Abstract
Research on batteries mostly focuses on electrodes and electrolytes while few activities regard separator membranes. However, they could be used as a toolbox for injecting chemical functionalities to capture unwanted species and enhance battery lifetime. Here, we report the use of biological membranes hosting a nanopore sensor for electrical single molecule detection and use aqueous sodium polysulfides encountered in sulfur-based batteries for proof of concept. By investigating the host-guest interaction between polysulfides of different chain-lengths and cyclodextrins, via combined chemical approaches and molecular docking simulations, and using a selective nanopore sensor inserted into a lipid membrane, we demonstrate that supramolecular polysulfide/cyclodextrin complexes only differing by one sulfur can be discriminated at the single molecule level. Our findings offer innovative perspectives to use nanopores as electrolyte sensors and chemically design membranes capable of selective speciation of parasitic molecules for battery applications and therefore pave the way towards smarter electrochemical storage systems.
Similar content being viewed by others
Introduction
Batteries, as one of the most versatile energy storage technologies, play a central role in the ongoing transition from fossil fuels to renewable energy1. Among them, the Li-ion batteries are the technology of choice as they offer the largest energy density while their cost is continuously decreasing2,3. Nevertheless, improvements are needed to increase their lifetime and sustainability. Hence the ongoing activities center on alternative technologies (Na-ion, Li-air, Li–S,…) but also in the quest of means to enhance present batteries lifespan, durability and reliability. The latter calls for the development of smart batteries embedded with intelligent sensing4 and curing chemical functionalities5,6.
Great advances have been made over the last few years to either restore the electrode conductivity7 or at the electrolyte-membrane component to regulate species migration when injured8. Not surprisingly, most of the auto-repairing approaches are inspired by biological systems and benefit from the general strategies and formalisms well established for most living creatures. Following nature’s strategy relying on the use of sacrificial weak bonds for self-repairing, battery scientists have developed materials based on bio-molecules or polymers, with auto-repairing properties6 relying on dynamic supramolecular self-assembly such as hydrogen bonding, ionic bonding or host-guest interaction5,7.
Among supramolecular materials, cyclodextrins have been extensively studied because of their rich molecular recognition that is temperature dependent and their wide range of functionalization. Thus the feasibility to use the temperature as a stimulus9 to regulate on demand the uptake or release of trapped species within their cavities, offers an interesting auto-repairing function10. Such properties have already been widely exploited for thermal switches11 and in medicine for drug delivery12 or as a molecular adapter to identify the DNA nucleotides13, or small organic molecules14. Cyclodextrins have recently entered the field of batteries as well, as witnessed by the design of highly stretchable binders integrating sliding ring polyrotaxanes15, e.g., self-assembled architectures of cyclodextrins to auto-repair fractured Si electrodes. Such supramolecular structures were also used as ionic conducting polymer membrane in Li-ion batteries16,17 and cyclodextrins trapping properties were exploited by placing cyclodextrins polymers in S-based electrodes to address the redox shuttle issue in Li–S batteries, e.g., migration of soluble polysulfides Li2Sn (3 ≤ n ≤ 8), intermediates back and forth between the two electrodes18. Alternatively, other strategies to tackle the polysulfide shuttle effect have focused on the separator to prevent (Sn)2− diffusion to the anode with either micro intrinsic porosity19,20 or using grafting chemistry to repeal (Sn)2− via repulsive electrostatic interactions21.
Up to now, existing analytical techniques to identify intermediate polysulfides and monitor Li–S batteries electrolyte composition, such as UV-vis, X-ray, mass spectrometry or cyclic voltammetry22 do not allow to detect and sequence at the single molecule level these parasitic redox species. Here, we focus on the high-resolution detection of sulfide-based species differing by one sulfur atom by using the nanopore technology. This powerful single molecule technique, based on electrical detection, is highly sensitive to the nature of the ion used, but more importantly to the size, conformation and chemical nature of molecules analyzed. Such single molecule approach using protein nanopores has already been successfully implemented for the ultrafast sequencing of nucleic acids23 together with the high-resolution size discrimination of biomolecules24,25, for biomarkers26,27 or pathogenic agent detection28. These nanopore-based sensors are now employed for the detection of subtle chemical modifications29,30 or to read the sequence of biomolecules31,32. Nanopores are also used to study protein conformational changes and transitions33,34, epigenetics by detection of DNA methylations35,36 or for metabolome analysis37. Further pushing this technique to its limits researchers have recently identified the twenty amino acids38, paving the way to protein sequencing. Thus, a burning question regards the implementation of this technique to the field of batteries.
Herein we report a protein channel, α-hemolysin (α-HL) nanopore, with a β-cyclodextrin adapter as a simplified model to understand the encapsulation of polysulfides by cyclodextrins to be extended to smart membrane design for Li–S batteries. As a proof of concept of both cyclodextrins affinity towards (Sn)2− and nanopore sensitivity, we have used Na-based polysulfide species Na2Sn (n = 3, 4, 5) in aqueous media that can be found as catholyte in aqueous Na-ion/polysulfide batteries39,40,41. In this contribution, the reversible equilibrium complexation between sodium polysulfides and cyclodextrins is demonstrated and their interaction characterized by complementary NMR spectroscopy, nanopore experiments and molecular docking calculations. Finally, we highlight the promising use of the nanopore technology as being able to detect and discriminate redox species at the single sulfur atom level.
Results
Cyclodextrins reversibly store and deliver polysulfides guest
Monitoring the outcome of polysulfides species in aqueous or non-aqueous (Li, Na) metal batteries has been of paramount importance. Few approaches have been tried. They consist in targeting polysulfide migration by trapping them at the positive electrode via confinement (mesoporous carbons) or surface adsorption (oxides) means42, or by tuning the electrolyte43 composition, or using chemically-grafted entangled separators20,21. Because of the limited success of these approaches we have looked for an alternative with enhanced selectivity in their entrapping. In light of a recent work18 on the supramolecular interaction between β-cyclodextrin and sodium or lithium S4 polysulfide, we further explore the cyclodextrin benefits towards polysulfide trapping. Our strategy consists in developing a methodology to quantify and understand the supramolecular interactions between different cyclodextrins (α, β, and γ) and different polysulfides chain lengths in aqueous media, which has not been achieved yet.
To do so, several aqueous polysulfides solutions were prepared in an aqueous buffer by mixing in stoichiometric ratio Na2S with S under Argon atmosphere. The sulfur chain lengths were varied in order to get Na2S2, Na2S3, Na2S4, Na2S5 via a solution process (see “Methods” section) and characterized by UV-vis spectroscopy as previously reported22 (Supplementary Fig. 1). The interaction between polysulfides and the different cyclodextrins was determined by 1H NMR following the chemical shifts of the different protons of cyclodextrins in presence of polysulfides. Figure 1bi shows the 1H NMR spectrum of β-cyclodextrin and Na2S5 in the deuterated buffer (red) with respect to the spectrum of β-cyclodextrin alone in the same buffer (blue). In addition, a 2D HSQC experiment was performed for signal attribution of the different protons of the cyclodextrins (Supplementary Fig. 2). The chemical shifts differences indicate the existence of a complexation equilibrium. As the exchange rate of this equilibrium is very fast compared to the NMR time scale, the peaks assigned to free and complexed cyclodextrin populations are merged and appear as the weight average chemical shift of both populations. It is worth noticing that the most shifted protons turn out to be the ones pointing inside the cavity of the host molecule such as H3 (Fig. 1bii and Supplementary Fig. 3). In this way, the host-guest interaction through the inclusion of Na2S5 inside the β-cyclodextrin cavity can be hypothesized. Therefore, we mainly focused on H3 to make the following quantitative analysis. Similarly, experiments with α-cyclodextrins and γ-cyclodextrins were performed and the existence of an equilibrium complexation was confirmed as well (Supplementary Figs. 2 and 3).
a Chemical structures and geometric dimensions of α-cyclodextrins, β-cyclodextrins, and γ-cyclodextrins52. As macrocyclic ring molecules composed of, respectively, 6, 7, or 8 glucopyranoside units, α-cyclodextrins, β-cyclodextrins, or γ-cyclodextrins can adopt their cone-truncated conformation whose inner and outer diameters vary, respectively, from 5.7 to 9.5 Å and 13.7 to 16.9 Å52. The primary hydroxyl groups are located on the narrower rim of the molecule and the secondary ones are on the wider one. b (i) Selected region of the 1H NMR spectra (600 MHz, D2O, 298 K, pH 10, 25 mM NaHCO3) of native β-cyclodextrin (10 mM, blue) and β-cyclodextrin in presence of Na2S5 (equimolar ratio 10 mM, red) illustrating the host-guest interaction. (ii) Schematic location in the cyclodextrins 3D-structures of the hydrogens shown in the NMR spectra. (iii) Schematic representation of the host-guest interaction equilibrium between β-cyclodextrin and Na2S5 (Na+ counter ions were not represented for better readability).
To go deeper in the characterization of the complex, continuous variation method, also known as Job’s plot, was performed to determine the stoichiometry of the studied supramolecular complexes44. Figure 2a shows the Job’s experiments for the different cyclodextrins (α, β, γ) with Na2S5. Thus, the (1:1) stoichiometry for the inclusion complexation which corresponds to the ∑ ratio of 0.5 was validated. We also performed molecular docking calculations in order to support our findings. Docking results show that in all cases polysulfides bind inside the cyclodextrins cavities, as shown in Supplementary Fig. 4 and Fig. 3c for β-cyclodextrin. Furthermore, regarding α-cyclodextrins and β-cyclodextrins, there is room for only one polysulfide guest per cyclodextrin host (see Supplementary Fig. 5). This is in full agreement with the NMR experiments. It can be recalled that reversible supramolecular aggregation relies on weak bonds : here both the curvature of the Job’s plots (Fig. 2a), indicative of weak binding, and the affinity scores evaluated in the docking runs (Supplementary Fig. 4) are consistent with this picture. At the same time these results, hinting strongly a one-to-one stoichiometry, provided us with a 2-states thermodynamic equilibrium model as illustrated in Fig. 1biii. Then we carried out 1H NMR titration experiments (see “Methods” section) by fitting the H3 NMR chemical shift of the different cyclodextrins with the corresponding binding isotherm model in order to calculate the different association constants for the complex formation of Na2S5 with α-cyclodextrins, β-cyclodextrins, and γ-cyclodextrins44 (Fig. 2b). Comparable values to the previous reports45 were found and interestingly, β-cyclodextrin turns out to have the best affinity towards Na2S5 with an association constant of Kβ5 = 181 ± 4 M−1, almost three times larger than with α-cyclodextrins and γ-cyclodextrins, Kα5 = 62 ± 11 M−1 and Kγ5 = 52 ± 9 M−1, respectively. Molecular docking affinity ranking over all the Na2S5/cyclodextrin inclusion complexes led to the same result highlighting the stronger affinity for the β-cyclodextrin (Supplementary Fig. 4). Hence our experimental and numerical results confirm previous literature reports10,45. Although the Van’t Hoff plots are useful for spotting an exothermic complexation phenomenon, they fall short in sensing the difference in the strength of complexation of α-cyclodextrins or β-cyclodextrins towards Na2S5 (Supplementary Fig. 6). This does not come as a surprise in light of the different dynamical conformations of α-cyclodextrins or β-cyclodextrins in water, noted by our Molecular Dynamics simulations, as well as the geometrical complementarity issues, to name a few, which likely play a role in this complex phenomenon (Supplementary Figs. 7 and 8).
a Experimental Job’s plot obtained following H3 shift for α-cyclodextrins, β-cyclodextrins, and γ-cyclodextrins in presence of Na2S5 (respectively, top-down). Second order polynomial fitting whose maximum is obtained for the stoichiometric ratio, ∑, (see “Methods” section) matches with the stoichiometry of the complex. b 1H NMR titration plots for the inclusion of Na2S5 in the different cyclodextrins cavities and fitted binding isotherms determined from H3 signal for α-cyclodextrins, β-cyclodextrins, γ-cyclodextrin with indication of the resulting association constants Kα5, Kβ5, and Kγ5 for the corresponding 1:1 inclusion complex. The statistic error relative to the chemical shift measurement was obtained by the calculation of the standard deviation of 10 similar and independent points and is estimated to 9%. The plotted error bars were calculated considering the uncertainties propagation (see “Methods” section).
a Experimental binding isotherms for β-cyclodextrin complexes with Na2S5 (red), Na2S4 (orange), Na2S3 (green), (See “Methods” section). The statistic error relative to the chemical shift measurement was obtained by the calculation of the standard deviation of 10 similar and independent points and is estimated to 9%. The plotted error bars were calculated considering the uncertainties propagation (see “Methods” section). b Best docking affinity scores (kcal mol−1) over all the calculated β-cyclodextrin inclusion complexes with Na2S5 (red), Na2S4 (orange), and Na2S3 (green). The first set of calculations used the cyclodextrin structures extracted from PDB as receptors (empty bars) and the second set used the cyclodextrin structures generated by Molecular Dynamics simulations (full bars) (See “Methods” section and Supplementary Fig. 4). c Corresponding best docking poses for (i) Na2S5/β-cyclodextrin, (ii) Na2S4/β-cyclodextrin, and (iii) Na2S3/β-cyclodextrin complexes. Receptors structures are those obtained from Molecular Dynamics simulations, top and side view; β-cyclodextrin in sticks, polysulfides in yellow space-fill representation, figure made with VMD64.
Tracking the most stable β-cyclodextrin inclusion complexes, we investigated different sizes of polysulfides. To do so, we coupled titration experiments with molecular docking calculations. As shown in Fig. 3a, a quite strong correlation is exhibited between the length of the sulfide chain and the corresponding association constants Kβ5 = 181 ± 4 M−1 and Kβ4 = 136 ± 4 M−1 for Na2S5 and Na2S4, respectively, with β-cyclodextrin. Kβ3 could not be estimated with as high reliability (152 ± 14 M−1) as the previous ones because of the uncertainties of the small H3 chemical shifts measured. On the other hand, considering H6 shifts validates the correlation hypothesis (Supplementary Fig. 9). This observation is consistent with the similar trend obtained by calculations (Fig. 3b), performed by docking the different polysulfides both on the cyclodextrin structures extracted from the Protein Data Bank46 (PDB) and on the ones generated by Molecular Dynamics simulations, which may better represent the cyclodextrin behavior in solution (Supplementary Figs. 7 and 8). In all cases, the general trend is a (1:1) complex formation, with (Sn)2− inside the cyclodextrin cavity (Fig. 3c). The calculated affinity scores increase with the length of the sulfide chain and thus confirm a weaker affinity for the Na2S3/β-cyclodextrin complex, explaining the experimental noisy curve for this pair (Fig. 3a, green points). Therefore, the results indicate a non-specific complex formation but highlighted a polysulfide’s size-dependent complexation towards β-cyclodextrin. Similar experiments performed with α-cyclodextrin exhibit the same trend (Supplementary Fig. 10). Thus, the number of sulfur atoms is supposed to play a major role in the host-guest interaction. Altogether, these experiments gather essential parameters pertaining to the β-cyclodextrin interactions with polysulfides that turn out to be the key to successfully implementing our nanopore sensing technique.
Detection of the different polysulfides species by nanopore
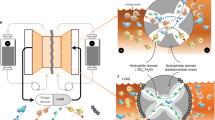
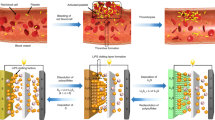
In order to fine-tune the characterization of Na2Sx/β-cyclodextrin complex, we used the nanopore approach to detect and discriminate at the single molecule level each species. With this approach, two compartments (cis and trans) immersed with an electrolyte are separated by a protein channel, α-hemolysin (α-HL), inserted into a lipid bilayer (Fig. 4a). Applying a constant potential difference between the two electrodes in absence of any species enables the measurement of a stable pA ionic current flowing through the pore (Fig. 4b, open pore current). Species passing through the nanopore, or clogging the nanopore without translocating, partially block ion transport inducing a detectable decrease of the open pore ionic current. These current blockades are characteristic of the species size, sequence and chemical modifications13,24,25,38. In the last years, β-cyclodextrins have been used to narrow the α-HL pore constriction to enhance the capability of the pore to discriminate nucleotides13 and organic compounds14. For protein pores such as α-HL, the ion selectivity of the channel creates an electro-osmotic flux (EOF). The EOF is the net water flow and it is associated to the ion selectivity of the pore47. Indeed, the selectivity results in an unbalance between positive and negative ion fluxes that gives rise to a net motion of water molecules. The EOF is known to affect the entrance of the neutral cyclodextrin through the stem of the nanopore and its lodging into the channel48. The interaction of β-cyclodextrin with α-HL has been studied as a function of a wide range of pH (5 to 11), transmembrane potentials (mV) and nature of electrolyte salt, in order to control the entry and dwell times of cyclodextrins into the nanopore48,49. We used this strategy to compare β-cyclodextrin and Na2Sx/β-cyclodextrin complex using an α-HL pore. We first tested the effect of pH 10 on the α-HL pore (Supplementary Fig. 11) and found similar currents as previously published49. The solution pH alters the protonation state of exposed amino acids and, hence, it can affect ion selectivity. A previous study50 indicates that α-HL selectivity inverts between pH 7.5 (anion selective) and pH 11 (cation selective). Here, we work at pH 10 where, to the best of our knowledge, no literature data provide information on the α-HL selectivity and on the direction of the EOF. In order to characterize the EOF, we hence performed a computational analysis via all-atom Molecular Dynamics simulation. We considered two different plausible titration states of α-HL at pH 10, a first one where only the seven N-Terminal (Ala1) are protonated and a second one where, in addition to Ala1, we also changed one (out of seven) Lys8 and one (out of seven) Tyr102, (Supplementary Fig. 12). For both cases, we explored transmembrane electric potentials ranging from −500 mV to 500 mV. We found that, in all the cases, the anion (Cl−) flow is much larger that the cation (Na+) flow and the electro-osmotic flux is directed as the anions (Supplementary Figs. 13 and 14). These results suggested us to perform experiments at a negative applied voltage so that the direction of negative ions and of the EOF is trans-to-cis, see Fig. 4a. In this condition, after addition of 1 mM β-cyclodextrin or Na2S5/β-cyclodextrin complex into the trans compartment, we detected partial transient ionic current blockades (Fig. 5c, d) due to interactions of the molecules within the α-HL stem. It implies that ions, here Na+ and Cl− are still transported through the Na2S5/β-cyclodextrin complex lodged in the α-HL while the complex cannot go through due to steric hindrance. In addition, we do not expect the polysulfides to cross the membrane via β-cyclodextrin constriction as well. This is based on the size of the (Sn)2− species51, ~9.1 Å (S3)2− or ~11.7 Å (S5)2−, which is greater than the narrower diameter (~7.8 Å) of the cone-like shape of cyclodextrin52. Nevertheless, when β-cyclodextrin or Na2S5/β-cyclodextrin complex is added to the cis compartment, no such current blockades were observed (data not shown). This result is indicative of an EOF from trans to cis compartment preventing molecules to enter the α-HL.
a Schematic of the ion−current measurement set-up. One protein nanopore is inserted into a suspended lipid bilayer. An electrical potential is applied via two Ag/AgCl electrodes, which induces an ionic current of Na+ and Cl− ions through the nanopore (1 M NaCl, 25 mM NaHCO3 pH 10, under argon atmosphere). At pH 10, the α-HL is anion selective (see Supplementary Fig. 15), consequently, an electro-osmotic flux (EOF) directed from trans to cis sets in favoring the entry of β-cyclodextrin and Na2Sx/β-cyclodextrin into the stem part of the α-HL. b Detail of a part of current trace blockades arising from β-cyclodextrin (blue) and from Na2S5/β-cyclodextrin complex (red) interaction with the pore at −100 mV. I0 is the ionic open pore current and Ib the blockade current. c Scatter plots of blockade ratio, given by the relation \((I_0 - I_b)/I_0\), versus dwell time in presence of 1 mM β-cyclodextrin (blue, 225 events) and 1 mM Na2S5/β-cyclodextrin complex solution (gray, 761 events) (5 independent experiments). d General distribution (gray) blockade ratio histogram in presence of 1 mM Na2S5/β-cyclodextrin complex (945 events). Deconvolution of the general distribution showing two populations attributed to β-cyclodextrin (blue, 64.0 ± 2.0%) and Na2S5/β-cyclodextrin (red, 74.0 ± 0.3%).
a Part of current versus time traces of an α-HL pore at −100 mV, in the presence of 1 mM β-cyclodextrin (blue), Na2S3/β-cyclodextrin (green), Na2S4/β-cyclodextrin (orange), Na2S5/β-cyclodextrin (red). All experiments are independent (6 in total), performed in 1 M NaCl, 25 mM NaHCO3 pH 10, at −100 mV, in an Argon filled glovebox. b Histograms of blockade ratio for β-cyclodextrin (blue, 61.8 ± 0.3%), Na2S3/β-cyclodextrin (green, 66.7 ± 0.2%), Na2S4/β-cyclodextrin (orange, 68.8 ± 0.1%), Na2S5/β-cyclodextrin (red, 74.0 ± 0.3%). c Accessible volume estimation from Molecular Dynamics simulations. α-HL constriction is highlighted in yellow, while β-cyclodextrin atoms are represented as balls-and-sticks. The plot reports the radius of the effective pore section available for ion passage. The effective pore section in the presence of β-cyclodextrin and Na2Sx/β-cyclodextrin are represented using the same color code of the other panels. d Experimental current blockades. Error bars are the confidence intervals of the average estimator (see “Methods” section). e Nanopore hindrance estimator calculated from atomistic model. Panel (c) was made using the VMD software.
The blockade ratio, given by the relation \((I_0 - I_b)/I_0\) where I0 is the ionic open pore current and Ib is the blockade current, versus individual event dwell times scatter plots (Fig. 4c) shows two distinct event populations for free β-cyclodextrin molecules compared to Na2S5/β-cyclodextrin complex. While dwell times cannot be used as a parameter for discrimination of β-cyclodextrin and Na2S5/β-cyclodextrin complex (Supplementary Fig. 11), we found that the general distribution of current blockade ratios for Na2S5/β-cyclodextrin complex can be deconvoluted (Fig. 4d) and shows two populations centered in 74.0 ± 0.3% and 64 ± 2%. These populations were attributed to the Na2S5/β-cyclodextrin complex (74.0 ± 0.3%) and the free β-cyclodextrin in solution (64 ± 2%), respectively. The ratio between number of events associated to the Na2S5/β-cyclodextrin complex (red distribution) and the total number of events (gray distribution) is 81 ± 2.5%. This is in very good agreement with the NMR calculated complexation fraction in the same conditions, namely 79 ± 4% (Supplementary Fig. 16).
We then characterized Na2S5/β-cyclodextrin, Na2S4/β-cyclodextrin, Na2S3/β-cyclodextrin complexes and compared our results to free β-cyclodextrin molecules (Fig. 5). The current traces of each species show two kinds of current blockades associated with two types of events: (i) “bumping” events, characterized by brief, low-level current blockades, which arise due to diffusion of molecules close to the pore; and (ii) interaction events, characterized by larger current blockades of longer duration. In order to characterize the interaction of β-cyclodextrin and Na2Sx/β-cyclodextrin complex with α-HL, each current trace was statistically analyzed to determine the event current amplitude and to separate bumping events from interaction events (see “Methods” section, the reported data refer only to interaction events). We can observe a statistical decrease in the event current amplitudes correlated to a decrease of sulfur number for each complex (Fig. 5a). We find 74.0 ± 0.3%, 68.8 ± 0.1%, 66.7 ± 0.2% for Na2S5/β-cyclodextrin, Na2S4/β-cyclodextrin and Na2S3/β-cyclodextrin complexes, respectively. Therefore, each complex is well discriminated from free β-cyclodextrin in solution with an average blockade ratio of 61.8 ± 0.3% (Fig. 5b). These results strongly suggest that polysulfide species are docked inside the β-cyclodextrin cavity and that we can discriminate each species at a single sulfur atom level. In order to support this finding, we also calculated from atomistic modeling the nanopore hindrance estimator introduced as a previous publication53. The calculation is based on an approximated quasi-1D continuum description of the nanopore electric resistance. As a first step, we calculated from Molecular Dynamics simulations for each pore section the effective area available to the electrolyte passage for both the open α-HL and the α-HL with the β-cyclodextrin in the pore stem (Fig. 5c). Then, we subtracted from the effective area of the β-cyclodextrin case a section corresponding to the occupancy of the different sulfides. This allowed us to calculate the nanopore hindrance estimator as \(b = 1 - R_0/R\) where R0 and R are the quasi-1D resistance estimators for open pore and clogged pore. As already discussed53, this method does not provide a quantitative estimation of the current blockade. Indeed, it is based on several simplified assumptions not satisfied at the nanoscale, the main being: (i) it is a continuum method where atomistic details enter only via the effective area profile, (ii) it is a quasi-1D method that, in principle, is valid only for smoothly varying pore section and (iii) it neglects electrical double layer effects. Nevertheless, it allows to catch the qualitative trends of the current blockades as shown by the good correlation with the experimentally observed average current blocked (see Fig. 5e and Supplementary Fig. 17).
Discussion
Via the use of a protein nanopore sensor inserted into a lipid membrane we have demonstrated the feasibility to discriminate molecules that solely differ by a single sulfur atom. Indeed, the different polysulfide/β-cyclodextrin complexes can be identified with great sensibility by a unique pattern of events in term of current blockade that increases with increasing the number of sulfur atoms, as intuitively expected. In addition, we could rationalize such an experimental work by molecular docking calculations and Molecular Dynamics simulations that provide evidence for the binding of the polysulfide inside the β-cyclodextrin cavity, as well as for the selective capture of the polysulfide/β-cyclodextrin complexes by the α-HL nanopore. In the present process, the polysulfide/β-cyclodextrin complexes formed in solution are specifically blocked by a nanopore embedded in a lipid membrane.
An extension of this work with respect to the Li–S technology is imminent and it enlists both the exploitation of cyclodextrin molecular recognition and its thermo-responsiveness9 for enhancing battery performances (Supplementary Figs. 18 and 19). To our knowledge, stimulus-responsive10 host-guest interaction properties have never been envisioned in batteries and we believe the described polysulfide/cyclodextrin interaction may offer some opportunities. At this stage with respect to practicality, we investigated the thermal stability of polysulfides/cyclodextrin complexes by 1H NMR experiments rising the temperature (Supplementary Fig. 20) and through in-situ temperature cycling experiments (Fig. 6a). We found that the complexes can dissociate and form with a fast kinetic over a narrow temperature range in a neatly reversible way by increasing and decreasing the temperature, respectively. This suggests that temperature can be used as a stimulus to regulate the capture and release of polysulfide species by cyclodextrins (Fig. 6). An envisioned scenario, as pictured in Fig. 6b, could be a separator membrane hosting cyclodextrins that could regulate polysulfides on demand. According to this scenario the diffusion of polysulfides intermediates towards the Li anode would be prevented while letting them participate in the redox process by thermal regeneration of the membrane.
a 1H NMR temperature cycling experiment (600 MHz, D2O, 25 mM NaHCO3, pH 10) on α-cyclodextrin (10 mM) in presence of Na2S5 (100 mM). Fraction of α-cyclodextrin complexed, given by the relation of \({\mathrm{\Delta}} {\mathrm{H}}_3/{\mathrm{\Delta}} {\mathrm{H}}_{3{\mathrm{max}}}\) (See “Methods” section), is represented as a function of temperature during heating up (60 °C, 55.1 ± 0.1%) and cooling down (25 °C, 84.3 ± 0.1%) cycles, where confidence intervals are calculated as reported in “Methods” section. Because of technical restriction of the NMR probe, we could not go to higher temperature. We expect an almost total dissociation of the complex being achieved increasing the temperature though. b Conceptual illustration of a thermo-responsive cyclodextrin material as an efficient regenerative Li–S battery separator.
Besides, in light of these results, the nanopore technology stands as being a powerful sensing tool to probe at the molecular level the sulfur speciation in aqueous Na-ion/polysulfide medias39,40,41 and to test the hypothesis of manufacturing such membrane proposed in Fig. 6. Hence it may be envisioned as a novel in-situ probe of the unwanted polysulfide shuttle effect with a reliable identification of the intermediate species. However, we must admit that implementing our developed methodology to discriminate polysulfides in organic solvent would not be straightforward. It will call first for a thorough study of the polysulfide/cyclodextrin complexation equilibrium in organic electrolytes that differs from the one in water, because of the different dielectric constants between both media54. Moreover, as non-aqueous electrolytes are not ideal for biological nanopores and lipid membranes, an alternative way to conduct nanopore sensing could consist, as being planned, in considering solid-state nanopores made of polymers or semi-conductors55 mimicking biological ionic channels such as the α-HL. Such tailored membranes will offer a nice playground for molecule discrimination near real battery conditions because of their chemical stability in organic solvents. Independently of the approach to be pursued, it remains that this nanopore technique could provide an extra analytical tool to the battery community for discriminating at the molecular level, any parasitic redox products diffusing in the electrolyte. Bearing in mind that the formation of such products can poison the positive electrode, an obvious extension of this work regards the discrimination of the soluble alkyl carbonates species well known as troublemakers in some battery systems56.
In summary, a comprehensive multidisciplinary approach combining spectroscopic and electrophysiological experiments was adopted to understand deeply and at the molecular level the host-guest complexation equilibrium between the cyclodextrins and polysulfides and this was complemented by Molecular Dynamics simulations and Docking calculations. We have used this rational strategy to enable single-molecule nanopore detection, and demonstrated this technique as being a powerful tool to discriminate in aqueous medium different polysulfides with a single sulfur atom resolution, hence specifically sequencing species migrating through a membrane. That perfectly fits with the need for increasing the selectivity of separators, which could completely prevent cross communication between two electrodes in various battery technologies. Finally, the thermally controlled polysulfide/cyclodextrin complex equilibrium offers opportunities to synthesize stimuli-responsive cyclodextrins materials for the development of novel regenerative separators. Although at its early stage and being aware that further developments are needed, we hope this work to pave the way towards smarter batteries with novel regenerative and sensing separators.
Methods
All materials were used as received without further purification step.
Preparation of sodium polysulfides solutions
Sodium polysulfides solutions (100 mM) were prepared according to a previously described method57. In brief, they are formed by adding, Na2S and S (purchased from Sigma-Aldrich) in stoichiometric ratios to a degassed aqueous solution buffered with 25 mM NaHCO3 and set to pH 10 under continuous stirring for a week. All the preparations are done in an Argon filled glovebox.
The characterization of the different polysulfides solutions was performed by UV-vis absorbance spectroscopy through the calculation of the different molar extinction coefficients (ε). Absorbance spectral profiles were measured over 200–1100 nm in a 1 cm airtight quartz cuvette for different diluted polysulfides solutions (0.3 mM to 1 mM) with a UV5 Bio Mettler Toledo spectrophotometer (Supplementary Fig. 1).
NMR titration experiments: polysulfide/cyclodextrin complexation
Both continuous variation method and titration experiments were performed as previously described44. Association constants (Ka) for the inclusion polysulfides/cyclodextrin complexes were determined in sodium bicarbonate deuterated buffer D2O (25 mM NaHCO3, pH 10) at 298 K by measuring the chemical shift variations in the 1H NMR spectra (600 MHz) of a solution of the native cyclodextrin (10 mM) in the absence and in the presence of increasing amounts of polysulfides. Polysulfides solutions were prepared as previously mentioned but in a deuterated buffer. Different diluted solutions of polysulfides (from 10 mM to 100 mM) were mixed with pre-weighted cyclodextrin (±1 mg) under Argon atmosphere. It is critically important to maintain a constant host concentration, pH and ionic strength throughout the titration experiment. Mixtures were vortexed for 1 min before acquisition of the 1H NMR spectrum on a 600 MHz Bruker spectrometer. The chemical shifts of selected host protons resonances obtained at approximately 10 different host-guest concentration ratios with Top Spin 3.6 were plotted against the concentration of polysulfide. An iterative least-squares fitting procedure using a 1:1 stoichiometry binding model was performed with Igor Pro 6.37 to estimate for each polysulfides/cyclodextrin complex the association constant (Ka) and the maximum chemical shift \(\left( {{\mathrm{\Delta }}_{{\mathrm{max}}}} \right)\) which is the chemical shift recorded if 100% complexation is achieved.
The one-to-one stoichiometry of the polysulfides/cyclodextrin complexes was determined using continuous variation method, also known as Job’s plot. The mole fraction of polysulfide was varied while keeping the total concentration of the cyclodextrin and polysulfide constant, respectively, 50 mM, 16 mM, and 50 mM for α-cyclodextrins, β-cyclodextrins, and γ-cyclodextrins. The chemical shifts of selected host protons (H3 essentially) resonances were obtained at approximately 8 different host-guest concentration ratios with Top Spin 3.6 and were plotted against the stoichiometric ratio:
According to the method, a second order polynomial fit was performed, and the stoichiometry of the studied complex was obtained from the x-coordinate at the maximum of the plot for which the concentration of the complexed cyclodextrin population is the highest.
The statistic error relative to the chemical shift measurement was obtained by the calculation of the standard deviation of 10 similar and independent points and is estimated to 9%. The plotted error bars were calculated considering the uncertainties propagation assuming the experimental errors (dilution and weighting) for each condition.
Electrical detection, data acquisition, and analysis
Membrane lipid bilayers were made according to previously described methods58,59. In brief, a film of a 1% solution of diphytanoyl-phosphatidylcholine-lecithin (Avanti) in anhydrous decane (Sigma) was spread across a 150 μm diameter hole drilled in a polysulfone wall separating the two compartments of a chamber. Each compartment contained 900 µL of 1 M NaCl, 25 mM NaHCO3, pH 10 in an Argon filled glovebox. After thinning of the decane film and formation of a planar lipid bilayer, a single α-hemolysin (α-HL) pore is inserted by adding monomeric α-HL (Sigma) from a stock solution into the cis compartment. Insertion and orientation of the α-HL pore was systematically checked to ensure transport of β-cyclodextrin or Na2Sx/β-cyclodextrin complex into the stem part of the pore. 100 µL of 10 mM β-cyclodextrin or 10 mM Na2Sx/β-cyclodextrin complex was added to the trans compartment. Each data set was collected using independent experiments and different pores.
Data were collected using Chimera Instruments VC100 at a sampling rate of 4.17 MHz and low-pass filtered at 10 kHz. Data were analyzed with a homemade macro using Igor software (Wavemetrics). The event measurements were based on a statistical analysis of the current traces: 425, 3904, 1254, and 1178 events analyzed for β-cyclodextrin, Na2S3/β-cyclodextrin, Na2S4/β-cyclodextrin and Na2S5/β-cyclodextrin, respectively. The statistical analysis of the current traces has been previously described60. As the blockades shorter than 100 µs are attributed to bumpings of β-cyclodexrin or complexes at the entrance of the channel, we removed them from the scatter plot to obtain the Fig. 4c. Nevertheless, in this figure, blockades could be attributed both to β-cyclodextrin or complexes. In order to discriminate both blockade types, we took the blockade ratio into account (Fig. 4d), which depends on the nature, shape of the analyte. From the blockade ratio distribution, we removed the part attributed to the β-cyclodextrin, which was previously characterized in Fig. 5b, left (blue). Finally, we obtained the blockade ratio distribution attributed to the complexes (Fig. 5b). The center of each blockade ratio distribution was calculated from the average of results obtained from 5 different positions of the interval fits.
Atomistic simulation of the α-HL nanopore
Molecular Dynamics simulation set-up and production runs for estimating the ionic and electro-osmotic flows through the α-HL nanopore were performed as previously described61. In brief, we employed the H++ server62, version 3.2, to determine the probability that each titratable residue of the α-HL is protonated or not. This analysis allowed us to select two different plausible protonation states for the α-HL at pH 10, see Supplementary Figs. 12, 13, and 14 and Supplementary Note 2. For each of the α-HL protonation states, a triperiodic simulations box containing the α-HL nanopore embedded in a lipid membrane was assembled by using protocols described in previous works61,63. The POPC lipid membrane, the water molecules and the ions were added using VMD64. The system is neutralized and solvated at 1 M NaCl and then equilibrated (T = 310 K, p = 1 atm). The equilibrated configuration was used as initial condition for non-equilibrium runs where a uniform and constant external electric field was applied perpendicularly to the lipid bilayer, ΔV = EzLz. Each simulation was run for 100 ns and frames were saved every 20 ps. Average currents and electroosmotic flow were calculated, similarly to previously reported61,63, after discarding a 20 ns transient. Errors were estimated using block average protocol, with block length 5 ns. All the Molecular Dynamics simulations were performed using the NAMD software65. The CHARMM36 force field66 was employed for the lipids membrane and the α-HL nanopore, while TIP3P model was used for water67. CUFIX corrections were applied to ions68.
Nanopore hindrance estimator
The pore resistance can be written as an integral function of the electrolyte accessible area profile along the pore axis A(z), as previously described53. The A(z) profiles for the empty pore, A0, and with the β-cyclodextrin inside the pore, AβCD, were estimated using Molecular Dynamics simulations. A0 was estimated through the simulation pH10a, discussed in Supplementary Note 2. AβCD was estimated through a second simulation, prepared locating a cyclodextrin molecule close to the α-HL constriction. The position of the cyclodextrin was chosen in accordance to the crystal structure of the M113F α-HL mutant complexed with β-cyclodextrin (PDB ID: 3M3R), by aligning the 3M3R α-HL structure upon the pre-equilibrated system pH10a. Alignment was performed minimizing the RMSD between the α-carbon of the protein. After alignment, the atom coordinates of the β-cyclodextrin were extracted from the 3M3R structure, and implemented into the pH10a system, deleting the overlapping water molecules. A first 1 ns NPT equilibration run was performed, keeping the cyclodextrin glycosidic oxygens (CHARMM type OG301) harmonically constrained (k = 2.00 kcal mol−1 Å−1) along the z-coordinates. Then, a second 16 ns NVT equilibration run was performed, with a constant electric field Ez applied along the z-axis, corresponding to a negative voltage of 0.5 V applied to the trans-side of the pore. Glycosidic oxygens of cyclodextrin, α-carbons of the protein and phosphorus of lipid heads were kept constrained (k = 0.5 kcal mol−1 Å−1). Finally, a 32-ns NVT production run were performed, letting the cyclodextrin free and keeping the other constrains same as the previous NVT. From the production run, the AβCD area profile was estimated. Regarding polysulfides, since, to the best of our knowledge, there was no reliable Molecular Dynamics force field to simulate the dynamical behavior of these molecules in our environment, we assumed in a first approximation that the volume of each single polysulfide (S3, S4, S5) species is simply spread in a cylinder of length L = 9 Å. So, we subtracted a constant effective area for each polysulfide to the area profile AβCD, from z0 = 36 Å. The molecular volume of each polysulfide was calculated treating atoms as spheres and measuring how many points of a dense grid (0.1 Å3) were within a Van der Waals radius from each atom center. The volumetric occupancy map was computed using VMD’s Volmap tool64.
Atomistic simulation of cyclodextrins in a NaCl electrolyte solution
Molecular Dynamics simulations of the three types of cyclodextrin, (α-cyclodextrin, β-cyclodextrin, and γ-cyclodextrin) were performed to study the flexibility and characteristic conformations of these molecules in water. As initial cyclodextrin structures we used the eight medoids resulting from the cluster analysis of the crystallographic structures extracted from the Protein Data Bank46 (see Supplementary Fig. 7 and Supplementary Note 1.1). For each medoid, we performed three independent Molecular Dynamics simulation replicas. Each simulation was composed of an ionized water box, 1 M NaCl, containing a single cyclodextrin molecule. The cyclodextrin molecules were modeled using the CHARMM General Force Field69. The TIP3 model was used for the water molecules67, and CUFIX corrections were used for ions68. The initial dimensions of the cell are 50 × 50 × 50 Å3. The system was first minimized for 100 steps with steepest descent gradient method and then equilibrated at constant pressure (1 atm) and temperature (300 K), for 2 ns. Then, a NVT production run of 160 ns was performed. Coordinates were sampled each 20 ps. Only the last 80 ns of each simulation were considered for the clustering analysis, resulting in a total of 4000 frames. All the simulations were performed with NAMD software65. A structural clustering analysis, similar to the one above described for the selection of the PDB structures and fully detailed in Supplementary Note 1, was applied to extract the most representative cyclodextrin conformations. The medoids of the clustered structures, showed in Supplementary Fig. 8, were then taken as receptors in the docking calculations.
Molecular Docking of polysulfides with cyclodextrins
To provide molecular information on the association of the inclusion complexes formed by polysulfides S3, S4, S5 and cyclodextrins, we performed a series of Molecular Docking calculations. In a first set of calculations we used as receptors all the cyclodextrins crystallographic structures extracted from the Protein Data Bank46, as described in Supplementary Note 1. In a second set, we used as receptors the structures generated by Molecular Dynamics simulations. The structures of the polysulfides ligands S3, S4, S5 were extracted from crystallized proteins in the Protein Data Bank46, corresponding to the query ID PS5, S4P and S3H, respectively. All the calculations were performed using Autodock Vina70. Ligands input files were prepared as described in Supplementary Note 1.2. The center of the box was aligned with the center of cyclodextrin glycosidic oxygen while the search space is 20 × 20 × 20 Å3. The results for each calculation are the affinity score (kcal mol−1) values for each ligand conformation in its respective complex.
Data availability
The authors declare that the main data supporting the findings of this study are available within the article and its Supporting Information file. Extra data are available from the corresponding authors on reasonable request.
Code availability
All MD simulation trajectories were generated using the NAMD2 software package http://www.ks.uiuc.edu/Research/namd. Trajectory analysis are performed using the VMD software https://www.ks.uiuc.edu/Research/vmd/. Nanopore hindrance estimation was carried out using the method described in ref. 53; the FORTRAN code used to compute the area profiles is available at https://github.com/giodimuccio/channelSearch.
References
Larcher, D. & Tarascon, J.-M. Towards greener and more sustainable batteries for electrical energy storage. Nat. Chem. 7, 19 (2015).
Tarascon, J.-M. & Armand, M. Materials for Sustainable Energy: A Collection of Peer-Reviewed Research and Review Articles from Nature Publishing Group. 171–179 (World Scientific, 2011).
Nitta, N., Wu, F., Lee, J. T. & Yushin, G. Li-ion battery materials: present and future. Mater.Today 18, 252–264 (2015).
Grey, C. & Tarascon, J. Sustainability and in situ monitoring in battery development. Nat. Mater. 16, 45 (2017).
Kwon, T.-W. Choi, J. W. & Coskun, A. Prospect for supramolecular chemistry in high-energy-density rechargeable batteries. Joule 3, 662–682 (2019).
Wang, H. et al. Recent advances on self‐healing materials and batteries. ChemElectroChem 6, 1605–1622 (2019).
Kwon, T.-w., Choi, J. W. & Coskun, A. The emerging era of supramolecular polymeric binders in silicon anodes. Chem. Soc. Rev. 47, 2145–2164 (2018).
Zhao, Y. et al. A self‐healing aqueous lithium‐ion battery. Angew. Chem. Int. Ed. 55, 14384–14388 (2016).
Zheng, Y., Hashidzume, A., Takashima, Y., Yamaguchi, H. & Harada, A. Temperature-sensitive macroscopic assembly based on molecular recognition. ACS Macro Lett. 1, 1083–1085 (2012).
Harada, A., Takashima, Y. & Nakahata, M. Supramolecular polymeric materials via cyclodextrin–guest interactions. Acc. Chem. Res. 47, 2128–2140 (2014).
Inoue, Y. et al. Thermal and photochemical switching of conformation of poly (ethylene glycol)-substituted cyclodextrin with an azobenzene group at the chain end. J. Am. Chem. Soc. 129, 6396–6397 (2007).
Davis, M. E. & Brewster, M. E. Cyclodextrin-based pharmaceutics: past, present and future. Nat. Rev. Drug Discov. 3, 1023 (2004).
Astier, Y., Braha, O. & Bayley, H. Toward single molecule DNA sequencing: direct identification of ribonucleoside and deoxyribonucleoside 5′-monophosphates by using an engineered protein nanopore equipped with a molecular adapter. J. Am. Chem. Soc. 128, 1705–1710 (2006).
Gu, L.-Q., Braha, O., Conlan, S., Cheley, S. & Bayley, H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398, 686 (1999).
Choi, S., Kwon, T.-w., Coskun, A. & Choi, J. W. Highly elastic binders integrating polyrotaxanes for silicon microparticle anodes in lithium ion batteries. Science 357, 279–283 (2017).
Imholt, L. et al. Supramolecular self-assembly of methylated rotaxanes for solid polymer electrolyte application. ACS Macro Lett. 7, 881–885 (2018).
Imholt, L. et al. Grafted polyrotaxanes as highly conductive electrolytes for lithium metal batteries. J. Power Sour. 409, 148–158 (2019).
Ni, L. et al. Supramolecular complexation of polysulfides by β-cyclodextrin polymer functionalized graphene hybrid cathode for high-performance lithium-sulfur batteries. Energy Storage Mater. 21, 378–389 (2019).
Bai, S., Liu, X., Zhu, K., Wu, S. & Zhou, H. Metal–organic framework-based separator for lithium–sulfur batteries. Nat. Energy 1, 16094 (2016).
Li, C. et al. Polysulfide-blocking microporous polymer membrane tailored for hybrid Li-sulfur flow batteries. Nano Lett. 15, 5724–5729 (2015).
Vizintin, A. et al. Fluorinated reduced graphene oxide as an interlayer in Li–S batteries. Chem. Mater. 27, 7070–7081 (2015).
Steudel, R. & Chivers, T. The role of polysulfide dianions and radical anions in the chemical, physical and biological sciences, including sulfur-based batteries. Chem. Soc. Rev. 48, 3279–3319 (2019).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nat. Biotechnol. 26, 1146–1153 (2008).
Cao, C. et al. Discrimination of oligonucleotides of different lengths with a wild-type aerolysin nanopore. Nat. Nanotechnol. 11, 713 (2016).
Piguet, F. et al. Identification of single amino acid differences in uniformly charged homopolymeric peptides with aerolysin nanopore. Nat. Commun. 9, 966 (2018).
Wang, Y., Zheng, D., Tan, Q., Wang, M. X. & Gu, L.-Q. Nanopore-based detection of circulating microRNAs in lung cancer patients. Nat. Nanotechnol. 6, 668 (2011).
Huang, G., Voet, A. & Maglia, G. FraC nanopores with adjustable diameter identify the mass of opposite-charge peptides with 44 dalton resolution. Nat. Commun. 10, 1–10 (2019).
Quick, J. et al. Real-time, portable genome sequencing for Ebola surveillance. Nature 530, 228 (2016).
Restrepo-Pérez, L. et al. Resolving chemical modifications to a single amino acid within a peptide using a biological nanopore. ACS Nano 13, 13668–13676 (2019).
Ouldali, H. et al. Electrical recognition of the twenty proteinogenic amino acids using an aerolysin nanopore. Nat. Biotechnol. 38, 176–181 (2020).
Restrepo-Pérez, L., Joo, C. & Dekker, C. Paving the way to single-molecule protein sequencing. Nat. Nanotechnol. 13, 786–796 (2018).
Wanunu, M. Nanopores: a journey towards DNA sequencing. Phys. Life Rev. 9, 125–158 (2012).
Galenkamp, N. S., Biesemans, A. & Maglia, G. Directional conformer exchange in dihydrofolate reductase revealed by single-molecule nanopore recordings. Nat. Chem. 12, 1–8 (2020).
Merstorf, C. L. et al. Wild type, mutant protein unfolding and phase transition detected by single-nanopore recording. ACS Chem. Biol. 7, 652–658 (2012).
Simpson, J. T. et al. Detecting DNA cytosine methylation using nanopore sequencing. Nat. Methods 14, 407 (2017).
Yu, J., Cao, C. & Long, Y.-T. Selective and sensitive detection of methylcytosine by aerolysin nanopore under serum condition. Anal. Chem. 89, 11685–11689 (2017).
Zernia, S., van der Heide, N. J., Galenkamp, N. S. P., Gouridis, G. & Maglia, G. Current blockades of proteins inside nanopores for real-time metabolome analysis. ACS Nano 14, 2296–2307 (2020).
Ouldali, H. et al. Electrical recognition of the twenty proteinogenic amino acids using an aerolysin nanopore. Nat. Biotechnol. 38, 1–6 (2019).
Tekin, B., Sevinc, S., Morcrette, M. & Demir‐Cakan, R. A new sodium‐based aqueous rechargeable battery system: the special case of Na0. 44MnO2/dissolved sodium polysulfide. Energy Technol. 5, 2182–2188 (2017).
Sevinc, S. et al. In-situ tracking of NaFePO4 formation in aqueous electrolytes and its electrochemical performances in Na-ion/polysulfide batteries. J. Power Sour. 412, 55–62 (2019).
Gross, M. M. & Manthiram, A. Development of low-cost sodium-aqueous polysulfide hybrid batteries. Energy Storage Mater. 19, 346–351 (2019).
Manthiram, A., Fu, Y. & Su, Y.-S. Challenges and prospects of lithium–sulfur batteries. Acc. Chem. Res. 46, 1125–1134 (2012).
Xiong, S., Xie, K., Diao, Y. & Hong, X. Characterization of the solid electrolyte interphase on lithium anode for preventing the shuttle mechanism in lithium–sulfur batteries. J. Power Sour. 246, 840–845 (2014).
Thordarson, P. Determining association constants from titration experiments in supramolecular chemistry. Chem. Soc. Rev. 40, 1305–1323 (2011).
Rekharsky, M. V. & Inoue, Y. Complexation thermodynamics of cyclodextrins. Chem. Rev. 98, 1875–1918 (1998).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Bhamidimarri, S. P., Prajapati, J. D., van den Berg, B., Winterhalter, M. & Kleinekathöfer, U. Role of electroosmosis in the permeation of neutral molecules: CymA and cyclodextrin as an example. Biophys. J. 110, 600–611 (2016).
Piguet, F. et al. Electroosmosis through α-hemolysin that depends on alkali cation type. J. Phys. Chem. Lett. 5, 4362–4367 (2014).
Gu, L.-Q. & Bayley, H. Interaction of the noncovalent molecular adapter, β-cyclodextrin, with the staphylococcal α-hemolysin pore. Biophys. J. 79, 1967–1975 (2000).
Gu, L.-Q., Cheley, S. & Bayley, H. Prolonged residence time of a noncovalent molecular adapter, β-cyclodextrin, within the lumen of mutant α-hemolysin pores. J. General Physiol. 118, 481–494 (2001).
Tegman, R. The crystal structure of sodium tetrasulphide, Na2S4. Acta Crystallogr. Sec. B 29, 1463–1469 (1973).
Szejtli, J. Introduction and general overview of cyclodextrin chemistry. Chem. Rev. 98, 1743–1754 (1998).
Di Muccio, G., Rossini, A. E., Di Marino, D., Zollo, G. & Chinappi, M. Insights into protein sequencing with an α-Hemolysin nanopore by atomistic simulations. Scie. Rep. 9, 6440 (2019).
Cuisinier, M., Hart, C., Balasubramanian, M., Garsuch, A. & Nazar, L. F. Radical or not radical: revisiting lithium–sulfur electrochemistry in nonaqueous electrolytes. Adv. Energy Mater. 5, 1401801 (2015).
Lepoitevin, M., Ma, T., Bechelany, M., Janot, J.-M. & Balme, S. Functionalization of single solid state nanopores to mimic biological ion channels: a review. Adv. Colloid Interface Sci. 250, 195–213 (2017).
Yan, G. et al. Assessment of the electrochemical stability of carbonate-based electrolytes in Na-ion batteries. J. Electrochem. Soci. 165, A1222–A1230 (2018).
Pringle, D. L. The Nature of the Polysulfide Anion. (1967).
Gutsmann, T., Heimburg, T., Keyser, U., Mahendran, K. R. & Winterhalter, M. Protein reconstitution into freestanding planar lipid membranes for electrophysiological characterization. Nat. Protocol. 10, 188 (2015).
Montal, M. & Mueller, P. Formation of bimolecular membranes from lipid monolayers and a study of their electrical properties. Proc. Natl Acad. Sci. USA 69, 3561–3566 (1972).
Cressiot, B. et al. Aerolysin, a powerful protein sensor for fundamental studies and development of upcoming applications. ACS Sensors. 4, 530–548 (2019).
Bonome, E. L., Cecconi, F. & Chinappi, M. Electroosmotic flow through an α-hemolysin nanopore. Microfluid. Nanofluid. 21, 96 (2017).
Anandakrishnan, R., Aguilar, B. & Onufriev, A. V. H++ 3.0: automating pK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations. Nucleic Acids Res. 40, W537–W541 (2012).
Aksimentiev, A. & Schulten, K. Imaging α-hemolysin with molecular dynamics: ionic conductance, osmotic permeability, and the electrostatic potential map. Biophys. J. 88, 3745–3761 (2005).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graphics 14, 33–38 (1996).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
MacKerell, A. D. Jr. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Yoo, J. & Aksimentiev, A. Improved parameterization of amine–carboxylate and amine–phosphate interactions for molecular dynamics simulations using the CHARMM and AMBER force fields. J. Chem. Theory Comput. 12, 430–443 (2015).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug‐like molecules compatible with the CHARMM all‐atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Acknowledgements
This work was supported by grant funding from the Region Ile-de-France in the framework of DIM ResPore and the Réseau sur le Stockage Electrochimique de l’Energie (RS2E) through F.B. PhD scholarship. F.B. and N.J. are grateful to the Conseil Regional d’Ile de France for the acquisition of the 600 MHz NMR spectrometer of University of Evry Val d’Essonne. J.-M.T. acknowledges the funding from the European Research Council (ERC) (FP/2014)/ERC Grant-Project 670116-ARPEMA. This research was supported by ANR-17-CE09-0044-02 (J.P., B.C., L.B.). We acknowledge support from Genopole for financing B.C. CSCS, the Swiss National Super-computing Centre, supported this work through the project ID s865 to M.C. and G.D.M.
Author information
Authors and Affiliations
Contributions
F.B., N.J., and J.-M.T. conceived the experiments regarding the polysulfides-cyclodextrin interactions that F.B. carried out. B.C. and J.P. designed and conducted the nanopore sensing experiments while L.B., B.C., and J.P. analyzed and interpreted the nanopore data. G.D.M., B.M.D.R., and M.C. performed the molecular docking calculations and molecular dynamics simulations. Finally, J.-M.T., F.B., and J.P. wrote the first draft of the manuscript. All authors contributed to editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bétermier, F., Cressiot, B., Di Muccio, G. et al. Single-sulfur atom discrimination of polysulfides with a protein nanopore for improved batteries. Commun Mater 1, 59 (2020). https://doi.org/10.1038/s43246-020-00056-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43246-020-00056-4