Abstract
Reliable metrics to monitor human impacts on biodiversity are essential for informing conservation policy. As insects are indicators of global change, whose declines profoundly affect ecosystems, insect diversity may predict biodiversity status. Here we present an unbiased and straightforward biodiversity status metric based on insect diversity (richness) and landscape naturalness. Insect diversity was estimated using spatially explicit earth observation data and insect species assemblages across microhabitats in two agro-ecological zones in Africa. Landscape naturalness was estimated using various human impact factors. Biodiversity status values differed considerably (p < 0.05) between protected and non-protected areas, while protected areas, regardless of agro-ecology, shared similar biodiversity status values. The metric is consistent when using richness from different indicator taxa (i.e., stingless bees, butterflies, dragonflies) and independent data for landscape naturalness. Our biodiversity status metric is applicable to data-scarce environments and practical for conservation actions and reporting the status of biodiversity targets.
Similar content being viewed by others
Introduction
Accelerated declines in biodiversity are of major concern globally1. As for most taxa, insect population declines are largely driven by habitat loss, including loss in habitat quality2. Moreover, losses in insect biodiversity can lead to declines in other species that feed on insects, as well as to crop yield losses due to scarcity of pollinators3,4. In some African countries, 15–40% of all calories, protein, and iron nutrient intakes come from pollinator dependent crops5. Thus, insect declines can result in yield gaps that can be 59% or more for some seed crops6. Overall, biodiversity in African ecosystems is increasingly imperiled, and likely more so in the future7. Spatially explicit indicators and metrics that make use of biotic inputs are needed to quantify the status of biodiversity at local levels throughout Africa8.
To varying degrees, depending on the taxonomic rank, insect diversity and abundance can be used to estimate overall ecosystem-level biodiversity and environmental integrity, especially when philopatric or indicator species are chosen for biomonitoring9,10. For instance, within the order Lepidoptera (butterflies and moths), some species may be generalists and hence adaptable, while others might become extinct or migrate to other places11,12. But the diversity and habitat suitability of many insect indicator families and/or orders is likely to correlate with overall ecosystem diversity and integrity13. In some cases, shifts in insect habitats can be used as early warning indicators of ecosystem-level environmental change14 before extinction risks or declines at upper trophic levels are measurable from other groups. Trophic responses to human transformations that affect species distributions and richness are more rapid at the micro-scale (i.e., insect mapping unit) than at national or regional scales15. A micro-scale for a species is characterized by a very specific vegetation structure as well as abiotic conditions (e.g., presence of raw soil, deadwood) and by a specific microclimate, whereby the spatial scale of a microhabitat is relative and species-specific2. Localized and/or sudden land transformation often abruptly affect insect life cycles, nesting (i.e., oviposition sites), and foraging behavior16,17.
The post-2000 Global Biodiversity Conservation (GBC) Framework of the UN proposes several indicators and corresponding targets that include species or ecosystem-specific biodiversity status information. These are set to guide member countries until 205018 and help to establish important biodiversity status baselines. The Essential Biodiversity Variables 2020 (EBV2020) Initiative19, which supports the post-2020 GBC initiative, has established that easy-to-use indicators of habitat size, resilience, connectivity, and biodiversity integrity must be developed and updated. The EBV2020 approach promotes the addition of new species groups and the need for metrics that consider functional traits and ecosystem composition20.
Most biodiversity measures use parametric species richness estimates for various ecosystem or land use types7,21. They often disregard structural landscape or explicit habitat patterns, including landscape fragmentation11 and actual ecosystem and habitat structural aspects of biodiversity22. Many biodiversity measures render different trends at local and regional levels. This leads to a hindrance in the adoption of relevant policies. In part, inconsistent biodiversity measures are due to the exclusion of localized biotic data, the effects of landscape patterns, and low sensitivity (or over-sensitivity) to biodiversity status over a range of human impacts23.
The insect-based biodiversity status (iBS) metric, introduced herein, exhibits various specific novelties, differences, and complementarities over existing composite biodiversity indicators such as the Biodiversity Habitat Index, the Living Planet Index, the PREDICTS assessment framework, or dissimilarity modeling methodologies for predicting beta-biodiversity15,19,21,24,25,26. The iBS metric does not consider the composition of species and their changes in time and space (as considered in beta biodiversity dissimilarly approaches), nor aspects related to interspecies interactions or population structure as determined by genetic processes and structure26,27. Unlike the case for dissimilarity/similarity model approaches28, the iBS does not aim to compare different species or species communities and quantify these differences between two sites and their ecological conditions. Rather, the metric aims to provide an earth observation (pixel-based) and straightforward measure (i.e., computationally easy) for local biodiversity based on insect assemblages as collective orders that represent indicator species for intact landscapes. Like many indicators, it combines biotic elements (habitat-based suitability) with various human impacts (for sensitivity to both biotic and human impact, see Supplementary Notes 1: Understand patterns of the contributing layers, illustrate visual patterns for the two components that make up the iBS and the metric result for each study area).However, it uses readily available and spatially explicit earth observation and insect data to help determine local-scale biodiversity patterns and status. These localized patterns are defined by landscape level disturbances and landscape type29, not directly by gene pools and their structure. Unlike most other indicators investigated, the iBS is not affected by broad taxonomic datasets that are ecologically restricted, not covering larger landscapes or ecologically diverse areas22. Given dense insect occurrence data is used for model training, our metric produces seamless localized outputs from pixel-based explicit geospatial data and models. Moreover, our pixel-based and data-driven metric does not use expert knowledge on land use intensity as a factor for biodiversity richness within one or several coarse-scale (>1 km pixel resolution) land use categories19. Essentially, most indicators and metrics investigated used generalized and published data with assumptions about the coordinates, often extracted from scientific papers or reports15.
Lastly, the iBS is uniquely tied to taxonomic insect orders (i.e., Lepidoptera) that function as known indicators of micro-habitats’ biodiversity status30. This differentiates it from composite indicators that use broad taxonomic groups with various interactions with human impacts, thereby reducing the appropriateness of the indicator as a measure of overall biodiversity31. The choice of assemblages as indicator species allows the pixel-based results to be averaged for ecosystems or key habitat areas (i.e., forests) within the area of interest, where sufficient insect data is available for model training. When using our metric approach, the prerequisite is that the area of interest lies within a common climate zone. Current trends show that regional to global biodiversity studies relying on indicators from big data (species and predictor variables), model biodiversity for narrow taxonomic ranges (i.e., all bird species). These results, however, are not applicable to determine the overall biodiversity status over broader scales32.
This work aimed to propose a practical, bottom-up, and easy-to-calculate metric that is sensitive to the status of local biodiversity, using indicator insect species as assemblages. As a result, the metric provides a comprehensive biodiversity status baseline that is applicable using existing data and responsive to incremental improvements when data sets are updated. This study does not aim to validate the metrics over a range of underlying socio-ecological conditions and ascertain its applicability over a broad scale. Data requirements and how users can apply the metrics and test its local scale validity and implementation risks are alluded to in the method protocol.
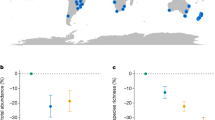
We found that irrespective of insect taxa, microhabitat investigated within a nested agro-ecological unit, and even over distinct climate zones, the proposed metric shows stability in value distributions and is thus suitable for localized biodiversity status mapping. The results also showed that the diversity of stingless bees, butterflies and dragonflies is higher in protected areas than in unprotected areas within various agro-ecological zones in Kenya and South Africa. In conclusion, the metric provides a biodiversity baseline measure that can be implemented in insect data dense regions to ascertain the micro-habitat-based biodiversity status of landscapes.
Results
In implementing the iBS metric for butterflies (BF) and moths (Moth) and rove beetles assemblages (termed BF/Moth/Rove_N) independently in two distinct agro-ecological climates (Kakamega in Kenya and St. Lucia in South Africa), we found that iBS values differed considerably between Protected (P) and Non-protected (NP) areas. The dissimilarity was confirmed using the Kruskal–Wallis multi-pairwise test, using class-specific variances in the iBS values (p = 0.001). However, we obtained value distributions that were comparable between the two agro-ecologies, both within P and within NP areas (Fig. 1). Specifically, in Kakamega (humid climate), the values for P ranged between 0.38 and 0.95, with a mean of 0.64 (n = 500 random samples); the value ranges for P were similar in semi-humid St. Lucia (0.49–0.89; mean = 0.53; n = 500).
Means (black dot at the center), standard errors (dark teal line intervals), and 95% confidence intervals (red intervals), standard deviation (gray intervals), for randomly sampled pixels are shown for land cover categories protected (P) (n = 500), non-protected (NP) (n = 500) and agroforestry (AF) (n = 57). The violin plot scores (numbers in parenthesis) illustrate how similar the distributional plots are, i.e., categories that can be grouped into one category.
Agroforestry (AF), only present in Kakamega, exhibits iBS values (0.2–0.51, mean = 0.31) that are closer to the protected category than the NP category (0.004–0.08; mean = 0.04) (Fig. 1). This is because agro-forestry sites provide a high degree of habitat heterogeneity, which results in more diverse ecological niches for various species33.
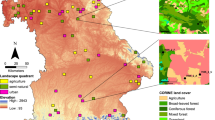
The spatial iBS pattern maps for BF/Moth/Rove_N (Fig. 2) showed interesting and realistic patterns of biodiversity status in both areas that clearly map sentinel habitats (i.e., gallery forests and wetlands) and how they have been affected by human activity. Moreover, the similarity/dissimilarity results (Fig. 1) are apparent; the stark differences between the Kakamega P (dark green shades; Fig. 2a) and NP areas (other shades) are well visible. Inset a illustrates that residual near-natural vegetation (gallery forests) and agro-forestry areas alongside the Kakamega P areas are clearly visible and have the same color shades (and iBS values) as the woody cover in the Kakamega P itself. In St. Lucia (Fig. 2b), low density settlement areas are characterized by a moderate to high insect-based iBS (0.2–0.82, n = 55). Although the insect assemblages used do not prefer transformed areas, low-density urban areas (buildings inter-mixed with urban gardens or urban agriculture) can sustain high insect species richness and diversity34. High iBS values in St. Lucia were also associated with both coastal forests and some wetland areas, often in proximity (inset b, Fig. 2). Although wetlands in St. Lucia are mostly regularly flooded grasslands, and not forested areas, the high biodiversity status of wetlands is due to the inclusion (in the species modeling) of some rove beetle families (e.g., Staphylinidae). These prefer waterline fringes and wetlands as habitat zones. This confirms that biodiversity status information should be comprehensive enough to include species that prefer various micro-habitats (when using assemblages).
Map a illustrates the iBS result for Kakamega in Kenya, area size approximately 2000 km2, and map b is the result for St. Lucia in South Africa, same approximate area size. The map insets show specific zoomed sub-sections within each area. The letters F (Forest) and W (Wetland) for the St. Lucia inset illustrate the location of these specific land cover features. iBS values are color coded according to respective value ranges, as shown. The violin plot scores (numbers in parenthesis) illustrate how similar the distributional plots are, i.e., categories that can be grouped into one category.
As a result, our biodiversity status metric considers the role insects can play as bio-indicators for the status of a whole landscape or region. Wetlands are considered highly important areas for biodiversity. Using the spatially explicit Earth Observation (EO) data, that includes wetness spectral features, helped in the identification of fine-scaled habitat zones most suitable for these semi-aquatic indicator species. This also confirms that biodiversity status is not only determined by forest cover, but also by habitat suitability.
Inset a for Kakamega shows associations of gallery forests and seepage lines with higher iBS values than the surrounding non-protected landscape. The inset b for St. Lucia shows a seasonally flooded grassland wetland (W) and an alongside forest (F). Both land cover features have similar iBS values. Google earth imagery is shown corresponding to each inset, respectively. Due to the skewness of the distribution, the iBS data are mapped with a linear color stretch, truncated at one standard deviation.
Spatially explicit and coherent data on insect biodiversity status is not available for cross comparison. However, an area-specific study that linked local butterfly assemblages (diversity/richness) to utilization intensity and land use in Miombo woodlands, Tanzania (semi-arid climate), found a mean butterfly species diversity of 88% for Forests (graded as low utilization) and 81% for Gallery Forests (slight utilization)35. The percentages in that study were expressed as the mean score attained compared to the maximum diversity value given for that land cover class. In our case, the values were favorably comparable; 88% for Forest and 79% for Gallery Forest. These percentage means were derived from the pixel means for each category in relation to the maximum value found in each category for the Kakamega site. We used Kakamega in the comparison, since it has larger and with comparable proportions of these two land cover categories.
The value distributions for P, NP, and AF were similar when based on three data sets using (i) BF/Moth/Rove_N in Kakamega and St. Lucia (ii) stingless bees (tribe level) (SB) amplified by the Forest Integrity (FI) index (SB_FI) in Kakamega and (iii) dragonfly (sub-orders) amplified by FI data (DF_FI) in St. Lucia (Fig. 3).
The distributions for BF/Moth/Rove_N (mean, range) for randomly selected pixels for protected (P) (n = 500), and non-protected (NP) (n = 500) and agroforestry (AF) (n = 57) are shown as black violin plots. Distributions for SB_FI (n = 50–500, random samples) for Kakamega are illustrated for P, NP, and AF in orange color. Distributions for DF_FI (n = 50–500, random samples) for St. Lucia are illustrated for each of the categories in blue color. Means (black dot at the center), standard errors (dark teal line intervals), and 95% confidence intervals (red intervals), standard deviation (gray intervals) are illustrated for each violin plot. The violin scores (numbers in parenthesis) exhibit how similar the distributional plots are, i.e., categories that can be grouped into one class.
The observed similarities were confirmed by the Kruskal–Wallis pairwise test. For the SB-FI and BF/Moth/Rove_N comparison, no class-specific variance discrimination (p > 0.05) was found for Kakamega. This same was true for the St. Lucia when comparing the independently predicted DF_FI to the BF/Moth/Rove_N outputs. For all D and N variable combinations, the value ranges were also higher in P than in the NP category. These differences in iBS values confirm the biodiversity integrity and human impacts, as realistically expected for P and NP areas.
The consistencies of the iBS across the two agro-ecological zones, despite dis-aggregating the metric along various taxonomic levels and using various human influence variables (i.e., FI and the Naturalness Index) shows the relevance of the metric as a reliable proxy for micro-scale biodiversity status. The metric values are comparable between taxa and sites despite different underlying landscape patterns. As stipulated, indicator insects can be used as a proxy for overall site-specific biodiversity status.
Discussion and conclusions
Biodiversity status measures or indicators need to be reproducible and moderately sensitive to drivers and consequences of biodiversity loss36. Moreover, these measures need to be easy to implement and robust enough to capture all inherent variations in data quality, even if not applicable to all species or purposes21. The metric developed herein can provide a framework to monitor localized micro-habitat-based biodiversity status conditions. As implemented, only one biodiversity component (insect assemblages) was considered.
The insect orders used herein (i.e., butterflies and moths) are considered sensitive to overall habitat changes as they require very specific habitats and microhabitat structures during larval development37. Micro-habitat quality and quantity can only be considered in a context where ecosystem structures are well discerned2. Our iBS metric made this possible by using EO techniques, since high-resolution and detailed ecosystem and landscape structure information could be considered and projected over wide areas. Furthermore, by pairing diversity-based species richness to spatially explicit naturalness information, we enhanced the iBS sensitivity to various tangible consequences of localized human impacts on biodiversity38. This is an advancement over sum of human pressure indicators such as the human footprint and/or the use of the Naturalness Index39 on its own as a proxy for biodiversity status.
Conceptually, the iBS is like the Biodiversity Intactness Index (BII)7 and the The Nature Index (NI) produced by the Norwegian Institute for Nature Research40. Both also relate human impact to species abundance data. Both the BII and the NI use biome-specific species abundances and relate this to average changes in biodiversity using high resolution land use data to predict biodiversity changes over time. As opposed to these species fraction estimates for specific land units (i.e., parametrically), the iBS has the advantage that it uses non parametric per pixel spectral features (i.e., abundances of greenness and wetness in each 20-m pixel) and high resolution tree heights data (from the Global Ecosystem Dynamics Investigation mission) to predict explicit insect species abundance and diversity patterns. This is then weighted by the human impact component using the 10-meter high-resolution naturalness index. The result (iBS) as a measure, and not indicator, for biodiversity status41. Our metric is suitable for characterizing the current localized biodiversity status baseline, without making use of an intact baseline reference site. Insect indicator species are specific to the overall biodiversity status of a localized site30. Furthermore, the use of taxonomic orders of indicator insects has the advantage that species-specific weights for human impact responses for certain ecosystems do not have to be pre-emptively determined40. Such species weights in ecosystem-based assessments can be biased. Some biases are even more problematic if they are based on expert knowledge and statistical confidence limits associated to expert’s reasoning42. Essentially, parametric assessments of relative species richness loss per ecosystem and expert judgment can lead to underestimation of biodiversity loss43.
In terms of applicability over any site and area coverage in Africa, our metric should be used with clear understanding that it relies on localized data. Regarding D, predictions would not be accurate in areas of low taxonomic data densities and evenness. Further, if used in areas with dramatic shift in climate regimes, the landscape-based species diversity models, as used herein, would produce inaccurate model outputs44. In our case, the assemblage diversity models are produced for each specific climate zone and area, and only then compared (Kakamega, humid climate; and St. Lucia, semi-humid). For both areas and climate zones, the Area Under Curve (AUC) prediction accuracies for D were >0.95 (+−0.02). The availability of the used EO data (spectral features from 10–20-meter Sentinel-2 data and 25-m GEDI tree heights) are not site-specific constraints. Due to this spatial explicitness (pixel-based), the metric values can be straightforwardly, and meaningfully averaged across various landscapes of interest (within a common agro-ecological zone) to effectively establish the overall zone-specific local biodiversity status. To increase its applicability and stability over wider regions, we recommend that the iBS be used in conjunction with regional and ecosystem-based biodiversity measures. For instance, it could be used as a site-specific reference for country-specific biodiversity intactness information such as the Natural Capital Index (NCI). The NCI uses region-specific bird and butterfly abundances (at various taxonomic levels) to estimate ecosystem quality45.
The replicability and the implementation of the iBS metric is aided using readily and freely available EO data and species assemblages from citizen science portals (i.e., iNaturalist.org). Both data sets are now more accessible than in the last decades. As more EO and species data becomes available, it can be effectively and increasingly integrated in spatial models for prospectively updating biodiversity status for various areas within common climate zones46. The development of big data processing platforms with advanced algorithms, such as Google Earth Engine, supports cloud-processing of EO data at planetary scales47. This is of particular importance for practitioners in developing countries, i.e., in so called data scarce environments and poorly studied areas of the world.
The policy domain has several demands regarding credible biodiversity status measures and indicators20. With the approach presented here, average values for a landscape unit or conservation area can be computed to determine its biodiversity status. This can be used to address policies that require address area under protection estimates or progress made in this regard48. The metric can be dis-aggregated and weighted to specific species of concern to assess vulnerability or extinction risks (e.g., Red Data Lists). As implemented herein, our results show that the iBS demonstrates a similar sensitivity to various insect sub-orders and families (e.g., stingless bees and dragonflies) (Fig. 3). Furthermore, given densely available species data, the metric allows for the rapid assessment of indicator groups or orders to provide a current baseline. This helps to ascertain conservation hot spots, including identifying last wild places49. Implementation of the Aichi Biodiversity Goals in Africa has shown that data deficiency gaps exist for undervalued species50, such as insects. Implementing the iBS can help to address these gaps in an effective way.
Methods
Characteristics and implementation of the metric
The metric is based on spatial distributions of insect assemblages—species, groups, or orders—as they occur within various habitats and land uses. As insect habitat demands are more-fine scaled than for plant or vertebrate communities, the measure is best implemented with data that more suitably characterizes insect micro-habitats. This includes data on vertical habitat structure, and spatially explicit data that can discern actual, as opposed to modeled, insect habitat suitability zones. Data would also have to consider habitat modifications by various human activities (including human impacts from past human activities). Ideally, spatially explicit habitat patterns would consider degraded versus non-degraded areas and land fragmentation patterns. Both are critical parameters of the micro-habitat status of insects and other taxa23. At the landscape level (<20-m grid cell or pixel resolution), spatial metrics can encompass micro-habitats of insect assemblages. For certain indicator species, generalists and species that are less tolerant to disturbances can be considered with appropriate spatial metrics3,12.
Within similar climate zones, and given that reliable data on taxa are used, biodiversity metrics or indicators should be scale appropriate in time and space. Moreover, the metric values should be comparable within a given local site, given the use of common measuring units and well-calibrated geospatial data. Given this scalability, taxonomic adaptability and computations include uncertainty; the measure can be a reliable proxy for overall and current localized biodiversity status (within common climate nesting zones). Thus, the iBS metric can help to address various conservation strategies at different political and administrative levels.
For insect assemblages i in agro-ecological zone j, the insect-based biodiversity status (iBS) metric is defined as;
Where iBS is the actual (current) insect-based biodiversity status and D denotes species diversity (species richness), predicted from 0 to 1, where 1 is the most species rich (diverse) space. Diversity/richness (D) includes local habitat structure and species composition. Landscape naturalness (N), which denotes various human impacts scaled from 0 to 1, where 1 refers the highest naturalness. This occurs through human impact manifestation from current and past human activities and disturbances. N is also sometimes referred to as human footprint51. Information from various existing sources can be used for both variables (i.e., for N, readily available human footprints data can also be used). The two variables can be expressed as percentages or proportions if common measuring scales are consistent. For insect micro-habitat monitoring, only high-resolution data for N is appropriate. We suggest the use of taxonomic orders that represent indicator species for ecosystem intactness. By using only two variables with given uncertainties, the metric is easy to understand.
The metric can be disaggregated for taxonomic levels of interests (i.e., International Union for Conservation of Nature red list species)52. This can be done by using insect or other occurrence data from any given taxonomic level, such as specific orders or families of interest. Moreover, weightings can also be straightforwardly applied by using exponents or multiplications for individual taxa of interest.
Climate variability is excluded by implementing the metric only within specific agro-ecological zones. Furthermore, local results from the iBS can be feasibly integrated into spatial species models that use bio-climate variables over wider areas for regional biodiversity pattern assessments, including future projections.
Sites and input data
Two sites, each about 2000 km2 in size, were selected to implement the metric. The sites lie in two agro-ecological zones (a semi-humid site in eastern South Africa, and a humid site in western Kenya). The iNaturalist platform (iNaturalist.org) was used to acquire open source and research graded (R) data on insect occurrence (species for a specific location). The export tool provided was used to export the tabular species data (as orders and sub-orders). The areas selected for this work show high species numbers (>1600 in all sites) and spatially balanced spreads (evenness) for the Lepidoptera order (butterflies and moths) and the Polyphaga sub-order (Rove beetles, some of which prefer semi-aquatic habitats). These (sub-) orders represent indicator species for overall landscape integrity.
For the two sites, freely available 10–20-m resolution European Space Agency (ESA) Sentinel-2 satellite data for 2019 to 2021 were acquired and cloud processed (see Description of methods and overall approach and Procedure sections below). Newly developed and freely available gridded 25-m tree height data metrics (Level 2A product) from the Global Ecosystem Dynamics Investigation (GEDI) mission (available only for 2019) were added as predictor bands53. Tree height and stand maturity is an important life cycle variable for the orders that were investigated54. All data sets are freely available through the cloud-based Google Earth Engine (GEE) data repository.
The Naturalness Index (N) consists of four human influence proxies namely, population density, land transformation, accessibility, and electrical power infrastructure that are measured with the geodata Gridded Population of the World v4.11, WorldCover V100, OpenStreetMap, and VIIRS Stray Light Corrected Nighttime Day/Night Band Composites Version 155, respectively. While the GPW, WorldCover and Night-time Lights data sets are available globally in the Google Earth Engine platform, OSM data were extracted manually from www.openstreetmap.org. For the two test sites and processed into the accessibility maps required by the fusion algorithm. The data were then combined using a heuristic data fusion approach described in39. The input geodata were converted to weights by applying preprocessing steps and summed up on a pixel level. The resulting product is then inversely normalized to the range [0;100] to form the Naturalness Index.
Description of the methods and overall approach
We implemented the metric using earth observation (EO) and other spatially explicit geo-spatial data for two agro-ecological/climate zones: Kakamega in western Kenya (humid climate) and St. Lucia in South Africa (semi-humid climate). Lepidoptera orders (butterflies (BF) and moths (Moth)) and Polyphaga sub-orders (rove beetles) were used as indicator species assemblages. The insect assemblages were sourced from the open source iNaturalist platform52. As predictors, 10–20-m resolution EO variable metrics on per pixel greenness (measurement value for chlorophyll active vegetation), wetness (measurement value for surface water), brightness (measurement value of interactions between soil and canopy moisture, usually sensitive to bare soil surfaces) were computed from Sentinel-2 time-line data. In addition, 25-m resolution per pixel tree heights were used to model D for the assemblages. For N, a 10-m Naturalness Index was predicted as described above. The Naturalness Index we produced measured the naturalness of land surfaces, defined not by biodiversity, flora, or fauna, but by the absence of human impacts. The spatial data frames corresponding to the D outputs were, respectively, used to compute the naturalness index.
For modeling D, a random forest species diversity regression model was used at both sites56. Random forest regressions are part of the ensemble learning concept, that uses decision trees to forecast values related to, in this case, species diversity of the used insect assemblages.. To avoid overfitting, we performed a k-fold cross validation technique57. Model performance was estimated for both sites using the Area Under Curve (AUC)58.The total species counts were separated into training and model evaluation data using a ratio of 30% (evaluation) to 70% (training data). The D and the N prediction results, as pixel raster layers, were then normalized using the formula described in the procedure section as number 10. Several arithmetic combinations of N and D were tested, and by visual inspection of the value distributions of the results (Supplementary Fig. 1 and Fig. 2) and descriptive histogram assessment (Supplementary Figs. 5, 6). Using visual inspection, a multiplication function rendered the most realistic results in terms of predicting biodiversity or conservation status. Multiplication was according to the mathematical formulae (1). The descriptive statistics showed that the product of the two variables produced results that included both the biotic component (D) and the human impact component, through N (Supplementary Fig. 3 and Fig. 4).
Our approach includes a statistical descriptive assessment of iBS value distributions over two different agro-ecological zones. Moreover, independent models were predicted for both sites using stingless bees (SB) (for Kakamega) and the dragonfly (DF) (for St. Lucia) insect assemblages. The comparison was performed to assess if our measure is realistic and useful to be used across various human impact and agro-ecological zones and robust enough when using various taxonomic rankings. Specifically, for Kakamega (humid agro-ecological zone), an existing pixel-explicit data set (250-meter resolution) predicting SB species distributional patterns was used59. The SB model exhibited AUC accuracies > 0.9 and various ecological predictors and tribe level insect occurrence data from 2017 to 2021 were used. The diversity model (variable D) for SB was amplified with a newly available global 30-m Forest Integrity (FI) data set60. SB_FI was produced according to the same mathematical function (1) that was used for the Lepidoptera/Polyphaga (sub-) orders. For St. Lucia, dragonflies (DF) sub-order assemblage data from the iNaturalist platform (N = 575) (collected from 2017 to current) were amplified with the FI data set, to produce DF_FI, also according to (1). Randomly sampled pixels (N = 50–500) were extracted from each model results, for Kakamega and St. Lucia, respectively. The random samples were extracted for functional land cover categories at each site. These were the protected (P) (national reserve with minimal human impact), the non-protected (NP) (areas around national reserves with various human impacts that are present in both St. Lucia wetland and Kakamega), and the agroforestry (AF, only for Kakamega) categories. AF constitutes a 5-km buffer around the Kakamega reserve. For each land cover category, site and insect assemblage grouping, diamond shaped distribution diagrams and associated statistics were computed. The statistics included land cover category means, standard deviation, standard errors, and confidence intervals. Moreover, the statistical differentiation between values attributed to these categories (from the category-based random sampling) was done using the Kruskal–Wallis rank sum test (with continuity correction)61 and by visualizing of the class-based variances. The Kruskal-Wallis test is used when the samples do not follow a normal distribution.
The statistical assessment per land cover category and taxonomic group aid to support the aggregation of the metric for functional categories or ecosystems. The error bars for the iBS values from various agro-ecological zones provide confidence in the credibility of the metric. Error bars in can be used to assess if biodiversity status ambitions can be reached, and with what associated uncertainty21.
Lastly, to furthermore, support aggregation over larger areas and create confidence in the scalability of the metric, various data gap assessments regarding the evenness of the insect occurrence data from the iNaturalist were performed (see Guidance on data and model requirements, and uncertainties of the approach). Specifically, for the (sub-) orders Lepidoptera/Polyphaga from the iNaturalist, point data densities from 2017 to current were computed for the two study sites using the total number of points divided by the entire area of the study site62. Using the same method, point weighted data density was also computed for the whole of Africa and going forward, assuming that more data will be available in future in the iNaturalist (current year + x). For the future projections, improvements of data density were assumed, within and alongside existing data points that are currently available in the iNaturalist. Furthermore, bootstrapped confidence limits for the used insect order Lepidoptera and the Polyphaga sub-order were computed to confirm the suitability of the two chosen sites in terms of insect occurrence data density. Lastly, 25% of the total insect occurrents data points were successively removed and the accuracy for D was re-computed with less data. This was done for both study sites to assess how data density affects model accuracies (for D). The data density computation and model sensitivity to the number of data point’s analysis were done to re-affirm the selection of the study sites and provide guidance on implementing the iBS elsewhere in Africa.
Procedure
-
1.
For one agro-ecological and/or climate zone, source orders, sub-orders or family level insect occurrence data from the iNaturalist platform (iNaturalist.org) (occurrence data from other portals can also be used). Assemblages of species that are indicators for landscape biodiversity and integrity should be used (e.g., butterflies, moths, beetles, and/or dragonflies)
-
2.
Review insect assemblages, orders or sub-orders, families or individual species data points and verify that these are well spread, and the area selected lies within one climate zone. An agro-ecological map can be overlaid to verify the climate nesting
-
3.
Using cloud-based services such as GEE (https://earthengine.google.com) or SEPAL (https://sepal.io/), source European Space Agency (ESA) 10–20-meter Sentinel-2 optical time-line data (at-sensor reflectance) that correspond (+−3 years) to the iNaturalist occurrence data
-
4.
Use the same cloud-platforms to compute tasseled cap spectral feature metrics on greenness, wetness, and brightness using the tasseled cap transformation coefficients for at-sensor data (see ref. 63), using 2–3 years of satellite timeline data. Use a median compositing approach and a cloud cover threshold of 25% to derive the spectral metrics representing the 2–3-year period64
-
5.
Source the L2A 25-m GEDI tree heights data65 set for the selected area in the cloud platforms, and geospatially resample the data to the Sentinel-2 spectral metrics.
-
6.
Perform a Variable Inflation Factor (VIF) and/or the Pearson correlation analysis to probe collinearity of the variables (10–20-m Sentinel-2 metrics and 25-m GEDI data)
-
7.
Use a machine learning algorithm, i.e., random forest with standard optimizations, to predict the order, sub-orders, species-based, or assemblages’ richness or diversity (D) using the non-collinear tasseled cap features and the 25-m GEDI tree heights as predictors for insect taxa occurrence. The results are insect diversity or richness data layers, termed D.
-
8.
Using a sub-set for the insect occurrence data (i.e., 30% of the training data), devise an accuracy score for the species-richness models (D), i.e., produce an area under curve (AUC) graph and statistics
-
9.
For the same area of interest, the Naturalness Index is computed (for N) using the approach described in ref. 39
-
10.
Rescale the D and N data layers using minimum and maximum value differential normalization; rescaled raster = [(raster − Min value from raster) / (Max value from raster − Min value from raster)]
-
11.
Apply multiplication on the normalized raster layers (D * N) for the area of interest to predict per pixel iBS
-
12.
Overlay geographical data layers for orientation and perform geo-statistical analysis as required, i.e., average iBS for a protected area or identify spatial biodiversity status patterns and conservation priority zones or help with identification of suitable sites for the establishment of protected areas
The following link also provides a link to the data and scripts used to implement the metrics at the two sites; https://dmmg.icipe.org/dataportal/dataset/measuring-insect-based-biodiversity-status-in-africa
Guidance on data and model requirements, and uncertainties of the approach
This section gives a stepwise and practical guideline on how to self-implement the iBS given data and model requirements, risks and applicability of the procedure over larger areas.
We recommend using four main data criteria and model requirement tests when implementing the model using butterfly, moths, and rove beetle orders (indicator taxa). The data requirements are according to the ecological requirements of the indicator taxa66.
Firstly, when using these indicator taxa/species, the iBS metric should only be implemented in the humid (the Kenyan site we used; Kakamega), and the semi-humid (the South African site we used; St. Lucia) agro-ecological zones according to67. Together with semi-arid and arid, these four zones represent major agro ecological zones in Africa. Numerous findings corroborate that the two selected agro-ecological zones are most suitable for the existence and survival of ecological indicator taxa used13. We illustrated that >90% of the area of the two sites we used lie within the two suitable agroecological zones (Supplementary Table 3, Supplementary notes 3). We compared our two sites to two other semi-arid areas that exhibit similar area sizes (Pendjari Park in Benin, and Kruger Park in South Africa). Although the two reference sites have permissible overall species data densities (determined through visual inspection), they lie outside the two suitable agroecological zones for the said taxa/species. Thus, the agro-ecological zoning criteria does not hold for them.
Secondly, we downloaded all available iNaturalists data for the respective assemblages in the study areas, including the above reference sites, and computed the data densities. We used the density scores to confirm if the species richness model (variable D according to 1) can feasibly be implemented. The occurrence datasets for butterflies, moth and rove beetles in csv format were converted to GIS point feature shapefiles. Point density calculation allows fitting two models where the first assumes that the process intensity is a function of covariates while the second (null hypotheses) assumes process intensity is not a function of any covariate68. The null hypothesis is calculated by;
For individual areas, points showing spatial locations are treated as the response variables and any other attribute information associated with them called marks are the types of data used. The outcome is basically the observed density of occurrence points within the study area. The R software (spatstat and maptools packages) was used for the point pattern analysis. The study area shapefiles were loaded, converted into a polygon feature (object of class owin) and attached with the point features to explicitly define point pattern boundaries. By default, point density analysis uses meters units. So, both points and polygons are rescaled to km.The null model is thus calculated as:
where H0 represents the null hypothesis, ρ signifies the point density, n is the number of observed points and A is the area of the study site.
The area weighted point data densities (Supplementary Table 4) further confirmed the suitability of the two study sites used.
The third data criteria were to test the effect the total number of occurrence points had on the data density. This serves to confirm the density scores (Supplementary Table 4) and re-affirm site suitability for implementing the diversity/richness model (variable D). For each site, the number of points for modeling was randomly reduced by 50% and 75%, expressed as percentage of points left (percentages, first column in Supplementary Table 5), and the point densities were calculated respectively (according to the above formulas). The first data row represents 100% of the insect data from the iNaturalist that was available and used in this study for insect assemblages
If 50% of the total number of points available in the iNaturalist for the said taxa assemblages for Kakamega (total n = 550) were to be removed, the data density score drops by a large degree (from 0.32 to 0.16). For St. Lucia, 50% of the points would still render a data density score of 0.42, however, for both reference areas the density scores decreased (to near zero) when 50% or 75% of the available data points are removed (Supplementary Table 5).
Point density criteria is important in this metric because of the small-scale nature of the habitats of the insect species being used. Thus, density scores are an important consideration in insect diversity/richness models as they are linked to localized habitat suitability. Insect habitat suitability (from species diversity models), furthermore, is linked to biodiversity or conservation status. Also, low data densities reduce species diversity model accuracies. For instance, in Kakamega in Kenya, a high AUC score was attained (AUC = 0.95) when 100% of occurrence points were used but diversity model performed was poor (AUC = 0.56) when only 25% of the occurrence data was used (Supplementary Table 6).
The fourth data criteria were canopy height and water availability. Butterflies and moths rely on trees in every phase of their entire life cycle. Majorly, they require mature trees during their larval stage development to increase their chances of survival, shelter, and protection69. Due to this reliance, tree height data was then factored in as a major factor of habitat suitability criteria. In addition, according to research done by70, insect species abundance and density were positively correlated with high canopy cover, high tree density and, closeness to water bodies. Rove beetles, specifically, require intact ecosystems often located next to freshwater habitats. As a result, study area suitability was furthermore assessed by the availability of water bodies within the area and if >50% of the area exhibits tree canopy heights > 3 m (using the GEDI tree heights data). Compared to the two reference sites, Kakamega and St Lucia fulfilled these criteria (Supplementary Table 7).
AUC accuracy scores for the insect assemblage-based diversity models (variable D) showed AUC > 0.95 for the two sites investigated, while the two reference sites, Pendjari National Park in Benin, and Kruger National Park in South Africa, had lower accuracies of 0.49 and 0.48 respectively. The lower accuracies for the two reference sites are due to lower iNaturalists data densities, the lower fractional coverages of mature trees at these sites (Supplementary Table 7), and the fact that the reference sites are characterized as semi-arid (Supplementary Table 3).
Finally, a filter encompassing the two most important input data criteria and requirements (agro-ecological zones and data densities from the iNaturalist), relevant to the used insect assemblages, was applied to Africa (Fig. 4). Using this, suitable areas in which the iBS can be implemented were identified on a map (iii in Figure a). In b, Africa-wide future suitability is illustrated (current year + x) (iii in Figure b), under the assumption that more insect occurrence data is collected in future (iNatulalist data density for insect species increases). Biodiversity data densities within citizens’ portals are expected to increase around or within areas already well sampled71. The suitability is shown separately as an inset for the two study sites and one of the reference areas (Kruger National Park in South Africa), respectively, for current (a) and for future data densities (b). Even with increasing data density, the Kruger National Park area will not be suitable since it is in the semi-arid agro-ecological zone. For all of Africa, the iBS implementation suitability using these insect taxa is currently 5.2% (iii in a, Fig. 4) of the total land mass (excluding Madagascar), while this may increase to 8.8% in the future scenario model (iii in b).
Part a is the current implementation suitability for the iBS (gray areas) and b is the future implementation suitability (gray areas) for various data criteria, that are shown in images i–iii, respectively. Specifically, images i, in both figures (a, b), illustrate the suitable agro-ecological zones (semi- humid and humid), and images ii illustrate the iNaturalist point data densities for the used assemblages (as per km2 grid cell). Images iii represent the overlay products of the two criteria given the current data densities (a), and based on anticipated (future) data densities (b). The zoomed areas, forth images in both (a) and (b), show the current and future implementation suitability results for the two study areas (extracted from iii) in comparison to the Kruger National Park reference area (also extracted as a zoomed inset from iii).
Other biases associated with the iNaturalist data and site selection that users should consider are dealt with from here onwards.
According to research28,72 concerns about potential biasness in collated biological datasets from sites such as the iNaturalist are mainly around uneven sampling efforts over space and time and uneven species detectability.
Apart from the data density assessment, the bias on uneven detectability was herein addressed by selecting a wide range of species within the order Lepidoptera and the sub-order Polyphaga. Grouped, they have been proven to be generally good indicator species instead of using a specific single species.
In this regard, proper selection of study sites is important. Both sites, selected herein, are important habitats for a wide range of butterflies, moths, and rove beetles73,74. Kakamega forest, is a natural reserve and one of the most continuous rainforests in Kenya. Due to its different habitat types with different ecological conditions, it harbors a large abundance of insect communities, including specifically butterflies and moths. Despite rapid human modification in the last decade, the Kakamega forest itself is still largely intact75. The St. Lucia site comprises areas of high human impact, coastal forests, and wetlands including Lake St Lucia which lies within the iSimangaliso wetland park. St, Lucia is an insect biodiversity hotspot according to UNESCO76.The St. Lucia site is characterized by a large beetle abundance and diversity, including rove beetles77.
Butterflies, moths, and rove beetles are suitable ecological indicator taxa for ecological intactness, and as stipulated found in great abundance in both sites78. According to ref. 79, areas highly endowed with butterflies and moths are highly likely also to be rich in other macroinvertebrates.
To further probe possible sampling bias in the iNaturalist insect data for both study sites, we calculated the species diversity scores and bootstrapped confidence limits for the used insect order Lepidoptera and the Polyphaga sub-order. The Shannon-Weiner Index (H’), Brillouin Index (HB) and the Simpson’s Index (λ) are three major indices published in measuring data specific species diversity in ecology. Based on randomness of insects’ data on both sites, the Brillouin Index, which is an improvement of the Shannon Index, was used to calculate the diversity scores as data collection randomness is unguaranteed.
The formula for Brillouin Index is;
For instance, the diversity scores for order Lepidoptera for sites Kakamega (Kenya) and St Lucia (South Africa) were 3.9 and 4.9, respectively (Supplementary Table 8). We also calculated uncertainty of diversity scores by calculating the bootstrap resampling method. We performed a 1000 resampling on diversity scores calculated using the Brilloiun’s index formula80. The bootstrap method estimates confidence intervals and uncertainty at site level (alpha diversity) without the need of assuming the normal or non-normal distribution of occurrence data from bio-collections such as the iNaturalist81. A 95% confidence interval, which indicates the range of values that are likely to include the true diversity value with a 95% probability was calculated. Our diversity scores for both insect taxa orders and sites lie within the confidence intervals insinuating high precision of estimates.
To overcome temporal bias, only the most recent iNaturalist insect occurrence data (from 2017 to 2022) were used for diversity modeling (D variable), for both sites. The assumption was made that in the last 5 years, ecological transformation in the two sites was minimal.
Other troubleshooting
Possible problems that can be encountered are listed, with possible reasons and solutions.
-
Problem: In areas with large areal coverage of managed forests, the species diversity results may be biased towards these sites, specifically in semi-arid areas. The reasons are that managed forest in singular year data (satellite metrics based on one or two years only), exhibit spectral responses like intact forest, thus the model would predict these areas as highly suitable or diverse. The solution is to increase the satellite data compositing period (>2 years). This would normalize forest management practices such as clearing and re-growth periods that are not found in intact forests.
-
Problem: For the calculation of the Naturalness Index (for N), accessibility maps processed from OSM data are a crucial input source. These OSM data need to be manually imported into Google Earth Engine, imperiling the end-to-end formation of N. Although this is a bottleneck, there are third-party libraries addressing this specific limitation and facilitate working pipeline in which the OSM data are seamlessly integrated, namely OSMnx (https://osmnx.readthedocs.io/en/stable/) and geemap (https://geemap.readthedocs.io/en/latest/).
Time taken
For predicting D, an experienced operator takes about 12 h for one site being about 25% of one Sentinel-2 tile (2000 km2). The most time is spent on geo-location of the 25-meter GEDI to the 10-m Sentinel-2 observations (metrics), and optimizing the tasseled cap transformations composites, based on cloud cover contamination over the observation/compositing period. For N, the processing time taken for areas as used in this study is estimated to be about 1 h. However, since the Naturalness Index is a measure of naturalness from various anthropogenic factors, the geo-data sources used to infer N are site-specific and vary in file size from area to area. This variability arises mainly due to the file size of the OSM data within the area of interest. Essentially, if the area lies in a less populated area (less human influence, thus less artificial structures), less processing time is used.
The iBS computation itself is performed in <1 min in any given cloud-processing or geospatial data processing software.
Anticipated results
The procedure is anticipated to produce accurate insect-micro-habitat-based biodiversity status results for individual areas across Africa, given insect data availability. Since some insect assemblages, like the once used herein, are indicator species for landscape integrity, the results are expected to ascertain actual and overall biodiversity status of a landscape. Also, there is little sense in monitoring individual species to assess the conservation status of specific ecosystems since some species may be less or more adaptable to global change effects. Within the local sites where the iBS is modeled, the metric values can then be aggregated for features of interest, such as protected areas. Between sites, landscape feature-specific biodiversity status can be compared and used to monitor progress towards conservation objectives or to identify current areas of concerns (i.e., land degradation sites within protected areas).
The main intermediate results, before computing the iBS, are the tasseled cap transformations modeled on the Sentinel-2 optical time series data, the species richness (D) outputs, based on species assemblages, and the Naturalness Index (N) for the corresponding area of interest. The number of insect occurrences points used in our case were as follows; n = 646 for the insect order assemblages (Lepidoptera and Polyphaga orders) at the Kakamega site in western Kenya and n > 1600 for the order assemblages (Lepidoptera order and Polyphaga orders) at the St. Lucia site in South Africa. For the diversity models (D), specifically, the VIF scores before performing the model run are the most important statistics needed. VIF is a measure of collinearity between the independent variables. In our case, VIF scores for all variable combinations were <5, meaning the variables were non-collinear. Regarding the overall accuracies of the random forest-based diversity models (for D), model AUC scores >0.95 could be attained for the two insect assemblages’ models. These scores indicate high statistical accuracy of the models60. The random forest model analysis also showed that the 25-m GEDI tree heights layer was the most relevant predictor, over the 10–20-m Sentinel-2 spectral tasseled cap predictors.
Through visual inspection, the dragonfly spatial diversity/richness patterns (D) matched well with the Lepidoptera/rove beetle diversity/richness results for the St. Lucia site (South Africa, semi humid climate). Likewise, the stingless tribe diversity models (D) matched well with the Lepidoptera/Polyphaga results for Kakamega (Kenya, humid climate). High accuracies (AUC > 0.95) were also attained for the two independent D outputs (SB_FI and DF_FI).
As evident, two different iBS reference insect diversity models were produced at the two areas, i.e., in Kakamega a SB-FI model was produced, while in St. Lucia the DF-FI model was produced. This is expected dueto unequal taxa data distributions across the two sites, i.e., in St. Lucia enough DF data points were available, while this site did not have sufficient data for SB. Thus, data availability and spread for D as well as model accuracy has ramifications for which sites to model and the scalability of the procedure. Furthermore, in our case, we anticipated that the D results would be moderately accurate if only Polyphaga sub-orders were used. Thus, we combined the occurrence data for the Polyphaga sub-order with the Lepidoptera order data (as assemblages). The Naturalness Index results (N) are less affected by input data variances, since the method relied on mostly globally homogeneous, not site-specific geodata, such as population density maps, land cover maps, and night-time lighting maps. The only exception is formed by the accessibility maps created from public OSM data. In this case, studies have shown increasing lack of quality and completeness for less developed regions of the world82.
It can be anticipated that, as evident from the iBS value distributions belonging to the P, NP and AF land cover categories, the iBS value distributions for the various land cover categories are statistically similar. This is irrespective of the agroecological zone.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The iBS outputs, input satellite feature maps and other intermediate data can be accessed by using the following Data Warehouse (CKAN) link–https://dmmg.icipe.org/dataportal/dataset/measuring-insect-based-biodiversity-status-in-africa. All insect taxa data can be sourced from the iNaturalist.org portal. The Naturalness Index data used in this study is freely available online at: https://doi.org/10.5281/zenodo.7323837.
Code availability
The custom produced code for the iBS is available through the following Data Warehouse (CKAN) link https://dmmg.icipe.org/dataportal/dataset/measuring-insect-based-biodiversity-status-in-africa. The tasseled cap processing code is available through the same warehouse link. QGIS version 3.16.16 was used to implement the iBS code; the tasseled cap code is implementable through the Google earth Engine platform, the current online version. No specific parameters are needed to run the codes.
References
Ceballos, G. et al. Accelerated modern human-induced species losses: entering the sixth mass extinction. Sci. Adv. 1, 9–13 (2015).
Habel, J. C., Schmitt, T., Gros, P. & Ulrich, W. Breakpoints in butterfly decline in Central Europe over the last century. Sci. Total Environ. 851, 158315 (2022).
Seibold, S. et al. Arthropod decline in grasslands and forests is associated with landscape-level drivers. Nature 574, 671–674 (2019).
Kumsa, T. & Ballantyne, G. I Nsect pollination and sustainable agriculture in S Ub. J. Pollinat. Ecol. 27, 36–46 (2021).
Teklewold, H., Kassie, M., Abro, Z., Mulungu, K. & Sevgan, S. The role of pollination services and disrupting cropping patterns in closing nutrition gap in Sub-Saharan Africa. Int. Assoc. Agric. Econ. https://doi.org/10.22004/ag.econ.315241 (2021).
Stein, K. et al. Bee pollination increases yield quantity and quality of cash crops in Burkina Faso, West Africa. Sci. Rep. 7, 1–10 (2017).
Hill, S. L. L. et al. Worldwide impacts of past and projected future land-use change on local species richness and the Biodiversity Intactness Index. Preprint at bioRxiv https://doi.org/10.1101/311787 (2018).
Frank, A. et al. Human actions alter tidal marsh seascapes and the provision of ecosystem services. Estuaries Coasts 44, 1628–1636 (2021).
Samways, M. J. et al. Solutions for humanity on how to conserve insects. Biol. Conserv. 242, 10847 (2020).
Talašová, A. et al. High degree of philopatry is required for mobile insects used as local indicators in biodiversity studies. Ecol. Indic. 94, 99–103 (2018).
Betts, M. G. et al. Extinction filters mediate the global effects of habitat fragmentation on animals. Science 366, 1236–1239 (2019).
Habel, J. C., Samways, M. J. & Schmitt, T. Mitigating the precipitous decline of terrestrial European insects: requirements for a new strategy. Biodivers. Conserv. https://doi.org/10.1007/s10531-019-01741-8 (2019).
de Bello, F. et al. Towards an assessment of multiple ecosystem processes and services via functional traits. Biodivers. Conserv. 19, 2873–2893 (2010).
Roque, F. D. O. et al. Warning signals of biodiversity collapse across gradients of tropical forest loss. Sci. Rep. 8, 1–7 (2018).
Hudson, L. N. et al. The database of the PREDICTS (Projecting Responses of Ecological Diversity In Changing Terrestrial Systems) project. Ecol. Evol. 7, 145–188 (2017).
Montgomery, G. A., Belitz, M. W., Guralnick, R. P. & Tingley, M. W. Standards and best practices for monitoring and benchmarking insects. Front. Ecol. Evol. 8, 579193 (2021).
Vickery, M. Butterflies as indicators of climate change. Sci. Prog. 91, 193–201 (2008).
Hoban, S. et al. Genetic diversity targets and indicators in the CBD post-2020 Global Biodiversity Framework must be improved. Biol. Conserv. 248, 108654 (2020).
GEO BON. What are EBVs? – GEO BON. https://geobon.org/ebvs/what-are-ebvs/.
Kissling, W. D. et al. Towards global data products of Essential Biodiversity Variables on species traits. Nat. Ecol. Evol. 2, 1531–1540 (2018).
Scholes, R. J. & Biggs, R. A biodiversity intactness index. Nature 434, 45–49 (2005).
Purvis, A. et al. Modelling and Projecting the Response of Local Terrestrial Biodiversity Worldwide to Land Use and Related Pressures: The PREDICTS Project. Adv. Ecol. Res. 58, 201–241 (2018).
Hill, S. L. L. et al. Reconciling biodiversity indicators to guide understanding and action. Conserv. Lett. 9, 405–412 (2016).
Ledger S. E. H. et al. Rewilding Europe. https://www.rewildingeurope.com/wp-content/uploads/publications/wildlife-comeback-in-europe-2022/index.html (2022).
Chase, J. M. et al. Embracing scale-dependence to achieve a deeper understanding of biodiversity and its change across communities. Ecol. Lett. 21, 1737–1751 (2018).
Mokany, K., Ware, C., Woolley, S. N. C., Ferrier, S. & Fitzpatrick, M. C. A working guide to harnessing generalized dissimilarity modelling for biodiversity analysis and conservation assessment. Glob. Ecol. Biogeogr. 31, 802–821 (2022).
Noss, R. F. & Cooperrider, A. Saving nature’s legacy: protecting and restoring biodiversity. Choice Rev. Online, https://doi.org/10.5860/choice.32-2131 (1994).
Ferrier, S., Manion, G., Elith, J. & Richardson, K. Using generalized dissimilarity modelling to analyse and predict patterns of beta diversity in regional biodiversity assessment. Divers. Distrib. 13, 252–264 (2007).
de Castro-Pardo, M., Martín Martín, J. M. & Azevedo, J. C. A new composite indicator to assess and monitor performance and drawbacks of the implementation of Aichi Biodiversity Targets. Ecol. Econ. 201, 107553 (2022).
Asbeck, T., Großmann, J., Paillet, Y., Winiger, N. & Bauhus, J. The use of tree-related microhabitats as forest biodiversity indicators and to guide integrated forest management. Curr. For. Rep. 7, 59–68 (2021).
Palma, A. De et al. Annual changes in the Biodiversity Intactness Index in tropical and subtropical forest biomes, 2001 – 2012. Sci. Rep. 1–13, https://doi.org/10.1038/s41598-021-98811-1 (2021).
Mammola, S. et al. Towards evidence-based conservation of subterranean ecosystems. Biol. Rev. 97, 1476–1510 (2022).
da Silva, P. M., Aguiar, C. A. S., de e Silva, I. F. & Serrano, A. R. M. Orchard and riparian habitats enhance ground dwelling beetle diversity in Mediterranean agro-forestry systems. Biodivers. Conserv. 20, 861–872 (2011).
Philpott, S. M. et al. Local and landscape drivers of carabid activity, species richness, and traits in urban gardens in coastal california. Insects 10, 112 (2019).
Jew, E. K. K., Loos, J., Dougill, A. J., Sallu, S. M. & Benton, T. G. Butterfly communities in miombo woodland: biodiversity declines with increasing woodland utilisation. Biol. Conserv. 192, 436–444 (2015).
van Hinsberg, A., van der Hoek, D. J., de Heer, M., & ten Brink, B. Informing Policy-makers about changes in Biodiversity. Mapping and Monitoring of Natural Areas in the Nordic Countries- Proceedings from the workshop, (Fuglsø, Denmark, 2002)
Habel, J. C., Teucher, M., Ulrich, W., Bauer, M. & Rödder, D. Drones for butterfly conservation: larval habitat assessment with an unmanned aerial vehicle. Landsc. Ecol. 31, 2385–2395 (2016).
García-Vega, D. & Newbold, T. Assessing the effects of land use on biodiversity in the world’s drylands and Mediterranean environments. Biodivers. Conserv. 29, 393–408 (2020).
Ekim, B., Dong, Z., Rashkovetsky, D. & Schmitt, M. The naturalness index for the identification of natural areas on regional scale. Int. J. Appl. Earth Obs. Geoinf. 105, 102622 (2021).
Certain, G. et al. The nature index: A general framework for synthesizing knowledge on the state of biodiversity. PLoS ONE 6, e18930 (2011).
Faith, D. P. Threatened species and the potential loss of phylogenetic diversity: conservation scenarios based on estimated extinction probabilities and phylogenetic risk analysis. Conserv. Biol 22, 1461–1470 (2008).
Nielsen, A. L., Shearer, P. W. & Hamilton, G. C. (b8) peach: (2007).
Martin, E. A. et al. The interplay of landscape composition and configuration: new pathways to manage functional biodiversity and agroecosystem services across Europe. Ecol. Lett. 22, 1083–1094 (2019).
Williams, B. K. & Brown, E. D. Technical challenges in the application of adaptive management. Biol. Conserv. 195, 255–263 (2016).
Czúcz, B. et al. Using the natural capital index framework as a scalable aggregation methodology for regional biodiversity indicators. J. Nat. Conserv. 20, 144–152 (2012).
Rapacciuolo, G., Young, A. & Johnson, R. Deriving indicators of biodiversity change from unstructured community-contributed data. Oikos 130, 1225–1239 (2021).
Zhang, X., Zhou, Y. & Luo, J. Deep learning for processing and analysis of remote sensing big data: a technical review. Big Earth Data 00, 1–34 (2021).
Carroll, C. & Noss, R. F. How percentage-protected targets can support positive biodiversity outcomes. Conserv. Biol. 36, 1–10 (2022).
D’Souza, M. L. et al. Biodiversity baselines: tracking insects in Kruger National Park with DNA barcodes. Biol. Conserv. 256, 109034 (2021).
Hochkirch, A. et al. A strategy for the next decade to address data deficiency in neglected biodiversity. Conserv. Biol. 35, 502–509 (2021).
Venter, O. et al. Global terrestrial human footprint maps for 1993 and 2009. Sci. Data 3, 1–10 (2016).
Bland, L. M. et al. Toward reassessing data-deficient species. Conserv. Biol. 31, 531–539 (2017).
Schneider, F. D. et al. Towards mapping the diversity of canopy structure from space with GEDI. Environ. Res. Lett. 15, 115006 (2020).
Nyafwono, M., Valtonen, A., Nyeko, P., Owiny, A. A. & Roininen, H. Tree community composition and vegetation structure predict butterfly community recovery in a restored Afrotropical rain forest. Biodivers. Conserv. 24, 1473–1485 (2015).
Elvidge, C. D., Baugh, K., Zhizhin, M., Hsu, F. C. & Ghosh, T. VIIRS night-time lights. Int. J. Remote Sens. 38, 5860–5879 (2017).
Rigatti, S. J. Random Forest. J. Insur. Med. 47, 31–39 (2017).
Manorathna, R. (PDF) k-fold cross-validation explained in plain English (For evaluating a model’s performance and hyperparameter tuning). https://www.researchgate.net/publication/348237224_k-fold_cross-validation_explained_in_plain_English_For_evaluating_a_model ’s_performance_and_hyperparameter_tuning (2020).
Mahendran, N. et al. Sensor-assisted weighted average ensemble model for detecting major depressive disorder. Sensors 19, 4822 (2019).
Makori, D. M. et al. The use of multisource spatial data for determining the proliferation of stingless bees in Kenya. GIScience Remote Sens. 59, 648–669 (2022).
Wu, S., Flach, P. & Ferri, C. An improved model selection heuristic for AUC. Lect. Notes Comput. Sci. (including Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinformatics) 4701 LNAI, 478–489 (2007).
Ostertagová, E., Ostertag, O. & Kováč, J. Methodology and application of the Kruskal-Wallis test. Appl. Mech. Mater. 611, 115–120 (2014).
Gómez-Rubio, V. Spatial point patterns: methodology and applications with R. J. Stat. Softw. 75, 1–6 (2016).
Shi, T. & Xu, H. Derivation of tasseled cap transformation coefficients for sentinel-2 MSI at-sensor reflectance data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 12, 4038–4048 (2019).
Corbane, C. et al. A global cloud free pixel- based image composite from Sentinel-2 data. Data Br. 31, 105737 (2020).
Potapov, P. et al. Mapping global forest canopy height through integration of GEDI and Landsat data. Remote Sens. Environ. 253, 112165 (2021).
Pearson, D. L. Selecting indicator taxa for the quantitative assessment of biodiversity on JSTOR. Philos. Trans. Biol. Sci. 345, 75–79 (1994).
RCMRD. Africa Agroecological Zones — GeoNode. http://geoportal.rcmrd.org/layers/servir%3Aafrica_agroecological_zoning.
Bivand, R. S., Pebesma, E. & Gómez-Rubio, V. Applied Spatial Data Analysis with R: Second Edition. Appl. Spat. Data Anal. with R Second Ed. 1–405, https://doi.org/10.1007/978-1-4614-7618-4 (2013).
Atlanta, T. Butterfly Garden: How Butterflies Rely on Trees | Trees Atlanta. https://www.treesatlanta.org/news/butterfly-garden-how-butterflies-rely-on-trees/.
Rija, A. A. Local habitat characteristics determine butterfly diversity and community structure in a threatened Kihansi gorge forest, Southern Udzungwa Mountains, Tanzania. Ecol. Process. 11, 13 (2022).
Knape, J., Coulson, S. J., van der Wal, R. & Arlt, D. Temporal trends in opportunistic citizen science reports across multiple taxa. Ambio 51, 183–198 (2022).
Isaac, N. J. B. & Pocock, M. J. O. Bias and information in biological records. Biol. J. Linn. Soc. 115, 522–531 (2015).
Mitchell, N., Schaab, G. & Wägele, J. W. Kakamega Forest ecosystem: an introduction to the natural history and the human context | Request PDF. Karlsruher. Geowiss. Schr. R 17, 5 (2009).
Perissinotto, R., Bird, M. S. & Bilton, D. T. Predaceous water beetles (Coleoptera, Hydradephaga) of the Lake St Lucia system, South Africa: biodiversity, community ecology and conservation implications. ZooKeys 595, 85–135 (2016).
Holstein, J. & Haas, F. Insects of Kakamega Forest. (2015).
KwaZulu-Natal. iSimangaliso Wetland Park - UNESCO World Heritage Centre. https://whc.unesco.org/en/list/914/ (1999).
Perissinotto, R., Miranda, N. A. F., Raw, J. L. & Peer, N. Biodiversity census of Lake St Lucia, iSimangaliso Wetland Park (South Africa): Gastropod molluscs. Zookeys 1, https://doi.org/10.3897/ZOOKEYS.440.7803 (2014).
Rákosy, L. & Schmitt, T. Are butterflies and moths suitable ecological indicator systems for restoration measures of semi-natural calcareous grassland habitats? Ecol. Indic. 11, 1040–1045 (2011).
Hoose, N. van. Butterflies and plants evolved in sync, but moth ‘ears’ predated bats – Research News. https://www.floridamuseum.ufl.edu/science/butterflies-plants-evolved-in-sync-but-moth-ears-predated-bats/ (2019).
Peet, R. K. Relative Diversity Indices. Ecology 56, 496–498 (1975).
Kehs, A. et al. From village to globe: a dynamic real-time map of african fields through plantvillage. Front. Sustain. Food Syst. 5, 124 (2021).
Ribeiro, A. & Fonte, C. C. A methodology for assessing openstreetmap degree of coverage for purposes of land cover mapping. ISPRS Ann. Photogramm. Remote Sens. Spat. Inf. Sci. 2, 297–303 (2015).
Acknowledgements
The authors gratefully acknowledge the financial support for this research by the following organizations and agencies: Center for International Migration and Development (CIM), Fund for Human Capacity Development with Partners from International Agricultural Research (PIAF), Deutsche Gesellschaft für Internationale Zusammenarbeit (GIZ) GmbH (18.2085.1-003.03); the Swedish International Development Cooperation Agency (Sida); the Swiss Agency for Development and Cooperation (SDC); the Australian Centre for International Agricultural Research (ACIAR); the Federal Democratic Republic of Ethiopia; and the Government of the Republic of Kenya. Michael Schmitt and Burak Ekim acknowledge the support by the German Research Foundation (Deutsche Forschungsgemeinschaft - DFG) as part of the project SCHM 3322/4-1 – MapInWild. We acknowledge the contributions made several individuals who contributed to the insect taxa data on the iNaturalist platform. The views expressed herein do not necessarily reflect the official opinion of the donors.
Author information
Authors and Affiliations
Contributions
T.L. conceptualized the idea and designed the experiment, analyzed data, and wrote and revised the manuscript. M.S. and B.E. performed coding, data analysis and map making to predict the Naturalist Index and edited the manuscript. J.V. helped in designing the experiment and the algorithm and edited the manuscript. F.A. performed coding, data collation and processing, analyses and map making. J.H. edited the manuscript regarding insect physiology and ecology and advised on the experiment set up. H.T. gave guidance on the overall concept, data science and performed writing and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth & Environment thanks Rob Cooke and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Aliénor Lavergne. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Landmann, T., Schmitt, M., Ekim, B. et al. Insect diversity is a good indicator of biodiversity status in Africa. Commun Earth Environ 4, 234 (2023). https://doi.org/10.1038/s43247-023-00896-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43247-023-00896-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.