Abstract
Cholinergic nerves are involved in tumor progression and dissemination. In contrast to other visceral tissues, cholinergic innervation in the hepatic parenchyma is poorly detected. It remains unclear whether there is any form of cholinergic regulation of liver cancer. Here, we show that cholinergic T cells curtail the development of liver cancer by supporting antitumor immune responses. In a mouse multihit model of hepatocellular carcinoma (HCC), we observed activation of the adaptive immune response and induction of two populations of CD4+ T cells expressing choline acetyltransferase (ChAT), including regulatory T cells and dysfunctional PD-1+ T cells. Tumor antigens drove the clonal expansion of these cholinergic T cells in HCC. Genetic ablation of Chat in T cells led to an increased prevalence of preneoplastic cells and exacerbated liver cancer due to compromised antitumor immunity. Mechanistically, the cholinergic activity intrinsic in T cells constrained Ca2+–NFAT signaling induced by T cell antigen receptor engagement. Without this cholinergic modulation, hyperactivated CD25+ T regulatory cells and dysregulated PD-1+ T cells impaired HCC immunosurveillance. Our results unveil a previously unappreciated role for cholinergic T cells in liver cancer immunobiology.
Similar content being viewed by others
Main
Hepatocellular carcinoma (HCC) is the most common primary liver malignancy in humans and the third most common cause of cancer-related deaths worldwide1. Risk factors for HCC include infection with hepatitis B virus or hepatitis C virus, excessive alcohol consumption, non-alcoholic fatty liver disease and aflatoxins2. These extrinsic risk factors cause chronic hepatitis that cooperates with intrinsic factors, particularly the accumulation of genetic mutations, to set the stage for HCC development3,4.
The immune system plays a dual role in liver cancer and can sense and eliminate preneoplastic and malignant hepatocytes5,6; it can also promote the selection of tumor cells and favor cancer progression in situations of chronic inflammation or immunosuppression7,8,9. This duality renders current immunotherapies that target immune checkpoints suboptimal for HCC8. Further exploration of the molecular determinants of immune responses in liver cancer is necessary to understand HCC biology and guide the design of effective therapeutic strategies.
The nervous system is involved in the development of cancer in multiple tissues10,11,12. For example, cholinergic fibers infiltrate prostate tumors, promoting their invasion and metastasis11. Engagement of nicotinic acetylcholine (ACh) receptors (nAChRs) mediates lung cancer growth13. Vagal innervation promotes gastric tumorigenesis through the M3 muscarinic ACh receptor (mAChR)10. Extensive efforts have been directed at delineating the hepatic nervous system. Despite some discrepancies among studies, sympathetic and parasympathetic neural markers have been detected in regions of the hepatic artery, portal vein and bile ducts in the majority of species investigated14. However, the liver parenchyma of rodents and humans appears to be devoid of vagal or cholinergic innervation, as determined by immunohistochemistry, retrograde tracing and advanced three-dimensional imaging15,16,17. Thus, whether and how cholinergic signaling plays a role in HCC regulation remains an open question.
In this study, we establish that subpopulations of CD4+ T cells expressing choline acetyltransferase (ChAT), the rate-limiting enzyme governing ACh synthesis, are induced during the development of liver cancer in mice. Importantly, we show that genetic ablation of Chat in T cells impairs HCC immunosurveillance. Examination of data from human HCC samples revealed parallels to our mouse findings. Our results demonstrate an unexpected aspect of the regulation of cancer immunosurveillance: mediation by an immune cell-derived neurotransmitter.
Results
Induction of HCC using CRISPR and transposon technology
We sought to model HCC in mice by combining genetic alterations recurrently observed in human disease. These changes included mutation of the TP53 and PTEN tumor suppressor genes and overexpression of the MYC oncogene18,19,20,21. CRISPR-mediated somatic knockout of tumor suppressor genes and transposon-based expression of oncogenes induce HCC in mice6,22,23. To mimic the multihit process of human liver carcinogenesis, we combined these two approaches by ablating Trp53 and Pten using duplex CRISPR and overexpressing Myc using the Sleeping Beauty transposon (Fig. 1a). These vectors were delivered in combination to mice via hydrodynamic injection, allowing for specific plasmid delivery to hepatocytes. Synergism between Myc expression and Trp53/Pten ablation induced rapid HCC development. Neoplasms were visible on the liver surface by 15 d after injection, with substantial tumor nodules present by day 25 (Fig. 1b). Immunostaining confirmed that the majority of these tumor clones were negative for p53 and PTEN and positive for MYC (Fig. 1c,d).
a, Schematic diagrams of the plasmids used to induce CRISPR–Cas9-mediated deletion of Trp53 and Pten (top) and transposon-mediated overexpression of Myc in mouse livers (bottom). b, Representative macroscopic views of livers from mice injected with the combination of plasmids shown in a at the indicated days after injection. Control mice received transposase vector only. Arrowheads indicate tumor nodules. c, Representative histological sections of tumor-burdened livers immunostained to detect MYC, p53 and PTEN. Images are representative of two independent experiments. d, Distribution of immunostained liver tumor nodules from the livers in c. Numbers of tumor nodules with the indicated immunostaining patterns are labeled in the pie plot, which is a summary of two independent experiments. Sections were resected from three mice per group on day 25 of HCC induction. e,f, Representative flow cytometry plots (e) and quantification (f) showing changes in the percentages of the indicated immune cell populations during the development of liver cancer in mice. In f, each dot represents an individual mouse (n = 6 mice per condition). Data are shown as mean ± s.e.m. Significance was assessed by unpaired, two-tailed t-test, and data are representative of three independent experiments. Control mice received transposase vector only. g, Representative histological sections from HCC-bearing livers that were immunostained to detect MYC or CD3 in areas of either preneoplastic cells (left) or HCC cells (right). Images represent immunostaining of liver sections from ten mice in one experiment. h, Survival of immunodeficient (NSG) and control wild-type (WT) mice (n = 10 per group) following injection of Trp53/Pten CRISPR and Myc overexpression plasmids to induce HCC development. P = 0.001 by log-rank test.
We engineered the CRISPR vector to also express Cre recombinase (Extended Data Fig. 1a). The transposon and Cre-encoding CRISPR vectors were delivered into Rosa26Confetti/+ mice, animals in which cells stochastically express one of four fluorescent proteins after Cre-mediated recombination24. We found that most tumor nodules expressed a single fluorescent marker, suggesting that these malignancies were monoclonal (Extended Data Fig. 1b,c).
Immunosurveillance is elicited in our HCC model
The immune microenvironment shapes the clonal evolution of tumor cells7,25. We investigated whether immune responses were evoked during tumorigenesis in our model. We found that hepatic CD4+ T cells and CD8+ T cells expanded as HCC development progressed, whereas the percentage of natural killer T (NKT) cells was reduced (Fig. 1e,f and Extended Data Fig. 1d). Analysis of OX40, a transient marker of T cell antigen receptor (TCR) activation, showed that antigen-stimulated CD4+ T cells were increased (Extended Data Fig. 1e). The infiltrating immune cells, including CD3+ T cells and CD11b+ antigen-presenting cells (APCs), were positioned around MYC+ preneoplastic cells and in established HCC (Fig. 1g and Extended Data Fig. 1f). Therefore, immune responses, particularly those mediated by T cells, are activated in our HCC model.
To determine the overall role of the immune system in our model, we induced HCC in severely immunodeficient NOD scid gamma (NSG) mice. Compared to immunocompetent animals, NSG mice developed more severe disease and died sooner, with tumor cells diffused throughout the liver instead of confined in discrete nodules (Fig. 1h and Extended Data Fig. 1g). Thus, immune cells participate in protection against liver cancer development in this setting.
ChAT-expressing T cells are induced during HCC development
Our group has been studying the function of cholinergic T cells in various contexts26,27,28. We analyzed the expression pattern of ChAT in liver tissues of reporter mice expressing green fluorescent protein (GFP) under the control of transcriptional regulatory elements for ChAT (Chat-GFP reporter mice). In contrast to the extensive cholinergic neural fibers and plexuses in the small intestine, we found no ChAT-expressing neural fibers in either the parenchyma of normal liver or in HCC (Extended Data Fig. 2a), findings in line with earlier reports15,16,17. However, we observed accumulation of lymphocyte-like ChAT-expressing cells in HCC (Extended Data Fig. 2a). These ChAT–GFP+CD4+ T cells were comprised mainly of CD44+ activated T cells that significantly increased during HCC progression (Fig. 2a,b). The percentages of ChAT–GFP+CD8+ T cells and NKT cells were also significantly elevated in HCC but to a lesser extent (Fig. 2b). In comparison, the percentage of ChAT–GFP+ B cells did not differ between control and HCC-bearing livers, and the expression of ChAT–GFP by NK cells and CD11b+ myeloid cells was negligible (Extended Data Fig. 2b). The overall level of Chat mRNA in bulk intrahepatic mononuclear cells (MNCs) from HCC-bearing livers was enhanced, reflecting the induction of ChAT-expressing T cells (Extended Data Fig. 2c). Therefore, lymphocytes are the dominant cholinergic cells in both healthy and HCC-bearing livers, and ChAT-expressing T cells are induced during tumorigenesis.
a,b, Representative flow cytometry plots (a) and quantification (b) of GFP expression in the indicated T cell subsets during HCC progression in Chat-GFP reporter mice. In b, each dot represents an individual mouse (n = 6 mice per condition). Data are shown as mean ± s.e.m. P values were determined by unpaired, two-tailed t-test, and data are representative of three independent experiments. c,d, Transcriptional landscape of ChAT–GFP+ and ChAT–GFP− CD4+ T cells in livers from control mice and mice with HCC. Uniform manifold approximation and projection (UMAP) representation of total hepatic CD4+ T cells (a total of 15,703 cells; c) and CD4+ T cells split according to GFP expression and HCC conditions (d) based on scRNA-seq analysis. ChAT–GFP+ and ChAT–GFP− CD4+ T cells were sorted from four mice with HCC and from four control mice. Cells from each mouse were stained with unique barcoded antibodies; ISG-T, interferon-stimulated gene-expressing T cells. e, Bubble plot comparing expression of Chat and the indicated marker genes across the 11 clusters in c and d. f, Heat map depicting the relative expression of the indicated genes in cluster C3 (Cxcr6+Pdcd1+) and cluster C7 (Cxcr6+Pdcd1–).
Transcriptional landscape of cholinergic CD4+ T cells in HCC
To delineate the heterogeneity of cholinergic CD4+ T cells in HCC, we conducted single-cell RNA sequencing (scRNA-seq) on sorted ChAT–GFP+ and ChAT–GFP− CD4+ T cells from four control and four HCC-bearing livers. Cells from individual mice were labeled with distinct antibody barcodes, pooled and processed for CITE-seq coupled with TCR-seq. In total, 11 clusters of CD4+ T cells were identified: clusters C1 and C8 were two naive T cell clusters; C2 was composed of T helper 17 (TH17) cells and cells expressing IL-18 receptor genes; C3 cells coexpressed Cxcr6 and Pdcd1; C4 cells contained follicular helper T (TFH) cells and follicular regulatory T (TFR) cells; C5 cells were actively cycling; C6 cells expressed Ccl5 and Nkg7 but few other markers; C7 cells expressed Cxcr6 but were negative for Pdcd1; C9 cells were canonical regulatory T (Treg) cells; C10 cells expressed Eomes, Prf1, Gzmk, Fasl, Gzmb and other cytotoxic genes; and C11 was a minor cluster expressing interferon (IFN)-stimulated genes (Fig. 2c–e). We observed a marked shift from naive T cells to effector T cells in the presence of HCC and, in particular, the induction of the C3 cluster.
Compared to ChAT–GFP− cells, the ChAT–GFP+ population lacked the naive T cell clusters as well as C7, with C2 cells also underrepresented (Fig. 2d). Cells in the C3, C4, C9 and C10 clusters were overrepresented among HCC ChAT–GFP+ T cells (Fig. 2d). We identified two subsets of Foxp3-expressing T cells, one in C4 and the other in C9. Comparing their transcriptomes, we discerned that the Foxp3+ cells in C4 were TFR cells that overexpressed Bcl6, Tcf7, Gpm6b and other TFR-associated genes and underexpressed Itgae and Gzmb (Extended Data Fig. 2d,e); this signature is consistent with previous reports29,30. Across all four HCC-bearing mice, Foxp3+ cells were significantly enriched among ChAT-expressing T cells (Extended Data Fig. 2f).
Cxcr6, a marker for resident T cells in the liver31, was enriched primarily in the C3 and C7 clusters. In HCC livers, the ChAT–GFP+ compartment contained a substantial number of C3 (Cxcr6+Pdcd1+) cells but was devoid of C7 (Cxcr6+Pdcd1–) cells (Fig. 2d). These C3 cells showed high expression of inhibitory immunoreceptors and exhaustion markers, such as Pdcd1, Havcr2, Ptpn11, Lag3, Tox and Tigit. These Pdcd1+ T cells also expressed Cd247, the gene encoding PD-L1, potentially providing an autologous ligand for PD-1 binding in addition to the PD-L1 expressed on APCs and HCC cells (Fig. 2f and Extended Data Fig. 2g). By contrast, C7 cells strongly expressed differentiation, cytotoxicity-related and other functional genes, such as Il2ra, Il4, Il2, Fasl, Gzmb and Csf2. These results suggest that Chat expression is associated with the appearance of dysfunctional Pdcd1+ T cells.
Examination of a published scRNA-seq dataset derived from immune cells isolated from individuals with HCC32 showed that ChAT-expressing T cells, including CD4+FOXP3+ Treg cells, CD8+MKI67+ proliferating T cells, CD8+GZMK+ T cells and CD8+PDCD1+ T cells, were present in liver tumor tissues but not in adjacent normal liver tissues (Extended Data Fig. 3a,b). Thus, the existence and phenotypes of human ChAT-expressing T cells are consistent with our scRNA-seq data on T cells from mouse HCC, indicating that a similar induction program of ChAT-expressing T cells also occurs in human HCC.
Cholinergic Treg cells and PD-1+CD4+ T cells are induced in HCC
To complement our scRNA-seq results, we profiled ChAT-expressing T cells in mouse HCC by flow cytometry. Treg cell accumulation is an immune hallmark of liver cancer33,34. In our model, we observed a marked increase in Foxp3+ChAT–GFP+ T cells alongside expansion of Treg cells during HCC development (Fig. 3a–c).
a–d, Representative flow cytometric plots (a) and quantification (b–d) of the percentages of the indicated CD4+ T cell subsets expressing Foxp3 and/or GFP in livers that were isolated on the indicated days from control or HCC-bearing Chat-GFP reporter mice. In b–d, each dot represents an individual mouse (n = 6 per condition). Data are shown as mean ± s.e.m. P values were determined by unpaired, two-tailed t-test, and data are representative of three independent experiments. e,f, Representative flow cytometric plots (e) and quantification (f) of the percentages of the indicated CD4+ T cell subsets expressing PD-1 and/or GFP in livers from control (n = 5) or HCC-bearing (n = 7) Chat-GFP reporter mice. In f, each dot represents an individual mouse (n = 5 mice in control and n = 7 mice in HCC). Data are shown as mean ± s.e.m. P values were determined by unpaired, two-tailed t-test. Data represent the summary of three independent experiments. g, Quantitation of flow cytometric determination of Ki67 expression in ChAT–GFP+ and ChAT–GFP– Tconv cells (Foxp3–) and Treg cells (Foxp3+) in livers from HCC-bearing Chat-GFP mice. Each set of dots represents an individual mouse. P values were determined by paired two-tailed t-tests, and data are representative of three independent experiments. For a–g, the control group received transposase vector only. h, Representative histological sections of livers from the mice in a–f immunostained to detect Foxp3 and GFP in normal liver tissue (control; left), liver areas containing preneoplastic cells (middle) or HCC cells (right). Images are representative of three independent experiments. The inset box on the right shows a higher magnification view of the smaller boxed area. Arrowheads denote cells with dual Foxp3 and GFP immunoreactivity, and arrows denote cells with GFP immunoreactivity only; T, tumor tissue; NT, non-tumor tissue.
HCC livers also exhibited a significant increase in Foxp3–ChAT–GFP+CD4+ T cells (Fig. 3d), and PD-1+CD4+ T cells that were induced in HCC preferentially expressed ChAT–GFP (Fig. 3e,f). Furthermore, significantly higher Ki67 expression occurred on ChAT–GFP+Foxp3+ Treg cells and Chat–GFP+Foxp3– conventional T (Tconv) cells (Fig. 3g), suggesting that proliferation underlies the induction of ChAT-expressing T cells in liver cancer.
Histologically, ChAT–GFP+Foxp3+ Treg cells and ChAT–GFP+ Tconv cells accumulated in HCC tissue, in particular at the tumor border, and were also present in immune cell clusters associated with neoplastic hepatocytes (Fig. 3h).
Tumor antigens drive cholinergic CD4+ T cell expansion
The proliferation characteristic of ChAT-expressing T cells in HCC livers led us to investigate the driving force behind their expansion. Single-cell TCR-seq revealed dominant TCR types across all animals (as defined by shared amino acid sequences for both the TCRα and TCRβ chains; Fig. 4a,b). Among the top 30 most prevalent TCR types, only 5 were present in control mice, while the remaining 25 were observed in their HCC-bearing littermates. Intriguingly, the prevalent TCRs in control mice preferentially belonged to ChAT–GFP– T cells, while those in HCC-bearing mice were more commonly found on ChAT–GFP+ T cells (Fig. 4a,b). These results demonstrate a TCR-specific expansion of ChAT–GFP+ T cells in liver cancer.
a,b, Circos plots showing the distribution of TCR types among GFP+ and GFP– T cells from control (a) and HCC-bearing (b) mice. T cells of the same TCR type share TCRα and TCRβ chains of the same amino acid sequences. The top 30 TCRs are numbered and highlighted with different colors. c, CDR3 sequences of clonotypes encoding TCR 1. V, D and J segments and N nucleotides and P nucleotides at V(D)J junctions are denoted with different colors. Nucleotides that are mismatched between clonotypes are shaded. d, Composition of ChAT–GFP+ and ChAT–GFP– cells among T cells bearing the indicated TCR 1 clonotypes color-coded by cell clusters. Each bar represents an individual clonotype and is labeled with the clone ID as shown in c. Horizontal axis labels indicate cell numbers. e, Percentage of ChAT–GFP+ cells among CD4+ T cells from normal or HCC-bearing livers of Chat-GFP or Chat-GFP; OT-II mice (n = 6 in the control, HCC and OT-II control groups; n = 5 in the OT-II HCC group). Statistical significance was assessed by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test, and data are representative of three independent experiments. f,g, Representative flow cytometry plots (f) and quantification (g) of percentages of ChAT–GFP+ cells among CD4+ T cells expressing TCR Vβ5+ (transgenic TCR) or TCR Vβ5– (natural TCRs) in livers of Chat-GFP; OT-II mice (n = 7 in control; n = 6 in HCC). h, Plasmids for simultaneous CRISPR–Cas9-mediated deletion of Trp53/Pten plus overexpression of Myc and tetracycline-on (Tet-On) inducible OVA. i, Experimental protocol for inducing OVA expression. Dox was added to drinking water following palpable HCC onset; FACS, fluorescence-activated cell sorting. j,k, Representative flow cytometry plots (j) and quantification (k) of percentages of ChAT–GFP+ cells among CD4+ T cells expressing TCR Vβ5+ or TCR Vβ5– in livers of mice that were left untreated or treated with Dox-containing drinking water following HCC onset. Data are the summary of two independent experiments (n = 3 mice per group). In e, g and k, each dot represents an individual mouse. Data are shown as mean ± s.e.m. In g and k, P values were determined by two-way ANOVA with Sidak’s multiple comparisons test.
The most dominant TCR type in HCC was denoted TCR 1, which was encoded by 25 clonotypes (as defined by shared mRNA sequences for both the TCRα and TCRβ chains). These clonotypes consisted of synonymous mRNA variants using the same combination of V, D and J genes. The variations arose from distinct junctions of the V, D and J segments occurring in different HCC-bearing mice, but the resulting amino acid sequences were identical due to codon redundancy (Fig. 4c). The majority of TCR 1 clonotypes were predominantly observed in ChAT–GFP+ cells (Fig. 4d). This TCR convergence across HCC-bearing mice suggests that TCRs specific for HCC antigens elicit the expansion of CD4+ T cells, particularly within the ChAT–GFP+CD4+ T cell compartment.
We next characterized the cells bearing particular TCR types. T cells carrying TCR 1 were mainly from the C3 cluster that harbored ChAT-expressing Pdcd1+ Tconv cells (Fig. 4d and Extended Data Fig. 4a). T cells carrying TCR 22 were chiefly ChAT-expressing Foxp3+ Treg cells (Extended Data Fig. 4b). These results reveal an accumulation of ChAT-expressing, clonally expanded PD-1+ Tconv cells and Foxp3+ Treg cells in HCC. Pertinently, clonal expansion of Treg cells has also been revealed by TCR usage analysis at the single-cell level in human HCC34. Notably, T cells bearing TCR 3 were exclusively ChAT–GFP– cells in HCC (Fig. 4b) and belonged primarily to cluster C7 (Cxcr6+Pdcd1–; Extended Data Fig. 4c). Among the 25 clonotypes of TCR 1, clones 14 and 25 were predominantly C3 cells but ChAT–GFP–, although they carried the same TCR and shared a similar composition of cell clusters with many other ChAT–GFP+ clonotypes of TCR 1 (Fig. 4c,d). We postulate that this clonal divergence in ChAT expression likely reflects the complexity of the tumor microenvironment.
To determine the role of tumor antigens in the induction of ChAT-expressing T cells, we crossed Chat-GFP mice with chicken ovalbumin (OVA)-specific TCR transgenic OT-II mice. First, we subjected Chat-GFP; OT-II mice to standard HCC induction, which does not involve OVA expression. We found that ChAT–GFP+ T cells did not become significantly elevated, and most were CD4+ T cells expressing a natural repertoire of TCRs that were Vβ5– (Fig. 4e–g).
Next, we used OVA to mimic a tumor antigen and devised a vector allowing the inducible expression of cytosolic OVA with constitutive expression of Myc. We introduced this vector alongside the CRISPR Trp53/Pten deletion vector into Chat-GFP; OT-II mice (Fig. 4h). When HCC tumors were palpable, doxycycline (Dox) was administered to these mice through their drinking water to induce OVA expression (Fig. 4i). When we compared ChAT expression in Dox-treated and untreated mice, we found that about 40% of OVA-specific (TCR Vβ5+) CD4+ T cells expressed ChAT after OVA induction in HCC (Fig. 4j,k). In untreated mice, only about 2% of Vβ5+CD4+ T cells expressed ChAT, comparable to the percentage in Chat-GFP; OT-II mice bearing OVA– HCC (Fig. 4f,g,j,k). Interestingly, we also observed an elevated percentage of ChAT–GFP+ cells among CD4+ T cells carrying natural TCRs (TCR Vβ5–) (Fig. 4j,k), suggesting that ChAT expression in T cells can also be induced through ‘antigen spreading’35.
We further characterized the OVA-specific ChAT+ T cells in our HCC model with inducible OVA expression and found that Dox-mediated activation of OVA expression markedly induced Treg cells, especially among OVA-specific T cells (Vβ5+; Extended Data Fig. 5a). There was also an induction of ChAT–GFP+ T cells among both Foxp3+ Treg cells and Foxp3– Tconv cells harboring OVA-specific TCRs (Extended Data Fig. 5a). PD-1 was highly expressed by these OVA-specific Tconv cells, and ~40% of these cells were ChAT–GFP+ (Extended Data Fig. 5a). Taken together, these data confirm that tumor antigens can induce ChAT-expressing Treg cells and PD-1+ Tconv cells and demonstrate that the expansion of ChAT-expressing T cells in liver cancer depends on TCR activation by tumor antigens.
Ablation of Chat in T cells dampens HCC immunosurveillance
To determine the role of ChAT-expressing T cells in the onset of liver cancer, we deleted Chat specifically in T cells by crossing mice carrying the conditional Chatfl allele with mice expressing the Cd4-cre transgene, thereby obtaining Chatfl/fl; Cd4-cre progeny. When we subjected these animals (and Chatfl/fl controls) to HCC induction, we found that Chatfl/fl; Cd4-cre mice developed liver cancer much faster than their Chatfl/fl littermates (Fig. 5a). The numbers of tumor nodules and liver weights were also significantly increased in Chatfl/fl; Cd4-cre mice (Fig. 5b–e).
a, Curves showing latency to palpable HCC development (left, black lines) and survival to humane endpoint (right, red lines) of Chatfl/fl (n = 14) and Chatfl/fl; Cd4-cre (n = 20) mice. P values were determined by log-rank (Mantel–Cox) test, and data are representative of two independent experiments. b–e, Representative images (b), numbers of tumor nodules (c), liver weights (d) and ratios of liver weight (LW) to body weight (BW; e) of mice on day 35 of HCC induction. For c–e, n = 11 mice in the Chatfl/fl group and n = 12 mice in Chatfl/fl; Cd4-cre group; data are representative of five independent experiments. f,g, Representative images (f) and quantification (g) of tumor incidence in mice fed for 15 months on a Western diet (WD). P values were determined by Fisher’s exact test. h,i, Representative histological sections (h) and quantification (i) of immunostaining to detect MYC+ preneoplastic cells in non-tumor areas of liver sections from the mice in b–e. In i, n = 11 Chatfl/fl mice and n = 9 Chatfl/fl; Cd4-cre mice. The inset box in h shows a higher-magnification view of the smaller boxed area. Arrowheads indicate immune cell clusters. j,k, Quantification of the number (j) and size (k) of immune cell clusters in hematoxylin and eosin (H&E)-stained sections of livers from the mice in b–e (n = 11 Chatfl/fl mice and n = 9 Chatfl/fl; Cd4-cre mice). l, Percentage of CD3+ T cells among MNCs isolated from HCC-bearing livers as assessed by flow cytometry (n = 11 Chatfl/fl mice and n = 12 Chatfl/fl; Cd4-cre mice); data are representative of three independent experiments. m–o, Representative flow cytometry plots (m) and quantification of IFNγ+CD4+ (n) and IL-17A+CD4+ T cells (o) in HCC-bearing livers (n = 11 mice per group); data are representative of two independent experiments. p, Quantitative PCR (qPCR) determination of mRNA levels (relative to Actb) of the indicated cytotoxicity genes and NK cell marker genes in HCC-bearing livers (n = 5 mice per group); data are representative of two independent experiments. For c–e, i–l and n–p, each dot represents an individual mouse. Data are shown as mean ± s.e.m., and P values were determined by unpaired, two-tailed t-tests.
To further substantiate the role of ChAT-expressing T cells in liver tumorigenesis, we used an alternative disease model in which Chatfl/fl; Cd4-cre and Chatfl/fl mice were fed long term on a Western diet (high fat, high cholesterol and high sugar) to induce non-alcoholic steatohepatitis (NASH). NASH sets the stage for liver cirrhosis, which eventually progresses to spontaneous HCC8,36. After 15 months on the Western diet, we found that the incidence of NASH-derived HCC was significantly higher in mice bearing T cells lacking Chat (Fig. 5f,g). These consistent results from two models of HCC development establish that a deficiency of Chat in T cells renders mice susceptible to liver tumorigenesis.
Before overt tumor nodules appeared in our HCC model, preneoplastic cells could be identified by their high MYC expression (Fig. 5h), reflecting successful transfection and oncogene expression. The proportion of preneoplastic hepatocytes showing high MYC expression was significantly elevated in livers of Chatfl/fl; Cd4-cre mice, and this increase was not due to a difference in vector delivery (Fig. 5h,i and Extended Data Fig. 6a–c). Immune cells are critical for clearing preneoplastic cells from the liver5. In Chatfl/fl mice, we observed immune cell clusters around MYC-expressing preneoplastic cells (Fig. 5h). These clusters were reduced in frequency and size in Chatfl/fl; Cd4-cre mice (Fig. 5h,j,k), suggesting a defect in immunosurveillance of preneoplastic cells. Accordingly, T cell infiltration into HCC-bearing livers was significantly decreased in the absence of Chat (Fig. 5l and Extended Data Fig. 6d,e).
IFNγ, a hallmark of the type 1 helper T cell immune response, has a pivotal function in antitumor immunity37. We found that IFNγ production by HCC-associated T cells was decreased in Chatfl/fl; Cd4-cre mice (Fig. 5m–p and Extended Data Fig. 6f). In addition to adaptive immune responses, innate cytotoxic NK cells play a crucial role in antitumor immune response in HCC38,39. We observed that mRNA levels of genes encoding IFNγ, granzymes and perforin, which are cytotoxic effectors shared by cytotoxic T cells and NK cells, were significantly decreased in Chatfl/fl; Cd4-cre mice (Fig. 5p, left). These deficits correlated with reduced expression of NK cell marker genes (Fig. 5p, right). Therefore, both adaptive and innate antitumor immune responses are hampered by the ablation of Chat in T cells.
Treg cells blunt HCC immunosurveillance in Chat fl/fl; Cd4-cre mice
To delve more deeply into the mechanism of antitumor immunity mediated by ChAT-expressing T cells, we devised a vector mediating the expression of OVA alongside MYC. We introduced this vector plus our Trp53/Pten deletion vector into OVA-immunized and non-immunized mice (Extended Data Fig. 7a). In non-immunized mice, tumor progression to endpoint was significantly accelerated by Chat ablation in T cells (Extended Data Fig. 7b,c), consistent with our previous observations (Fig. 5a). However, OVA-immunized Chatfl/fl and Chatfl/fl; Cd4-cre mice were equally protected against HCC development (Extended Data Fig. 7b,c). These results showed that the antitumor immune response elicited by a potently immunogenic tumor antigen was not compromised in the absence of ChAT in T cells. In addition, depletion of cytotoxic T lymphocytes (CTLs) by treatment with anti-CD8 had little effect on liver tumor burden (Extended Data Fig. 7d,e). Thus, CTLs do not play a non-redundant role in the immunosurveillance of liver cancer in this setting.
To determine if loss of adaptive immune cells in general would compromise antitumor immunity in our model, we compared the onset of HCC in lymphocyte-deficient Rag1–/– mice to that in Chatfl/fl; Cd4-cre mice. In contrast to the significantly increased tumor burden in Chatfl/fl; Cd4-cre mice, HCC progression was not exacerbated in Rag1–/– mice but instead was alleviated (Extended Data Fig. 7f,g). We reasoned that this unexpected observation could be attributed to the absence of Treg cells and dysfunctional effector T cells in Rag1–/– mice. This loss of control by adaptive immune cells (especially Treg cells) causes Rag1–/– mice to exhibit an excessive innate immune response by NK cells40. The fact that HCC progression was aggravated in NSG mice (which lack both adaptive immune cells and NK cells; Fig. 1h) bolsters our contention that the antitumor activity of NK cells in our model is curbed by adaptive immune cells, particularly Treg cells.
To investigate our hypothesis that the accelerated tumor onset linked to T cell-specific Chat ablation could be due to suppression of antitumor responses by elevated Treg cell activity and/or Tconv cell dysfunction, we examined how Treg cells modulate the antitumor activity of ChAT-expressing T cells. We observed no difference in the abundance of Foxp3+ Treg cells in liver tumors of Chatfl/fl and Chatfl/fl; Cd4-cre mice (Extended Data Fig. 8a). However, CD25 expression by Foxp3+ Treg cells in Chatfl/fl; Cd4-cre mice was substantially increased compared to in control mice (Fig. 6a–c), whereas CTLA-4 expression levels were similar (Extended Data Fig. 8b–d). We then purified CD25–CD4+ T cells from spleens of Chatfl/fl and Chatfl/fl; Cd4-cre mice and analyzed CD25 induction following TCR stimulation by anti-CD3/CD28 beads. CD25 expression in Foxp3+ Treg cells from Chatfl/fl; Cd4-cre mice was significantly higher than in Treg cells from Chatfl/fl mice (Extended Data Fig. 8e). This induction of CD25 by TCR activation was much stronger on Treg cells than on Tconv cells (Extended Data Fig. 8e). Therefore, TCR-induced expression of CD25 in Treg cells is modulated by ChAT in T cells. When we used anti-CD25 to deplete CD25-expressing Treg cells (Fig. 6d and Extended Data Fig. 8f), we observed that the enhanced tumor burden in Chatfl/fl; Cd4-cre mice was partially decreased (Fig. 6e,f). Together, these results point toward the involvement of Treg cells in the suppression of antitumor immune responses in Chatfl/fl; Cd4-cre mice.
a–c, Representative flow cytometry plots (a), quantification of the percentages of Foxp3+CD4+ T cells expressing CD25 (b) and the CD25 mean fluorescent intensity (MFI; c) in HCC-bearing livers from Chatfl/fl and Chatfl/fl; Cd4-cre mice on day 35 of HCC induction (n = 9 mice per group). P values were determined by unpaired, two-tailed t-test; data are representative of two independent experiments. d, Experimental protocol used to deplete CD25-expressing Treg cells during HCC induction. e,f, Representative low-magnification histological images of H&E-stained sections (e) and quantification of numbers of tumor nodules per mm2 in these sections (f) of livers from Chatfl/fl and Chatfl/fl; Cd4-cre mice treated as in d. In f, n = 7 mice in the Chatfl/fl + IgG, Chatfl/fl + anti-CD25 and Chatfl/fl; Cd4-cre + anti-CD25 groups, and n = 9 mice in the Chatfl/fl + anti-CD25 group. P values were determined by two-way ANOVA with Tukey’s multiple comparisons tests. The ‘X’ symbols indicate animals that reached the humane endpoint before day 20. g, Experimental protocol used to deplete CD4+ T cells and NK cells during HCC induction. Each mouse received 100 μg of depleting antibody per injection. Mice in the control group received either 100 μg of rat IgG or PBS. h,i, Quantification of numbers of tumor nodules per mm2 in H&E-stained sections (h) and ratios of liver weight to body weight (i) of Chatfl/fl and Chatfl/fl; Cd4-cre mice treated as in g. The ‘X’ symbols indicate animals that reached the humane endpoint before day 20. In h, n = 8 mice in the Chatfl/fl + IgG/PBS and anti-CD4 groups; n = 7 mice in the Chatfl/fl; Cd4-cre + IgG/PBS group; n = 9 mice in the Chatfl/fl + anti-NK1.1 group; and n = 5 mice in the Chatfl/fl; Cd4-cre + anti-NK1.1 group. In i, n = 8 mice in the IgG/PBS and anti-CD4 groups; n = 9 mice in the Chatfl/fl + anti-NK1.1 group; and n = 5 mice in the Chatfl/fl; Cd4-cre + anti-NK1.1 group. In b, c, f, h and i, each dot represents an individual mouse. Data are shown as means ± s.e.m. P values in h were determined by paired, two-tailed t-tests. P values in i were determined by two-tailed Mann–Whitney test because the Chatfl/fl group did not pass the normality test; NS, not significant.
CD25-expressing Treg cells inhibit the antitumor activities of cytotoxic T cells and NK cells41. Chatfl/fl; Cd4-cre mice showed significantly increased expression of CD25 on Treg cells (Fig. 6a–c) and reduced numbers of IFNγ+CD4+ T cells and NK cells (Fig. 5m–p). When we removed CD4+ T cells during HCC induction using anti-CD4 (Fig. 6g and Extended Data Fig. 8g), we observed that CD4+ T cell depletion did not affect HCC progression in Chatfl/fl mice but tended to reduce it in Chatfl/fl; Cd4-cre mice (Fig. 6h,i). This result suggested that the collective functions of Treg cells and Tconv cells in HCC are neutral in Chatfl/fl mice, whereas Chat deletion in T cells tilts the balance toward HCC promotion. Because CD4+ T cell depletion abolished the difference in HCC development between Chatfl/fl mice and Chatfl/fl; Cd4-cre mice, the independent contributions to HCC of other ChAT-expressing T lineage cells, including CD8+ T cells and NKT cells, appear to be minor. When we used anti-NK1.1 to deplete NK cells from our model (Fig. 6g and Extended Data Fig. 8h), HCC development was promoted in Chatfl/fl mice but not in Chatfl/fl; Cd4-cre mice, essentially eliminating the differences in tumor progression (Fig. 6h,i). Thus, NK cells are indispensable for HCC immunosurveillance in our model, and the antitumor functions of NK cells are impeded in the absence of cholinergic T cells.
ChAT loss in T cells exacerbates T cell dysfunction in HCC
To investigate the role of ChAT specifically in Treg cells, we crossed Chatfl/fl mice with Foxp3Cre mice and induced HCC in Foxp3Cre and Foxp3Cre; Chatfl/fl littermates. We found that deleting Chat only in Treg cells had a milder effect on HCC progression than did deleting Chat in all T cells (Extended Data Fig. 8i,j), suggesting that ChAT-expressing Tconv cells are indispensable for a full-fledged anti-HCC immune response. ChAT-expressing Tconv cells induced in HCC are primarily PD-1+ T cells (Fig. 3e,f) that coexpress inhibitory immunoreceptors, such as Tim-3 (Havcr2), Lag-3, CTLA-4 and other molecules characteristic of T cell exhaustion and dysfunction (Fig. 2f). PD-1 expression was significantly higher in Foxp3–CD4+ Tconv cells from Chatfl/fl; Cd4-cre mice than in those from Chatfl/fl mice (Fig. 7a,b). Notably, PD-1 levels strongly correlated with HCC grade in Chatfl/fl; Cd4-cre mice but not in Chatfl/fl mice (Fig. 7c). We previously reported that loss of ChAT in T cells promotes the expression of PD-1, Tim-3 and Lag-3 during chronic viral infection27. In the present study, both CD4+ T cells and CD8+ T cells in HCC-bearing livers showed a broad trend of upregulation of these inhibitory receptors in the absence of cholinergic T cells (Extended Data Fig. 9a–f). Thus, in the absence of Chat in T cells, an unleashing of PD-1 inhibitory activity occurs that may restrict the functions of antitumor Tconv cells, allowing HCC progression.
a,b, Representative flow cytometry histogram overlay plot (a) and quantification of the percentages of Foxp3–CD4+ Tconv cells expressing PD-1 (b) in HCC-bearing livers from Chatfl/fl and Chatfl/fl; Cd4-cre mice. Each dot represents an individual mouse (n = 8 Chatfl/fl mice and n = 9 Chatfl/fl; Cd4-cre mice). Data are shown as mean ± s.e.m. The P value was determined by unpaired, two-tailed t-test. c, Correlation of PD-1 expression in Tconv cells with HCC grade in Chatfl/fl and Chatfl/fl; Cd4-cre mice. HCC grade is represented by the ratio of liver weight to body weight. P values were determined by two-tailed Pearson correlation. d, Experimental protocol used for PD-1 blockade in vivo in Chatfl/fl and Chatfl/fl; Cd4-cre mice subjected to standard HCC induction. e,f, Representative low-magnification histological images of H&E-stained sections (e) and quantification of numbers of tumor nodules per mm2 in these sections (f) of livers from Chatfl/fl and Chatfl/fl; Cd4-cre mice treated as in d. In f, each dot represents an individual mouse (n = 12 mice in the Chatfl/fl + IgG group, n = 10 mice in the Chatfl/fl + anti-PD-1 and Chatfl/fl; Cd4-cre + IgG groups, and n = 11 mice in the Chatfl/fl + anit-PD-1 group). Data are shown as mean ± s.e.m. P values were determined by two-way ANOVA with Tukey’s multiple comparisons test.
To test this hypothesis, we administered PD-1 blockade antibodies to Chatfl/fl and Chatfl/fl; Cd4-cre mice during HCC development (Fig. 7d). As reported for NASH-induced HCC8, we did not observe a therapeutic effect of PD-1 blockade in control Chatfl/fl mice, perhaps due to ChAT expression by PD-1+ Tconv cells and the negative effects of cholinergic activity on PD-1 expression. However, PD-1 blockade significantly reduced HCC development in Chatfl/fl; Cd4-cre mice, substantially eliminating the differences in tumor progression (Fig. 7e,f). We did observe two tumor-free animals among anti-PD-1-treated Chatfl/fl; Cd4-cre mice (Fig. 7f), a status rarely seen for this genotype. These data suggest that ChAT invigorates dysfunctional T cells in HCC.
Cholinergic modulation of TCR-induced Ca2+–NFAT signaling
CD25 expression is controlled by the transcription factor NFAT, which is regulated by Ca2+ signaling42. TCR-induced Ca2+–NFAT signaling is indispensable for T cell exhaustion and induces the expression of inhibitory surface receptors, such as PD-1, Lag-3 and Tim-3 (ref. 43). NFAT-induced transcription factors, including NR4A and TOX, drive T cell exhaustion/dysfunction44,45,46,47. We therefore sought to determine if cholinergic activity affects Ca2+ signaling in T cells. Ca2+ influx elicited by TCR engagement was significantly stronger in Chatfl/fl; Cd4-cre T cells than in their Chatfl/fl counterparts (Fig. 8a–c). Furthermore, TCR activation-induced nuclear translocation of NFAT was higher in Chatfl/fl; Cd4-cre T cells than in Chatfl/fl T cells (Fig. 8d,e). Interestingly, when Chatfl/fl; Cd4-cre T cells were prestimulated with ACh, NFAT nuclear translocation was no longer inducible by TCR activation (Fig. 8d,e). Thus, cholinergic activity in T cells constrains TCR-induced Ca2+–NFAT signaling.
a–c, Representative overlaid kinetics plots of flow cytometry curves (a), quantification of Ca2+ influx peaks (b) and ‘area under curve’ (AUC; c) for the Ca2+ flux occurring in Chatfl/fl and Chatfl/fl; Cd4-cre CD4+ T cells. In b and c, each dot represents T cells from an individual mouse. Chatfl/fl and Chatfl/fl; Cd4-cre littermates of both sexes across various ages were paired for analysis, with at least two measurements taken per mouse. P values were determined by paired, two-tailed t-test; data are a summary of two independent experiments. d,e, Representative fluorescence micrographs (d) and quantification (e) of NFAT immunofluorescent staining of splenic CD4+ T cells purified from Chatfl/fl and Chatfl/fl; Cd4-cre mice. Ratios of nuclear NFAT to cytoplasmic NFAT are displayed in each image in d and are statistically compared in e. Each dot in e represents one cell (n = 160 control Chatfl/fl T cells; n = 180 anti-CD3-treated Chatfl/fl T cells; n = 101 control Chatfl/fl; Cd4-cre T cells; and n = 269, 128, 119, 246, 114, 157 and 212 Chatfl/fl; Cd4-cre T cells treated with anti-CD3, ACh, ACh + anti-CD3, nicotine (Nic), nicotine + anti-CD3, Oxo-M or Oxo-M + anti-CD3). Data are shown as mean ± s.e.m. P values were determined by one-way ANOVA with Tukey’s multiple comparisons test and are representative of two independent experiments. f, qPCR determination of mRNA levels (relative to Actb) of the indicated nAChR (Chrna1–Chrna9) and mAChR (Chrm1–Chrm5) genes in Tconv and Treg CD4+ T cells sorted from livers of HCC-bearing Foxp3-YFP mice (n = 4); data are representative of three independent experiments. g, Diagram summarizing our proposed model of ChAT function in T cells during HCC. In wild-type mice, HCC antigens induce the expression of ChAT in T cells. Autocrine/paracrine cholinergic signaling by ChAT-expressing T cells influences T cell Ca2+ homeostasis and regulates TCR-induced Ca2+ signaling. Without such cholinergic modulation (as occurs in Chatfl/fl; Cd4-cre mice), TCR-induced Ca2+ signaling is hyperactivated, leading to T cell exhaustion, overexpression of PD-1 in Tconv cells and increased CD25 in Treg cells. The inhibitory activity of PD-1 is unleashed, and Treg-mediated suppression is enhanced, compromising antitumor responses mounted by NK cells and Tconv cells. In the absence of ChAT, HCC progression proceeds unabated; Ag, antigen; MHC-II, major histocompatibility complex class II.
ACh regulates intracellular Ca2+ levels by binding to nAChRs and mAChRs48. We found that Treg cells and Tconv cells in livers of HCC-bearing mice expressed similar arrays of AChRs (Fig. 8f). We then applied either a nicotinic agonist or a muscarinic agonist to Chatfl/fl; Cd4-cre T cells in vitro. Nicotine, which activates nAChRs, had no detectable influence on TCR-induced NFAT translocation. By contrast, oxotremorine methiodide (Oxo-M), an mAChR agonist, by itself induced an increase in NFAT nuclear translocation and abolished subsequent TCR-induced NFAT translocation (Fig. 8d,e).
The Gq/11-coupled M1, M3 and M5 mAChRs elicit Ca2+ release from the endoplasmic reticulum (ER) by activating the downstream phospholipase C/inositol 1,4,5-trisphosphate/inositol 1,4,5-trisphosphate receptor (PLC/IP3/IP3R) cascade49, the same pathway triggered by TCR signaling. Mobilization of Ca2+ in T cells is a biphasic event divided into the initial releasing of intracellular ER Ca2+ stores and the subsequent extracellular Ca2+ influx from ‘Ca2+ release-activated Ca2+’ (CRAC) channels50. The CRAC channels can also be triggered by mAChR activation in T cells51,52. However, the activation of mAChR depletes the IP3-sensitive Ca2+ pool (ER Ca2+) and renders the T cells unresponsive to further CRAC channel-dependent stimulation52. In line with these findings, our results indicate that cholinergic signaling via muscarinic receptors desensitizes T cells to TCR-induced Ca2+–NFAT signaling.
Parallels to human HCC
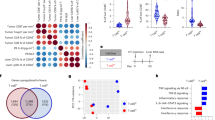
We examined an scRNA-seq dataset34 and noted that T cells from individuals with HCC expressed CHAT and an array of mAChRs and nAChRs (Extended Data Fig. 10a). To further study this association between cholinergic signaling and human HCC, we examined HCC cases profiled by The Cancer Genome Atlas (TCGA). High expression of CHRM3 or CHRM5 in samples from individuals with HCC was positively correlated with a favorable prognosis (Extended Data Fig. 10b–d). These observations prompted us to use our mouse model to determine if Chrm3 and Chrm5 expressed by HCC cells were acting to directly suppress HCC development. To this end, we designed CRISPR vectors to target M3 and M5 mAChRs and induced Chrm3/Chrm5-knockout HCC. We found that the knockout of M3 and M5 AChRs in mouse HCC cells did not significantly affect HCC development (Extended Data Fig. 10e,f). Therefore, our data do not support the direct suppression of HCC by CHRM3 and CHRM5 in HCC cells. We do acknowledge that there could be compensation by other receptors or interspecies differences.
Collectively, our study supports a model (Fig. 8g) whereby tumor antigens induce the expansion of ChAT-expressing Treg cells and PD-1+ Tconv cells. ACh produced by these T cells modulates TCR-induced Ca2+ signaling to prevent its hyperactivity. In the absence of such cholinergic modulation, the Ca2+–NFAT pathway becomes hyperactivated to increase immunosuppression by Treg cells and to impose dysfunction on Tconv cells, resulting in compromised antitumor immunity.
Discussion
The immune milieu of the liver at steady state favors tolerance over immune responses to prevent overreaction to innocuous antigens from food, microbial substances or by-products of metabolism. Liver Treg cells have a crucial function in the maintenance of this peripheral tolerance state53. In the livers of individuals with HCC, Treg cell infiltration is prominently elevated, alongside the exhaustion and impaired function of CD8+ T cells34,54. The clonal expansion of Treg cells and exhausted T cells in individuals with HCC has been revealed by scRNA-seq34. Using scRNA-seq and flow cytometric analysis, we have demonstrated the induction of Foxp3+ Treg cells and PD-1+ Tconv cells in our mouse HCC model and have shown that ChAT-expressing T cells predominantly belong to these two populations.
The clonal expansion of ChAT-expressing Treg cells and PD-1+ Tconv cells in our model appears to be driven by tumor antigens. It should be stressed that our HCC model does not involve viral vectors or mutant proteins; as such, the tumor antigens involved here are likely ‘self’ proteins. Such proteins are postulated to be weakly immunogenic, particularly within the tolerogenic immune milieu in the liver. Indeed, the T cell immune response to HCC antigens is reportedly dysfunctional and unreactive to HCC55. Our data suggest that ChAT deficiency in T cells exacerbates the dysfunction of Tconv cells and reinforces the immunosuppressive function of Treg cells, significantly compromising immunosurveillance against liver cancer. We used OVA to mimic a ‘foreign’ tumor antigen by inducing OVA HCC in immunized mice. We expected that HCC onset in this situation would induce a potent antitumor immune response rather than the accumulation of Treg cells and dysfunctional Tconv cells. Indeed, OVA HCC development was prevented to the same degree in Chatfl/fl and Chatfl/fl; Cd4-cre mice. Thus, the cholinergic activity of T cells is most relevant when triggered by ‘self’ tumor antigens.
Many actions of ACh in the nervous system are mediated through Ca2+ signaling via either G-protein-coupled mAChRs or ionotropic nAChRs48. nAChRs are ACh-gated ionic channels with a variable range of permeability to Ca2+. The M1, M3 and M5 mAChRs are coupled with Gq/11 and elicit the PLC/IP3/IP3R cascade to trigger ER Ca2+ release, whereas M2 and M4 mAChRs are generally linked via Gi/o to cAMP production48. We demonstrated that these three classes of AChRs are expressed by both Tconv cells and Treg cells in HCC-bearing livers. In T lymphocytes, M1 mAChR activation and TCR engagement rely on the same molecular pathway to trigger Ca2+ influx. Due to this overlap, Ca2+ signaling events triggered by these pathways are mutually exclusive52. In the Premack study, the maximal cholinergic stimulation mediated by overexpressed M1 receptor completely emptied the ER Ca2+ store, the Ca2+ pool essential for TCR-induced Ca2+ signaling. Moreover, the ER Ca2+ release mediated by cholinergic signaling occurred in a quantal manner: a submaximal concentration of cholinergic agonist rapidly released a fraction of the ER Ca2+ store, followed by a slower or terminated Ca2+ release56,57. Certain mechanisms, including the inactivation of IP3 receptors, can be triggered to attenuate ER Ca2+ release57,58. During persistent submaximal cholinergic stimulation, the ER Ca2+ release elicited by other IP3-dependent agonists is dampened56. Due to the modest expression levels of AChRs by T cells and the ubiquitous presence of acetylcholinesterase59, we speculate that autocrine/paracrine ACh signaling by T cells might operate in such a submaximal way. Accordingly, although we did not observe significant differences in basal levels of Ca2+ influx and NFAT nuclear translocation between Chatfl/fl and Chatfl/fl; Cd4-cre T cells, both TCR-induced Ca2+ influx and NFAT nuclear translocation were elevated in the absence of cholinergic activity in T cells. Moreover, ACh pretreatment inhibited TCR-induced NFAT nuclear translocation. Collectively, these data allow us to propose a model in which autocrine/paracrine ACh produced by ChAT-expressing T cells affects their Ca2+ homeostasis. This altered Ca2+ status restrains the expression of CD25 by Treg cells and the expression of PD-1 and other exhaustion markers by Tconv cells. In the absence of ChAT in T cells, hyperimmunosuppressive Treg cells and dysfunctional Tconv cells interfere with adaptive and innate antitumor responses and permit HCC progression.
In conclusion, our study shows that lymphocytes are the dominant cholinergic cells in HCC-bearing livers and that ChAT-expressing T cells orchestrate immune responses against HCC. Our results may prompt investigation of how lymphocyte-mediated cholinergic regulation of liver carcinogenesis can be exploited to enhance antitumor immune responses in individuals with liver cancer.
Methods
Mice
Chat-GFP (B6.Cg-Tg(RP23-268L19-EGFP)2Mik/J), Chatfl (B6.129-Chattm1Jrs/J), Cd4-cre (Tg(Cd4-cre)1Cwi/BfluJ), Il21r–/– (B6.129-Il21rtm1Kopf/J), OT-II (B6.Cg-Tg(TcraTcrb)425Cbn/J), Confetti (Gt(ROSA)26Sortm1(CAG-Brainbow2.1)Cle/J), Foxp3YFP/Cre (B6.129(Cg)-Foxp3tm4(YFP/icre)Ayr/J), NSG (NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ) and control NOD/ShiLtJ mice were all purchased from the Jackson Laboratory and bred in the animal facility at the Princess Margaret Cancer Centre. The mice were housed on ventilated racks supplied with autoclaved microisolator cages. Reverse osmosis water was supplied through an automatic watering system. The light cycle was lights off at 1800 h and lights on at 0600 h. The ambient temperature was held between 22 and 23 °C with a humidity of 40–60%. Mice were routinely fed on the irradiated 7012 Teklad LM-485 Mouse/Rat Sterilizable Diet. All animal experiments were approved by the University Health Network Animal Care Committee.
CRISPR and transposon vectors
Single guide RNAs (sgRNAs) targeting Trp53 and Pten were as previously described21. The guide oligonucleotides were designed to express mouse Pten sgRNA and Trp53 sgRNA (Supplementary Table 1). The annealed double-stranded guide oligonucleotides were cloned into the BbsI cut sites of the pX330 vector60. The Trp53 sgRNA cassette in pX330-p53 was amplified by PCR with primers (Supplementary Table 1). This additional sgRNA cassette was then cut with NheI and XbaI and subcloned into the NheI site of pX330-Pten to obtain the duplex CRISPR vector pX330-p53-Pten.
The transposon system using SB100X and pT2/BH was as described previously61. The mouse Myc coding sequence was cloned into pT2/BH with EcoR1 and NotI restriction enzymes to obtain the pT2-Myc plasmid.
A modified puromycin-T2A-NLS-Cre version of the pX330 vector was generated. In brief, this vector contained an additional expression cassette under the control of the mouse PGK promoter (driving expression of the puromycin-resistance gene), a T2A self-cleaving peptide flanked by flexible GSG linkers and a P1 bacteriophage Cre recombinase engineered with an N-terminal nuclear localization site and an HSVpA signal. This vector was further modified to express both mouse Pten and Trp53 U6 promoter-driven gRNAs following the process used to build pX330-p53-Pten (see above).
For the pT2-OVA-P2A-Myc plasmid, a 660-base pair (bp) fragment of the cytosolic region (amino acids 173–386, containing both major histocompatibility class I- and II-restricted epitopes) of OVA cDNA was amplified by PCR and engineered with a Kozak consensus methionine and EcoRI and NheI cloning sites using the following primers: OVA_ERI_U1 and OVA_Nhe1_L1 (Supplementary Table 1). A P2A self-cleaving 2A peptide cassette, flanked by a flexible GSG linker region, was then added in-frame to the 3′ end of the OVA region using NheI and XhoI restriction sites. Finally, the mouse Myc cDNA was subcloned in-frame 3′ of the P2A cassette using PCR primers Cmyc_XhoI_LE_U1 and Cmyc_BstB1_L1 (Supplementary Table 1). This primer set removed the mouse c-Myc start methionine and engineered an additional BstBI site at the 3′ end of mouse Myc that allowed subcloning of the entire Kozak OVA-P2A-Myc cassette into the pT2/BH-CAG-GS-Myc plasmid using EcoRI and BstBI restriction enzyme cloning sites.
For the pT2-EF1a-rtTA-P2A-Myc+TRE-OVA plasmid, the entire ~1,700-bp CAG-CMV promoter and enhancer region in the pT2-OVA-P2A-Myc plasmid was removed and replaced with the 289-bp human elongation factor 1-α (EF1a) core promoter to create the pT2-EF1a-OVA-P2A-Myc plasmid. The pT2-EF1a-OVA-P2A-Myc plasmid was then digested with EcoRI and NheI to remove the chicken OVA region and replace it with a reverse tetracycline-controlled transactivator (rtTA) coding sequence (rtTA-advanced) to create the pT2-EF1a-rtTA-P2A-Myc plasmid. TRE tight Promoter from pTRE-Tight caspase-3 (p12)::nz [TU#817] (a gift from M. Chalfie (Columbia University), Addgene plasmid 16084) was subcloned upstream of the BGH 3′ untranslated region and poly(A) signal of pcDNA3.1-Zeo (Invitrogen) via XhoI and EcoRI restriction sites to generate pcDNA3.1(–)Zeo-TRE-BGHpA. Next, OVA cDNA was subcloned into pcDNA3.1(–)Zeo-TRE-BGHpA using EcoRI and HindIII restriction sites to generate pcDNA3.1(–)Zeo-TRE-OVA-BGHpA. The entire 1,256-bp TRE-OVA-BGHpA fragment was PCR amplified using primers BGHpA_IF_Fwd and TRE_IF_Rev (Supplementary Table 1). The amplified fragment was then cloned in the reverse ‘trans’ orientation into HindIII-linearized pT2-EF1a-rtTA-P2A-Myc plasmid using In-Fusion (Takara Bio), generating the final pT2-EF1a-rtTA-P2A-Myc+TRE-OVA plasmid.
To induce Chrm3/Chrm5-knockout HCC, the guide oligonucleotides were designed to express mouse Chrm3 sgRNA and Chrm5 sgRNA (Supplementary Table 1). The annealed double-stranded guide oligonucleotides were cloned into the BbsI cut sites of a modified version of the pX330 vector from which the Cas9 cassette had been removed. The Chrm3 and Chrm5 sgRNA expression vectors were combined with pX330-p53-Pten, SB100X and pT2-Myc for HCC induction; in this way, the Chrm3 and Chrm5 sgRNAs only targeted the cells receiving the pX330-p53-Pten vector.
Hydrodynamic injections to induce HCC
For delivery of transposon and CRISPR vectors, mice (8–20 weeks old) were injected with a volume of 100 ml per kg (body weight) containing 25 μg of pX330-p53-Pten (or its modified form) plus 0.66 μg of SB100X and 5 μg of pT2-Myc (or its modified forms). The molar ratio of SB100X to pT2-Myc was 1 to 5. Hydrodynamic injection into the lateral tail vein took 5–7 s. Blinding was achieved during injection by putting littermates of different genotypes into new cages lacking mouse information.
To evaluate and quantify HCC development, mice were monitored daily for the appearance of palpable tumors, which were defined as a discernable enlargement of the abdomen. The humane endpoint was reached when one of the following criteria was met: a 10% increase in body weight from baseline, an estimated liver weight of over 5 g determined by the degree of abdominal distension, moribund condition or persistent facial displays of pain and distress. For the survival curve, the exact dates when mice reached the humane endpoint were determined after killing mice bearing significant HCC. Livers collected at this stage were 3–7 g in weight, and the exact endpoints (liver weight over 5 g) were adjusted by 1 d per g (live weight). Early mortalities (<5 d) were considered to be due to injection-associated death and were removed from the analysis. The number of liver tumors was quantified by counting tumor nodules on the surface of the liver or by normalizing the number of tumor clones to the area of liver sections. Blinding was performed in quantification of liver sections but not surface tumor nodules.
Antibodies and treatments
Antibodies used to deplete mice of NK cells, CD4+ T cells, CD8+ T cells or CD25+ Treg cells or antibodies used for PD-1 blockade (and isotype control antibodies) were from BioXCell. Briefly, anti-NK1.1 (clone PK136) and anti-CD4 (clone GK1.5) were injected intraperitoneally (i.p.) at 100 μg per mouse on days −1, 2, 7, 12 and 16 of standard HCC induction. Anti-CD8 (clone 2.43) was injected i.p. at 100 μg per mouse on days −1, 2, 7, 12, 16 and 21 of standard HCC induction. Anti-CD25 (clone PC-61.5.3) was injected i.p. at 150 μg per mouse on days 1, 6, 11 and 17 of standard HCC induction. To analyze the efficiency of Treg cell depletion, Foxp3-YFP mice were injected at the same time, and anti-mouse CD25–Alexa Fluor 647 (clone 7D4, BD) was used for flow cytometry. Anti-PD-1 (clone RMP1-14) was injected i.p. at 100 μg per mouse on days 3, 7, 11 and 15 of standard HCC induction.
To induce NASH-derived HCC, male mice (4–6 weeks old) were fed a Western diet (Research Diets, D16022301i) for 15 months. The diet contained 40 kcal% fat, 20 kcal% fructose and 2% cholesterol.
For OVA immunization, mice were injected into the base of their tails with 50 μl of OVA/complete Freund’s adjuvant emulsion (Hooke Laboratories, EK-0301).
To induce OVA expression in HCC, HCC was induced in mice by injection of pT2-EF1a-rtTA-P2A-Myc+TRE-OVA. When mice showed a discernable enlargement of the abdomen, they received Dox-containing drinking water (600 mg liter–1; Sigma) for 15 d, followed by euthanasia and fluorescence-activated cell sorting analysis of liver cells isolated as described below.
Hepatic MNC isolation
Mice were killed by CO2 asphyxiation and immediately subjected to whole-body perfusion with ice-cold PBS containing 10 mM EDTA. Liver tissues were collected, disrupted and passed through 70-μm sieves to obtain single-cell suspensions. MNCs were enriched by centrifugation through a 40%/80% Percoll gradient for 20 min at 2,000 r.p.m.
Flow cytometry
Antibodies included anti-mouse CD4–BUV737 (612843), anti-mouse CD8–PerCP-Cy5.5 (551162), anti-mouse CD19–BUV395 (563557), anti-mouse CD25–Alexa Fluor 647 (clone 7D4, 563598) and anti-mouse CD62L–BUV737 (612833) from BD; anti-mouse CD45.2–Alexa Fluor 700 (109822), anti-mouse NK1.1–BV605 (108740), anti-mouse CD11b–BV510 (101263), anti-mouse CD44–Alexa Fluor 700 (103026), anti-mouse OX40–PE (119409), anti-mouse CD4–BV510 (100559), anti-mouse CD45–PerCP-Cy5.5 (103132), TCR Vβ5.1/Vβ5.2–PE-Cy7 (139508), anti-mouse CD25–PE-Cy7 (clone PC-61, 102016), anti-mouse CD4–PE (100408), anti-mouse CD62L–FITC (104406), anti-mouse CD44–APC (103012), anti-mouse CD8–APC-Cy7 (100714), anti-mouse PD-1–APC (109112), anti-mouse Tim-3–PE (119704), anti-mouse Lag-3–PerCP-Cy5.5 (125219), anti-mouse PD-L1–PE-Cy7 (124314), anti-mouse CTLA-4–PE (106306), anti-mouse IFNγ–APC (505810) and anti-mouse IL-17A–BV605 (506927) from BioLegend and anti-mouse FOXP3–PE (clone FJK-16S, 12-5773-82) from Thermo Fisher. These antibodies were used at a 1:200 dilution. Mouse CD1d–PBS-57 BV421-labeled tetramer was from the NIH Tetramer Facility. We determined the appropriate concentration of the CD1d tetramer by conducting a pilot experiment on each lot, and either a 1:1,000 or 1:400 dilution was used.
For intracellular cytokine staining, MNCs were stimulated for 4 h with 1 mg ml–1 ionomycin plus 25 μg ml–1 phorbol myristate acetate in the presence of BD GolgiPlug. Cytokine staining was performed with Cytofix/Cytoperm kits (BD) following the manufacturer’s instructions. For staining of transcription factors, the eBioscience Foxp3/Transcription Factor Staining Buffer Set was used following the manufacturer’s instructions. Anti-GFP Alexa Fluor 488 (Thermo Fisher, A21311) were used to label GFP in intracellular staining analyses.
Flow cytometric analyses were performed using BD LSRFortessa cell analyzers at the Princess Margaret Flow Facility.
scRNA-seq and data analysis
HCC was induced in Chat–GFP mice using pX330-p53-Pten plus SB100X and pT2-Myc. HCC livers (from two male and two female mice) and control livers (from sex-matched littermates) were collected for hepatic MNC isolation on day 26 of HCC induction. Hepatic MNCs were stained with antibodies to CD4, CD8, CD19, TCRβ, NK1.1, CD45 and CD1d tetramer as well as with barcode antibodies (TotalSeq-C0304, C0305, C0306 or C0307) to hashtag cells from individual mice of the HCC or control group. CD45+DAPI–NK1.1–CD1dTetramer–CD19–TCRβ+CD4+CD8–GFP+ (ChAT–GFP+CD4+ T cells) and CD45+DAPI–NK1.1–CD1dTetramer–CD19−TCRβ+CD4+CD8–GFP–(ChAT–GFP–CD4+ T cells) populations were sorted on a FACSAria Fusion cell sorter (BD) at the Princess Margaret Flow Facility, with the two CD4+ T cell compartments individually sorted from each mouse. The same CD4+ T cell compartments from mice receiving the same treatment were pooled into four samples: control ChAT–GFP+, control ChAT–GFP–, HCC ChAT–GFP+ and HCC ChAT–GFP–. After sorting and pooling, the samples were immediately submitted to the Princess Margaret Genomics Centre for downstream processing. The four samples were loaded onto a 10x Chromium Controller, and libraries were prepared using a Chromium Next GEM Single Cell 5′ HT Reagent kit v2 (Dual Index, 10x Genomics). The libraries were sequenced on an Illumina NovaSeq 6000 instrument. The sequencing depths were GEX ~50,000 reads per cell, TCR ~5,000 reads per cell and cell hashing TotalSeq C ~2,000 reads per cell.
For scRNA-seq data analysis, sequencing data were processed using Cell Ranger (version 7.0.0) and aligned to the annotated mouse genome (mm10). The Cell Ranger VDJ pipeline was used to call TCR sequences. The clonotype analysis was performed on the merged contig annotations of four samples. The junctions of the V, D and J segments were determined with the IMGT database. The filtered feature barcode matrices in the hierarchical data format (.h5 files) and .csv files of filtered contig annotations from the four samples were analyzed with Partek Flow software (version 10.0.23.0214, license from the Centre for PanorOmic Sciences Bioinformatics Core) and analyzed together. With the ‘split by feature type’ tool, the single-cell counts were split into two data nodes: gene expression and antibody capture. After excluding low-quality cells (counts of <30,000; percentage of mitochondrial counts of <30), gene expression was normalized using the recommended counts per million method. Antibody capture was normalized with the recommended method (add 1.0, divide by geometric mean, add 1.0 and log 2.0), and the multiplets and cells with ambiguous hashing were filtered out. These two sets of data were then merged with the ‘merge matrices’ tool to obtain the filtered, uniquely hashtagged and normalized counts (15,703 cells in total). This counts data node was resplit to generate new gene expression and antibody capture nodes. Dimensionality reduction and visualization were performed on the new gene expression data node using ‘PCA’ (number of principal components: 100; features contribute: by variance; split by sample: no), ‘graph-based clusters’ (with default parameters except the resolution was set to 1.0) and ‘UMAP’ (with default parameters) tools. ‘Compute biomarkers’ was performed on graph-based clustering results to identify marker genes of each cell cluster. Differential analyses were performed using ‘GSA’ and visualized using heat maps and volcano plots.
Immunohistological analyses
Sections cut from formalin-fixed, paraffin-embedded blocks of mouse livers were used for immunohistochemistry (IHC). After dewaxing and rehydration, endogenous peroxidase was deactivated in 3% hydrogen peroxide (20 ml of 30% hydrogen peroxide + 180 ml of PBS) for 15 min at room temperature. Antigen retrieval with 10 mM sodium citrate buffer (pH 6.0) was performed before immunostaining.
Primary antibodies used for IHC included goat anti-GFP (Novus, NB100-1678), rat anti-Foxp3 (Thermo Fisher, clone FJK-16s), rabbit anti-CD3 (Abcam, ab5690), rabbit anti-CD11b (Abcam, ab133357), rabbit anti-p53 (Vector Labs, VP-P956), rabbit anti-c-MYC (Cell Signaling, 5605) and rabbit anti-PTEN (Cell Signaling, 9559). Polymer-conjugated secondary antibodies included alkaline phosphatase (AP) goat anti-rat IgG (MP-544415), horseradish peroxidase (HRP) goat anti-rat IgG (MP-5444), HRP horse anti-goat IgG (MP-7405), HRP horse anti-rabbit IgG (MP-7405) and AP horse anti-rabbit IgG (MP-5401; all from Vector Laboratories). ImmPACT Vector Red Substrate kits, including AP substrate (SK-5105) and DAB peroxidase substrate (SK-4100; both from Vector Laboratories), were used for chromogenic detection.
Immunostained histological sections were scanned using a NanoZoomer 2.0-HT slide scanner from Hamamatsu. Quantification of immune cell clusters in scans of H&E-stained liver sections and determination of c-MYC, p53 and PTEN expression in tumor clones present in scans of IHC-stained liver sections were performed using NDP.view2 (Hamamatsu) in a blinded fashion. For analysis of c-MYC expression in liver sections, random 20× non-tumor fields of the c-MYC IHC scans from liver sections of individual mice were output using NDP.view2 and blindly analyzed using ImageJ (https://imagej.nih.gov/ij/) with the plugin ‘IHC Profiler’62.
For the immunofluorescence staining shown in Extended Data Fig. 1f, Alexa Fluor 568-conjugated goat anti-rabbit IgG (Thermo Fisher, A-11036) was used.
For detection of fluorescent proteins in livers of Rosa26Confetti mice, frozen liver sections were fixed with 2% paraformaldehyde for 8 min at room temperature and directly observed using an Olympus FluoView FV1000 confocal laser-scanning microscope.
Measurement of Ca2+ flux
CD4+ T cells were purified from spleens of Chatfl/fl and Chatfl/fl; Cd4-cre mice using a CD4+ T Cell Isolation kit (Miltenyi Biotec) and autoMACS following the manufacturer’s instructions. Purified CD4+ T cells were loaded with 1 μM Indo-1 Ca2+ indicator (Thermo Fisher) in a 37 °C water bath for 45 min. After washing, CD4+ T cells were stained with anti-CD4–PE, anti-CD62L–FITC and anti-CD44–APC at 4 °C for 30 min and stained with 1 μg ml–1 hamster anti-CD3 (Biolegend, 100359, clone 145-2C11) at 4 °C for 30 min. To activate TCR signaling, rabbit anti-hamster (Jackson ImmunoResearch, 307-005-003) was added at 10 μg ml–1 at the indicated time during flow cytometric analysis. CD4+CD62L+CD44– naive CD4+ T cells were gated for comparison.
Analysis of NFAT nuclear translocation
CD4+ T cells were purified from spleens of Chatfl/fl and Chatfl/fl; Cd4-cre mice using a CD4+ T Cell Isolation kit (Miltenyi Biotec) and autoMACS following the manufacturer’s instructions. Purified CD4+ T cells were resuspended in serum-free RPMI medium at 5 × 105 cells per ml. These CD4+ T cell suspensions were incubated with ACh (Sigma), nicotine (Tocris Bioscience) or Oxo-M (Tocris Bioscience) at 37 °C for 15 min. Anti-CD3/CD28 microbeads (Thermo Fisher) were then added to the treated T cell suspensions at a 1:1 bead:cell ratio, and the cells were cultured for another 15 min. After removing the anti-CD3/CD28 microbeads with a magnetic block, CD4+ T cells were fixed and prepared for cytospins with a cytocentrifuge (Thermo Fisher). The cytospins were permeabilized with 1% Triton X-100 for 30 min, blocked with 3% fetal bovine serum, 1% bovine serum albumin and 0.3% Triton X-100 in PBS for 30 min and stained with Alexa Fluor 488-conjugated anti-NFAT1 diluted 1:50 (clone D43B1, Cell Signaling) overnight at 4 °C. After washing and counterstaining with DAPI, the cytospins were mounted with cover slides and examined with a Nikon A1R confocal microscope. Fluorescence micrograms were acquired using a ×40 objective lens. Consecutive images across the diameter of each cytospin (about 16 shots) were collected and quantified. CellProfiler (v4.2.1) was used for the quantification of NFAT nuclear translocation. The minimum cross-entropy thresholding method was used to identify the objects. The MeasureObjectIntensityDistribution module was used for the analysis of NFAT distribution. Briefly, the radial distribution of NFAT staining was measured with three concentric rings (bins) starting from the center of DAPI staining. The ratio of the fraction of total NFAT intensity in the first bin (innermost ring) versus the third bin (outermost ring) was designated as the parameter for NFAT nuclear translocation. The CellProfiler project file will be shared upon inquiry.
Analysis of human HCC datasets
Expression levels of CHAT in various clusters of T cells isolated from tumor tissue and adjacent liver tissue of individuals with HCC were determined by examining the scRNA-seq dataset of immune cells of human HCC previously published by Zhang et al.32. Violin plots were generated using their interactive web-based tool (http://cancer-pku.cn:3838/HCC/). Expression levels of CHAT, mAChRs and nAChRs (α-subunits) in T cells isolated from samples from individuals with HCC were determined by extracting data from a published scRNA-seq dataset (GSE98638) on T cells from individuals with HCC34. Survival curves of individuals with HCC with high expression of CHRM3 (cutoff was set to log2 (fragments per kilobase of transcript per million mapped reads upper quartile (FPKM-UQ) + 1) = 11) and CHRM5 (cutoff was set to log2 (FPKM-UQ + 1) = 9.6) were generated using data obtained from the TCGA Research Network (https://www.cancer.gov/tcga) and analyzed with University of California, Santa Cruz, Xena (http://xena.ucsc.edu/).
Statistics and reproducibility
Pilot experiments were used to estimate the sample size necessary to generate statistically significant results using the appropriate statistical tests. Genetically modified mice and their littermate controls were used for all experiments where possible. For comparing liver weights and nodule numbers, five to ten mice per group were sufficient to achieve statistical significance. For survival curves, a cohort of 10–20 mice per group was used. To account for potential technical failures, including missed hydrodynamic injection and early mortality associated with injection, we usually included an extra 10% of mice per group. Early mortalities (<5 d) were considered to be due to injection-associated death and were removed from the analysis. The numbers of replicates and independent experiments have been stated in the figure legends. The attempts at replication were successful.
To generate statistically appropriate numbers, it was usually necessary to use more than three litters of mice for each experiment. To control for the treatments (including plasmids and antibodies), mice from each litter were randomly divided into groups to guarantee that sex-matched and genotype-matched individuals obtained different treatments. This grouping was performed ahead of each experiment. For vector delivery by hydrodynamic injection, blinding was achieved during injection by placing littermates of different genotypes into new cages lacking mouse information. Blinding was also performed for quantitative analyses of liver sections.
Statistical analyses
The Kolmogorov–Smirnov test was used to evaluate normality. Pair-wise comparisons were assessed using two-tailed unpaired Student’s t-tests unless otherwise denoted in the figure legends. Data are presented as mean ± s.e.m. unless otherwise indicated. GraphPad Prism 8 and Microsoft Excel (version 2019) were used for the statistical analyses. P values of <0.05 were considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
scRNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession code GSE231322. The scRNA-seq datasets of human HCC analyzed in this study include those published by Zheng et al.34 and Zhang et al.32. The accession code of the Zheng et al. data is GSE98638. The accession codes of the Zhang et al. data are GSE140228 and EGAS00001003449.
The human liver HCC (TCGA-LIHC) data analyzed in this study were derived from the TCGA Research Network (http://cancergenome.nih.gov/).
All other data supporting the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
Details of software versions, plugins, modules and parameters are specified in relevant sections in the Methods. The code for Circos plots showing the distribution of TCR types among GFP+ and GFP– T cells from control and HCC-bearing mice has been deposited in a Zenodo repository (https://zenodo.org/record/8058337)63.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Mittal, S. & El-Serag, H. B. Epidemiology of hepatocellular carcinoma: consider the population. J. Clin. Gastroenterol. 47, S2–S6 (2013).
Yu, L.-X., Ling, Y. & Wang, H.-Y. Role of nonresolving inflammation in hepatocellular carcinoma development and progression. NPJ Precis. Oncol. 2, 6 (2018).
Cancer Genome Atlas Research Network. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell 169, 1327–1341 (2017).
Kang, T.-W. et al. Senescence surveillance of pre-malignant hepatocytes limits liver cancer development. Nature 479, 547–551 (2011).
Xue, W. et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 445, 656–660 (2007).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Pfister, D. et al. NASH limits anti-tumour surveillance in immunotherapy-treated HCC. Nature 592, 450–456 (2021).
Zamarron, B. F. & Chen, W. Dual roles of immune cells and their factors in cancer development and progression. Int. J. Biol. Sci. 7, 651–658 (2011).
Zhao, C.-M. et al. Denervation suppresses gastric tumorigenesis. Sci. Transl. Med. 6, 250ra115 (2014).
Magnon, C. et al. Autonomic nerve development contributes to prostate cancer progression. Science 341, 1236361 (2013).
Saloman, J. L. et al. Ablation of sensory neurons in a genetic model of pancreatic ductal adenocarcinoma slows initiation and progression of cancer. Proc. Natl Acad. Sci. USA 113, 3078–3083 (2016).
Schuller, H. M. Is cancer triggered by altered signalling of nicotinic acetylcholine receptors? Nat. Rev. Cancer 9, 195–205 (2009).
Yi, C.-X., la Fleur, S. E., Fliers, E. & Kalsbeek, A. The role of the autonomic nervous liver innervation in the control of energy metabolism. Biochim. Biophys. Acta 1802, 416–431 (2010).
Berthoud, H., Kressel, M. & Neuhuber, W. L. An anterograde tracing study of the vagal innervation of rat liver, portal vein and biliary system. Anat. Embryol. 186, 431–442 (1992).
Akiyoshi, H., Gonda, T. & Terada, T. A comparative histochemical and immunohistochemical study of aminergic, cholinergic and peptidergic innervation in rat, hamster, guinea pig, dog and human livers. Liver 18, 352–359 (1998).
Liu, K. et al. Metabolic stress drives sympathetic neuropathy within the liver. Cell Metab. 33, 666–675 (2021).
Zucman-Rossi, J., Villanueva, A., Nault, J.-C. & Llovet, J. M. Genetic landscape and biomarkers of hepatocellular carcinoma. Gastroenterology 149, 1226–1239 (2015).
Hu, T.-H. et al. Expression and prognostic role of tumor suppressor gene PTEN/MMAC1/TEP1 in hepatocellular carcinoma. Cancer 97, 1929–1940 (2003).
Horie, Y. et al. Hepatocyte-specific Pten deficiency results in steatohepatitis and hepatocellular carcinomas. J. Clin. Invest. 113, 1774–1783 (2004).
Kaposi-Novak, P. et al. Central role of c-Myc during malignant conversion in human hepatocarcinogenesis. Cancer Res. 69, 2775–2782 (2009).
Xue, W. et al. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature 514, 380–384 (2014).
Chow, E. K.-H., Fan, L., Chen, X. & Bishop, J. M. Oncogene-specific formation of chemoresistant murine hepatic cancer stem cells. Hepatology 56, 1331–1341 (2012).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
McGranahan, N. & Swanton, C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168, 613–628 (2017).
Rosas-Ballina, M. et al. Acetylcholine-synthesizing T cells relay neural signals in a vagus nerve circuit. Science 334, 98–101 (2011).
Cox, M. A. et al. Choline acetyltransferase-expressing T cells are required to control chronic viral infection. Science 363, 639–644 (2019).
Nechanitzky, R. et al. Cholinergic control of TH17 cell pathogenicity in experimental autoimmune encephalomyelitis. Cell Death Differ. 30, 407–416 (2023).
Linterman, M. A. et al. Foxp3+ follicular regulatory T cells control T follicular helper cells and the germinal center response. Nat. Med. 17, 975–982 (2011).
Śledzińska, A. et al. Regulatory T cells restrain interleukin-2- and blimp-1-dependent acquisition of cytotoxic function by CD4+ T cells. Immunity 52, 151–166 (2020).
Dudek, M. et al. Auto-aggressive CXCR6+ CD8 T cells cause liver immune pathology in NASH. Nature 592, 444–449 (2021).
Zhang, Q. et al. Landscape and dynamics of single immune cells in hepatocellular carcinoma. Cell 179, 829–845 (2019).
Kobayashi, N. et al. FOXP3+ regulatory T cells affect the development and progression of hepatocarcinogenesis. Clin. Cancer Res. 13, 902–911 (2007).
Zheng, C. et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169, 1342–1356 (2017).
Kreiter, S. et al. Mutant MHC class II epitopes drive therapeutic immune responses to cancer. Nature 520, 692–696 (2015).
Wolf, M. J. et al. Metabolic activation of intrahepatic CD8+ T cells and NKT cells causes nonalcoholic steatohepatitis and liver cancer via cross-talk with hepatocytes. Cancer Cell 26, 549–564 (2014).
Castro, F., Cardoso, A. P., Gonçalves, R. M., Serre, K. & Oliveira, M. J. Interferon-γ at the crossroads of tumor immune surveillance or evasion. Front. Immunol. 9, 847 (2018).
Mikulak, J., Bruni, E., Oriolo, F., Di Vito, C. & Mavilio, D. Hepatic natural killer cells: organ-specific sentinels of liver immune homeostasis and physiopathology. Front. Immunol. 10, 946 (2019).
Cheng, M., Chen, Y., Xiao, W., Sun, R. & Tian, Z. NK cell-based immunotherapy for malignant diseases. Cell. Mol. Immunol. 10, 230–252 (2013).
Kim, K. D. et al. Adaptive immune cells temper initial innate responses. Nat. Med. 13, 1248–1252 (2007).
Shimizu, J., Yamazaki, S. & Sakaguchi, S. Induction of tumor immunity by removing CD25+CD4+ T cells: a common basis between tumor immunity and autoimmunity. J. Immunol. 163, 5211–5218 (1999).
Hermann-Kleiter, N. & Baier, G. NFAT pulls the strings during CD4+ T helper cell effector functions. Blood 115, 2989–2997 (2010).
Martinez, G. J. et al. The transcription factor NFAT promotes exhaustion of activated CD8+ T cells. Immunity 42, 265–278 (2015).
Chen, J. et al. NR4A transcription factors limit CAR T cell function in solid tumours. Nature 567, 530–534 (2019).
Liu, X. et al. Genome-wide analysis identifies NR4A1 as a key mediator of T cell dysfunction. Nature 567, 525–529 (2019).
Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8+ T cell exhaustion. Proc. Natl Acad. Sci. USA 116, 12410–12415 (2019).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Kandel, E. R., Schwartz, J. H., Jessell, T. M., Siegelbaum, S. A. & Hudspeth, A. J. Principles of Neural Science 5th edn (McGraw-Hill Education, 2013).
Wess, J., Eglen, R. M. & Gautam, D. Muscarinic acetylcholine receptors: mutant mice provide new insights for drug development. Nat. Rev. Drug Discov. 6, 721–733 (2007).
Feske, S., Skolnik, E. Y. & Prakriya, M. Ion channels and transporters in lymphocyte function and immunity. Nat. Rev. Immunol. 12, 532–547 (2012).
Mashimo, M., Yurie, Y., Kawashima, K. & Fujii, T. CRAC channels are required for [Ca2+]i oscillations and c-FOS gene expression after muscarinic acetylcholine receptor activation in leukemic T cells. Life Sci. 161, 45–50 (2016).
Premack, B. A., McDonald, T. V. & Gardner, P. Activation of Ca2+ current in Jurkat T cells following the depletion of Ca2+ stores by microsomal Ca2+-ATPase inhibitors. J. Immunol. 152, 5226–5240 (1994).
Tiegs, G. & Lohse, A. W. Immune tolerance: what is unique about the liver. J. Autoimmun. 34, 1–6 (2010).
Fu, J. et al. Increased regulatory T cells correlate with CD8 T-cell impairment and poor survival in hepatocellular carcinoma patients. Gastroenterology 132, 2328–2339 (2007).
Schietinger, A. et al. Tumor-specific T cell dysfunction is a dynamic antigen-driven differentiation program initiated early during tumorigenesis. Immunity 45, 389–401 (2016).
Muallem, S., Pandol, S. J. & Beeker, T. G. Hormone-evoked calcium release from intracellular stores is a quantal process. J. Biol. Chem. 264, 205–212 (1989).
Yamashita, M. ‘Quantal’ Ca2+ release reassessed—a clue to oscillation and synchronization. FEBS Lett. 580, 4979–4983 (2006).
Rossi, A. M. et al. Quantal Ca2+ release mediated by very few IP3 receptors that rapidly inactivate allows graded responses to IP3. Cell Rep. 37, 109932 (2021).
Fujii, T. et al. Expression and function of the cholinergic system in immune cells. Front. Immunol. 8, 1085 (2017).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Varghese, F., Bukhari, A. B., Malhotra, R. & De, A. IHC Profiler: an open source plugin for the quantitative evaluation and automated scoring of immunohistochemistry images of human tissue samples. PLoS ONE 9, e96801 (2014).
Yin, T. Code for Circos plots showing the distribution of TCR types. Zenodo https://doi.org/10.5281/zenodo.8058337 (2023).
Acknowledgements
We express our gratitude to the Princess Margaret Flow Facility and the Advanced Optical Microscopy Facility at University Health Network for their exceptional training and services. We extend our thanks to S. J. Han for technical advice and to the Animal Resource Centre at the Princess Margaret Cancer Centre for animal care. We are also grateful to G. Basi, A. Daghrach, T. Ketela and Z. Lu from the Princess Margaret Genomics Centre for providing scRNA-seq services. We would like to acknowledge S. Fang from the Centre for PanorOmic Sciences in the University of Hong Kong for her training in Partek Flow. Lastly, our appreciation goes to P. Ramachandran, Z. Zheng and L. Hendrikse for their input on the scRNA-seq experiment.
This work was supported by grants from the Canadian Institutes of Health Research (to T.W.M.), including grant 143268 and grant 488053, the Centre for Oncology and Immunology under the Health@InnoHK Initiative funded by the Innovation and Technology Commission, the Government of Hong Kong SAR, China, and Cancer Research Institute Irvington Postdoctoral Fellowships (to C.Z. and S.L.).
Author information
Authors and Affiliations
Contributions
T.W.M. and C.Z. conceptualized the study. C.Z. designed the experiments and wrote the draft of the manuscript. C.Z., B.E.S., A.J.E., S.L., J.H., C.T., K.H. and K.T.G. performed experiments and/or data analysis. R.N., C.D.-B., S.L., M.A.C., A.C.W., W.M., J.F., T.B., E.F. and M.H. provided critical reagents and/or insights. C.Z., Y.T. and S.Y.L. analyzed the scRNA-seq data. J.F. and M.E.S. edited the manuscript. T.W.M. supervised the research.
Corresponding author
Ethics declarations
Competing interests
T.M. owns equity in Treadwell Therapeutics, Inc., and Agios Pharmaceuticals and is a consultant for AstraZeneca. All other authors have no competing interests.
Peer review
Peer review information
Nature Cancer thanks Hiroyoshi Nishikawa, Lars Zender and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The immunosurveillance of murine liver cancer.
a Schematic diagrams of the plasmids used to simultaneously induce CRISPR/Cas9-mediated deletion of Trp53 and Pten, Cre expression, and Myc overexpression in Rosa26Confetti/+ reporter mice. b Representative fluorescence microscopy images depicting (left) a monoclonal and (right) a polyclonal tumor from the livers of the mice in (a). Clonality was determined based on the expression of one or multiple fluorescent markers encoded by the Rosa26Confetti reporter, representative of one experiment. c Quantification of monoclonal and polyclonal tumors expressing the indicated fluorescent markers in livers from the mice in (a). Numbers of tumor nodules expressing the indicated fluorescent marker(s) are labeled in the pie plot. Data are from examination of 113 tumor nodules in liver sections from 10 Rosa26Confetti reporter mice. d Gating strategy used for flow cytometric analysis of murine hepatic immune cells. e Percentage of OX40+ CD4+ T cells in livers on the indicated days of liver cancer development. Each dot represents an individual mouse (n=6 mice per group). Data are the mean ± SEM. P values were determined by unpaired, two-tailed t-test. f Representative immunofluorescence images showing CD3 and CD11b expression in adjacent sections of an HCC-bearing liver, representative of one experiment. g Representative macroscopic images (upper) and microscopic images (lower) of tumor-bearing livers resected at the humane endpoint from NSG and control mice subjected to HCC induction, representative of one experiment. Related to Fig. 1.
Extended Data Fig. 2 Identification of cholinergic cells in liver cancer.
a Representative histological sections showing GFP immunoreactivity in (left) small intestine, (middle) normal liver, and (right) HCC-bearing liver resected from Chat-GFP mice. Arrowheads, cells with GFP immunoreactivity. Inset boxes show higher magnification views of the GFP+ cells in the small boxes. Scale bars, 100 μm. T, tumor tissue. NT, non-tumor tissue. The micrographs displayed are representative of sections obtained from three independent experiments. b Percentage of GFP+ cells among the indicated cell subsets in livers of Chat-GFP mice at the indicated time points following HCC induction. Each dot represents an individual mouse (n=6 mice per group). Data are the mean ± SEM. n.s., not significant by unpaired, two-tailed t-test was performed. Results are representative of three independent experiments. c qPCR determinations of Chat mRNA levels (relative to Actb) in hepatic mononuclear cells of Chat-GFP mice at the indicated time points following standard HCC induction. Each dot represents an individual mouse. P values were determined by unpaired, two-tailed t-test. d Expression of Foxp3 in CD4+ T cells from control and HCC-bearing livers as determined in the scRNAseq UMAP plot shown in Fig. 2c. e Volcano plot comparing transcripts between Foxp3 mRNA-expressing cells from cluster 4 and those from cluster 9. Gene-set analysis (GSA) was performed to identify sets of differentially expressed genes. f Percentages of Foxp3 mRNA+ cells among Chat mRNA+ CD4+ T cells and Chat mRNA- CD4+ T cells from HCC-bearing livers as determined from scRNAseq data. Cells from each mouse were identified using antibody barcodes. Each dot represents an individual mouse (n=4 mice per group). Data are the mean ± SEM. P values were determined by unpaired, two-tailed t-test. g Representative flow cytometry plots showing the expression of PD-L1 by CD4+ T cells, CD45+CD3- immune cells and CD45- cells from HCC-bearing liver. Related to Fig. 2.
Extended Data Fig. 3 Identification of CHAT-expressing T cells in human HCC.
Extended Data Fig. 4 Expression of Chat by clonally expanded PD-1+ Tconvs and Foxp3+ Tregs in HCC.
a UMAP representation of Chat-GFP+ and Chat-GFP- CD4+ T cells bearing TCR #1 induced in HCC, color-coded by cell clusters and shape-coded by mouse ID. b UMAP representation of Chat-GFP+ and Chat-GFP- CD4+ T cells bearing TCR #22 induced in HCC, color-coded by expression level of Foxp3 and shape-coded by mouse ID. c UMAP representation of Chat-GFP+ and Chat-GFP- CD4+ T cells bearing TCR #3 induced in HCC, color-coded by cell clusters (upper) and expression levels of Cxcr6, Pdcd1, and Foxp3 (lower), and shape-coded by mouse ID. Related to Fig. 4.
Extended Data Fig. 5 Induction of OVA specific Chat-GFP+ Tregs and PD-1+ Tconvs in HCC with inducible OVA expression.
a Representative flow cytometry plots showing the expression of Foxp3 and PD-1 in OVA-specific Chat-GFP+CD4+ T cells in livers of mice that were left untreated or treated with Dox-containing drinking water to induce OVA expression following HCC onset, representing two independent experiments. Related to Fig. 4.
Extended Data Fig. 6 Analysis of the efficiency of vector delivery and T cell activities in HCC.
a,b Representative fluorescence microscopy images (a), and quantification of the proportion of hepatocytes positive for the Rosa26Confetti RFP marker (b), in liver sections from Rosa26Confetti/+;Chatfl/fl and Rosa26Confetti/+;Chatfl/fl;CD4-Cre mice on the indicated days following plasmid injection. In (b), each dot represents a microscopy image (n=6, 11, and 9 acquired on days 1, 3, and 10 from Rosa26Confetti/+;Chatfl/fl mice; n=5, 12, and 11 acquired on days 1, 3, and 10 from Rosa26Confetti/+;Chatfl/fl;CD4-Cre mice. Images were collected from 2-4 mice/group). Data are the mean ± SEM. P values were determined by two-way ANOVA with Tukey’s multiple comparisons test. c qPCR determination of transposon-derived Myc mRNA expression (relative to Actb) in livers of Chatfl/fl and CD4-Cre;Chatfl/fl mice at the indicated time points following HCC induction. The forward primer used was within the Myc ORF, with the reverse primer in the transposon vector. Each dot represents an individual mouse (n=6 Chatfl/fl mice in Control group, and n=2, 4, 3 and 5 Chatfl/fl mice on days 1, 3, 10, and 23; n=2, 4, 4, and 5 Chatfl/fl;CD4-Cre mice on days 1, 3, 10 and 23). Data are the mean ± SEM. P values were determined by two-way ANOVA with Sidak’s multiple comparisons test. d,e Percentages of CD4+ T cells (d) and CD8+ T cells (e) among mononuclear cells (MNCs) isolated from HCC-bearing livers of Chatfl/fl and CD4-Cre;Chatfl/fl mice as assessed by flow cytometry, representative of three independent experiments. Each dot represents an individual mouse (n=11 mice in Chatfl/fl and n=12 mice in Chatfl/fl;CD4-Cre). Data are the mean ± SEM. P values were determined by unpaired, two-tailed t-test. f Percentage of IFN-γ+ CD8+ T cells in HCC-bearing livers of Chatfl/fl and Chatfl/fl;CD4-Cre mice as analyzed by flow cytometry. Each dot represents an individual mouse. Each dot represents an individual mouse (n=11 mice in each group). Data are the mean ± SEM. P values were determined by unpaired, two-tailed t-test. Related to Fig. 5.
Extended Data Fig. 7 Delineating the role of the adaptive immune response in HCC development.
a Schematic diagrams of (left) the plasmids used to induce simultaneous CRISPR/Cas9-mediated deletion of Trp53 and Pten plus overexpression of Myc and chicken ovalbumin (OVA), and (right) the experimental protocol for OVA immunization and OVA-HCC induction in mice. See main text for details. b,c Curves showing the latency to palpable HCC development (b), and survival to humane endpoint (c), of Chatfl/fl and Chatfl/fl;CD4-Cre mice that were left unimmunized, or immunized as depicted in panel (a), and subjected to OVA-HCC induction (n=14, 13, 21, and 21 mice in the Chatfl/fl, Chatfl/fl;CD4-Cre, Chatfl/fl immunized and Chatfl/fl;CD4-Cre immunized groups, respectively). P values were determined by log-rank test. d,e Percentage of CD8+ T cells among MNCs (d), and number of tumor nodules observed in histological sections (e), in HCC-bearing livers of mice treated with anti-CD8 antibodies (α-CD8) or IgG control. Each dot represents an individual mouse (n=8 mice per group). Data are the mean ± SEM. In (d), P value was determined by unpaired, two-tailed t-test. f,g Number of HCC tumor nodules (f), and weights (g) of livers from mice of the indicated genotypes on day 23 of HCC induction. Each dot represents an individual mouse (n=4, 5, 5, and 6 mice in the Rag1+/-, Rag1-/-, Chatfl/fl and Chatfl/fl;CD4-Cre groups, respectively). Data are the mean ± SEM. P values were determined by paired, two-tailed t-test. Related to Fig. 6.
Extended Data Fig. 8 Delineating the effects of Chat on Tregs in HCC.
a Percentage of Foxp3+ Tregs among CD4+ T cells in HCC-bearing livers of Chatfl/fl and CD4-Cre;Chatfl/fl mice, representative of three independent experiments. Each dot represents an individual animal (n=11 mice in Chatfl/fl and n=12 mice in Chatfl/fl;CD4-Cre). b-d Representative flow cytometry plots (b), and quantification of the percentages of CTLA-4+ cells among Tregs (c) and Tconvs (d), in HCC-bearing livers from Chatfl/fl and Chatfl/fl;CD4-Cre mice on day 16 of HCC induction. In (c) and (d), each dot represents an individual mouse (n=6 mice per group). e Quantification of the percentages of induced CD25+ cells among Foxp3+CD4+ Tregs and Foxp3-CD4+ Tconvs isolated from purified splenic CD4+CD25- T cells of Chatfl/fl and Chatfl/fl;CD4-Cre mice. CD4+CD25- T cells were enriched with negative selection microbeads and stimulated with the indicated ratios of anti-CD3/28 microbeads. CD25 expression was analyzed by flow cytometry at the indicated time points. P values were determined by unpaired, two-tailed t-test, three replicates for each condition, representative of two independent experiments. f Representative flow cytometric plots of CD25+ Foxp3+ CD4+ Tregs in blood of Foxp3-YFP mice that were treated with either anti-CD25 antibody (α-CD25) or control IgG. Data are representative of 2 mice/group analyzed on day 18 of antibody treatment. g,h Percentage of CD4+ T cells (g) and NK cells (h) among MNCs from HCC-bearing livers of mice treated with anti-CD4 antibodies (α-CD4), anti-NK1.1 antibodies (α-NK1.1) or IgG/PBS control. Each dot represents an individual mouse (n=8 mice in the IgG/PBS group; n=9 mice in α-CD4 and α-NK1.1 groups). i,j Quantification of numbers of tumor nodules/mm2 in H&E-stained sections (i), and percentages of CD25+ Tregs in HCC-bearing livers (j), of Foxp3Cre and Chatfl/fl;Foxp3Cre mice on day 22 of HCC induction. Each dot represents an individual mouse. In (i), n=8 mice in the Foxp3Cre group and n=7 mice in the Chatfl/fl;Foxp3Cre group. In (j), n=8 mice in Foxp3Cre and n=6 mice in Chatfl/fl;Foxp3Cre. In (a), (c), (d), and (g-j), data are the mean ± SEM. P values were determined by unpaired, two-tailed t-test. Related to Figs. 6 and 7.
Extended Data Fig. 9 Elucidating the effects of Chat on inhibitory immunoreceptor expression by T cells in HCC.
a-f Expression levels of the immune inhibitory receptors PD-1, Tim-3 and Lag-3 in CD4+ T cells (a-c), and in CD8+ T cells (d-f), from HCC-bearing livers of Chatfl/fl and Chatfl/fl;CD4-Cre mice on day 35 of HCC induction. Each dot represents an individual mouse (n=11 mice in Chatfl/fl and n=12 mice in Chatfl/fl;CD4-Cre). Data are the mean ± SEM. P values were determined by unpaired, two-tailed t-test. Related to Fig. 7.
Extended Data Fig. 10 Cholinergic signaling is associated with human HCC.
a Violin plots showing the expression of CHAT and the indicated muscarinic acetylcholine receptors and nicotinic acetylcholine receptors (alpha subunits) in T cells isolated from patient HCC samples (GSE98638). b-d Survival of HCC patients with high expression of CHRM3 (b), CHRM5 (c), or both CHRM3 and CHRM5 (d). Data are from the TCGA database. e,f Ratios of liver weight to body weight (e) and numbers of tumor nodules (f) of Chatfl/fl and Chatfl/fl;CD4-Cre mice on day 20 of HCC induction with standard vectors or plus vectors carrying gRNAs for Chrm3 and Chrm5. Each dot represents an individual mouse (n=7 mice per group). Data are the mean ± SEM. n.s., not significant, determined by unpaired, two-tailed t-test.
Supplementary information
Supplementary Information
Supplementary Table 1. Oligonucleotide sequences for sgRNAs and PCR amplifying fragments.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zheng, C., Snow, B.E., Elia, A.J. et al. Tumor-specific cholinergic CD4+ T lymphocytes guide immunosurveillance of hepatocellular carcinoma. Nat Cancer 4, 1437–1454 (2023). https://doi.org/10.1038/s43018-023-00624-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-023-00624-w