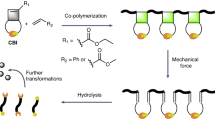
Abstract
Improving the efficiency of enzymes towards decomposing substrates has been one of the central goals in the biotechnology industry. However, the modification of enzymes for upgrading natural materials to high-value performant materials is largely unexplored. Here, we demonstrate that the ancestral form of a Cel5A bacterial endoglucanase, unlike its modern descendant from Bacillus subtilis, was able to generate cellulose nanocrystals (EnCNC) chemically pure, maintaining native cellulose structure and displaying higher thermal stability and crystallinity than standard CNC obtained by acidic treatment. We demonstrate that EnCNC alone is a suitable matrix to grow cells in 2D and 3D cultures. Importantly, EnCNC accepts well graphene derivatives to fabricate conductive hybrids inks forming a stable flat surface where cells also attach and proliferate. Our results demonstrate that EnCNC has physicochemical properties unattainable with standard CNC, making it a unique material ideal as a matrix for the design of biocompatible advanced materials for tissue engineering and other applications.
Similar content being viewed by others
Introduction
Over the past decades, the biotechnology industry has incorporated a plethora of enzymes to numerous bioconversion processes, ranging from bioenergy1 to food industry2. Generally, most enzymes are improved to achieve high catalytic yields under the desired conditions3. Towards this end, several techniques for enzyme design have been central for the development of the biotechnology industry4,5,6. Interestingly, a less investigated aspect in biotechnology is how to modify enzymes in order to upgrade or transform substrates into high-value performants to be used as platform materials7. Such development would expand the range of applicability of enzymes, making them useful not only for downgrading but also upgrading new biomaterials derived from enzymatic transformations. This can contribute to the development of a sustainable bio-based technology.
Biomass is a traditional target for enzyme decomposition due to the numerous products that can be obtained from bioresources8,9. However, given the polymeric nature of bioresources such as chitin, starch or cellulose, it would be desirable to transform them into high-value materials for application in material science and biotechnology. One such material is nanocellulose, a nano-sized form of cellulose. The extraordinary physicochemical features of nanocellulose have made it an attractive material for applications in material science and biotechnology10,11. Nanocellulose is obtained from cellulose whose hierarchical organization renders different types of nanocellulose (Fig. 1). One of the most sought-after forms is cellulose nanocrystals (CNC)12, which are small crystalline particles with lengths ranging from 100 to 500 nm. Currently, CNC production is mainly achieved through chemical treatments using sulfuric acid that swells cellulose amorphous regions keeping the crystalline part. This process leaves sulfate groups attached to the CNCs surface affecting their physicochemical properties13. Enzyme hydrolysis has been proposed for nanocellulose production6. However, CNC obtention has only been possible through the use of complex enzyme systems9. Obtention of CNC by a single enzyme ex profeso has remained challenging.
The hierarchical structure of cellulose fiber permits its degradation into nano scale particles. Enzymatic hydrolysis was performed in water at 50 °C at different times in agitation with different endoglucanase. Here, we show how endoglucanase with CBM can attach to the cellulose fiber placing the catalytic domain in the fiber surface and improving its activity. Enzymes degradation can produce nanocellulose with different sizes depending on the hydrolysis time and enzyme usage. Cellulose nanocrystals (CNC) are small crystalline particles with lengths between 100 and 500 nm and diameters between 3 and 40 nm.
Ancestral sequence reconstruction (ASR) has been proposed for enzyme engineering14,15. This technique is commonly used in molecular evolution studies16,17, but previous studies reported that ancient enzymes show extraordinary physicochemical properties18,19,20,21, making them potentially suitable for biotechnology. We have recently reconstructed a 2.8 billion-year-old ancestral endoglucanase (ANC EG) from bacteria species that showed superior performance in comparison with modern ones14. Here, we show that ANC EG alone can produce pure nanocellulose (EnCNC). The same CNC form could not be obtained by using a modern descendant from Bacillus subtilis, or other endoglucanase forms. EnCNC is chemically pure, displays higher thermal stability, crystallinity, and mechanical strength than standard CNC isolated by acid hydrolysis (AcCNC), being a grenner and sustainable process. Unlike AcCNC, EnCNC can be used as stable matrix for cell growth for 2D and 3D cultures. Also, EnCNC mixes well with graphene to fabricate hybrid conductive biomaterials where cells attach and proliferate. We present EnCNC as a potential platform for bioinks, biosensors, and other biocompatible advanced materials.
Results
EnCNC production and characterization
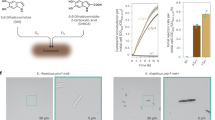
ANC EG belongs to the last common ancestor of firmicutes (LFCA) and has been fully characterized previously, including its substrate-interaction mechanism14, demonstrating high activity over a wide range of conditions and in several substrates. This enzyme can be used in two forms, only the catalytic domain or linked to a carbohydrate-binding module (CBM). In nature, most bacterial EG have a CBM that helps in the degradation of cellulose over recalcitrant substrates. In this work, we have obtained the catalytic domain ANC EG as well as the form containing the CBM, i.e., ANC EG+CBM, given the impossibility of constructing an ancestral CBM, we used the CBM from B. subtilis. Both variants were used to hydrolyze pure cellulose at 50 °C, monitoring nanocellulose production versus hydrolysis time (Fig. 2a). We compared the yield to that of the modern descendant from B. subtilis, used as query sequence for the ancestral reconstruction with its own CBM, therefore the difference between ANC EG and B. subtilis EG can be only attributed to the catalytic domain. We also used a modern bacterial EG from Thermotoga maritima, that does not have a native CBM. In all cases the ancestral variants showed a higher yield, with the variant ANC EG+CBM reaching eight-fold that of T. maritima.
a Nanocellulose conversion from filter paper using ANC EG, ANC EG+CBM, and EG from T. maritima, B. subtillis, and T. reesei at 50 °C at different times. The average yield and S.D. were calculated from three independent experiments. b AFM images of nanoparticles produced by EGs hydrolysis at different time points (5 µm × 5 µm). The images show the nanoparticles that the EGs produced at 1, 5, 24, and 48 h using filter paper as a substrate. c Average size of nanocellulose from filter paper using ANC EG, ANC EG+CBM, and EG from T. maritima, B. subtillis, and T. reesei at 50 °C at different times. The smaller length was measured after 24 h ANC EG+CBM hydrolysis. The average yield and SEM were calculated from three independent experiments. d Comparison of size distribution of nanocellulose produced with enzymatic hydrolysis and AcCNC. The distribution was calculated with the length of 100 particles from each condition. e AFM images of nanoparticles produced with ANC EG+CBM hydrolysis and AcCNC (3 µm × 3 µm). Height images (I) of nanocellulose produced by ancestral endoglucanase with CBM at 24 h hydrolysis using filter paper as substrate and (II) of nanocellulose produced by sulfuric acid hydrolysis purchased from Maine University. (III) A 3D image of a single EnCNC, showing the needle-like shape and (IV) a 3D image from the AcCNC sample with ribbon-like shape.
After hydrolysis we analyzed nanocellulose morphology using atomic force microscopy (AFM) (Fig. 2b and Supplementary Table 1). ANC EG+CBM hydrolysis produced smaller particles compared to ANC EG (Fig. 2c). At 1-h hydrolysis we observed a mixture of EnCNC and cellulose nanofibers (CNF) of ~1 μm and above. After 24 h hydrolysis we obtained a homogenous EnCNC suspension with average length of 408 nm and diameters of 12.5 nm, measurements described as CNC22. After 48 h hydrolysis we observed bigger particles in both cases probably due to CNCs aggregations (Fig. 2c). Also, the activity of ancestral endoglucanase deceased after 24 h, favoring the agglomeration of the crystals at longer times. In the case of EGs from B. subtilis and T. maritima EG, they mostly provided CNF of ~800 nm and above (Fig. 2c). We have been able to obtain smaller particles (~600 nm) using an enzyme preparation from Trichoderma reesei but still with lower efficiency than that of ANC EG+CBM (Fig. 2c). We have been able to obtain smaller particles (~600 nm) using an enzyme preparation from T. reesei but still with lower efficiency than that of ANC EG+CBM (Fig. 2c). We plotted the size population distribution from all enzymatic hydrolysis to analyze the suspensions homogeneity (Supplementary Fig. 1). The aspect ratio of EnCNC obtained with ANC EG corresponded to CNC with cellulose I polymorph23 and it is within the interval usually measured in CNC (<100)24.
We also compared commercial AcCNC with EnCNC produced after 24 h hydrolysis (Fig. 2d, e), and EnCNC obtained using ANC EG+CBM was longer but maintained similar diameters to AcCNC (Supplementary Table 1). EnCNC showed a needle-like morphology due to its high aspect ratio. Conversely, AcCNC has a ribbon-like morphology with lower aspect ratio, which is in agreement with previous reports, and allows to classify EnCNC as cellulose I and AcCNC as cellulose II25. Analysis using FTIR and its second derivatives spectra confirm the native cellulose structure of EnCNC when compared to AcCNC26 (Supplementary Figs. 2–5). AcCNC spectra also display the sulfate groups from the acidic hydrolysis measured at 0.95% (Supplementary Fig. 6). Sulfate groups have a direct impact on the physicochemical properties of nanocrystals, allowing AcCNC to generate a stable suspension in water due to negatively charged OSO3− groups. Meanwhile, EnCNC generates an opaque dispersion in which aggregates and sedimentation are apparent due to the absence of charged groups (blue diagram in Fig. 3a). The stability of EnCNC and AcCNC water dispersions were analyzed by measuring the zeta potencial (ζ). The zeta potential values for EnCNC was −32 mV and for AcCNC was −54 mV, highlighting the good stability of both suspension27. This difference showed the higher surface charge of AcCNC due to the sulfate groups which is reflected in the higher stability. Although, the lower zeta potential of EnCNC suspension is desired for biomedical applications28.
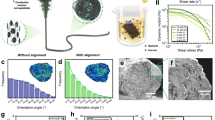
a Aqueous suspension of EnCNCs and AcCNCs. Illustration of free –OH groups in EnCNC surface and the random –OSO3− groups anchored by the sulfuric acid treatment in the AcCNC surface. b Topography images from EnCNC and AcCNC samples used for nano-FTIR measurement. Color circles mark the analyzed nanocrystals. c, d Spectra from EnCNC and AcCNC nanocrystals selected in two different regions, 3600 to 3200 cm−1 and 1800 to 750 cm−1. Spectra form EnCNC and AcCNC showed different peaks due to its different polymorph structure and AcCNC sulfate content. e X-ray diffraction analysis of the filter paper, EnCNCs produced by 24 h hydrolysis of ANC EG and ANC EG+CBM and AcCNCs. Figure shows the diffractograms of cellulose I in the filter paper and EnCNCs. The different planes are indicated. f Thermogravimetric analysis curves of filter paper, EnCNCs produced by 24 h hydrolysis of ANC EG and ANC EG+CBM and AcCNCs. Weight loss curves and derivative from TGA curves.
We further analyzed single EnCNC particles using nano-FTIR, which allows infrared analysis with nanoscale resolution (Fig. 3b)29. The color circles in the topography images in Fig. 3 mark the crystals selected for the measurement, and the corresponding nano-FTIR spectra are shown in Fig. 3c, d. We observe that only the nano-FTIR spectra of AcCNC fibers show the shoulders at around 3486 and 3441 cm−1 (marked with orange lines) that confirm the cellulose II structure. The peak at 814 cm−1 (marked with orange lines) indicates the sulfur present in individual crystals. Nano-FTIR offers improved sensitivity compared to FTIR, showing intensified O–H peaks at 3300, 3345, and 3400 cm−1 (marked with blue lines). Furthermore, only the spectra of AcCNC show a clear peak at around 1250 cm−1 (marked with orange line), which could be attributed to the asymmetrical vibration associated to sulfate groups30. Nano-FTIR spectra of AcCNC are reproducible; however, the spectra of EnCNC vary significantly suggesting that the EnCNC nanocellulose crystals have higher spatio-chemical variations (Supplementary Fig. 7).
The previous analysis probes that EnCNC is structurally and chemically different than AcCNC. We also investigate the physical properties of EnCNC. One of the most important features of CNC is its crystallinity because it correlates to its thermal and mechanical properties. The measurements of the crystallinity index (CI) using X-ray, demonstrate that EnCNC has a higher CI than AcCNC. Determined as 88% and 85% for EnCNC using ANC EG and ANC EG+CBM, respectively, versus 80% for AcCNC. The CI value of EnCNC is in the order of the value for bacterial CNC, which are considered the purest and more crystalline31. EnCNC also displays higher crystallite size, determined as 5 vs. 4 nm for EnCNC and AcCNC, respectively (Supplementary Table 2). This difference also correlates with the transformation from cellulose I to cellulose II during the acidic treatment, forming a less crystalline polymer32. This transformation as well as the lower crystallinity are also corroborated in the CP/MAS 13C NMR experiments reported in Supplementary Fig. 8. EnCNC produced by T. reesei EG hydrolysis was also analyzed to study the ability of modern EG in CNC production (Supplementary Fig. 9a, b). FTIR spectrum was similar to that of EnCNC obtained with the ancestral form, as expected. However, EnCNc produced with T. ressei EG displayed a reduction in CI%, 77.2%, in comparison with EnCNC produced by ANC EG+CBM.
We hypothesized that the observed chemical and structural differences may be reflected in physical properties such as thermal stability. CNC thermostability can be determined using TGA analysis that provides a measure of the weight loss and its derivative (DTG) (Fig. 3f and inset). In Supplementary Table 3, the onset degradation temperature (To), the maximum degradation temperature (Td), and char weight (%) are shown for EnCNCs and AcCNC and also for filter paper for comparison purposes. Maximum degradation temperature corresponded to filter paper at 361 °C33 closely followed by ANC EG+CBM with Td of 356 and 337 °C for ANC EG. This variation between the two ANC EGs could be explained to the fact that ANC EG acts randomly in the fiber in comparison with ANC EG+CBM that is processive. These values contrast with the thermal stability of AcCNC that has a To of 269 °C and a Td of 298 °C. Although it has been described that cellulose II is more thermostable than cellulose I32,34, the sulfate groups on the crystal surface of AcCNC seem to decrease the thermal stability35. Besides, this analysis was performed in EnCNC sample obtained with T. reesei EG (Supplementary Fig. 9c, d). This sample showed lower To (223 °C) than EnCNC produced by ANC EG+CBM, and similar Td. The lower crystallinity could catalyze the thermal degradation of this material. The lower To and CI% from of T. reesei EG EnCNC highlight the different physicochemical properties of EnCNC obtained with the more efficient ancestral EG.
EnCNC as matrix for cell culture
All the above determinations corroborate that EnCNC obtained with the ancestral EG is indeed a unique material with physical and chemical features distinguishable from those of standard CNC. To probe the importance of that uniqueness, we envisioned applications of EnCNC where standard AcCNC shows limitations. Both CNF and CNC are currently used in bionkins for tissue engineering due to their mechanical stability, biocompatibility, and surface chemistry possibilities; however, in the case of CNC it is mostly used as additive to prepare hydrogels as it lacks the ability to form stable matrices for 2D and 3D cultures of living cells36. A stable pure CNC hydrogel would greatly expand the possibilities for tissue engineering as it would provide a single platform for chemical modifications. We suspected that the properties of EnCNC would make it possible to have a pure CNC hydrogel for 2D and 3D printing. We first studied the printability of EnCNC to form stable 3D pieces (Fig. 4a), we concentrated EnCNC suspension at 10 wt% and printed a piece resembling a human ear that holds shape fidelity (Fig. 4b), EnCNC showed similar printability as modified CNFs alone37. We also printed a scaffold for cell growth using only EnCNC as ink (Fig. 4c) that acted as matrix for living cell to growth between the nanocellulose (Fig. 4d). HEK293T were seeded onto the scaffold and let then in the incubator for 3 days (Fig. 4e), cells were dyed with DAPI to observe the nucleus of the living cells that migrated and proliferated within the EnCNC matrix (Fig. 4f). AcCNC could be printed at a concentration of 20 wt% but the scaffolds were unable to hold the shape in the culture medium without introducing other components such as alginate or collagen.
a Schematic representation of 3D printer using EnCNC suspension as ink. b Image of a human ear-like piece of ~1 cm3 printed with 10 wt% EnCNC suspension. c Scaffold of 1.5 cm3 made with 10 wt% EnCNC suspension for 3D cell culture. d Representation of cells embedded on EnCNC matrix. e Portion of a 24-well plate containing a 3D-printed scaffold of EnCNC bioink similar to the one in c and HEK293 in cell culture medium. f Confocal microscopy image at ×63 of HEK293T-stained nuclei with DAPI. Cells proliferated embedded on EnCNC matrix after 3 days of culture. g Schematic representation of 2D EnCNC for 2D cell culture. h SEM images of HEK293T living over EnCNC film after 3 days of culture. Cells were colored in pink and EnCNC film in blue. i Close-up image of two HEK293 cells dividing over EnCNC film.
We also analyzed the ability of EnCNC and AcCNC to produce nanopaper-like materials as 2D substrate for cell proliferation (Fig. 4g). Unlike AcCNC, EnCNC was able to form a thin film with transparency and integrity to be submerged in aqueous solutions (Supplementary Fig. 10). HEK293T were seeded over an EnCNC film that maintained its shape in the culture medium for 3 days and cells were able to attach to its surface (Supplementary Fig. 11). We use scanning electron microscopy (SEM) to observe that cell attached properly (Fig. 4h), and that they proliferate and divide well attached to the EnCNC film (Fig. 4i). These experiments demonstrate that EnCNC could serve as unique platform to make bioinks, in which other components, such as proteins, polymers, or surface groups could be incorporated to create a tailored matrix for tissue engineering or drug delivery.
EnCNC/GO materials fabrication and characterization
Another application where CNC has been suggested to be of interest is in the preparation of substrates and matrices for nanocomposites38 with controlled electrical and mechanical properties39. These nanocomposites also need structural consistency. In this vein, we investigate the fabrication of a biomaterial combining EnCNC with graphene forms. Our reasoning for such combination resides in the ability of EnCNC to form stable matrices in which the physical properties of graphene could be combined to those of EnCNC, thus generating an advanced hybrid material suitable for biocompatible devices. The first integration of graphene in EnCNC was the fabrication of conductive inks in the form of nanopapers. Thereafter, we fabricate nanopapers of EnCNC as a matrix containing graphene oxide (GO) in different proportions, that were chemically reduced to rGO when the paper was assembled (Fig. 5a, b). We measured the zeta potential of these EnCNC/GO suspension and all the values were very similar to the EnCNC suspension, which shows that stability was maintained after GO addition (Supplementary Table 4). We analyzed the morphology and chemical composition of the nanopapers, demonstrating the homogeneity of the GO/EnCNC mix (Supplementary Figs. 12–17).
a Illustration of hybrid nanopaper, the conductive nanopapers were produced by mixing graphene oxide (GO) with nanocellulose and its posterior reduction once the paper was assembled. b Photographs of films prepared with EnCNC and EnCNC with different reduced graphene content: 2, 5, and 10 wt% rGO addition produced a black appearance to the films. c Conductive of EnCNC films at different rGO content (volumetric %), inset shows a log–log plot for determination of electrical percolation threshold of the (ρc). d Young’s modulus of the EnCNC film and EnCNC films with different rGO concentrations. e Water contact angle measurement for neat EnCNC and nanocomposites with different rGO content. The average values and S.D. were calculated from five independent experiments (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001). f Confocal microscopy image at ×10 and zoom at ×63 of adhered HEK293T dyed with DAPI in a substrate made of 10 wt% EnCNC suspension with 10% rGO. Cells proliferate in the composite substrate after 3 days.
An important new feature that rGO gave to the EnCNC film is the ability to conduct electricity. We measured the conductivity of our films observing and increasing conductivity as the content of rGO increases (Supplementary Table 5). We fabricated nanopapers with a rGO content of 0.3–15 wt% that were used to measure conductivity. We measured conductivity in all samples, but the conductivity reached a plateau of 25.2 S/m at around 10 wt% rGO content. Using these measurements we determined the electrical percolation threshold (ρc) following the percolation theory40. We plotted the conductivity against the volumetric rGO content to apply the theory (Fig. 5c) and calculated a ρc of 0.16%, that in our system corresponds to 0.25 wt% rGO, which is in the range of that determined for other graphene nanocomposites41. These nanopapers show higher conductivity than other composites using CNF42,43, and bacterial cellulose (BC)44 at similar concentration of rGO.
Similarly, we analyzed the mechanical behavior of EnCNC films with different rGO contents (Supplementary Fig. 18) by measuring tensile strength and Young’s modulus (Fig. 5d and Supplementary Table 6). When rGO content increased tensile strength decreased for films containing 2 and 5 wt% rGO and increased at 10 wt% rGO concentration, reaching 49.6 MPa (Supplementary Table 6). Young’s modulus changed similarly with rGO addition. The improvement at 10 wt% rGO could be the result of an enhanced interaction between graphene sheets in the matrix. This is in agreement with other studies performed using CNF42,45. In addition, we used WCA technique and determined that the contact angle increased with rGO content due to the graphene sheets hydrophobicity (Fig. 5e). Finally, the addition of rGO does not change the thermal stability of EnCNC as measured by TGA (Supplementary Fig. 19 and Supplementary Table 7). The DTG curves are similar in all cases. These conductive nanopapers could be used in applications at high temperatures and humid environment where plastic materials fail. Also, this composite may serve as the basis for obtaining conductive printable inks. In addition, we prepared a mixture of EnCNC 10 wt% suspension and added 10% rGO. We seeded HEK293T cells into the material to test the viability of the hybrid material for 2D cultures. After 3 days of culture, we dyed cells with DAPI and observe by confocal microscopy the nucleus of living cells in this material. This shows that rGO at this concentration allows cell proliferation (Fig. 5f). We hypothesize that this hybrid material could be useful for studies where neuron or muscle cells are grown and studied using electrical impulses. For comparative purposes, we fabricated nanopapers with EnCNC from T. reesei EG+10% rGO. We measured similar conductivity, 20 vs. 25.2 S/m for EnCNC made with ANC EG+CBM. However, nanopapers produced with T. reesei EG EnCNC had lower thermal stability (Supplementary Fig. 20), a To, 193 °C, was measured in this sample. This is likely related to the faster thermal degradation of this EnCNC produced with T. reesei EG. This demonstrates the important role of CNC when used as matrix for nanomaterials.
Finally, we tested the ability of EnCNC to serve as substrate by transferring graphene sheets synthesized by chemical vapor deposition (CVD) (Fig. 6a, b). Graphene monolayer (EnCNC ML) was successfully transferred, as confirmed by Raman spectroscopy (Supplementary Fig. 21), showing high transparency and transmittance (Supplementary Fig. 22). We also demonstrate the good conductivity of the material (Fig. 6c). EnCNC ML shows a resistance of 949 ± 300 Ω per sq, a value that increases to 11873 ± 300 Ω per sq in EnCNC BL, probably due to defects (Supplementary Fig. 23). The EnCNC ML value is in the range of other CVD deposited over polymeric materials, but EnCNC ML resistivity was higher in comparison with metal substrates, such as nickel46 or copper47, likely due to the roughness of the surface48. The two preparations present a similar topology upon application of a bias voltage between a conductive AFM tip and the sample (Fig. 6b). Resolution and detail increased with voltage due to higher conductive response of CVD layers. Phase images with a constant voltage were taken at 5, 0, and −5 V, from both EnCNC ML and EnCNC BL (Supplementary Fig. 24), showing that they are uniformly conductive. The topography profile measured in the AFM height images show similar pattern, highlighting the good contact and adhesion of graphene over the EnCNC film (Supplementary Fig. 25).
a Illustration of CVD graphene monolayer transferred over nanocellulose transparent film. b Photographs and EFM images of EnCNC film, CVD graphene monolayer (EnCNC ML), and CVD graphene bilayer (EnCNC BL) over EnCNC. EFM height and phase images from EnCNC, EnCNC ML, and EnCNC BL. Topography of the films was shown in the height images and the phase showed the response of the films to different voltage bias 0, 2, 4, and 6 V, from the AFM tip to the sample. c Sheet resistance measurement of EnCNC ML and EnCNC BL. The average values and SD were calculated from two independent experiments (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001).
Discussion
In this work, we report that the ancestral form of an endoglucanase Cel5A is able to upgrade cellulose to a nanomaterial suitable for high-performance applications, EnCNC. This material could not be obtained by the two other bacterial endoglucanases tested, including the modern descending of ANC EG. Interestingly, EnCNC differs also from other CNC obtained using complex enzyme systems and preparations. To the best of our knowledge, this is the first attempt to generate, in an efficient manner, CNC using a single enzyme. Although some reports can be found in the literature reporting the use of enzymes or organisms to hydrolyze cellulose49,50,51,52,53, no method reports the design of a specific enzyme that offers special capabilities towards crystalline nanocellulose obtention. The evolutionary reasons behind the ability of ANC EG to hydrolyze cellulose to crystalline particles below 500 nm, unlike its modern counterpart, are unknown; but we hypothesize that ancient cellulose may have had a high proportion of crystalline cellulose that required a more efficient enzyme to hydrolyze it. Therefore, ASR captures a chemical phenotype of Cel5A enzymes that has been lost, as lignocellulosic biomass became more complex and other enzymes, such as xylanases, exoglucanases and even laccases were required for biomass decomposition, thus specializing the job of each enzyme. Luckily, ASR brings back the ability to generate CNC in one single reaction by one enzyme alone. This may be related to the processive activity reported for the ancestral EG, that demonstrated endo-activity and exo-activity, unlike its modern counterparts14.
From the physicochemical characterization studies, we conclude that EnCNC is a unique nanocellulose that conserves cellulose I structure with elevated stability, crystallinity, and controlled aspect ratio. The enzymatic obtention of nanocellulose is a green and sustainable method that renders EnCNC as an empty canvas for further surface modification as opposed to AcCNC that contains sulfate groups on the surface. Moreover, this enzymatic method for nanocrystals obtention could be applied to other natural polysaccharides such as chitin and starch54. The features of EnCNC make it a good platform for numerous applications, but we envision a good potential implementation in tissue engineering due to its good printability, shape fidelity, surface modification possibilities, and good proliferation of cells. We also foresee a good implementation of EnCNC for conductive inks due to the integration of graphene derivatives. These applications combined allow envisioning the design of matrices for artificial biointelligence where the ability of printing circuits could be integrated with different types of cells that can feel and react to electrical and optical stimuli. In the same direction, these materials may pave the way towards fabrication of biocompatible biosensors for wearable or implant devices for diagnosis55.
Methods
Protein expression and purification
ANC EG, ANC EG+CBM, T. maritima EGs protein-encoding genes were synthesized and codon optimized for E. coli cell expression. ANC EG, ANC EG+CBM, T. maritima were cloned in pQE80L expression vector (Qiagen) and transformed in E. coli BL21 (DE3) (Life Technologies) for protein expression. B.subtillis EG gene cloned in pET-28a vector was kindly supplied by Dr. Mário Tyago Murakami. The CBM used belongs to B. subtilis, the species used as query sequence in the reconstruction, due to the impossibility of reconstructing an ancestral CBM14. Cells were incubated in LB medium at 37 °C until OD600 reached 0.6, IPTG was added to the medium to 1 mM concentration for protein induction overnight. Cells were pelleted by centrifugation at 4000 rpm. Pellets were resuspended in extraction buffer (50 mM sodium phosphate, pH 7.0, 300 mM NaCl) and mechanically lysed using French Press. Cell debris was separated by ultracentrifugation at 33,000×g for 1 h. For purification, the supernatants were mixed with His GraviTrap affinity column (GE Healthcare) and eluted in elution buffer (50 mM sodium phosphate, pH 7.0, 300 mM NaCl, 150 mM imidazole). Proteins were further purified by size exclusion chromatography using a Superdex 200 HR column (GE Helthcare) and eluted in 50 mM citrate buffer pH 4.8. For protein purification verification sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) was used with 12% gels. The protein concentration was calculated by measuring the absorbance at 280 nm in Nanodrop 2000C. Cellulase from Trichoderma reseei was purchase from Sigma Aldrich (C2730).
Enzymatic nanocellulose isolation
Whatman® filter paper was used as substrate; 0.4 g of filter paper was used for each hydrolysis prepared in small squares of 1 cm2. The reaction was conducted in 40 ml of water and 5 mg of each EG was used (ANC EG, ANC EG+CBM, and T. maritima, B. subtillis, and T. reesei EG) per gram of substrate. Hydrolysis was incubated at 50 °C with agitation for different times (1, 5, 24, and 48 h). Reactions were stopped by incubating the hydrolysis on ice and the mixtures were sonicated with a micro-tip sonicator UPH100H ultrasonic processor (Hielscher) for 25 min at 75%. Nanocellulose was isolated by gradual centrifugation steps and concentrated by ultracentrifugation at 33,000×g for 1 h. Pellets were resuspended in water and lyophilized in a Telstar Lyoquest for physicochemical characterization by freeze-drying for 24 h. CNC produced by sulfuric acid treatment and used for comparison were purchased from Maine University (Lot# 2014-FPL-CNC-065).
CNC/graphene nanopapers films
The nanocellulose nanopapers were prepared by casting method using an EnCNC suspension with 1 wt% of concentration. Nanocellulose suspension was sonicated for 1 h and then casted in a Teflon mold and dried for 2 days at 37 °C. The conductive nanopapers were produced by mixing GO with EnCNC and its posterior reduction once the paper was assembled. GO (4 wt% water suspension) was kindly supplied by Graphenea (San Sebastian, Spain). Different concentrations of GO were added to an EnCNC water suspension with 1 wt% concentration, to prepare nanopapers with 0.3, 1, 2, 5, 10, and 15 wt% of graphene. EnCNC and GO were mixed by sonication for 1 h and then casted in Teflon molds and dried for 2 days at 37 °C, obtaining films with a thickness around 100 μm. The GO in the nanopapers was then reduced by submerging the films in an ascorbic acid solution (30 g/l) for 2 h at 95 °C that kept nanocellulose unaltered. The reduced films were washed with miliQ water for ascorbic acid removal and dried at room temperature overnight. The nanopapers were named as “EnCNC+x% rGO”, where x was the content of rGO in the total mass of the nanopaper. Monolayer and bilayer graphene were grown on copper (Cu) foils and deposited by CVD. The graphene synthesis was carried out in a cold-walled CVD reactor (Aixtron BM) at 1000 °C and at low pressure using methane as the carbon source. Prior to the growth the Cu foils were annealed at 1000 °C under a hydrogen and argon flow. After the synthesis, a poly(methyl methacrylate) (PMMA) support layer was spin coated onto the graphene covered Cu foil. The Cu was etched using a ferric chloride-containing solution and the layer of graphene transferred onto the nanocellulose film. Finally, the PMMA layer was removed by dipping into acetone and isopropanol. The EnCNC papers covered with CVD graphene monolayer and bilayer were designed as EnCNC ML and EnCNC BL, respectively.
Characterization of materials
Nanocellulose conversion was calculated by weighting the freeze-dried nanocellulose from each hydrolysis and using the following Eq. (1) to calculate the yield:
AFM was used for the study of nanocellulose and GO morphology. Images were captured at room temperature, in tapping mode, using a Nanoscope V scanning probe microscope (Multimode 8 Bruker Digital Instruments) with an integrated force generated by cantilever/silicon probes. The applied resonance frequency was 320 kHz. The cantilever had a tip radius of 5–10 nm and was 125 µm long. The samples were prepared by spin-coating (Spincoater P6700) at 2000 rpm for 130 s by casting a droplet of nanocellulose suspension on mica substrate. AFM height and phase images were collected simultaneously from all the samples. From nanocellulose images we measured the length and the diameter of 100 nanoparticles for each sample to calculate the average of the length and the diameter and the aspect ratio (length/diameter).
Fourier transform infrared (FTIR) spectroscopy was used to study cellulose, GO, and rGO functional groups and their evolution in hybrid nanopapers spectra. FTIR spectra were measured by a Nicolet Nexus spectrometer provided with a MKII Golden Gate accessory (Specac) with a diamond crystal at a nominal incidence angle of 45° and ZnSe lens. Spectra were measured in attenuated reflection (ATR) mode between 4000 and 650 cm−1, with averaging 32 scans and a resolution of 4 cm−1. Scattering-type scanning near-field optical microscopy (IR s-SNOM) and nanoscale-resolved FTIR (nano-FTIR) permit to study the infrared spectra from a single nanocellulose particle of a suspension. The IR s-SNOM images and nano-FTIR spectra, as well as the corresponding AFM images, were recorded with a neaSNOM system (Neaspec GmbH, Germany) comprising both s-SNOM and nano-FTIR capabilities. Nano-FTIR spectroscopy of the nanocellulose suspension was performed with illumination from a mid-infrared laser supercontinuum, using Au-coated AFM tips. The total acquisition time of each spectrum was 15 min and the spectral resolution was 16 cm−1. The spectra were normalized to that obtained on a clean silicon surface (reference measurements).
Sulfate content on AcCNC was measured by conductometric titration at 25 °C with a Crison ECMeter GLP 31 conductometer that was calibrated with 147, 1413, and 12.88 µS/cm standards. We diluted the AcCNC powder in 25 ml of water and sonicated for 30 min, then we added 10 ml of 10 mM HCl. The dispersion was continuously titrated with 10 mM NaOH56. We measured the NaOH consumption for the sample titration and the blank, and we calculated the sulfur content using Eq. (2):
where M is the NaOH concentration, V is the volume of NaOH solution that we need to titrate the sample (without the volume used to titrate the blank) and ω is the weight of AcCNC used.
The Zeta potential (ζ, mV) of EnCNC and AcCNC water suspensions (0.1 wt%) were measured using a Zetasizer Ultra (Malvern Intstrument).
X-ray diffraction (XRD) was used for the analysis of the cellulose crystalline structure, crystallinity, and crystallite size. X-ray powder diffraction patterns were collected by using a Philips X’pert PRO automatic diffractometer operating at 40 kV and 40 mA, in theta–theta configuration, a secondary monochromator with Cu-Kα radiation (λ = 1.5418 Å) and a PIXcel solid state detector (active length 2θ = 3.347°). Data were collected from 5° to 50° 2θ (step size 0.026 and time per step 80 s) at room temperature. A fixed divergence and antiscattering slit, which give a constant volume of sample illumination were used. From the diffractograms, we calculated the CI% using the Segal Eq. (3)57:
where I200 is the intensity of the cellulose crystalline peak and Iam is the intensity of the amorphous peak. The crystallite size was measured from the diffractograms using the Scherrer’s Eq. (4)58:
where λ is the wavelength of the incident X-ray, θ is the angle of the (200) plane, β is the full width at half maximum of the (200) peak, τ is the crystallite size and κ is a constant value.
Solid-state cross-polarization magic angle spinning 13C nuclear magnetic resonance (13C CP/MAS NMR) was used to study cellulose structure. 13C CP/MAS NMR spectra were recorded on a 400 MHz BRUKER system equipped with a 4 mm MASDVT TRIPLE resonance HYX MAS probe. Larmor frequencies were 400.17 and 100.63 MHz for 1H and 13C nuclei, respectively. Chemical shifts were reported relative to the signals of 13C nuclei in glycine. Sample rotation frequency was 12 kHz and relaxation delay was 5 s. The number of scans was 2k. Polarization transfer was achieved with RAMP cross-polarization (ramp on the proton channel) with a contact time of 5 ms. High-power SPINAL 64 heteronuclear proton decoupling was applied during acquisition.
Thermogravimetric analysis (TGA) was performed to determine thermal stability of nanocellulose and hybrid nanopapers. The analysis was performed using TGA/SDTA 851 Mettler Toledo equipment. Up to 10 mg of the samples were used and were heated from 30 to 800 °C in a nitrogen atmosphere at a scanning rate of 10 °C/min. The initial degradation temperature (To) is described as the loss of 5% of the weight of the total sample and the maximum degradation temperature (Td) is the minimum of the degradation peak in the derivative of thermogravimetric curves (DTG).
Raman spectra were measured using the Raman microscope alpha 300R (WITec GmbH) with a laser in an excitation wavelength of 532 nm. 100 accumulations were recorded and the laser intensity was 1 mW. The spectra were normalized by the highest peak in the range of 800–3600 cm−1.
The mechanical properties of the nanopapers were measured using a Miniature materials tester Minimat 2000, Rheometric Scientific, equipment, at room temperature with a 200 N load cell and an elongation rate of 2 mm/min. We cut the samples to a width of 0.5 cm and a length of 2 cm. The stress–strain curves, the Young’s modulus, tensile strength, and strain at break were calculated from five samples of each nanopaper. The hydrophilicity of the surface of the nanopapers was measured by static water contact angle (WCA) using a Dataphysics OCA20 equipment at room temperature. In this technique, we deposited a deionized water drop in the film surface to analyze the contact angle value that the water drops formed, that depend on the chemical interactions between the water and the material surface. When the material is hydrophilic the contact angle is small and increases with the hydrophobicity. The contact angle (θc) values of 10 water drops of 2 µl deposited by a syringe tip were averaged for each sample. The transmittance of EnCNC and CVD graphene containing EnCNC films was evaluated by UV–Vis using UV–Vis–NIR spectroscopy (3600 Shimadzu). The morphology of the nanopaper surface and the CVD graphene deposited over nanocellulose film was analyzed by SEM using a FEI ESEM Quanta 200 microscope operating at 5–20 kV. Nanopapers and CVD samples were put on a carbon tape for adhesion. Electrical conductivity was measured by a four-point probe method using a Probe Station 4 Everbeing. The specific resistances (q) were calculated with the sheet resistances (Rs, Ω/squares) and the thickness of the nanopapers (t, cm) in Eq. (5):
We used the specific resistance calculated to infer the corresponded conductivity (S/cm) with the following Eq. (6) that was transformed to S/m.
The percolation threshold, ρc, was measured using a power-law equation based on the percolation theory40:
where σ is the conductivity of the nanopaper, ρf is the rGO conductivity, ρ is the rGO content expressed at volume fraction, and n is the exponent describing the rapid variation of the conductivity near the percolation threshold (ρc).
Electrostatic force microscopy (EFM) permits us to measure the electrostatic force between the AFM tip and the surface of a sample. CVD graphene samples measurement were carried out in a dimension icon scanning probe microscope operating in the lift mode (100 nm) in ambient conditions and equipped with a Pt/Ir-coated tip (ca. 75 kHz resonant frequency). The secondary imaging mode derived from the tapping mode that measures the electric field gradient distribution above the sample surface was detected by applying different voltages to the cantilever tip.
3D printing of EnCNC-based scaffold
Pieces were printed with a Voladora 3D printer (Tumaker, S.L. Spain) which has been modified for layer by layer syringe extrusion 3D printing. Scaffold (human ear-like) were printed directly on polytetrafluoroethylene slides at room temperature with a needle of 0.8 mm in diameter and speed of 5 mm/s.
2D and 3D cell cultures
HEK293T cells were a kind gift of Dr. Maria Muñoz Caffarel. Cells were maintained in DMEN+10% FBS medium supplemented with 1% (w/v) l-glutamine and penicillin–streptomycin (100 IU/ml). EnCNC film and EnCNC scaffold were sterilized by UV for 3 h. 20,000 cells were seeded on each substrate and let to growth for 3 days in the incubator at 37 °C in 5% CO2. Cells were dyed with DAPI solution (5 mg/ml) diluted in PBS (1:2000) after washing each sample three times with PBS. DAPI-stained cells were observed after 5 min of incubation by confocal microscopy. EnCNC films were fixed for SEM examination in 4% formaldehyde for 30 min and washed with PBS. Films were dehydrated by increasing concentration of ethanol solution until 100% concentration of ethanol was reached.
Data availability
Data, images, samples of EnCNC, and other material are available upon reasonable request.
References
Rollin, J. A. et al. High-yield hydrogen production from biomass by in vitro metabolic engineering: mixed sugars coutilization and kinetic modeling. Proc. Natl Acad. Sci. USA 112, 4964–4969 (2015).
You, C. et al. Enzymatic transformation of nonfood biomass to starch. Proc. Natl Acad. Sci. USA 110, 7182–7187 (2013).
Liszka, M. J., Clark, M. E., Schneider, E. & Clark, D. S. Nature versus nurture: developing enzymes that function under extreme conditions. Annu. Rev. Chem. Biomol. Eng. 3, 77–102 (2012).
Yang, H. et al. Evolving artificial metalloenzymes via random mutagenesis. Nat. Chem. 10, 318 (2018).
Longwell, C. K., Labanieh, L. & Cochran, J. R. High-throughput screening technologies for enzyme engineering. Curr. Opin. Biotechnol. 48, 196–202 (2017).
Zeymer, C. & Hilvert, D. Directed evolution of protein catalysts. Annu. Rev. Biochem. 87, 131–157 (2018).
Michael, R., Cindy, S., Dagmara, J., Tobias, H. & Greta, F. Novel materials through nature’s catalysts. Mater. Today 18, 459–467 (2015).
Himmel, M. E. et al. Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science 315, 804–807 (2007).
Martin-Martinez, F. J., Jin, K., López Barreiro, D. & Buehler, M. J. The rise of hierarchical nanostructured materials from renewable sources: learning from nature. ACS Nano 12, 7425–7433 (2018).
Thomas, B. et al. Nanocellulose, a versatile green platform: from biosources to materials and their applications. Chem. Rev. 118, 11575–11625 (2018).
Yang, X., Reid, M.S., Olsén, P. & Berglund, L.A. Eco-friendly cellulose nanofibrils designed by nature—effects from preserving native state. ACS Nano 14, 724–735 (2019).
Habibi, Y., Lucia, L. A. & Rojas, O. J. Cellulose nanocrystals: chemistry, self-assembly, and applications. Chem. Rev. 110, 3479–3500 (2010).
Iwamoto, S., Nakagaito, A. & Yano, H. Nano-fibrillation of pulp fibers for the processing of transparent nanocomposites. Appl. Phys. A 89, 461–466 (2007).
Barruetabeña, N. et al. Resurrection of efficient Precambrian endoglucanases for lignocellulosic biomass hydrolysis. Commun. Chem. 2, 76 (2019).
Zakas, P. M. et al. Enhancing the pharmaceutical properties of protein drugs by ancestral sequence reconstruction. Nat. Biotechnol. 35, 35–37 (2017).
Manteca, A. et al. Mechanochemical evolution of the giant muscle protein titin as inferred from resurrected proteins. Nat. Struct. Mol. Biol. 24, 652 (2017).
Randall, R. N., Radford, C. E., Roof, K. A., Natarajan, D. K. & Gaucher, E. A. An experimental phylogeny to benchmark ancestral sequence reconstruction. Nat. Commun. 7, 12847 (2016).
Risso, V. A., Gavira, J. A., Mejia-Carmona, D. F., Gaucher, E. A. & Sanchez-Ruiz, J. M. Hyperstability and substrate promiscuity in laboratory resurrections of Precambrian β-lactamases. J. Am. Chem. Soc. 135, 2899–2902 (2013).
Perez-Jimenez, R. et al. Single-molecule paleoenzymology probes the chemistry of resurrected enzymes. Nat. Struct. Mol. Biol. 18, 592 (2011).
Gaucher, E. A., Thomson, J. M., Burgan, M. F. & Benner, S. A. Inferring the palaeoenvironment of ancient bacteria on the basis of resurrected proteins. Nature 425, 285 (2003).
Gumulya, Y. et al. Engineering highly functional thermostable proteins using ancestral sequence reconstruction. Nat. Catal. 1, 878 (2018).
de Rodriguez, N. L. G., Thielemans, W. & Dufresne, A. Sisal cellulose whiskers reinforced polyvinyl acetate nanocomposites. Cellulose 13, 261–270 (2006).
Sèbe, G., Ham-Pichavant, F. D. R., Ibarboure, E., Koffi, A. L. C. & Tingaut, P. Supramolecular structure characterization of cellulose II nanowhiskers produced by acid hydrolysis of cellulose I substrates. Biomacromolecules 13, 570–578 (2012).
Panchal, P., Ogunsona, E. & Mekonnen, T. Trends in advanced functional material applications of nanocellulose. Processes 7, 10 (2019).
de Souza Lima, M. M. & Borsali, R. Rodlike cellulose microcrystals: structure, properties, and applications. Macromol. Rapid Commun. 25, 771–787 (2004).
Dinand, E., Vignon, M., Chanzy, H. & Heux, L. Mercerization of primary wall cellulose and its implication for the conversion of cellulose I→ cellulose II. Cellulose 9, 7–18 (2002).
Niu, F. et al. The characteristic and dispersion stability of nanocellulose produced by mixed acid hydrolysis and ultrasonic assistance. Carbohydr. Polym. 165, 197–204 (2017).
Bondeson, D., Mathew, A. & Oksman, K. Optimization of the isolation of nanocrystals from microcrystalline cellulose by acid hydrolysis. Cellulose 13, 171 (2006).
Huth, F. et al. Nano-FTIR absorption spectroscopy of molecular fingerprints at 20 nm spatial resolution. Nano Lett. 12, 3973–3978 (2012).
Chi, K. & Catchmark, J. M. The influences of added polysaccharides on the properties of bacterial crystalline nanocellulose. Nanoscale 9, 15144–15158 (2017).
Urbina, L., Corcuera, M. Á., Eceiza, A. & Retegi, A. Stiff all-bacterial cellulose nanopaper with enhanced mechanical and barrier properties. Mater. Lett. 246, 67–70 (2019).
Yue, Y. et al. Comparative properties of cellulose nano-crystals from native and mercerized cotton fibers. Cellulose 19, 1173–1187 (2012).
Poletto, M., Ornaghi, H. L. & Zattera, A. J. Native cellulose: structure, characterization and thermal properties. Materials 7, 6105–6119 (2014).
Gong, J., Mo, L. & Li, J. A comparative study on the preparation and characterization of cellulose nanocrystals with various polymorphs. Carbohydr. Polym. 195, 18–28 (2018).
Lin, N. & Dufresne, A. Surface chemistry, morphological analysis and properties of cellulose nanocrystals with gradiented sulfation degrees. Nanoscale 6, 5384–5393 (2014).
Du, H. et al. Cellulose nanocrystals and cellulose nanofibrils based hydrogels for biomedical applications. Carbohydr. Polym. 209, 130–144 (2019).
Ajdary, R. et al. Acetylated nanocellulose for single-component bioinks and cell proliferation on 3D-printed scaffolds. Biomacromolecules 20, 2770–2778 (2019).
Zhu, C., Monti, S. & Mathew, A. P. Cellulose nanofiber–graphene oxide biohybrids: disclosing the self-assembly and copper-ion adsorption using advanced microscopy and reaxff simulations. ACS Nano 12, 7028–7038 (2018).
Hamedi, M. M. et al. Highly conducting, strong nanocomposites based on nanocellulose-assisted aqueous dispersions of single-wall carbon nanotubes. ACS Nano 8, 2467–2476 (2014).
Kirkpatrick, S. Percolation and conduction. Rev. Mod. Phys. 45, 574 (1973).
Marsden, A. J. et al. Electrical percolation in graphene–polymer composites. 2D Materials 5, 032003 (2018).
Dang, L. N. & Seppälä, J. J. C. Electrically conductive nanocellulose/graphene composites exhibiting improved mechanical properties in high-moisture condition. Cellulose 22, 1799–1812 (2015).
Chen, Y., Pötschke, P., Pionteck, J., Voit, B. & Qi, H. Smart cellulose/graphene composites fabricated by in situ chemical reduction of graphene oxide for multiple sensing applications. J. Mater. Chem. A 6, 7777–7785 (2018).
Feng, Y., Zhang, X., Shen, Y., Yoshino, K. & Feng, W. J. C. P. A mechanically strong, flexible and conductive film based on bacterial cellulose/graphene nanocomposite. Carbohydr. Polym. 87, 644–649 (2012).
Beeran, Y. et al. Mechanically strong, flexible and thermally stable graphene oxide/nanocellulosic films with enhanced dielectric properties. RSC Adv. 6, 49138–49149 (2016).
Kim, K. S. et al. Large-scale pattern growth of graphene films for stretchable transparent electrodes. Nature 457, 706 (2009).
Bae, S. et al. Roll-to-roll production of 30-inch graphene films for transparent electrodes. Nat. Nanotechnol. 5, 574 (2010).
Kafiah, F., Khan, Z., Ibrahim, A., Atieh, M. & Laoui, T. Synthesis of graphene based membranes: effect of substrate surface properties on monolayer graphene transfer. Materials 10, 86 (2017).
Zhu, J. Y., Sabo, R. & Luo, X. J. G. C. Integrated production of nano-fibrillated cellulose and cellulosic biofuel (ethanol) by enzymatic fractionation of wood fibers. Green Chem. 13, 1339–1344 (2011).
Satyamurthy, P., Jain, P., Balasubramanya, R. H. & Vigneshwaran, N. J. C. P. Preparation and characterization of cellulose nanowhiskers from cotton fibres by controlled microbial hydrolysis. Carbohydr. Polym. 83, 122–129 (2011).
Yarbrough, J. M. et al. Multifunctional cellulolytic enzymes outperform processive fungal cellulases for coproduction of nanocellulose and biofuels. Acs Nano 11, 3101–3109 (2017).
Wang, W. et al. Endoglucanase post-milling treatment for producing cellulose nanofibers from bleached eucalyptus fibers by a supermasscolloider. Cellulose 23, 1859–1870 (2016).
Tang, Y. et al. Extraction of cellulose nano-crystals from old corrugated container fiber using phosphoric acid and enzymatic hydrolysis followed by sonication. Carbohydr. Polym. 125, 360–366 (2015).
Lin, N., Huang, J. & Dufresne, A. Preparation, properties and applications of polysaccharide nanocrystals in advanced functional nanomaterials: a review. Nanoscale 4, 3274–3294 (2012).
Golmohammadi, H., Morales-Narvaez, E., Naghdi, T. & Merkoci, A. Nanocellulose in sensing and biosensing. Chem. Mater. 29, 5426–5446 (2017).
Camarero Espinosa, S., Kuhnt, T., Foster, E. J. & Weder, C. J. B. Isolation of thermally stable cellulose nanocrystals by phosphoric acid hydrolysis. Biomacromolecules 14, 1223–1230 (2013).
Segal, L., Creely, J., Martin, A. Jr & Conrad, C. An empirical method for estimating the degree of crystallinity of native cellulose using the X-ray diffractometer. Text. Res. J. 29, 786–794 (1959).
Cullity, B. D. Elements of X-ray Diffraction. Addison-Wesley Publishing (1978).
Acknowledgements
Research was financially supported by the Basque Government in the frame of ELKARTEK program to R.P.-J. and A.E., and partially by the Ministry of Economy and Competitiveness (MINECO) in the frame of BIO2016-77390, PID2019-109087RB-I00, and BFU2015-71964 to R.P.-J. ‘Materials+Technologies’ Research Group also acknowledges UPV/EHU and the Basque Government in the frame of “Research Group” (GIU 18/216) and “Grupos Consolidados” (IT776-13), respectively. We also thank Gipuzkoako Foru Aldundia for financial support. Finally, we thank SGIker (UPV/EHU/ERDF, EU) for technical and scientific support.
Author information
Authors and Affiliations
Contributions
R.P.-J. and A.E. conceived the project and designed research. B.A.-L., L.U., L.B., I.L., A.R, N.B., R.O.-J., I.A., R.H., carried out all experimental measurements and preparations including protein and sample preparation, activity assays, and instrumental measurements. B.A.-L., R.H., A.E., and R.P.-J. performed data analysis. B.A.-L., A.E., and R.P-J. wrote the manuscript and all authors contributed in revising and editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alonso-Lerma, B., Barandiaran, L., Ugarte, L. et al. High performance crystalline nanocellulose using an ancestral endoglucanase. Commun Mater 1, 57 (2020). https://doi.org/10.1038/s43246-020-00055-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43246-020-00055-5
This article is cited by
-
Preparation of cellulose nanocrystals from commercial dissolving pulp using an engineered cellulase system
Bioresources and Bioprocessing (2023)
-
A facile, low-thermal, and environmentally friendly method to improve the properties of lignin-containing cellulose nanocrystals (LCNCs) and cellulose nanofibrils (LCNFs) from bagasse unbleached soda pulp
Biomass Conversion and Biorefinery (2023)
-
Exploration of a novel and efficient source for production of bacterial nanocellulose, bioprocess optimization and characterization
Scientific Reports (2022)
-
Enzymatic upgrading of nanochitin using an ancient lytic polysaccharide monooxygenase
Communications Materials (2022)
-
Enzymatic coupled mechanical defibrillation process for the production of corn (Zea mays) cob nanofibrillated cellulose: preparation, characterization and evaluation as Pickering emulsifier for oil-in-water emulsion
Cellulose (2022)