. 6b and Table 2). A first cluster grouped the sea surface height (C_Dyn_077) and historical seawater temperature anomalies linked to minimal recorded seawater temperature (H_Cli_001), cold waves (in frequency and duration, respectively H_Cli_060, H_Cli_065) but also to the recovery time after heat wave (H_Cli_043, H_Cli_044). In the second cluster, we identified biomarker correlations with three physico-chemistry variables (corresponding to high oxygen, turbidity and sun exposure duration; C_Phy_081, C_Phy_090, and C_Phy_101) and 11 historical climate variables depicting recurrent past and recent seawater cold temperature anomalies (H_Cli_047, H_Cli_048, R_Cli_049, H_Cli_052, H_Cli_073) and strong past fluctuation of seawater temperatures (H_Cli_004, H_Cli_012, H_Cli_013, H_Cli_014, H_Cli_015, H_Cli_019). Finally, a third cluster grouped only climate variables linked with past and recent heatwaves events (H_Cli_020, H_Cli_021, H_Cli_025, H_Cli_026, H_Cli_027, H_Cli_028, R_Cli_029, H_Cli_030, H_Cli_031, H_Cli_032, H_Cli_033, H_Cli_035, H_Cli_037, H_Cli_039). In Porites, we also observed anti-correlation between TOSC and carbonyl content biomarkers versus animal biomass. However, contrasting with Pocillopora results, animal and symbiont biomasses did not correlate in the same way with the variables. In addition, even when the protein ubiquitination content of Porites passed the sPLS filters and showed correlations with the environment variables, the correlation values were far weaker.
Although the intensity of the correlations between biomarkers and environmental variables and the type of environmental variables mostly depended on the genus, 7 environmental variables were common to Pocillopora and Porites (Fig. 7a) and concerned the sun exposition duration (C_Phy_101) and O2 concentration (C_Phy_081), frequency of recent (R_Cli_029) and historical sea temperature anomalies (H_Cli_013, H_Cli_015), the sea surface height (C_Dyn_077), and the turbidity (C_Phy_090). Comparing the correlation pattern of these biomarkers (Fig. 7b–e and Supplementary Table S7), the animal biomass (Fig. 7b) appeared to be the biomarker that responded most similarly in the analysed Pocillopora and Porites.
Number of common and private environmental variables with a correlation coefficient higher than 0.3 for at least one biomarker in Pocillopora (purple-pink) or Porites (cyan) (a). Details of the correlations between Pocillopora and Porites’ biomarkers with common environmental variables: animal biomass (b), symbiont biomass (c), protein carbonylation (d), and antioxidant capacity (e). The symbols on the left codes indicate the interpretation of the environmental variables also presented in Table 2 and Fig. 6. The flame points out warm anomaly (R_Cli_029: TSA_DHW_freq_snapshot_sampling_day_day), the snowflake points out temperature anomaly frequencies (H_Cli_013: SST_anomaly_freq_max_day, H_Cli_015: SST_anomaly_freq_std_day), fork & knife point out nutrition proxy variables (C_Phy_081: o2_copernicus, C_Phy_090: KD490_Sat, C_Phy_101: Sun),, and the waves point out an upwelling proxy variable (C_Dyn_077: ssh_copernicus).
Discussion
Characterising multiple biomarker variations in the widespread and co-occurring Pocillopora and Porites species in the Pacific Ocean, we identified diverse phenotypes, reflecting discrete physiological states. In addition, comparing their phenotypic signatures, we highlighted two different strategies of environmental response that could be synthesised as 'the Oak and the Reed' strategies. Indeed, those two genera respond to the environment by a contrasting gradient of phenotypic plasticity. Porites exhibited more robust phenotypes (with limited changes in physiological traits) mainly influenced by past climate events while Pocillopora harboured a greater phenotypic plasticity responding to a large variety of environmental variables.
In this study, we validated the use of five different biomarkers, never measured in combination at such large species and geographic scales, to determine the phenotypic profiling of coral colonies. The analysis of the diverse phenotypes corroborated in both genera a significant and strong anti-correlation between the animal biomass, and to a lesser extent the symbiont biomass, with the redox state. A coral with thick tissue and high symbiont density hence presents a low redox state with reduced oxidative damage and reduced activation of antioxidant defences.
Previous studies have observed that variations of coral tissue thickness and symbiont density could be as well species-specific, related to seasonality, geography or environmental perturbations34,35,36,37,58,59 and/or considered to be a proxy of the trophic state of the colony58. In addition, high tissue thickness would be characteristic of stress-tolerant species with high survival rates after bleaching7,60. Taking into account those considerations, our results from a large geographic and species scale show that coral colonies with high animal and symbiont biomasses present a resistant phenotype, less susceptible to stressors.
The redox baseline of corals can reflect host or symbiont specificities61,62 and/or environmental perturbations that are responsible for reactive oxygen species overproduction in coral tissues, which in turn activates antioxidant systems preventing cellular damages63. The extended ability of corals to activate antioxidant capacities and/or to eliminate oxidised cellular compounds can then be considered as a crucial point that allows corals to respond to the environment64. Associated with high biomasses, a low redox state measured in the sampled colonies could then reflect a healthy physiological status not challenged by any exogenous stressors.
The contribution of the protein ubiquitination biomarker is less resolutive at the scale of the entire transect. In corals, protein ubiquitination level has been linked to thermal stress in the field38,39 or laboratory-controlled conditions43,65,66 and associated to tissue thickness decrease, bleaching process and high redox state. The incongruency here observed is however not surprising since the ubiquitination profile could also be linked not only to damaged proteins degradation under stress conditions but also to other cellular processes in normal physiological conditions. Therefore, the ubiquitination profile remains more difficult to decipher. It is however interesting to note that in Gambier and Ducie, the coral colonies harbouring high protein ubiquitination presented concomitantly some of the lowest bacterial diversity67. This observation suggests that the presence of highly ubiquitinated proteins in corals would be an indicator of weakness of the colonies by a general bacterial dysbiosis and a signal of immune response68,69. Assuredly, our work points out that further efforts should be made to identify the drivers that lead to the diverse ubiquitination signature in order to gain a better insight into the biochemical coral response.
In our study, we observed phenotypes significantly linked to the island of origin for Pocillopora spp. and for the island of origin and the species for Porites spp. The analysis of Pocillopora samples showed that the analysed Eastern Pacific Ocean colonies (from Las Perlas and Coiba) had a strong phenotypic differentiation with high biomasses and low redox state, in contrast with the Central Pacific ones (especially Niue and Samoa), which harboured low biomasses and high redox state (Figs. 1 and 4). In the other islands, the corals harboured intermediate phenotypes defined by diverse biomarker signatures. As all Las Perlas and Coiba colonies belonged to species 4 (identified as P. grandis synonym of P. eydouxi57; Table 3), it is tempting to attribute a species-specific phenotype to P. grandis as a result of environmental selection pressure experienced in this area. On the other hand, P. meandrina and P. verrucosa (species 2 and 3, respectively, Table 3) displayed undifferentiated phenotypes wherever they have been sampled. The absence of a genetic link in Pocillopora spp. is also confirmed by the insignificant explanation of variance by species assignment (Fig. 5a) and the lack of phenotype–genetic correlation (Supplementary Fig. S3).
As the coral sampling has been performed during the Tara Pacific expedition within a 6-month frame, it is important to point the impact of time of sampling as a confounding factor. From July 2016 (Panama Islands) to January 2017 (Guam), the expedition spanned several seasons, making it difficult to tease apart the seasonal from the geographical effect. However, if we restrict our comparison of island phenotypes to islands sampled on a reduced time scale, we still find statistically significant differences. For example, the phenotypes of the Pocillopora colonies of Moorea and Aitutaki were different from those of Niue and Samoa (Fig. 4) even though sampling was carried out in less than a month within the same season (between November 6th and December 3rd, Supplementary Table S8). On the other hand, the similarity of Easter Pocillopora phenotypes to the ones from Aitutaki was also observed over different months (September to November) which correspond to seasonal changes (from winter to spring). Nevertheless, to definitively unravel the impact of the seasons on an expedition involving the collection of samples in time and space, it will be necessary in the future to carry out additional studies resampling the same colonies at different times (seasons and years) and also to have a larger geographical sampling with more seasonal replicates.
In conclusion, the environmental characteristics of each island seem to be the main factors structuring the Pocillopora physiological state with an extended phenotypic plasticity reflecting the diversity of the colonised habitats. This result is in accordance with the definition of Pocillopora as a weedy or competitive group of species11,70,71 and allows the comparison to be made in terrestrial habitat with the reed lifestyle. Weedy species are indeed characterised by small morphotypes with fast growth and short lifespan. They fit in the r-selection strategy in which rapid reproduction allows to colonise rapidly a large variety of environments. As we observed in most of the Pocillopora species here studied, weedy corals are also characterised by a large phenotypic plasticity which can provide acclimatisation potential and represent intrinsic advantages for their survival to future environmental changes. In addition, high phenotypic plasticity can stimulate diversification and speciation by allowing adaptation to diverse conditions72. This interpretation is in line with the finding that species differentiation is linked to past temperature variation in Pocillopora57.
Our work depicts a complex structuring of the physiological status in Porites, shaped not only by the environment but also by a genetic component. Porites spp. assuredly present less diverse phenotypes, but some phenotypic peculiarities have been found along the transect. Among them, as observed in Pocillopora, the Las Perlas Porites colonies displayed a physiological status with high animal biomass and low redox state. Again, this specific phenotype is not diagnostic of the unique genetic lineage sampled in Las Perlas (species 1 identified as P. evermanni, Table 3), as in Malpelo the same lineage displayed different phenotypic signatures. Finally, a strong phenotypic species specificity has been found in species 3 (SSH3_plob, Table 3), widely distributed from Easter Island to Guam, which demonstrated strong phenotypic homeostasis independent of habitat diversity. Limited range of phenotypic plasticity, slow growth and longer generation time is in accordance with the definition of Porites as a genus of stress-tolerant species11.
While Pocillopora and Porites species show different strategies of environmental response, we observed in some islands some convergence in phenotypes. More specifically, we measured in both genera high biomasses and low redox states in Las Perlas and Coiba islands. These samplings were performed in the two largest marine protected areas of Panama (the Coiba National Park and the Archipiélago de Las Perla), both inscribed from 2004 in the UNESCO World Heritage List of Natural Sites. Among the reasons for this protection, the presence of extreme environmental disturbances, from short-term variability in sea temperature and seasonal upwelling to long-term temperature excursions was considered73. The convergent phenotypes could then be attributed to the success of the ecosystem protection and/or to the result of the natural selection of less susceptible corals due to repetitive perturbation episodes, suggesting that this specific physiological status (high biomasses and low redox state) is a trait of low susceptibility (or high capacity for survival) to local perturbations. It is interesting to note that the sampling in the Panama island has been performed during the wetter and warmer season (July 2016) with poorly nutrient-rich seawater74. In this environmental context, corals tend to have low tissue thickness (reflecting low nutrient reserves) and a seasonal decrease in symbiont density75. The observation of a completely opposite phenotype reinforces our conclusion about the particularly pristine state of the Panama colonies.
Pocillopora colonies, and to a lesser extent Porites colonies sampled in Niue and Upolu (Samoa Archipelago) islands, showed phenotypic specificities. More precisely, they accumulated low biomasses and a high redox state, suggesting a fragile physiological state. Few data are available concerning the health status of these coral reefs, but recent reports underlined low to extremely low coral coverage (down to <1% in some Upolu sites) with coral communities that remain under stress from recurrent storms and coral bleaching76,77. In the light of climate predictions for the coming years, our observations could then constitute a solid argument to reinforce coral conservation in suffering reefs. Indeed, stronger marine protected area policy, as set up in Las Perlas and Coiba, can reduce the direct anthropogenic pressure, mitigating the impact of the more difficult to control climate change.
In both coral genera, Symbiodiniaceae and bacterial compositions were not detected as explanatory factors of the diverse phenotypes. However, previous studies focused on some coral populations and reefs have highlighted that low bacterial diversity may be indicative of a less resilient status of the reef68. It is plausible that the discrepancy with our findings stem from our field approach, as the analysis of colonies under natural and non-extreme conditions reveals various stable physiological states well adapted to their environment and therefore without any signs of dysbiosis (excepted for few highly ubiquitinated colonies as discussed previously).
Furthermore, enrichment with specific thermotolerant Symbiodiniaceae (i.e., Durusdinium spp.)61,78,79 in some Pocillopora colonies from the most warm equatorial waters (such as Panama and Guam) did not confer a unique phenotype. For example, two hypotheses on the Pocillopora-Durusdinium couple could then be made: (i) this couple can harbour diverse warm adapted/acclimatised phenotypes; (ii) this couple in warm water cannot be systematically an acclimatisation sign of the holobiont but reflect the opportunistic capacity of Durusdinium to proliferate in warm water. The latter hypothesis is supported by experiments on symbiotic cnidarian models which have shown that hosts harbouring Durusdinium can be unproductive and highly stressed80.
Our results, summed up in Table 2, show that the characteristics of the environment experienced by the colonies at the time of sampling, but also the past climatic events, shape the coral phenotypes of both genera. Nevertheless, Pocillopora phenotypes were linked to a large diversity of contemporary environmental factors including biological, physico-chemical, dynamic and climate variables, while Porites was mainly only linked to the past climate. Extreme past and recent cold anomalies (in intensity, duration and frequency) seem to fragilize Pocillopora and Porites phenotypes that systematically presented low biomass and high redox state (Table 2). Even if, in the context of global ocean warming, the impact of heatwaves on coral reefs was strongly documented, several studies highlighted the impact of severe cold waves on corals, demonstrating an even stronger effect on modulation of gene expressions81 on bleaching or mortality susceptibility82. The strong common correlation with sea surface height can also be linked to strong cold upwelled waves reaching the reefs that can increase coral stress susceptibility83.
In contrast, the frequency of recent or past sea surface temperature anomalies (including heat and cold waves) was among the strongest explanatory factors that correlated equally in both genera with almost all biomarkers and leading to colonies with low redox state and high biomasses. This link was more expected as the analysed biomarkers have been shown to be sensitive to temperature increases in the laboratory or in the field. It can however reinforce the statement that recurrent exposures to sea surface temperature anomalies play an important role in coral acclimatisation84, reducing the cost of stress response (i.e., low redox state) and enhancing coral tissue reserves (i.e., high animal biomass). Our results were also in accordance with the conclusions of ref. 70 that stressed the importance of the past climate and especially repeated marine heat and cold waves in the sensitivity of massive corals, like Poritidae25. This hormesis phenomenon can also explain the overall more stable phenotypes observed in Porites spp.. On a different note, this behaviour could also be a resultant of their long-life strategy, as environmental selection pressures could have shaped Porites spp. to deal with frequent environmental fluctuations without acting on their sensitivity to extreme events.
In addition, Porites spp. and Pocillopora spp. shared similar responses to turbidity, light exposure duration and oxygen (Fig. 7 and Table 2). The positive correlation between those factors and the animal biomass low redox state revealed an important weight of the nutrient-enriched water in the trophic state of the corals. Several studies have demonstrated that naturally turbid coral reefs exhibit high coral cover with fast-growing coral species85,86,87. Indeed, an increase of suspended particulate matter and light exposure can very likely favour the nutrition state of the corals, thus enhancing their mixotrophic nutrition. This is even more notable in Pocillopora phenotypes that are strongly and positively linked with the seawater richness in zoo- and phytoplankton, suggesting that Pocillopora spp. are able to adjust the autotrophic/heterotrophic balance in response to the environment variability with the capacity to maintain or even increase their energy reserves. In addition, some studies have already linked large polyp phenotypes with increased dependence on heterotrophy and stress resistance88,89,90,91. We can therefore suggest that this capacity of trophic modulation could explain the extended distribution of Pocillopora genus in the Pacific Ocean and its stress resistance.
Finally, our work highlighted the sensitivity of Pocillopora colonies to phosphate and acidification. Indeed, in our data, phosphate accumulation in the environment was associated with high redox state and small polyps with low symbiont density confirming the sensitivity of Pocillopora physiology to an imbalance in nutrient level92. This is in accordance with previous studies performed in other coral species, showing that phosphate contamination (mainly due to eutrophication through the addition of fertilisers) negatively affects coral traits as skeleton production, reproduction and symbiont photosynthesis93 but can also provoke increase in DNA damages and lipid peroxidation94. Finally, carbonate chemistry also exhibited a link with the Pocillopora phenotypes harbouring high redox state and low biomass. Branched corals, such as the Pocilloporidae, are very sensitive to acidification that provokes a decrease in growth and/or increase in skeletal porosity95 but also oxidative stress response96,97.
Conclusions
In this work, by comparing the phenotypic signatures of the widespread Pocillopora and Porites genera, we confirmed that those two genera present two different strategies of environmental response that we have synthesised as 'the Oak and the Reed' strategies. This terrestrial and poetic metaphor helps to illustrate the diverse phenotypic plasticities and sensitivities to environmental factors observed in the genera here studied and their implication to future environmental changes. The reduced plasticity quantified in Porites species is consistent with stress tolerance in a wide range of habitats through a strong investment in homeostasis regulation. However, this robustness can potentially limit, as in the case of an oak tree, their rapid response to drastic and rapid environmental changes. In contrast, under severe environmental changes, Pocillopora species would have, like a reed, an extended ability to rapidly modify its physiology (i.e., activating antioxidant capacities) allowing it to be resilient to stress. This shows unequivocally that natural populations of corals sharing the same habitat can have diverse responses that are influenced by different environmental parameters. It also allows for a reconsideration of the subdivision into loser and winner species, suggesting that it may not be the largest and strongest species of today that will be the most resilient tomorrow.
Methods
Unless otherwise specified, all chemicals were obtained from Merck (France).
Animal sampling
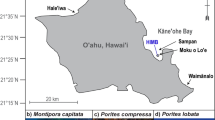
Coral colonies of Pocillopora spp. and Porites spp.57 were sampled between July 2016 and February 2017 across the Pacific Ocean during the Tara Pacific expedition (see Supplementary Table S8 for sampling sites, GPS coordinates and sampling date). They have been sampled according to UNCLOS and CITES permits (see Supplementary Note S1) and to the sampling protocol described in ref. 98. Briefly, 40 g of coral fragments from 155 Pocillopora spp. branched colonies and 166 Porites spp. massive colonies were sampled from 11 islands (but unfortunately Pocillopora samples from Malpelo island have been lost during the shipping), in at least three different sites at around 5 metres depth, and frozen onboard in liquid nitrogen. The fragments remained then stored at −20 °C until analysed. The time before freezing varied in respect to the distance between the Tara shooner and the sampling site, we performed a preliminary and complementary experiment to assess any artifactual results due to the freezing time lag (Supplementary Fig. S5).
Phenotypic biomarkers assays
Each coral sample was fragmented into two parts, a 10 g fragment was used for the biochemical biomarker analyses and a fragment of about 1 cm was used for the biomass analyses.
Animal and Symbiont biomasses: To assess animal and symbiont biomasses, the individual coral fragments were incubated at 37 °C in 4 M sodium hydroxide solution for 1–3 h99. Following the NaOH incubation, aliquots of the lysate were used for Symbiobiodiniaceae cell counting using the improved Neubauer hemocytometer and for protein concentration determined by Bradford protein assay100. The surface area of the remaining coral fragments was then measured using the aluminium foil technique101. Animal biomass was expressed in mg of protein per cm2, and symbiont biomass was expressed in number of cells per mg of protein and per cm2.
Proteins extraction
Each fragmented coral sample was mechanically grinded to powder with a Qiagen Tissue LyserII at 25HZ for 10 s. Soluble cytoplasmic protein were extracted by resuspending the obtained powder in a hypo-osmotic lysis buffer consisting of 25 mM 2-[4-(2-hydroxyethyl)piperazin-1-yl]ethanesulfonic acid (HEPES), 5 mM MgCl2, 5 mM 4-dithiothreitol (DTT), 5 mM ethylenediaminetetraaceticacid (EDTA), 2 mM phenylmethylsulfonylfluoride (PMSF), protease inhibitor cocktail (1 µl ml−1) and anti-phosphatase (10 µl ml−1) at pH 7.5 (ThermoScientific). Lysates were centrifuged at 12,000×g for 5 min, and only the supernatant was recovered containing the cytosoluble protein extracts. All extraction steps were performed under constant cooling. A Bradford protocol was followed to determine supernatant protein concentration using bovine serum albumin (BSA) solution (2 mg ml−1) as standard100.
Protein ubiquitination
Ubiquitinated proteins were assayed by dot-blotting 3 μg of Pocillopora protein samples and 0.5 µg of Porites protein samples on nitrocellulose membranes following ref. 102. The membranes were blocked in 5% milk in PBS-Tween (0.05%) overnight and then incubated with primary antibody (1:1000; mouse mono- and polyubiquitinylated conjugate recombinant, Enzo Life science) and Goat anti-mouse IgG HRP-conjugated antibody (1:5000; Bio-Rad) as secondary antibodies. HRP signals were visualised using chemiluminescent substrates (Immobilon classico, Merck) and acquired using a FusionSolo imaging system (Vilber). Levels of spot density were measured using the image analysis system G:Box (SynGene). To ensure the comparison of blotted membranes, a complementary normalisation has been performed with a standard curve of known protein concentration blotted in each membrane. The known proteins came from a unique extraction of protein from the sea anemone Anemonia viridis. The standard curve included a range of 0.5–4 µg of A. viridis protein extracts. The amount of ubiquitinated protein has been defined as arbitrary units (AU) per milligrams of protein.
Protein carbonylation
Carbonylated protein contents were measured using ELISA assay followed by spectrophotometric quantification according to ref. 103. A total of 0.5 mg ml−1 of protein were derivatized using dinitrophenylhydrazine (DNP) solution (10 mM in 6 M guanidine hydrochloride and 0.5 M potassium phosphate, pH 2.5). Antibody against DNP component (anti-rabbit DNP; 1:2000) was used as primary antibody and a Goat anti-rabbit IgG peroxidase conjugate (1:3000) was used for detection as secondary antibody. A standard curve of reduced and oxidised BSA was included in each microplate. Derivatized proteins were finally revealed by incubation of the microplate with a solution containing o-phenylenediamine (0.6 mg ml−1) and hydrogen peroxide (1:2500) in 50 mM Na2HPO4 plus 24 mM citric acid. Absorbances were read at 490 nm by spectrophotometer (Safas, Monaco). Carbonyl contents were expressed as nanomoles per mg of protein.
Antioxidant capacity
The total antioxidant capacities of protein samples were tested with the total oxidant scavenging capacity (TOSC) assay according to Naguib104, with modifications allowing measurement in black 96-well microplates (96-wells Grener Bio-One). Protein extracts were diluted in a 75 mM phosphate buffer (pH 7.0) to obtain protein concentrations in the reactive medium between 0.01 and 0.045 mg mL−1. The fluorescence signal was obtained by the oxidation of the fluorescent probe (180 nM; 6-carboxylfluorescein) by the peroxyl radical generator (36 mM AAPH (2,2′-azobis (2-amidinopropane) dihydrochloride), and Trolox (6-hydroxy-2,5,7,8-tetramethyl-1-chroman-2-carboxylic acid; 20 μM) was used as antioxidant calibrator for the assay. The fluorescence decay was measured by a spectrofluorometer (Safas, Monaco) at an excitation/emission wavelength of 520/495 nm every 2 min for a total duration of 1 h. Relative antioxidant activities of protein samples were measured by comparison with Trolox standard and results are reported as µmol Trolox equivalents mg−1.
Tara Pacific metadata
To explore the relationships between the measured phenotype data and the genetics and the associated microorganism diversity from the samples and the environment variables from the sampled sites, we used metadata from the Tara Pacific consortium (Table 3). The coral-species identification was performed on genome-wide markers57 and sequencing of divergent genomic fragments56 and resulted in five different identified species within the Pocillopora genus and three species within the Porites genus. The geographical distribution of the species analysed in the present work is shown in Supplementary Fig. S6. For the coral-associated microorganisms, on the one hand, we used the Symbiodiniaceae symbiont composition identified by the analysis of ribosomal nuclear ITS2105. For the colonies analysed in our study colonies, Symbiodiniaceae composition was divided into 10 and 11 clusters for Pocillopora and Porites respectively. Concerning the microbiome, we used the bacterial diversity obtained thanks to ribosomal 16 S analysis for bacterial microbiome67. For the colonies analysed in our study, microbiome composition was divided into five and seven clusters for Pocillopora and Porites, respectively. Finally, we also considered 101 contemporary (abbreviated to C), recent (R) and historical (H) environmental variables collected during the Tara Pacific expedition98. Contemporary data correspond to a rich collection of measurements of environmental variables collected onboard the day of sampling or recovered by satellite imagery and operational models. Recent and historical data correspond to past climate recordings extracted from climate satellite measurements, from respectively one year before sampling or from 2002. All the environmental variables were classified in four variables categories: Biological, Climate, Dynamism, or Physico-chemical (abbreviated to Bio, Cli, Dyn and Phy, respectively). The complete code-names of the environmental variables used in the present study are available in Supplementary Table S9.
Phenotypic analysis
The phenotypic signature from Pocillopora and Porites’ colonies were analysed through the pipeline summarised in Supplementary Fig. S7.
Data visualisation and exploration
The specific distribution of variance by biomarker was displayed as boxplots by islands or coral species which were generated with the ggplot2 v3.3.5 R package106. The integrative phenotypic signal, combining the different biomarkers, was visualised by PCA with a comparison either between the two genera or within a given genus and displaying island or species belonging, with FactoMineR v2.4 package107 and factoextra v1.0.7108.
Statistical analysis
Statistical pipelines and figures were made with R v4.1.1109.
Global variation of biomarkers through the Pacific Ocean
Biomarker by biomarker, distributions were compared by island or by species with Kruskall–Wallis’ test and a Dunn’s test as post hoc when necessary with the dunn.test v1.3.5 package110, after filtering individuals with missing data. Because of the complementary and eventual redundancy of some of the biomarkers used, we tested their correlation with Kendall’s tau before using all the biomarkers into the integrative phenotypic signal (Supplementary Fig. S2). We hence conserved all of them except the symbiont biomass reported by the surface of tissue.
Identification of putative drivers of coral phenotypic responses
The comparison between the coral genera, or among the islands and the coral species of a whole phenotypic signature was tested with a PERMANOVA (vegan v2.5-7111 and pairwiseAdonis v0.4112 packages using Bray–Curtis distance). This last analysis was run three times to ensure the repeatability of the outputs. To limit the number of false positive comparisons in the pairwise analyses, we applied a false discovery rate correction using the method described in ref. 113, implemented in the adonis function. The comparison at the coral genus scale was completed with a test of homogeneity of dispersion with betadisper function in vegan v2.5-7111. To estimate the influence of the origin island, coral species belonging or the symbiotic diversity of coral, the percentage of explained variance by each was measured with variancePartition v1.23.4 package114. We considered only coral colonies from islands containing more than one coral species per genus to limit bias (i.e., 87 Pocillopora colonies and 96 Porites colonies). An additional analysis was performed within coral species to determine a possible correlation between phenotypic and genetic distances (Mantel’s tests computed with the package vegan v2.5-7111).
Biomarkers and environment variables correlation
The influence of the environment on the phenotypic signal was assessed using 101 environmental variables. To identify correlations among the biomarkers and environmental variables, an sPLS (sparse Partial Least Squares) analysis was performed (mixOmics v6.17.29 package115). The sPLS analysis, designed to treat multicollinear variables and noise, allowed us to model multiple correlated response variables, considering the biomarkers as multiple responses and the environmental variables as multiple predictors of the coral phenotypes. Briefly, the values (from biomarkers and environmental data) were first turned into absolute values and then transformed with BoxCox transformation (caret v6.0-90116), tending toward normal distribution. The correlation among the predictor variables (here the environmental data) and the response variables (here the biomarkers) was measured in regression mode. Only 20% of the most correlated variables for the two components of the analysis were retained, i.e., a maximum of 20 for each component. We also added a correlation coefficient cut-off of 0.3 to highlight the strongest correlations. Then the common environmental variables between Pocillopora spp. and Porites spp. were identified with ggvenn v0.1.9117 and compared for each biomarker.
Data availability
Data generated during the study are available in the public repository (https://doi.org/10.5281/zenodo.7148413). The additional interpreted data, generated by the Tara consortium are the environmental variables98,118, the Symbiodiniaceae (ITS2)105 and bacterial (16 S)119 communities, and species delimitation56,57.
References
Johnson, N. A. et al. Integrative taxonomy resolves taxonomic uncertainty for freshwater mussels being considered for protection under the U.S. Endangered Species Act. Sci. Rep. 8, 15892 (2018).
Cooley, S. et al. Oceans and coastal ecosystems and their services. In Climate Change 2022: Impacts, Adaptation and Vulnerability. Contribution of Working Group II to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change (eds Pörtner, H.-O. et al.) pp. 379–550 (Cambridge University Press, Cambridge, UK and New York, NY, USA, 2022).
Bulgin, C. E., Merchant, C. J. & Ferreira, D. Tendencies, variability and persistence of sea surface temperature anomalies. Sci. Rep. 10, 7986 (2020).
Alexander, M. A., Shin, S.-I. & Battisti, D. S. The influence of the trend, basin interactions, and ocean dynamics on tropical ocean prediction. Geophys. Res. Lett. 49, e2021GL096120 (2022).
Muir, P. R., Marshall, P. A., Abdulla, A. & Aguirre, J. D. Species identity and depth predict bleaching severity in reef-building corals: shall the deep inherit the reef? Proc. R. Soc. B Biol. Sci. 284, 20171551 (2017).
McClanahan, T. R. et al. Large geographic variability in the resistance of corals to thermal stress. Glob. Ecol. Biogeogr. 29, 2229–2247 (2020).
Loya, Y. et al. Coral bleaching: the winners and the losers. Ecol. Lett 4, 122–131 (2001).
Hoegh-Guldberg, O., Kennedy, E. V., Beyer, H. L., McClennen, C. & Possingham, H. P. Securing a long-term future for Coral Reefs. Trends Ecol. Evol. 33, 936–944 (2018).
McClanahan, T. R., Weil, E., Cortés, J., Baird, A. H. & Ateweberhan, M. Consequences of coral bleaching for sessile reef organisms. in Coral Bleaching: Patterns, Processes, Causes and Consequences (eds van Oppen, M. J. H. & Lough, J. M.) 121–138 (Springer, 2009).
van Woesik, R., Sakai, K., Ganase, A. & Loya, Y. Revisiting the winners and the losers a decade after coral bleaching. Mar. Ecol. Prog. Ser. 434, 67–76 (2011).
Darling, E. S., Alvarez-Filip, L., Oliver, T. A., McClanahan, T. R. & Côté, I. M. Evaluating life-history strategies of reef corals from species traits. Ecol. Lett. 15, 1378–1386 (2012).
van Woesik, R. et al. Coral-bleaching responses to climate change across biological scales. Glob. Change Biol. 28, 4229–4250 (2022).
Barshis, D. J. et al. Protein expression and genetic structure of the coral Porites lobata in an environmentally extreme Samoan back reef: does host genotype limit phenotypic plasticity? Mol. Ecol 19, 1705–1720 (2010).
Polato, N. R., Altman, N. S. & Baums, I. B. Variation in the transcriptional response of threatened coral larvae to elevated temperatures. Mol. Ecol. 22, 1366–1382 (2013).
Pinzón, C. J. H., Beach-Letendre, J., Weil, E. & Mydlarz, L. D. Relationship between phylogeny and immunity suggests older Caribbean coral lineages are more resistant to disease. PLoS ONE 9, e104787 (2014).
Thinesh, T., Meenatchi, R., Jose, P. A., Kiran, G. S. & Selvin, J. Differential bleaching and recovery pattern of southeast Indian coral reef to 2016 global mass bleaching event: occurrence of stress-tolerant symbiont Durusdinium (Clade D) in corals of Palk Bay. Mar. Pollut. Bull. 145, 287–294 (2019).
Williams, S. D. & Patterson, M. R. Resistance and robustness of the global coral–symbiont network. Ecology 101, e02990 (2020).
Forsman, Z. H., Ritson-Williams, R., Tisthammer, K. H., Knapp, I. S. S. & Toonen, R. J. Host-symbiont coevolution, cryptic structure, and bleaching susceptibility, in a coral species complex (Scleractinia; Poritidae). Sci. Rep. 10, 16995 (2020).
Vega-Thurber, R. et al. Deciphering coral disease dynamics: integrating host, microbiome, and the changing environment. Front. Ecol. Evol. 8, 575927 (2020).
Wiedenmann, J. et al. Nutrient enrichment can increase the susceptibility of reef corals to bleaching. Nat. Clim. Change 3, 160–164 (2012).
Morris, L. A., Voolstra, C. R., Quigley, K. M., Bourne, D. G. & Bay, L. K. Nutrient availability and metabolism affect the stability of coral–Symbiodiniaceae symbioses. Trends Microbiol 27, 678–689 (2019).
DeCarlo, T. M. et al. Nutrient-supplying ocean currents modulate coral bleaching susceptibility. Sci. Adv. 6, eabc5493 (2020).
Alderdice, R. et al. Divergent expression of hypoxia response systems under deoxygenation in reef-forming corals aligns with bleaching susceptibility. Glob. Change Biol. 27, 312–326 (2021).
Altieri, A. H., Johnson, M. D., Swaminathan, S. D., Nelson, H. R. & Gedan, K. B. Resilience of tropical ecosystems to ocean deoxygenation. Trends Ecol. Evol. 36, 227–238 (2021).
Grottoli, A. G. et al. The cumulative impact of annual coral bleaching can turn some coral species winners into losers. Glob. Change Biol. 20, 3823–3833 (2014).
Sully, S., Burkepile, D. E., Donovan, M. K., Hodgson, G. & van Woesik, R. A global analysis of coral bleaching over the past two decades. Nat. Commun. 10, 1264 (2019).
McClanahan, T. R. et al. Temperature patterns and mechanisms influencing coral bleaching during the 2016 El Niño. Nat. Clim. Change 9, 845–851 (2019).
Madin, J. S. et al. A trait-based approach to advance coral reef science. Trends Ecol. Evol. 31, 419–428 (2016).
Todd, P. A. Morphological plasticity in scleractinian corals. Biol. Rev 83, 315–337 (2008).
Anthony, K. R. & Hoegh-Guldberg, O. Kinetics of photoacclimation in corals. Oecologia 134, 23–31 (2003).
Iluz, D. & Dubinsky, Z. Coral photobiology: new light on old views. Zoology 118, 71–78 (2015).
Muscatine, L., R. McCloskey, L. & E. Marian, R. Estimating the daily contribution of carbon from zooxanthellae to coral animal respiration 1. Limnol. Oceanogr. 26, 601–611 (1981).
Suggett, D. J. & Smith, D. J. Coral bleaching patterns are the outcome of complex biological and environmental networking. Glob. Change Biol. 26, 68–79 (2020).
Fitt, W. K. et al. Response of two species of Indo-Pacific corals, Porites cylindrica and Stylophora pistillata, to short-term thermal stress: the host does matter in determining the tolerance of corals to bleaching. J. Exp. Mar. Biol. Ecol. 373, 102–110 (2009).
Grottoli, A. G., Rodrigues, L. J. & Palardy, J. E. Heterotrophic plasticity and resilience in bleached corals. Nature 440, 1186–1189 (2006).
Qin, Z., Yu, K., Liang, Y., Chen, B. & Huang, X. Latitudinal variation in reef coral tissue thickness in the South China Sea: potential linkage with coral tolerance to environmental stress. Sci. Total Environ. 711, 134610 (2020).
Wooldridge, S. A. Excess seawater nutrients, enlarged algal symbiont densities and bleaching sensitive reef locations: 1. identifying thresholds of concern for the Great Barrier Reef, Australia. Mar. Pollut. Bull. 152, 107667 (2020).
Brown, B., Dunne, R., Goodson, M. & Douglas, A. Experience shapes the susceptibility of a reef coral to bleaching. Coral Reefs 21, 119–126 (2002).
Downs, C. A. et al. Oxidative stress and seasonal coral bleaching. Free Radic. Biol. Med. 33, 533–543 (2002).
Lesser, M. P. & Farrell, J. H. Exposure to solar radiation increases damage to both host tissues and algal symbionts of corals during thermal stress. Coral Reefs 23, 367–377 (2004).
Richier, S. et al. Oxidative stress and apoptotic events during thermal stress in the symbiotic sea anemone, Anemonia viridis. FEBS J 273, 4186–4198 (2006).
Richier, S. et al. Depth-dependant response to light of the reef building coral, Pocillopora verrucosa: implication of oxidative stress. J. Exp. Mar. Biol. Ecol. 357, 48–56 (2008).
Pey, A., Zamoum, T., Allemand, D., Furla, P. & Merle, P.-L. Depth-dependant thermotolerance of the symbiotic Mediterranean gorgonian Eunicella singularis: evidence from cellular stress markers. J. Exp. Mar. Biol. Ecol. 404, 73–78 (2011).
Krueger, T. et al. Differential coral bleaching—contrasting the activity and response of enzymatic antioxidants in symbiotic partners under thermal stress. Comp. Biochem. Physiol. A. Mol. Integr. Physiol. 190, 15–25 (2015).
Dias, M. et al. Oxidative stress on scleractinian coral fragments following exposure to high temperature and low salinity. Ecol. Indic. 107, 105586 (2019).
Marangoni, L. F. et al. Peroxynitrite generation and increased heterotrophic capacity are linked to the disruption of the coral–dinoflagellate symbiosis in a scleractinian and hydrocoral species. Microorganisms 7, 426 (2019).
Planes, S. et al. The Tara Pacific expedition-A pan-ecosystemic approach of the ‘-omics’ complexity of coral reef holobionts across the Pacific Ocean. PLoS Biol 17, e3000483 (2019).
Bythell, J. C., Brown, B. E. & Kirkwood, T. B. L. Do reef corals age? Biol. Rev. 93, 1192–1202 (2018).
Armstrong, E. J. et al. Host transcriptomic plasticity and photosymbiotic fidelity underpin Pocillopora acclimatization across thermal regimes in the Pacific Ocean Nat. Commun. 14, 3056 (2023).
Cunning, R., Glynn, P. W. & Baker, A. C. Flexible associations between Pocillopora corals and Symbiodinium limit utility of symbiosis ecology in defining species. Coral Reefs 32, 795–801 (2013).
Rouzé, H., Lecellier, G., Pochon, X., Torda, G. & Berteaux-Lecellier, V. Unique quantitative Symbiodiniaceae signature of coral colonies revealed through spatio-temporal survey in Moorea. Sci. Rep. 9, 1–11 (2019).
Stat, M., Pochon, X., Cowie, R. O. M. & Gates, R. D. Specificity in communities of Symbiodinium in corals from Johnston Atoll. Mar. Ecol. Prog. Ser. 386, 83–96 (2009).
Turnham, K. E., Wham, D. C., Sampayo, E. & LaJeunesse, T. C. Mutualistic microalgae co-diversify with reef corals that acquire symbionts during egg development. ISME J 15, 3271–3285 (2021).
Ziegler, M., Roder, C., Büchel, C. & Voolstra, C. R. Niche acclimatization in Red Sea corals is dependent on flexibility of host-symbiont association. Mar. Ecol. Prog. Ser. 533, 149–161 (2015).
Smith, A. et al. Field measurements of a massive Porites coral at Goolboodi (Orpheus Island), Great Barrier Reef. Sci. Rep. 11, 15334 (2021).
Deshuraud, R. et al. From genome wide SNPs to genomic islands of differentiation: the quest for species diagnostic markers in two scleractinian corals, Pocillopora and Porites https://doi.org/10.1101/2022.10.21.513203 (2023).
Voolstra, C. R. et al. Disparate genetic divergence patterns in three corals across a pan-Pacific environmental gradient highlight species-specific adaptation. npj Biodiversity 2, 15 (2023).
Barnes, D. J. & Lough, J. M. Systematic variations in the depth of skeleton occupied by coral tissue in massive colonies of Porites from the Great barrier reef. J. Exp. Mar. Biol. Ecol. 159, 113–128 (1992).
Tivey, T. R., Parkinson, J. E. & Weis, V. M. Host and symbiont cell cycle coordination is mediated by symbiotic state, nutrition, and partner identity in a model cnidarian-dinoflagellate symbiosis. mBio 11, e02626–19 (2020).
Thornhill, D. J. et al. A connection between colony biomass and death in Caribbean reef-building corals. PLoS ONE 6, e29535 (2011).
Krueger, T. et al. Antioxidant plasticity and thermal sensitivity in four types of Symbiodinium sp. J. Phycol. 50, 1035–1047 (2014).
Richier, S., Furla, P., Plantivaux, A., Merle, P.-L. & Allemand, D. Symbiosis-induced adaptation to oxidative stress. J. Exp. Biol 208, 277–285 (2005).
Baird, A. H., Guest, J. R. & Willis, B. L. Systematic and biogeographical patterns in the reproductive biology of scleractinian corals. Annu. Rev. Ecol. Evol. Syst. 40, 551–571 (2009).
Finkel, T. & Holbrook, N. J. Oxidants, oxidative stress and the biology of ageing. Nature 408, 239–247 (2000).
Ezzat, L., Merle, P.-L., Furla, P., Buttler, A. & Ferrier-Pagès, C. The response of the Mediterranean Gorgonian Eunicella singularis to thermal stress is independent of its nutritional regime. PLoS ONE 8, e64370 (2013).
Zhou, Z. et al. Suppression of NF-κB signal pathway by NLRC3-like protein in stony coral Acropora aculeus under heat stress. Fish Shellfish Immunol. 67, 322–330 (2017).
Galand, P. E. et al. Diversity of the Pacific Ocean coral reef microbiome. Nat. Commun. 14, 3039 (2023).
Santoro, E. P. et al. Coral microbiome manipulation elicits metabolic and genetic restructuring to mitigate heat stress and evade mortality. Sci. Adv. 7, eabg3088 (2021).
Wright, R. M. et al. Intraspecific differences in molecular stress responses and coral pathobiome contribute to mortality under bacterial challenge in Acropora millepora. Sci. Rep. 7, 2609 (2017).
Hackerott, S., Martell, H. A. & Eirin-Lopez, J. M. Coral environmental memory: causes, mechanisms, and consequences for future reefs. Trends Ecol. Evol. 36, 1011–1023 (2021).
van Woesik, R. et al. Hosts of the Plio-Pleistocene past reflect modern-day coral vulnerability. Proc. R. Soc. B Biol. Sci. 279, 2448–2456 (2012).
Johnston, E. C. et al. A genomic glance through the fog of plasticity and diversification in Pocillopora. Sci. Rep. 7, 5991 (2017).
Glynn, P. W., Manzello, D. P. & Enochs, I. C. Coral Reefs of the Eastern Tropical Pacific: Persistence and Loss in a Dynamic Environment (Springer, 2016).
D’Croz, L. & O’Dea, A. Variability in upwelling along the Pacific shelf of Panama and implications for the distribution of nutrients and chlorophyll. Estuar. Coast. Shelf Sci. 73, 325–340 (2007).
Stimson, J. The annual cycle of density of zooxanthellae in the tissues of field and laboratory-held Pocillopora damicornis (Linnaeus). J. Exp. Mar. Biol. Ecol. 214, 35–48 (1997).
Bosserelle, P. et al. Niue marine ecological surveys, 2016 and 2017. Pacific Community, Noumea, New Caledonia. pp. 74. https://doi.org/10.1101/2021.11.12.468330 (2018).
Ziegler, M. et al. Status of coral reefs of Upolu (Independent State of Samoa) in the South West Pacific and recommendations to promote resilience and recovery of coastal ecosystems. Mar. Pollut. Bull. 129, 392–398 (2018).
Berkelmans, R. & van Oppen, M. J. The role of zooxanthellae in the thermal tolerance of corals: a ‘nugget of hope’ for coral reefs in an era of climate change. Proc. R. Soc. Lond. B Biol. Sci. 273, 2305–2312 (2006).
McGinty, E. S., Pieczonka, J. & Mydlarz, L. D. Variations in reactive oxygen release and antioxidant activity in multiple symbiodinium types in response to elevated temperature. Microb. Ecol. 64, 1000–1007 (2012).
Matthews Jennifer, L. et al. Partner switching and metabolic flux in a model cnidarian–dinoflagellate symbiosis. Proc. R. Soc. B Biol. Sci. 285, 20182336 (2018).
Wuitchik, D. M. et al. Characterizing environmental stress responses of aposymbiotic Astrangia poculata to divergent thermal challenges. Mol. Ecol. 30, 5064–5079 (2021).
Saxby, T., Dennison, W. C. & Hoegh-Guldberg, O. Photosynthetic responses of the coral Montipora digitata to cold temperature stress. Mar. Ecol. Prog. Ser. 248, 85–97 (2003).
DeCarlo, T. M. & Harrison, H. B. An enigmatic decoupling between heat stress and coral bleaching on the Great Barrier Reef. PeerJ 7, e7473 (2019).
Ip, J. C.-H., Zhang, Y., Xie, J. Y., Yeung, Y. H. & Qiu, J.-W. Comparative transcriptomics of two coral holobionts collected during the 2017 El Niño heat wave reveal differential stress response mechanisms. Mar. Pollut. Bull. 182, 114017 (2022).
Browne, N. K., Smithers, S. G. & Perry, C. T. Coral reefs of the turbid inner-shelf of the Great Barrier Reef, Australia: an environmental and geomorphic perspective on their occurrence, composition and growth. Earth-Sci. Rev. 115, 1–20 (2012).
Browne, N. K., Precht, E., Last, K. S. & Todd, P. A. Photo-physiological costs associated with acute sediment stress events in three near-shore turbid water corals. Mar. Ecol. Prog. Ser. 502, 129–143 (2014).
Morgan, K. M., Perry, C. T., Smithers, S. G., Johnson, J. A. & Daniell, J. J. Evidence of extensive reef development and high coral cover in nearshore environments: implications for understanding coral adaptation in turbid settings. Sci. Rep. 6, 29616 (2016).
Alamaru, A., Loya, Y., Brokovich, E., Yam, R. & Shemesh, A. Carbon and nitrogen utilization in two species of Red Sea corals along a depth gradient: insights from stable isotope analysis of total organic material and lipids. Geochim. Cosmochim. Acta 73, 5333–5342 (2009).
Conti-Jerpe, I. E. et al. Trophic strategy and bleaching resistance in reef-building corals. Sci. Adv. 6, eaaz5443 (2020).
Houlbrèque, F. & Ferrier-Pagès, C. Heterotrophy in tropical scleractinian corals. Biol. Rev 84, 1–17 (2009).
Levas, S. J., Grottoli, A. G., Hughes, A., Osburn, C. L. & Matsui, Y. Physiological and biogeochemical traits of bleaching and recovery in the mounding species of coral porites lobata: implications for resilience in mounding corals. PLoS ONE 8, e63267 (2013).
D’Angelo, C. & Wiedenmann, J. Impacts of nutrient enrichment on coral reefs: new perspectives and implications for coastal management and reef survival. Curr. Opin. Environ. Sustain. 7, 82–93 (2014).
Dunn, J. G., Sammarco, P. W. & LaFleur, G. Effects of phosphate on growth and skeletal density in the scleractinian coral Acropora muricata: a controlled experimental approach. J. Exp. Mar. Biol. Ecol. 411, 34–44 (2012).
Kteifan, M., Wahsha, M. & Al-Horani, F. A. Assessing stress response of Stylophora pistillata towards oil and phosphate pollution in the Gulf of Aqaba, using molecular and biochemical markers. Chem. Ecol. 33, 281–294 (2017).
Allemand, D. & Osborn, D. Ocean acidification impacts on coral reefs: from sciences to solutions. Reg. Stud. Mar. Sci. 28, 100558 (2019).
Luz, D. C. et al. Oxidative stress in the hydrocoral Millepora alcicornis exposed to CO2-driven seawater acidification. Coral Reefs 37, 571–579 (2018).
Soriano-Santiago, O. S., Liñán-Cabello, M. A., Delgadillo-Nuño, M. A., Ortega-Ortiz, C. & Cuevas-Venegas, S. Physiological responses to oxidative stress associated with pH variations in host tissue and zooxanthellae of hermatypic coral Pocillopora capitata. Mar. Freshw. Behav. Physiol. 46, 275–286 (2013).
Lombard, F. et al. Open science resources from the Tara Pacific expedition across coral reef and surface ocean ecosystems. Sci. Data 10, 324 (2023).
Zamoum, T. & Furla, P. Symbiodinium isolation by NaOH treatment. J. Exp. Biol. 215, 3875–3880 (2012).
Jackson, A. P. A reconciliation analysis of host switching in plant-fungal symbioses. Evolution 58, 1909–1923 (2004).
Hoegh-Guldberg, O. A method for determining the surface area of corals. Coral Reefs 7, 113–116 (1988).
Haas, A. L. & Bright, P. M. The immunochemical detection and quantitation of intracellular ubiquitin-protein conjugates. J. Biol. Chem. 260, 12464–12473 (1985).
Buss, H., Chan, T. P., Sluis, K. B., Domigan, N. M. & Winterbourn, C. C. Protein carbonyl measurement by a sensitive ELISA method. Free Radic. Biol. Med. 23, 361–366 (1997).
Naguib, Y. M. A. A fluorometric method for measurement of oxygen radical-scavenging activity of water-soluble antioxidants. Anal. Biochem. 284, 93–98 (2000).
Hume, B. C. C. et al. Tara Pacific ITS2 Symbiodiniaceae data release version 1. https://doi.org/10.5281/zenodo.4061797 (2020).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis R package version 3.3.5 (Springer, 2016).
Lê, S., Josse, J. & Husson, F. FactoMineR: an R package for multivariate analysis. J. Stat. Softw. 25, 1–18 (2008).
Kassambara, A. & Mundt, F. factoextra: extract and visualize the results of multivariate data analyses. R package version 1.0.7. (2020).
R Core Team. R: A Language and Environment for Statistical Computing R Foundation for Statistical Computing, Vienna. https://www.R-project.org (2022).
Dinno, A. dunn.test: Dunn’s test of multiple comparisons using rank sums. R package version 1.3.5. (2017).
Oksanen, J. et al. vegan: community ecology package. R Package Version 2, 5–7 (2020).
Martinez Arbizu, P. pairwiseAdonis. R package version 0.4. (2021).
Benjamini, Y. & Hochberg, Y. Controlling false discovery rate: a practical and powerful approach to multiple testing. J. R. Soc. Ser. B 57, 289–300 (1995).
Hoffman, G. E. & Schadt, E. E. variancePartition: interpreting drivers of variation in complex gene expression studies. BMC Bioinforma. 17, 483 (2016).
Rohart, F., Gautier, B., Singh, A. & Cao, K.-A. L. mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput. Biol. 13, e1005752 (2017).
Kuhn, M. Caret: Classification and Regression Training. R Package Version 6.0-88. https://CRAN.Rproject.org/package=caret (2021).
Yan, L. ggvenn: draw Venn diagram by ‘ggplot2’. R package version 0.1.9. (2021).
Pesant, S. et al. Tara Pacific samples provenance and environmental context-version 2. https://doi.org/10.5281/zenodo.6299409 (2020).
Ruscheweyh, H.-J. et al. Tara Pacific 16S rRNA data analysis release. https://doi.org/10.5281/zenodo.6598864 (2022).
Acknowledgements
This work would not have been possible without the support, help and dedication of Etienne Bourgeois, the Tara Ocean Foundation’s teams and crew and all the Tara Pacific Expedition Participants (https://doi.org/10.5281/zenodo.3777760). We are keen to thank the commitment of the following institutions for their financial and scientific support that made this unique Tara Pacific Expedition possible: CNRS, PSL, CSM, EPHE, Genoscope, CEA, Inserm, Université Côte d’Azur, ANR, agnès b., UNESCO-IOC, the Veolia Foundation, the Prince Albert II de Monaco Foundation, Région Bretagne, Billerudkorsnas, AmerisourceBergen Company, Lorient Agglomération, Oceans by Disney, L’Oréal, Biotherm, France Collectivités, Fonds Français pour l’Environnement Mondial (FFEM), the Ministère des Affaires Européennes et Etrangères, the Museum National d’Histoire Naturelle. For the experimental and analytic support, we thank Noemie Dechaux, Nina Luckas, Alexia Oechsel, Brigitte Poderini, Guillaume Signoret, Ludovic Toisoul and Marion Tredez. For the statistical discussions and advice, we warmly thank Diane Bailleul, Marc Bailly-Bechet, Jérôme Laurent, Mélanie Pousse and Florentine Riquet. For the sample storage and management, we also thank Alice Rouan. The work was supported by the ANR CORALGENE (ANR-17-CE02-0020), the 'Investments for the Future' programs LABEX SIGNALIFE (ANR-11-LABX-0028) and IDEX UCAJedi (ANR-15-IDEX-01) as well as the master fellowship from the french 'Fondation pour la recherche sur la biodiversité'. This is publication number #30 of the Tara Pacific Consortium. We are keen to thank the local people and authorities who made this study possible. Specifically, we thank the people of Colombia and the following local institutions for their administrative and scientific support that made this singular expedition possible, the Prime Minister Office and their teams; the Ministry of Environment and National Natural Parks of Colombia with Guillermo Alberto Santos Ceballos, Edna Carolina Jarro Fajardo, and their teams; la Fundacion Malpelo y Otros Ecosistemas Marinos with their Executive Director and Founder Sandra Bessudo, with Felipe Orlando Ladino and their teams; el Santuario de Fauna y Flora Malpelo and their teams; la Direccion General Maritima with Ivan Fernando Castro Mercado and their teams; The Pacific Territorial Directorate and their teams; the Colombian biodiversity information system and their teams; and all other locally supports. We are keen to thank the commitment of the people of Panama and the following local institutions for their administrative and scientific support that made this singular expedition possible: The Ministry of Environment of Panama with Samuel Valdès Diaz, Lissette Trejos, Patricia Hernandez and their teams; the Smithsonian Tropical Research Institute with Rachel Page, Zurenayka Alain, Juan Mate and their teams; the Head of MPA at Coiba Park Didiel Nunez and their teams; and all other locally supports. We are keen to thank the commitment of the people of Pitcairn Islands and the following local institutions for their administrative and scientific support that made this singular expedition possible: the Government of Pitcairn Islands with Melva Evans and their teams; the Office of the Mayor and Council for Pitcairn, Henderson, Ducie and Oeno Islands with Shawn Christian and their teams; the Environmental, Conservation & Natural Resources Division with her Manager Michele Christian, and their teams; the Police & Immigration Officer with Christian Brenda, and their teams; and all other locally supports. We are keen to thank the commitment of the people of French Polynesia and the following local institutions for their administrative and scientific support that made this singular expedition possible: Le Haut-commissaire de la République en Polynésie française and their teams; The City of Papeete with its Mayor, its Port Commander, and their teams; le Centre de Recherches Insulaires et Observatoire de l’Environnement (CRIOBE) and their teams; and all other locally supports. We are keen to thank the commitment of the people of the Cook Islands and the following local institutions for their administrative and scientific support that made this singular expedition possible: the Government of the Cook Islands and their teams; the Office of the Prime Minister with Tina Samson, Elizabeth Wright-Koteka from the Cook Islands Research Committee, and their teams; the Ministry of Marine Resources with its Director Dorothy Solomona, and their teams; the National Environment Service with its Director Joseph Brider, Elizabeth Munro, Bobby Bishop, and their teams; the City of Aitutaki, its mayor, and their teams; the Aitutaki Marine Research Station with its station manager Richard Story, and their teams; the Cawthron Institute with Lesley Rhodes, Kirsty Smith, and their teams; and all other locally supports. We are keen to thank the commitment of the people of Niue and the following local institutions for their administrative and scientific support that made this singular expedition possible: the Government of Niue with Richard Hipa, James Tatafu, Aldric Hipa, and their teams; the Office of External Affairs of the Government of Niue with its Director Emi Hipa, and their teams; the Cabinet and Parliamentary Services with its Director Christine External; the Ministry of Infrastructure and its Department of Transport, with Lynsey Talagi, and their teams; the Ministry of Natural Resources and its Director-General Josie Tamate, and their teams; the Department of Agriculture, Forestry and Fisheries at the Ministry of Natural Resources with its Director Brendon Pasisi, and their teams; the Department of Environment of the Ministry of Natural Resources and its Director Sauni Tongatule, and their teams; the Pacific Community with its Deputy Director-General Cameroun Diver, Coral Pasisi, and their teams; and all other locally supports.We are keen to thank the commitment of the people of Samoa Islands and the following local institutions for their administrative and scientific support that made this singular expedition possible: the Government of the Independent State of Samoa, and their teams; the Ministry of Foreign Affairs and Trade with its CEO Peseta Noumea Simi, its Acting CEO Leroy E. Hunkin-Mamae, Sapeti Titiii, and their teams; the Ministry of Natural Resources and Environment with its Principal Marine Conservation Officer Maria R. Satoa Peni, and their teams; the Ministry of Agriculture and Fisheries with its Assistant CEO Magele Etuati Ropeti, and their teams; and all other locally supports. We are keen to thank the commitment of the people of Guam and the following local institutions for their administrative and scientific support that made this singular expedition possible: the Department of Agriculture Division of Aquatic and Wildlife Resources (DAWR), its Director Matthew L.G. Sablan, its Assistant Chief Jay Gutierrez, and their teams; the Division of Management Authority Branch of Permits, the Senior Biologist Anna Barry, and their teams; the U.S. Fish and Wildlife Service—Office of Law Enforcement, its Wildlife Inspector Arthur T. Taimanglo, and their teams; and all other locally supports.
Author information
Authors and Affiliations
Contributions
Conceptualisation: P.F., B.P., T.Z., S.A., D.A., B.B., E.Bio., E.Bos., C.B., C.d.V., C.V., D.F., E.G., S.B.V., F.L., M.F., P.G., S.Pe., S.P.l., S.Re., M.S., S.S., O.T., R.T., R.V.T., P.W. and D.Z.; sample management: C.M., E.Boi., G.B., G.I., J.P. and S.Ro.; methodology and investigation: T.Z., P.F., E.D.F. and A.P.; formal analysis: B.P., P.F., D.F. and F.L.; data curation: T.Z., B.P., P.F., B.H., C.V., P.G., F.L. and G.B.; visualisation: B.P.; project administration and supervision: P.F.; writing— original draft preparation: P.F., B.P. and T.Z.; funding acquisition: P.F., E.G., E.R., S.B.V., S.P.l., D.A., S.A., B.B., E.Boi., E.Bos., C.B., C.d.V., E.D.F., E.D., M.F., P.G., S.pe., S.Re., M.S., S.S., O.T., R.T., R.V.T., C.V., P.W. and D.Z. All authors have read and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth & Environment thanks Raphael Ritson-Williams, Marina Tonetti Botana and Thierry Work for their contribution to the peer review of this work. Primary Handling Editors: Christopher Cornwall and Clare Davis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Porro, B., Zamoum, T., Forcioli, D. et al. Different environmental response strategies in sympatric corals from Pacific Islands. Commun Earth Environ 4, 311 (2023). https://doi.org/10.1038/s43247-023-00946-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43247-023-00946-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.