Abstract
Nitrous oxide (N2O) is important to the global radiative budget of the atmosphere and contributes to the depletion of stratospheric ozone. Globally the ocean represents a large net flux of N2O to the atmosphere but the direction of this flux varies regionally. Our understanding of N2O production and consumption processes in the ocean remains incomplete. Traditional understanding tells us that anaerobic denitrification, the reduction of NO3− to N2 with N2O as an intermediate step, is the sole biological means of reducing N2O, a process known to occur in anoxic environments only. Here we present experimental evidence of N2O removal under fully oxygenated conditions, coupled with observations of bacterial communities with novel, atypical gene sequences for N2O reduction. The focus of this work was on the high latitude Atlantic Ocean where we show bacterial consumption sufficient to account for oceanic N2O depletion and the occurrence of regional sinks for atmospheric N2O.
Similar content being viewed by others
Introduction
N2O is now the 3rd most important greenhouse gas1 and number one stratospheric ozone depleting compound2. Constraining the magnitude of production and consumption processes is therefore essential to understand how the biotic environment contributes to atmospheric concentrations3. Sixty percent of the increasing atmospheric concentration has a natural source, approximately 30% of which is produced in coastal and oceanic waters4. Canonical understanding of the nitrogen cycle tells us that the only biological process that consumes N2O is the final stage of denitrification5 where N2O is reduced to N2 under anoxic conditions by the enzyme nitrous oxide reductase (N2OR), encoded by the nosZ gene6.
Whilst the global ocean is considered on balance to represent a strong source of N2O to the atmosphere4, this source is not uniformly distributed. Regions of the northern and southern polar waters are often undersaturated and provide a sink for atmospheric N2O7,8,9,10. Indeed from the limited number of observations made, it appears that undersaturation is the prevailing condition for the ice-free surface waters of the Arctic11,12,13. Zhan et al.10,13 argued that the extant undersaturation must be driven solely by physical processes, whilst Verdugo et al.14 proposed potential biological mechanisms.
Experimental evidence indicates the potential for dissolved N2O to be biologically consumed, even when oxygen is replete. The bacteria Paracoccus pantotrophus (previously Thiosphaera pantotropha) and Pseudomonas stutzeri have been shown under laboratory conditions to utilise both nitrate and oxygen as terminal electron acceptors in respiration during a process termed aerobic denitrification15,16. Aerobic denitrifiers have now been observed in diverse environments which include freshwater17 and saline18 wastewater treatments, in soils19 and in coastal marine sediments20. Whilst there are no reported observations of N2O consumption in oxygenated ocean waters, several authors have indicated the presence of microbial communities with the potential to reduce N2O21,22,23,24,25,26. In the Arabian Sea, Wyman et al.24 found a close affiliation between a group of alphaproteobacteria expressing nosZ and Trichodesmium, a colonial, nitrogen fixing cyanobacterium. Coates and Wyman21 extended the geographical coverage of these observations to include the tropical and sub-tropical Red Sea, Atlantic and Indian Oceans, and hypothesised denitrifying activity associated with anoxic microsites within the cyanobacterial colony. Raes et al.23 reported nosZ genes associated with Rhodobacteraceae in oxic waters of the south-eastern Indian Ocean whilst Farias et al. reported the potential for cultured and natural diazotrophs to consume N2O, though under conditions far in excess of ambient concentrations22.
The nosZ gene occurs as two equally abundant but distinct clades, one of which had previously been unaccounted for with respect to the reduction of N2O27. The well characterised conventional denitrifiers of the alpha-, beta- and gamma- proteobacteria are grouped as Clade I, whereas Clade II organisms possess the novel, atypical nosZ gene sequences that are found in organisms previously not associated with denitrification, including Firmicutes, CFB (Cytophaga-Flavobacteria-Bacteroides) supergroup and Verrumicrobia. Sun et al.26 showed that Clade II genes were particularly abundant in oxygen minimum zones (OMZs), even in surface waters, and suggested that they might be associated with aerobic N2O reduction. Both Clades I and II of the nosZ gene were detected in several geographical regions associated with OMZ of the Eastern Tropical Pacific and the Arabian Sea, and were also observed in oxygenated surface waters of the Arctic and Southern Oceans28. Neither N2O production rates nor the distribution of N2O consuming microbes has been investigated in the high latitude waters represented by this study.
There are regions of the polar oceans which are undersaturated in N2O relative to the atmosphere and therefore offer a sink to atmospheric N2O, this is recognised10,13 but not wholly explained. During two high latitude research expeditions in northern and southern waters of the Atlantic Ocean, including those presented in28 we performed incubation experiments and collected samples which indicated that N2O was consumed and that both Clade I and II denitrifiers were present in near surface waters. In the North Atlantic, measurements of dissolved N2O confirmed the existence of an N2O sink throughout the upper 100 m of the water column, with saturations in the upper 50 m of 90 ± 1% for areas north of 58.44° N. The consumption of N2O was observed at the base of the upper mixed layer (40 to 85 m) where rates were comparable to rates of net production. In near surface waters, production of N2O was not detected and the mean rates of N2O consumption were greater than the sum of the N2O supply from diapycnal transport of deep waters and surface exchange with the atmosphere. The determination of significant rates of N2O consumption coincident with the detection of organisms possessing atypical gene sequences for N2O reduction provides evidence of a biological mechanism that contributes to N2O sink conditions in these high latitude waters.
Results
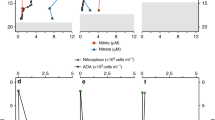
Research cruise JR271 onboard the RRS James Clark Ross during June 2012 made observations at 14 stations on a track between the northern North Sea, Greenland Sea, Fram Straits and the Barents Sea (Supplementary Fig. 1; Table 1). At five of these stations (EO1 – EO5), dissolved N2O was assayed at time of collection and then at 48 and 96 h in seawater from the near surface (10–20 m), which had been incubated under in-situ conditions of light and temperature for four days. At four out of the five stations N2O concentration decreased over the 96 h period at rates of between 0.17 and 0.31 nM d−1 (linear regression of concentration over time: r2 = 0.97, 0.99, 0.72, 0.78 for E01 to E04 respectively, Table 1). Logistical constraints meant that there was no sample replication within these experimental treatments and so data from the 5 stations were pooled and treated as replicates. To account for the heterogeneity of near-surface conditions, N2O concentrations were normalised to the initial ambient concentration and presented as percentage of the time zero value (Fig. 1a). The linear regression of N2O through time, averaged across the whole set of observations reveals a significant loss of N2O (adjusted r2 = 0.439, p < 0.005). At stations E01 to E04 the decrease equates to removal of 1.4–2.4% of the ambient N2O concentration per day. There was no indication of N2O removal at station E05 in the Barents Sea, here the mean N2O concentration remained at 10.37 ± 0.04 nM over the four-day period. Removing this station from the linear regression of decreasing N2O over time improved the fit to an adjusted r2 of 0.721 (p < 0.0005).
a, b From the North Atlantic during research cruise JR271 from near surface and base of the upper mixed layer respectively; c from cruise JR274 to the South Atlantic, samples collected from the base of the upper mixed layer at stations B03 and B04. In a due to the large environmental variability individual station data is presented as percentage of the original N2O concentration. Stations were located at North Sea (E01 – filled black circle), South of Iceland (E02 – open black circle), Greenland Sea – ice edge (E03 – filled black square), Greenland Sea – in ice (E04 – open black square), Barents Sea (E05 – filled red diamond). In b mean N2O concentrations at each time point, for each station are joined by the grey lines as indicated. In b and c data from all stations were combined, data points (filled black circle) represent mean concentrations ± 1 standard deviation. In a and c the linear regression line representing loss of N2O over time is shown as the solid black line, in c production of N2O over the first 48 h is represented by the dotted line, the red dashed line represents the mean expected value of N2O concentration assuming complete equilibration with the atmosphere.
At the same five stations, seawater was also collected and incubated, this time in triplicate, from the base of the seasonal upper mixed layer (40–60 m), depths that were generally coincident with, or close to, maxima in the N2O concentration profile. All incubated samples showed an increase in N2O concentration over the first 48 h with net rates of N2O production (change in concentration over time) from 0.2–3.3 nMd−1 followed by a decrease in concentration (net consumption) of 1.0−3.7 nMd−1 over the subsequent 48 h (Fig. 1b, Table 1). Again using the pooled measurements from the 5 experiments as replicates, a one-way ANOVA showed that the mean N2O concentrations at each time point were significantly different (F2,12 = 10.2, p < 0.003). After 96 h of incubation, N2O had decreased in four (E01−E04) out of five incubations to a level below that of the initial concentration.
In order to interrogate the pattern of production and apparent consumption of N2O, further incubations were performed at Stations BO3 and BO4 in the South Atlantic during research expedition JR274 onboard RRS James Clark Ross (Supplementary Fig. 1). Similar incubations performed in triplicate on water from the base of the mixed layer were performed with an increased sampling resolution over a 5 day period (Fig. 1c, Table 1). The same pattern of production (linear regression analysis of the combined dataset over the first 48 h; slope = +0.035 ± 0.007, adjusted r2 = 0.83, p = 0.007) followed by consumption (slope = −0.041 ± 0.01, adjusted r2 = 0.741, p < 0.02) as in northern waters was observed. Production and consumption rates were comparable to those in the north (Table 1) and consumption continued up to 120 h after the start of the incubation. At 120 h the observed concentration of N2O at both stations was lower than both the start point and the atmospheric equilibration value. 2.0 and 0.9 nM of N2O were removed from sites in the vicinity of South Georgia and the South Sandwich Islands, equivalent to 8.9% and 4.4% per day of ambient concentrations respectively.
The first step of nitrification, the aerobic oxidation of ammonium to nitrite, is the dominant source of N2O in the ocean except for in surface water overlying oxygen minimum zones29, but measurements of nitrification are limited in the polar regions. The observations of N2O production presented in Table 1 for samples collected from the base of the upper mixed layer are higher than those reported previously elsewhere30, which are typically less than 1.7% of nitrite production. These rates of N2O production are not entirely unexpected if associated with high rates of nitrification, such as reported for the Southern Ocean31 and the North Sea32, and considering that N2O yield in culture of up to 10% have been recorded33. A N2O yield of 1.7% would imply nitrification rates of 10–195 nmol L−1d−1 at these stations. The pattern of reversal observed between production and consumption (Fig. 1b, c) during incubations argues that production is limited by a finite substrate not identified here, but likely NH4+:NH3, and that the observed consumption maintains a tight balance between the two processes.
nosZ gene abundance and diversity
Diverse nosZ genes, both in DNA and cDNA and including both conventional Clade I and atypical Clade II sequences, were detected in every sample (Fig. 2). Thus a surprising range of microbes, many of which were actively transcribing the functional gene for N2O reduction, were detected in oxygenated surface waters of these high latitude regions. The species richness of cDNA Archetypes (an Archetype represents organisms possessing a nosZ gene sequence with ≥ 87% identity to the probe sequence) exceeded that of DNA Archetypes at all stations except EO1 and geographically cDNA and DNA species richness was significantly higher at the northern vs southern stations (p < 0.023) (diversity measures are reported in Supplementary Table 1). cDNA and DNA community compositions were very similar within stations, with the exception of Station BO4. Multi-response permutation procedure analysis indicated that community composition was significantly different between the North Atlantic and Southern Ocean regions (p < 0.002). The community compositions represented by cDNA and DNA were not different (p < 0.875). Among the Clade I nosZ genes, Archetype NosZ20 (representing cultivated denitrifying strains Pseudomonas stutzeri and P. fluorescens) was among the most abundant archetypes in the northern samples, but was a minor component of those from the Southern Ocean. Clade II Archetypes, representing atypical nosZ genes with greatest identity to Anaeromyxobacter genes, were present in all samples but dominated the nosZ assemblage in the southern samples. Archetypes WnosZ16 and WnosZ21 had the highest relative abundance of all WnosZ archetypes in all samples.
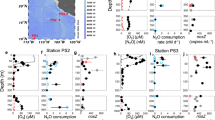
The suite of environmental and experimental variables collected for all samples from all stations is presented in Table 1 and Supplementary Table 2 and the Pearson correlation coefficient for these in Supplementary Table 3. Highly correlated variables were omitted from further ordination analysis. The ambient N2O concentration was negatively correlated with temperature (p < 0.0001) although both N2O production and consumption rates were positively correlated with temperature (p < 0.0001). Further study is required, not only to investigate the extent and distribution of these processes on geographic and seasonal scales, but also within the context of our warming environment. N2O concentration is largely governed by temperature dependent solubility, and as water temperatures increase there is potential for greater consumption (and production) which in concert with decreased solubilty indicate an increase in the polar ocean N2O sink. Despite the geographical differences in species richness and community composition noted above, N2O production and consumption were also correlated with nosZ copy number in both DNA and cDNA (p < 0.0003). Normalising the consumption rate to cDNA count (Fig. 3a) reveals a positive exponential relationship with temperature for Clade II archetypes which is not evident for the Clade I community.
a N2O consumption rate normalised to cDNA count (Red = Clade II, Blue = Clade I, Black = total), b RDA plot showing relationships between environmental variables (T = temperature, BottomZ = water column depth, SampleZ = depth sample collected) and NosZ/WnosZ community compositions based on Archetypes (red cross) from the microarray analysis. Circles = DNA, triangles = cDNA. Green = EO1, orange = EO2, cyan = EO3, blue = EO4, purple = EO5, pink = BO3, yellow = BO4.
In the redundancy analysis (Fig. 3b) combining community composition and the six most important environmental variables, the first two axes explained 49% and 37% of the variability, respectively. The two southern stations clustered closely and were distinct from those from the north. NosZ65 (Marinobacter-like), and NosZ −20, −22 -and −42 (all derived from saltmarsh sediments) signals were positively correlated with qPCR_cDNA and were prominent in the North Sea station EO1 samples, which can explain why station EO1 was the only one that exhibited a strong correlation with gene copy number in cDNA (qPCR-cDNA). The other northern stations at which NosZ20 was a major signal in both the DNA and cDNA clustered in the upper left quadrant and were not highly correlated with any of the environmental variables.
Discussion
At both of the polar stations (E04 and B04) presented in28 there was a notable lack of the nirS gene, which is responsible for the reduction of NO2− during the denitrification process, though the two dominant Clade II nosZ archetypes were abundant. This suggests a decoupling of the denitrification pathway and provides the potential for an efficient N2O scavenging mechanism in the absence of complete denitrification. The dominance of the WnosZ Archetypes at both of the southern stations, as well as their high relative abundance in all northern samples for both DNA and cDNA, suggests an important role for the Clade II microbes. This form of nosZ is preferentially associated with microbes that do not possess a full denitrification pathway, i.e., they do not possess the upstream enzymes to respire the soluble N oxides, and have been implicated in N2O consumption in soils27,34. It seems unlikely that the uncultivated microbes from agricultural soil and the cultivated Anaeromyxobacter strains, from which Archetypes WnosZ16 and WnosZ21, respectively, are derived, are present in the ocean. Rather, we suggest that N2O consuming marine microbes with highly similar enzymes are performing a similar function in the ocean.
The prevalence of the Clade II nosZ genes and the observed consumption of N2O at all geographical positions sampled argues very strongly that this is an important process, which had previously been unrecognised, in these high latitude waters. At the base of the upper mixed layer, the removal of N2O by the WnosZ archetypes moderates the flux of N2O from the deeper ocean to the near surface. In the near surface waters the rate of consumption is equivalent to the sum of fluxes from the atmosphere and deeper water (Fig. 4a) and thus provides a mechanism to contribute to the undersaturation of surface waters, which is prevalent in the polar regions7,11,12,13 and which was observed at all stations north of 58°N during this study (Fig. 4b).
a Mean (± 1 s.d.) rates of production, consumption & fluxes of N2O observed for stations E01 to E05 inclusive. Gross production assumes that consumption processes are constant throughout, which the experimental design did not resolve. b Mean N2O saturation (red line) ± 1 s.d. (dashed lines) for 13 stations north of 58.44°N during the same cruise. Grey line represents N2O saturation at the North Sea station E01 (56.16°N).
The melting of sea-ice has been shown to contribute to undersaturation of N2O in surface waters7,11 whilst the convection of low N2O water from the deep ocean coupled with air-sea exchange is used by Zhan et al.13 to account for what they describe as a permanent sink in the Nordic Seas. Neither of these mechanisms is necessarily exclusive to our observations, indeed all three are likely to contribute to N2O undersaturation. This current study presents coincident observations, at both poles of the Atlantic Ocean, of Clade II archetype WnosZ abundance and robust measures of apparent N2O consumption. This is a novel observation of the diverse presence and activity of an atypical gene sequence which enables an N2O removal mechanism not previously observed in oxygenated seawater. Although our measurements were restricted to summertime observations in both hemispheres it is clear that N2O consumption is occurring in oxygenated surface waters and is contributing to the presence of N2O understaturation in high latitude waters. However, more rigorous investigations are required over greater scales of time and space before incorporating this process into budgetary exercises at regional or global scales.
Methods
Seawater samples were collected from Niskin bottles deployed on a titanium frame at the near surface (10–20 m) and at the base of the surface mixed-layer (40 to 85 m) directly into ~4.5 L polycarbonate incubation bottles (NalgeneTM) (Table 1, Supplementary Fig. 1). Bottles from the near surface were sealed and transferred into incubators at ambient temperature and simulated light conditions to match those of collection. N2O concentration was determined on single bottles immediately and then at several time points up to 96 h later. Sample bottles collected from the base of the surface mixed-layer were sealed and incubated in the dark at ambient temperature within a purpose-built experimental laboratory container allowing precise temperature control. N2O concentration was determined on triplicate bottles immediately and then at several time points up to 120 h later. Temperature within a dummy incubation bottle was monitored using a traceable thermometer, while two recording thermometers were used to monitor air temperature in the incubator. Oxygen was determined by Winkler titration at the beginning of the experiment and for selected stations E05, B03 and B04 after 48 and 96 h. Samples were also collected for N2O analysis from CTD casts performed at 14 stations including the five experimental stations during JR271 only (Supplementary Fig. 1).
At six positions on a transect through the Atlantic Ocean between the UK and the Falkland Islands (Supplementray Fig. 1), tests were performed to confirm the validity of this approach and to test the integrity of the polycarbonate bottles to N2O diffusion. At each station up to 24 bottles were filled with seawater, sealed and and incubated in the dark at collection temperature. A second set of bottles were collected which were poisoned with the addition of 1 ml saturated HgCl2. N2O concentration was determined on collection, and thereafter triplicate analyses of N2O were made at several time points over 6 days. N2O concentration remained stable during each of these storage tests (Supplementary Fig. 2), the coefficient of variation varied between 1.3 and 5.2% (mean 3.9%, n = 16 time points × 3 analyses). F tests between N2O in killed samples and initial concentrations, and between killed samples and the expected (atmospheric equilibrium) concentration confirmed 86% and 87% similarity in variance respectively.
N2O analysis
Samples were collected using acid cleaned Tygon tubing directly from CTD Niskin bottles or by siphoning from 4.5 L incubation bottles into 1 L borosilicate flasks. Single samples were taken from CTD bottles and triplicates from the incubated sample. Samples were overfilled in order to expel air bubbles, poisoned with 200 µL of saturated HgCl2 solution and temperature equilibrated at 25.0 ± 0.5 °C. In all cases samples were analysed within 8 h of collection. N2O was determined by single‐phase equilibration gas chromatography with electron capture detection similar to that described by Upstill-Goddard et al.35. Each individual sample was calibrated against three certified (± 5%) reference standards of 287, 402 and 511 ppb (Air Products Ltd) which are traceable to NOAA WMO-N2O-X2006A scale for N2O mole fractions. Mean instrument precision from daily, triplicate analyses of the three calibration standards (n = 81) was 0.95%. Concentrations of N2O in seawater were calculated from solubility tables36 at equilibration temperature (~25 °C) and salinity. To capture variability within experimental treatments at the appropriate scale, mean values from each location-time combination were used as replicates in standard statistical analyses (linear regression and analysis of variance), which were performed using the Data Analysis ToolPak add-in module in Excel for Microsoft 365.
Air-sea flux
The exchange of N2O between the ocean and the atmosphere, the sea–air flux density (FN2O), was estimated from:
Where Kw is the gas exchange coefficient adjusted for Sc the Schmidt number for N2O as described in ref. 31. Cw is the measured seawater concentration and Ca is the equilibrium concentration of N2O in seawater based on the measured atmospheric value.
Diapycnal flux
The flux of N2O from below the pycnocline to the surface layer (QN) was estimated from32:
where: δN/δz is the vertical gradient of N2O across the pycnocline, and Kz is the vertical turbulent diffusion coefficient, estimated from
where ε is the turbulent kinetic energy dissipation, and N is the buoyancy (Brunt-Vaisala) frequency
where g is gravitational acceleration, ρ is the density and Δρ/Δz is the vertical density gradient across the pycnocline.
Nucleic acid manipulations and quantitative PCR analysis
Seawater samples (up to 8 L) were filtered onto 0.2 µm pore size Sterivex filters (Millipore, Billerica, MA) using a peristaltic pump, and filters were flash frozen in liquid nitrogen and stored at –80 °C. Total DNA and RNA were both extracted from the same Sterivex capsule filters using the ALLPrep DNA/RNA Mini Kit (Qiagen Sciences, Maryland, USA). Only one filter was extracted for each depth, i.e., no biological replicates. Reverse transcription from RNA to cDNA was performed using SuperScript R III First-Strand Synthesis System for RT-PCR (InvitrogenTM by Life TechnologiesTM). Any remaining RNA was removed by RNase at the end of the synthesis. PCR and qPCR using SYBR Green and the Qiagen master mix (Qiagen Sciences, Maryland, USA) (as described for seawater samples37) were used to amplify a 259 bp fragment of the nosZ gene using primers nosZ1F (WCSYTGTTCMTCGACAGCCAG) and nosZ1R (ATGTCGATCARCTGVKCRTTYTC)38. Standardisation and verification of specificity for qPCR assays was performed as described in28. Primers nosZ1F and nosZ1R targeted the Clade I genes; Clade II genes were not amplified separately. Rhodopseudomonas palustris (Clade I), was used as a positive control to optimise the reaction and to construct a plasmid for use as a standard in the qPCR assays. The amplified products were visualised after electrophoresis in 1.0% agarose gels stained with ethidium bromide. Assays were carried out within a single assay plate39. Each assay included triplicates of the no template controls (NTC), no primer control (NPR), four or more serial dilutions for the standard curve, and triplicates of known quantity of the environmental DNA samples (20–25 ng). DNA was quantified using Pico Green fluorescence (MolecularProbes, Eugene, OR) calibrated with several dilutions of phage lambda standards. Quantitative PCR was performed using a Stratagene MX3000P (AgilentTechnologies, LaJolla, CA, USA). Automatic analysis settings were used to determine the threshold cycle (Ct) values. The copy numbers were calculated according to:
and then converted to copy number ml−1 seawater filtered assuming 100% extraction efficiency. To maintain continuity and consistency among qPCR assays a subset of samples from the first assay was included in subsequent assays, as well as fresh dilution series for standard curves on every assay. Template DNA and plasmid DNA were quantified prior to every assay as above using Pico Green fluorescence to account for DNA loss that occurs upon repeated freeze-thaw cycles.
Microarray
The microarray (BC016) was developed following the archetype array approach described and employed previously (e.g.40,41). An archetype is defined as all sequences that hybridise with an individual probe, here within 87% identify to the probe sequence. Each 90-mer oligonucleotide probe included a nosZ-specific 70-mer region and a 20-mer control region (5′-GTACTACTAGCCTAGGCTAG-3′) bound to a glass slide. The design and spotting of the probes has been described previously40. Microarray BC016 contained two probe sets (NosZ and WnosZ) for the nosZ gene29. Clade I nosZ, most commonly found in marine and terrestrial heterotrophic denitrifying bacteria and associated with cultured strains representing alpha-, beta-,and gamma-Proteobacteria, was represented by 71 NosZ probes derived from whole genome sequences in public databases plus sequences obtained from clone libraries made using DNA extracts from the Great Sippewissett Marsh in Falmouth, MA, USA42. An additional 43 WnosZ probes were included to capture the atypical Clade II nosZ sequences27,34. Cultivated members of the atypical WnosZ probe set include alpha-, beta- and delta-Proteobacteria, CFB supergroup, Firmicutes and Verrumicrobia. The probe accession numbers and sequences are listed in Table S2 and the phylogenetic trees of the probe sequences are found in28. Arrays were printed on glass slides43 by Microarrays, Inc. (Huntsville, AL, USA). Targets were prepared from the amplified DNA produced in the qPCR assays above according to41. DNA concentration of the targets were measured on a Nanodrop spectrophotometer and the volume required for 200 ng of DNA was hybridised to duplicate arrays overnight in sealed tubes and then washed according to41. Arrays were scanned with an Agilent laser scanner 4300 (Agilent, Palo Alto, CA) and analysed using GenePix 6.0 software. Replicate features on the same array were averaged to calculate the Cy3/Cy5 ratio for each probe. Relative fluorescence ratio (RFR, the fraction that each probe fluorescence (Cy3/Cy5 ratio) contributes to the total fluorescence of the probe group) and normalised fluorescence ratio (FRn, the Cy3/Cy5 ratio of each probe normalised to the maximum Cy3/Cy5 detected on that array for the probe group) were used for plotting and statistical analysis. The vegan package in R (CRAN website; http://www.R-project.org)44 was used for ordination and diversity analysis of the array data. FRn values were transformed (Arcsin(Square root)) in order to normalise the proportional data. Archetypes with FRn < 0.01 were considered absent. Environmental data were standardised around zero (decostand in vegan). The transformed data were used in redundancy analysis (RDA) using the vegan package in R45.
Data availability
All N2O data from this study are available from the British Oceanographic Data Centre according to https://doi.org/10.5285/268dfd3b-dcc6-3f4a-e053-6c86abc0c2f946. The original nosZ array data are available at Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/projects/geo/) at the National Center for Biotechnology Information under GEO Accession No. GSE121473. Further supporting datasets from both cruises (JR271 and JR274) can be found at: https://www.bodc.ac.uk/projects/data_management/uk/ukoa/data_inventories/cruise. All data used in Figs. 1–4 are available either from the locations above or are found within Table 1 and supplementary information.
References
Butler, J. H. A. S. A. M. The NOAA Annual Greenhouse Gas Index (AGGI). 2018; Available from: http://www.esrl.noaa.gov/gmd/aggi/aggi.html.
Ravishankara, A. R., Daniel, J. S. & Portmann, R. W. Nitrous oxide (N(2)O): the dominant ozone-depleting substance emitted in the 21st century. Science 326, 123–125 (2009).
Zamora, L. M. & Oschlies, A. Surface nitrification: a major uncertainty in marine N2O emissions. Geophys. Res. Lett. 41, 4247–4253 (2014).
Bange, H. W. New directions: the importance of oceanic nitrous oxide emissions. Atmos. Environ. 40, 198–199 (2006).
Devol, A. H. Denitrification Including Anammox, in Nitrogen in the Marine Environment (Second Edition), (eds Douglas, D.A.B., Capone, G., Mulholland, M.R. & Carpenter, E.J.) 263–301 (Academic Press, 2008).
Zumft, W. G. Cell biology and molecular basis of denitrification. Microbiol. Mol. Biol. Rev. 61, 533 (1997).
Rees, A. P., Owens, N. J. P. & UpstillGoddard, R. C. Nitrous oxide in the Bellingshausen Sea and Drake Passage. J. Geophys. Res. 102, 3383–3391 (1997).
Zhan, L. Y. and L. Q. Chen, Distributions of N2O and its air-sea fluxes in seawater along cruise tracks between 30 degrees S-67 degrees S and in Prydz Bay, Antarctica. J. Geophys. Res., 114 (2009).
Zhan, L. Y. et al. A vertical gradient of nitrous oxide below the subsurface of the Canada Basin and its formation mechanisms. J. Geophys. Res. 120, 2401–2411 (2015).
Zhan, L. Y. et al. Austral summer N2O sink and source characteristics and their impact factors in Prydz Bay, Antarctica. J. Geophys. Res. 120, 5836–5849 (2015).
Fenwick, L. et al. Methane and nitrous oxide distributions across the North American Arctic Ocean during summer, 2015. J. Geophys. Res. 122, 390–412 (2017).
Randall, K. et al. First measurements of nitrous oxide in Arctic sea ice. J. Geophys. Res., 117 (2012).
Zhan, L. et al. A permanent N2O sink in the Nordic Seas and its strength and possible variability over the past four decades. J. Geophys. Res. 121, 5608–5621 (2016).
Verdugo, J. et al. Climate relevant trace gases (N2O and CH4) in the Eurasian Basin (Arctic Ocean). Deep-Sea Res. 117, 84–94 (2016).
Robertson, L. A. et al. Confirmation of ‘aerobic denitrification’ in batch cultures, using gas chromatography and 15N mass spectrometry. FEMS Microbiol. Ecol. 18, 113–119 (1995).
Su, J.-J., Liu, B.-Y. & Liu, C.-Y. Comparison of aerobic denitrification under high oxygen atmosphere by Thiosphaera pantotropha ATCC 35512 and Pseudomonas stutzeri SU2 newly isolated from the activated sludge of a piggery wastewater treatment system. J. Appl. Microbiol. 90, 457–462 (2001).
Frette, L., Gejlsbjerg, B. & Westermann, P. Aerobic denitrifiers isolated from an alternating activated sludge system. FEMS Microbiol. Ecol. 24, 363–370 (1997).
Fu, G. P. et al. Isolation and identification of a salt-tolerant aerobic denitrifying bacterial strain and its application to saline wastewater treatment in constructed wetlands. Bioresource Technol., 290 (2019).
Cavigelli, M. A. & Robertson, G. P. Role of denitrifier diversity in rates of nitrous oxide consumption in a terrestrial ecosystem. Soil Biol. Biochem. 33, 297–310 (2001).
Marchant, H. K. et al. Denitrifying community in coastal sediments performs aerobic and anaerobic respiration simultaneously. ISME J. 11, 1799 (2017).
Coates, C. J. & Wyman, M. A denitrifying community associated with a major, marine nitrogen fixer. Environ. Microbiol. 19, 4978–4992 (2017).
Farias, L. et al. Biological N2O fixation in the Eastern South Pacific Ocean and Marine Cyanobacterial cultures. PLoS ONE, 8 (2013).
Raes, E. J. et al. Reduction of the powerful greenhouse gas N2O in the South-Eastern Indian Ocean. PLoS ONE, 11 (2016).
Wyman, M., Hodgson, S., Bird, C. Denitrifying alphaproteobacteria from the Arabian Sea that express nosZ, the gene encoding nitrous oxide reductase, in oxic and suboxic waters. Appl. Environ. Microbiol. 79, 2670–2681 (2013).
Cornejo, M., Murillo, A. A. & Farias, L. An unaccounted for N2O sink in the surface water of the eastern subtropical South Pacific: Physical versus biological mechanisms. Prog. Oceanogr. 137, 12–23 (2015).
Sun, X., A. Jayakumar, and B. B. Ward, Community composition of nitrous oxide consuming bacteria in the oxygen minimum zone of the Eastern Tropical South Pacific. Front. Microbiol., 8 (2017).
Jones, C. M. et al. The unaccounted yet abundant nitrous oxide-reducing microbial community: a potential nitrous oxide sink. ISME J 7, 417–426 (2013).
Jayakumar, A. et al. Community composition of nitrous oxide reducing bacteria investigated using a functional gene microarray. Deep Sea Res. 156, 44–50 (2018).
Ji, Q. X. et al. Global nitrous oxide production determined by oxygen sensitivity of nitrification and denitrification. Global Biogeochem. Cycles 32, 1790–1802 (2018).
Ji, Q. X. & Ward, B. B. Nitrous oxide production in surface waters of the mid-latitude North Atlantic Ocean. J. Geophys. Res. 122, 2612–2621 (2017).
Nightingale, P. D. et al. In situ evaluation of air-sea gas exchange parameterizations using novel conservative and volatile tracers. Global Biogeochem. Cycles 14, 373–387 (2000).
Slawyk, G. et al. Isotopic and enzymatic analyses of planktonic nitrogen utilisation in the vicinity of Cape Sines (Portugal) during weak upwelling activity. Deep-Sea Res. 44, 1–25 (1997).
Goreau, T. J. et al. Production of NO2- and N2O by nitrifying bacteria at reduced concentrations of oxygen. Appl Environ Microbiol. 40, 526–532 (1980).
Sanford, R. A. et al. Unexpected nondenitrifier nitrous oxide reductase gene diversity and abundance in soils. Proc. Natl Acad. Sci. USA 109, 19709–19714 (2012).
Upstill-Goddard, R. C., Rees, A. P. & Owens, N. J. P. Simultaneous high-precision measurements of methane and nitrous oxide in water and seawater by single phase equilibration gas chromatography. Deep-Sea Res. 43, 1669–1682 (1996).
Weiss, R. F. & Price, B. A. Nitrous-oxide solubility in water and seawater. Marine Chem. 8, 347–359 (1980).
Jayakumar, A., Peng, X. F. & Ward, B. B. Community composition of bacteria involved in fixed nitrogen loss in the water column of two major oxygen minimum zones in the ocean. Aquatic Microb. Ecol. 70, 245 (2013).
Henry, S. et al. Quantitative detection of the nosZ gene, encoding nitrous oxide reductase, and comparison of the abundances of 16S rRNA, narG, nirK, and nosZ genes in soils. Appl. Environ. Microbiol. 72, 5181–5189 (2006).
Smith, C. J. et al. Evaluation of quantitative polymerase chain reaction-based approaches for determining gene copy and gene transcript numbers in environmental samples. Environ. Microbiol. 8, 804–815 (2006).
Bulow, S. E. et al. Sediment denitrifier community composition and nirS gene expression investigated with functional gene microarrays. Environ. Microbiol. 10, 3057–3069 (2008).
Ward, B. B. and N. J. Bouskill, The utility of functional gene arrays for assessing community composition, relative abundance, and distribution of ammonia-oxidising bacteria and archaea, in Methods in Enzymology, Vol 46: Research on Nitrification and Related Processes, Pt B, (eds Klotz, M.G. & Stein, L.Y.) 2011. p. 373–396.
Kearns, P. J. et al. Long-term nutrient addition differentially alters community composition and diversity of genes that control nitrous oxide flux from salt marsh sediments. Estuarine Coast. Shelf Sci. 154, 39–47 (2015).
DeRisi, J. L., Iyer, V. R. & Brown, P. O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997).
Borcard, D., R. Gillet, and P. Legendre, Numerical Ecology with R. Use R! 2011: Springer. 306.
Veuger, B. et al. Nitrification and growth of autotrophic nitrifying bacteria and Thaumarchaeota in the coastal North Sea. Biogeosciences 10, 1775–1785 (2013).
Brown, I. and A. P. Rees, Nitrous Oxide (N2O) concentration data from CTD casts and Bioassay experiments during cruises JR271 (North Sea to Arctic Ocean) in June 2012 and JR274 (Scotia Sea, Southern Ocean) in January-February 2013, U.K. British Oceanographic Data Centre - Natural Environment Research Council, Editor. 2015.
Acknowledgements
We thank the captains and crew of the RRS James Clark Ross during research cruises JR271, JR274 and JR300. We are grateful to Eric Acterberg, Richard Sanders, Mark Moore and Mark Stinchcombe for nutrient and oxygen measurements and acknowledge the contributions made to this work by Vas Kitidis, Darren Clark and Phil Nightingale. This work was funded by the UK Natural Environment Research Council Grant, UKOA-Ocean Acidification impacts on sea surface biology, biogeochemistry & climate (NE/H017259/1) and further by NERC National Capability funding as part of the Climate Linked Atlantic Section Science (CLASS) programme (NE/R015953/1).
Author information
Authors and Affiliations
Contributions
A.R., I.B., and B.W. designed the research; A.R. and I.B. measured and analysed the N2O; A.J. and B.W. measured and analysed nosZ; Statistical analysis of the N2O experiments was by PS; Interpretation of data and writing of manuscript was led by A.R. with contributions from B.W., G.L., P.S., I.B. and A.J.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Primary handling editor: Teresa Ortner.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rees, A.P., Brown, I.J., Jayakumar, A. et al. Biological nitrous oxide consumption in oxygenated waters of the high latitude Atlantic Ocean. Commun Earth Environ 2, 36 (2021). https://doi.org/10.1038/s43247-021-00104-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43247-021-00104-y
This article is cited by
-
Direct biological fixation provides a freshwater sink for N2O
Nature Communications (2023)
-
Unexpectedly minor nitrous oxide emissions from fluvial networks draining permafrost catchments of the East Qinghai-Tibet Plateau
Nature Communications (2022)
-
Nitrous oxide and methane in a changing Arctic Ocean
Ambio (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.