Abstract
Cancer-testis (CT) genes participate in the initiation and progression of cancer, but the role of CT-associated long non-coding RNAs (CT-lncRNAs) in hepatocellular carcinoma (HCC) is still elusive. Here, we discovered a conserved CT-lncRNA, named lnc-CTHCC, which was highly expressed in the testes and HCC. A lnc-CTHCC-knockout (KO) mouse model further confirmed that the global loss of lnc-CTHCC inhibited the occurrence and development of HCC. In vitro and in vivo assays also showed that lnc-CTHCC promoted HCC growth and metastasis. Mechanistically, lnc-CTHCC bound to heterogeneous nuclear ribonucleoprotein K (hnRNP K), which was recruited to the YAP1 promoter for its activation. Additionally, the N6-methyladenosine (m6A) modification was mediated by N6-adenosine-methyltransferase 70-kDa subunit (METTL3) and recognized by insulin-like growth factor 2 mRNA-binding protein 1 (IGF2BP1)/IGF2BP3, which maintained lnc-CTHCC stability and increased its expression in HCC. Together, our results show that lnc-CTHCC directly binds to hnRNP K and promotes hepatocellular carcinogenesis and progression by activating YAP1 transcription, suggesting that lnc-CTHCC is a potential biomarker and therapeutic target of HCC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNA-sequencing data have been deposited in GEO under accession codes GSE160912 and GSE160913. All other data supporting the findings of this study are available from the corresponding author upon reasonable request. GTEx is available at https://gtexportal.org/home/, and TCGA is available at http://cancergenome.nih.gov/. Source data are provided with this paper.
References
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Primers 2, 16018 (2016).
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Global Burden of Disease Cancer Collaboration et al. Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 29 cancer groups, 1990 to 2016: a systematic analysis for the global burden of disease study. JAMA Oncol. 4, 1553–1568 (2018).
Villanueva, A. Hepatocellular carcinoma. N. Engl. J. Med. 380, 1450–1462 (2019).
Simpson, A. J., Caballero, O. L., Jungbluth, A., Chen, Y. T. & Old, L. J. Cancer/testis antigens, gametogenesis and cancer. Nat. Rev. Cancer 5, 615–625 (2005).
Wang, C. et al. Systematic identification of genes with a cancer-testis expression pattern in 19 cancer types. Nat. Commun. 7, 10499 (2016).
Gibbs, Z. A. & Whitehurst, A. W. Emerging contributions of cancer/testis antigens to neoplastic behaviors. Trends Cancer 4, 701–712 (2018).
Dang, E. et al. BAP31, a newly defined cancer/testis antigen, regulates proliferation, migration, and invasion to promote cervical cancer progression. Cell Death Dis. 9, 791 (2018).
Li, Y. et al. Cancer/testis antigen-Plac1 promotes invasion and metastasis of breast cancer through Furin/NICD/PTEN signaling pathway. Mol. Oncol. 12, 1233–1248 (2018).
Wang, C. et al. A cancer-testis non-coding RNA LIN28B-AS1 activates driver gene LIN28B by interacting with IGF2BP1 in lung adenocarcinoma. Oncogene 38, 1611–1624 (2019).
Wang, S. et al. JAK2-binding long noncoding RNA promotes breast cancer brain metastasis. J. Clin. Invest. 127, 4498–4515 (2017).
Hu, Q. et al. LncRNAs-directed PTEN enzymatic switch governs epithelial–mesenchymal transition. Cell Res. 29, 286–304 (2019).
Li, Z. et al. Gain of LINC00624 enhances liver cancer progression by disrupting the HDAC6–TRIM28–ZNF354C corepressor complex. Hepatology 73, 1764–1782 (2021).
Schmitt, A. M. & Chang, H. Y. Long noncoding RNAs in cancer pathways. Cancer Cell 29, 452–463 (2016).
Liu, S. J., Dang, H. X., Lim, D. A., Feng, F. Y. & Maher, C. A. Long noncoding RNAs in cancer metastasis. Nat. Rev. Cancer 21, 446–460 (2021).
Li, S. Y. et al. LncRNA lnc-APUE is repressed by HNF4α and promotes G1/S phase transition and tumor growth by regulating miR-20b/E2F1 axis. Adv. Sci. 8, 2003094 (2021).
Hosono, Y. et al. Oncogenic role of THOR, a conserved cancer/testis long non-coding RNA. Cell 171, 1559–1572 (2017).
Sharma, A. et al. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell 183, 377–394 (2020).
Qin, G. et al. Long noncoding RNA p53-stabilizing and activating RNA promotes p53 signaling by inhibiting heterogeneous nuclear ribonucleoprotein K deSUMOylation and suppresses hepatocellular carcinoma. Hepatology 71, 112–129 (2020).
Wu, Y. et al. m6A-induced lncRNA RP11 triggers the dissemination of colorectal cancer cells via upregulation of Zeb1. Mol. Cancer 18, 87 (2019).
Wang, Q. et al. METTL3-mediated m6A modification of HDGF mRNA promotes gastric cancer progression and has prognostic significance. Gut 69, 1193–1205 (2020).
Zhu, S. et al. An oncopeptide regulates m6A recognition by the m6A reader IGF2BP1 and tumorigenesis. Nat. Commun. 11, 1685 (2020).
Huang, H., Weng, H. & Chen, J. m6A modification in coding and non-coding RNAs: roles and therapeutic implications in cancer. Cancer Cell 37, 270–288 (2020).
Batista, P. J. & Chang, H. Y. Long noncoding RNAs: cellular address codes in development and disease. Cell 152, 1298–1307 (2013).
Luo, H. et al. HOTTIP lncRNA promotes hematopoietic stem cell self-renewal leading to AML-like disease in mice. Cancer Cell 36, 645–659 (2019).
Kim, J. et al. Long noncoding RNA MALAT1 suppresses breast cancer metastasis. Nat. Genet. 50, 1705–1715 (2018).
Yao, R. W., Wang, Y. & Chen, L. L. Cellular functions of long noncoding RNAs. Nat. Cell Biol. 21, 542–551 (2019).
Zhu, S., Wang, Z. & Xu, J. Connecting versatile lncRNAs with heterogeneous nuclear ribonucleoprotein K and pathogenic disorders. Trends Biochem. Sci 44, 733–736 (2019).
Li, D. et al. Long noncoding RNA pancEts-1 promotes neuroblastoma progression through hnRNPK-mediated β-catenin stabilization. Cancer Res. 78, 1169–1183 (2018).
Liu, L. et al. MRPL33 and its splicing regulator hnRNPK are required for mitochondria function and implicated in tumor progression. Oncogene 37, 86–94 (2018).
Gallardo, M. et al. hnRNP K is a haploinsufficient tumor suppressor that regulates proliferation and differentiation programs in hematologic malignancies. Cancer Cell 28, 486–499 (2015).
Jeong, S. H. et al. Hippo-mediated suppression of IRS2/AKT signaling prevents hepatic steatosis and liver cancer. J. Clin. Invest. 128, 1010–1025 (2018).
Cheung, P. et al. Regenerative reprogramming of the intestinal stem cell state via hippo signaling suppresses metastatic colorectal cancer. Cell Stem Cell 27, 590–604 (2020).
Dey, A., Varelas, X. & Guan, K. L. Targeting the Hippo pathway in cancer, fibrosis, wound healing and regenerative medicine. Nat. Rev. Drug Discov. 19, 480–494 (2020).
Chen, R. et al. High mobility group protein B1 controls liver cancer initiation through yes-associated protein-dependent aerobic glycolysis. Hepatology 67, 1823–1841 (2018).
Xu, Y. et al. LncRNA PVT1 up-regulation is a poor prognosticator and serves as a therapeutic target in esophageal adenocarcinoma. Mol. Cancer 18, 141 (2019).
Wang, Q. et al. Emerging role of RNA methyltransferase METTL3 in gastrointestinal cancer. J. Hematol. Oncol. 13, 57 (2020).
Huang, H. et al. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat. Cell Biol. 20, 285–295 (2018).
Muller, S. et al. The oncofetal RNA-binding protein IGF2BP1 is a druggable, post-transcriptional super-enhancer of E2F-driven gene expression in cancer. Nucleic Acids Res. 48, 8576–8590 (2020).
Lederer, M., Bley, N., Schleifer, C. & Huttelmaier, S. The role of the oncofetal IGF2 mRNA-binding protein 3 (IGF2BP3) in cancer. Semin. Cancer Biol. 29, 3–12 (2014).
Xu, J. et al. Long noncoding RNA GMAN promotes hepatocellular carcinoma progression by interacting with eIF4B. Cancer Lett. 473, 1–12 (2020).
Wang, S. et al. CHIP functions as a novel suppressor of tumour angiogenesis with prognostic significance in human gastric cancer. Gut 62, 496–508 (2013).
Wang, S. et al. Prognostic and predictive role of JWA and XRCC1 expressions in gastric cancer. Clin. Cancer Res. 18, 2987–2996 (2012).
Duran, A. et al. p62/SQSTM1 by binding to vitamin D receptor inhibits hepatic stellate cell activity, fibrosis, and liver cancer. Cancer Cell 30, 595–609 (2016).
Shen, B. et al. Efficient genome modification by CRISPR–Cas9 nickase with minimal off-target effects. Nat. Methods 11, 399–402 (2014).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743 (2016).
Broutier, L. et al. Human primary liver cancer-derived organoid cultures for disease modeling and drug screening. Nat. Med. 23, 1424–1435 (2017).
Sun, L. et al. Modelling liver cancer initiation with organoids derived from directly reprogrammed human hepatocytes. Nat. Cell Biol. 21, 1015–1026 (2019).
Dhar, D. et al. Liver cancer initiation requires p53 inhibition by CD44-enhanced growth factor signaling. Cancer Cell 33, 1061–1077 (2018).
Acknowledgements
This work was supported by grants from the National Key Research and Development Program of China (2016YFC0905900 to B.S.), Funds for International Cooperation and Exchange of the National Natural Science Foundation of China (82120108012 to B.S.), State Key Program of the National Natural Science Foundation (81930086 to B.S.; 31530047 to Z.H.), National Natural Science Foundation (82073114 and 81773383 to S.W.; 81902836 to Y.G.) and Innovative Research Groups of the National Natural Science Foundation (81521004 to B.S.). B.S. is a Distinguished Professor Yangtze River Scholar.
Author information
Authors and Affiliations
Contributions
A.X. performed the experiments and wrote the paper. A.X., Q.W., W.Y., Y.G., L.Z. and Z.W. analyzed the data. Y.G., W.Y. and Z.H. generated the lnc-CTHCC-KO mouse model using the CRISPR–Cas9 system. W.Y. and Y.G. performed some data analysis from TCGA and GTEx databases. J.X., C.C., Y.G., D.W., Q.H., W.Y., F.W., C.X., Y.Z., G.B., X.T. and S.L. provided the samples. A.X., B.S., S.W. and Z.H. designed the study and revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review information
Nature Cancer thanks Sven Diederichs and Lars Zender for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
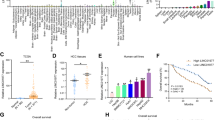
Extended Data Fig. 1 The characteristics of cancer-testis-associated lncRNA (lnc-CTHCC).
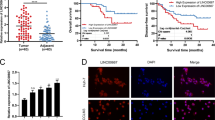
(a) Expression in TPM (Transcripts Per Kilobase Million) of lnc-CTHCC among the GTEx normal tissue RNA-seq database (n = 17382 samples). The lower bound of the box is 25% quantile of the data, the upper bound of the box is 75% quantile of the data and the centre bound of the box is 50% quantile of the data. The IQR (Inter-Quartile Range) is the distance between the lower and upper bound of the box. The lower whisker extends from the lower bound to the lowest value within 1.5×IQR and the upper whisker extends from the upper bound to the highest value within 1.5×IQR. (b) Expression in FPKM (Fragments Per Kilobase Million) of lncCTHCC among the various tumors of TCGA (n = 9669 samples). The box plots are defined in the same terms as (A). (c-d) Representative images of PCR products from the human (C) and the mouse (D) 5’ RACE and 3’ RACE and sequencing of RACE products. The results are representative of three independent experiments. (e) The expression of the long versus short lnc-CTHCC transcripts in HCC tissues (n = 18 tumors, P = 0.0144). Two-tailed paired t-test was performed. (f) The aligned sequences of human and mouse lnc-CTHCC transcripts. (g-i) The protein-coding potential of lnc-CTHCC using the Coding Potential Assessment Tool (http://lilab.research.bcm.edu/cpat/), Coding Potential Calculator (http://cpc.cbi.pku.edu.cn/) and ORF finder software from the National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/orffinder/). GAPDH and β-actin served as the positive controls of coding genes, and PVT1 and HOTAIR served as the positive control of non-coding genes. (j) The secondary structure of lnc-CTHCC by the Vienna RNA Web Services (http://rna.tbi.univie.ac.at/). (k) Univariate Cox regression analysis in the HCC cohort (n = 64 patients). All the bars corresponded to 95% confidence intervals. (l) Multivariate Cox regression analysis from TCGA with complete clinical data (n = 240 patients). All the bars corresponded to 95% confidence intervals. *P < 0.05.
Extended Data Fig. 2 The expression and prognoses of lnc-CTHCC in cholangiocarcinoma and other types of cancer from TCGA.
(a) The RNA levels of lnc-CTHCC in cholangiocarcinoma and paired non-tumor tissues (n = 18 patients). Data are shown as mean ± SEM. Two-tailed paired t-test was performed. (b) The RNA levels of lnc-CTHCC in tumors from HCC (n = 64 patients) and cholangiocarcinoma (n = 18 patients, P = 0.0353, two-tailed unpaired t-test). Data are shown as mean ± SEM. (c-l) Kaplan–Meier survival curves of overall survival among other types of cancer using TCGA database. Log-rank test was performed. CHOL: cholangiocarcinoma (n = 36 patients), LUAD: lung adenocarcinoma (n = 471 patients), THYM: thymoma (n = 118 patients), BRCA: breast invasive carcinoma (n = 1061 patients), OV: ovarian serous cystadenocarcinoma (n = 422 patients), COAD: colon adenocarcinoma (n = 265 patients), ESCA: esophageal carcinoma (n = 178 patients), KIRC: kidney renal clear cell carcinoma (n = 508 patients), PAAD: pancreatic adenocarcinoma (n = 178 patients), STAD: stomach adenocarcinoma (n = 379 patients). *P < 0.05.
Extended Data Fig. 3 The effect of lnc-CTHCC KO in mice HCC and the localization of lnc-CTHCC in single cells from mouse liver cancer.
(a) The expression of adjacent genes, such as MAPK3, GDPD3, INO80E, SEZ6L2, and MVP, in the livers of WT and KO mice (n = 3 mice per group). Data are shown as mean ± SEM. Two-tailed unpaired t-test was performed. (b) Schematic representation of the DEN-induced HCC model. Two-week-old mice were intraperitoneally injected with DEN (25 mg/kg) and then fed a normal diet for 34 weeks (n = 6 mice per group). (c) Representative images of gross morphology from the livers of WT and KO mice. The images are representative of n = 6 animals. (d) The ratio of liver to body weight (P = 0.0003), the number of tumors (P = 0.0001) and the largest tumor diameter (P = 0.0022) from WT and KO mice. (e) ALT and AST levels in the serums of WT and KO mice. P values are as follows: P = 0.0003 (ALT); P = 0.0171 (AST). (f-g) HE, Ki67, TUNEL, and CD31 staining of the livers from WT and KO mice (scale bars = 50 μm). P values are as follows: P < 0.0001 (Ki67); P < 0.0001 (TUNEL); P < 0.0001 (CD31). For D-G, data are shown as mean ± SEM (n = 6 mice per group). Two-tailed unpaired t-test was performed. (h) FISH of lnc-CTHCC in the H22 cell line (scale bars = 5 μm). The results are representative of three independent experiments. (i-j) Liver cancer tissues from WT mice were dissociated into single cells for IF staining and FISH (scale bars = 5 μm). The markers of macrophages, epithelial cells, endothelial cells, and immune cells were F4/80, KRT, CD31, and CD45, respectively. The results are representative of three biologically independent samples. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 4 Overexpression of lnc-CTHCC promotes cell growth in HCC.
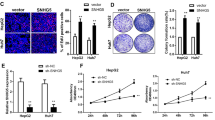
(a) The knockdown and overexpression efficiencies at the RNA levels in different HCC cells. P values are as follows: P < 0.0001 (Hep3B-sh1); P = 0.0001 (Hep3B-sh2); P = 0.0002(Hep3B-sh3); P < 0.0001 (MHCC97L-sh1); P = 0.0001 (MHCC97L-sh2); P = 0.0006(MHCC97L-sh3); P = 0.0006 (SMMC7721); P = 0.0003 (MHCC97H); P = 0.0004 (L02). (b) CCK8 assays in different HCC cells with the knockdown or overexpression of lnc-CTHCC. For Hep3B cells, P values are as follows: P = 0.0057 (48 h); P = 0.0029 (72 h); P = 0.0037 (96 h). For MHCC97L cells, P values are as follows: P = 0.0022 (48 h); P = 0.0011(72 h); P = 0.0003 (96 h). For SMMC7721 cells, P values are as follows: P = 0.0013 (72 h); P = 0.0002 (96 h); For MHCC97H cells, P values are as follows: P = 0.0011 (48 h); P = 0.0021 (72 h); P = 0.0011 (96 h). (c) Colony formation assays in different HCC cells with the knockdown or overexpression of lnc-CTHCC. P values are as follows: P = 0.0006 (Hep3B); P = 0.0013 (MHCC97L); P = 0.0008 (SMMC7721); P = 0.0026 (MHCC97H). (d) EdU assays in SMMC7721 and MHCC97H cells with lncCTHCC overexpression (scale bars = 50 μm). P values are as follows: P = 0.0010(SMMC7721); P < 0.0001 (MHCC97H). (e) Soft agar assays in SMMC7721 cells with lnc-CTHCC overexpression (scale bars = 50 μm, P = 0.0080). (f) Angiogenesis assays in HUVECs cultured with medium from SMMC7721 cells with lnc-CTHCC overexpression or control cells (scale bars = 50 μm). P values are as follows: P = 0.0022 (number of tubes); P = 0.0412 (cell growth). (g) The cytokine array analysis of the conditioned medium from the control groups and lnc-CTHCC-overexpressing SMMC7721 cells. (h) Heat map showing the differentially expressed proangiogenic factors, such as MCP-1, EGF, VEGF, Angiogenin, FGF-6, and MIG, in SMMC7721 cells with lnc-CTHCC overexpression. These experiments were performed twice. For A-F, data are from three biologically independent experiments and shown as mean ± SEM. Two-tailed paired t-test was performed. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 5 Overexpression of lnc-CTHCC promotes cell metastasis in HCC.
(a) Wound-healing assays in different HCC cells with the knockdown of lnc-CTHCC (scale bars = 50 μm). P values are as follows: P = 0.0002 (Hep3B); P = 0.0007 (MHCC97L). (b) Wound-healing assays in different HCC cells with the overexpression of lnc-CTHCC (scale bars = 50 μm). P values are as follows: P < 0.0001 (SMMC7721); P = 0.0033 (MHCC97H). (c) Representative images of cell migration and invasion assays in different HCC cells with the knockdown of lnc-CTHCC (scale bars = 50 μm). P values are as follows: P = 0.0051 (Hep3B-migration); P = 0.0022 (Hep3B-invasion); P = 0.0013 (MHCC97L-migration); P = 0.0042 (MHCC97L-invasion). (d) Representative images of cell migration and invasion assays in different HCC cells with the overexpression of lnc-CTHCC (scale bars = 50 μm). P values are as follows: P = 0.0012 (SMMC7721-migration); P = 0.0019 (SMMC7721-invasion); P = 0.0011 (MHCC97H-migration); P = 0.0034 (MHCC97H-invasion). (e-g) Overexpression of lnc-CTHCC significantly promoted HCC lung metastasis in nude mice (n = 10 mice per group). (E) Representative bioluminescent images in lung regions (P < 0.0001). (F) Representative images of the metastatic nodes in the lungs (n = 10 mice per group, P = 0.0007). For E-F, data are shown as mean ± SEM and two-tailed unpaired t-test was performed. (G) HE-stained lung sections (scale bars = 50 μm). (h-i) The expressions of EMT-related markers were analyzed by western blots, which indicated that lnc-CTHCC regulated EMT genes specifically via hnRNP K or YAP. The results are representative of three independent experiments. For A-D, data are from three biologically independent experiments and shown as mean ± SEM. Two-tailed paired t-test was performed. **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 6 Lnc-CTHCC has no effect on the expression and localization of hnRNP K.
(a,b) The knockdown efficiencies of hnRNP K at the RNA and protein levels in Hep3B cells. P values are as follows: P = 0.0008 (si-1); P = 0.0004 (si-2); P = 0.0001 (si-3). (c) Overexpression of hnRNP K rescued the colony formation ability of Hep3B cells with lnc-CTHCC knockdown. P values are as follows: P = 0.0004 (sh-NC + NC and sh-lnc-CTHCC + NC); P = 0.0017 (sh-lnc-CTHCC + NC and sh-lnc-CTHCC + hnRNP K). (d) Knockdown of hnRNP K rescued the colony formation ability of SMMC7721 cells with lnc-CTHCC overexpression. P values are as follows: P < 0.0001 (NC + si-NC and lnc-CTHCC + si-NC); P = 0.0006 (lnc-CTHCC + si-NC and lnc-CTHCC + sihnRNP K). (e) Overexpression of hnRNP K rescued the invasion ability of Hep3B cells with lnc-CTHCC knockdown (scale bars = 50 μm). P values are as follows: P = 0.0004 (sh-NC + NC and sh-lncCTHCC + NC); P = 0.0005 (sh-lnc-CTHCC + NC and sh-lnc-CTHCC + hnRNP K). (f) Knockdown of hnRNP K rescued the invasion ability of SMMC7721 cells with lnc-CTHCC overexpression (scale bars = 50 μm). P values are as follows: P < 0.0001 (NC + si-NC and lncCTHCC + si-NC); P = 0.0018 (lnc-CTHCC + si-NC and lnc-CTHCC + si-hnRNP K). (g) The RNA levels of hnRNP K in different HCC cells with knockdown or overexpression of lnc-CTHCC. (h) Western blot assays of hnRNP K in different HCC cells with knockdown or overexpression of lnc-CTHCC. (i) The hnRNP K staining of the xenograft tumors with lnc-CTHCC knockdown or overexpression (scale bars = 50 μm). (j) Cellular immunofluorescence assays of hnRNP K in different HCC cells with knockdown or overexpression of lnc-CTHCC (scale bars = 25 μm). For B and H-J, the results are representative of three independent experiments. For A and C-G, data are from three biologically independent experiments and shown as mean ± SEM. Two-tailed paired t-test was performed. **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 7 Transcriptome sequencing analysis after knockdown or knockout of lnc-CTHCC in HCC.
(a,b) Heat map showing the differentially expressed genes in Hep3B cells with lnc-CTHCC knockdown (A) and in liver cancer tissues from wild-type and knockout mice (B). (c) The RNA levels of YAP in Hep3B cells with lnc-CTHCC knockdown (n = 3 independent experiments, P = 0.0007, two-tailed paired t-test) or in the tumors from WT and KO mice (n = 3 mice per group, P < 0.0001, two-tailed unpaired t-test). Data are shown as mean ± SEM. (d) IHC staining of YAP in the xenograft tumors with lnc-CTHCC overexpression (scale bars = 50 μm). The results are representative of three biologically independent samples. (e) The RNA levels of YAP target genes (FGF2, CTGF, CYR61, CDC20 and SERPINE1) in MHCC97L cells with lnc-CTHCC knockdown. P values are as follows: P = 0.0006 (SERPINE1); P = 0.0048 (CDC20); P = 0.0035 (CTGF); P = 0.0004 (CYR61); P = 0.0004 (FGF2). (f) Luciferase reporter assays for YAP activity in Hep3B cells with sh-lnc-CTHCC and hnRNP K overexpression. P values are as follows: P < 0.0001 (sh-NC + NC and sh-lnc-CTHCC + NC); P = 0.0017 (sh-lnc-CTHCC + NC and sh-lnc-CTHCC + hnRNP K); P = 0.0008 (sh-NC + NC and sh-NC + hnRNP K). (g) The protein levels of YAP in Hep3B cells with sh-lnc-CTHCC and hnRNP K overexpression. The results are representative of three independent experiments. (h) The RNA levels of YAP in HCC and normal tissues from TCGA (P = 0.0001, two-tailed unpaired t-test) or in HCC and paired non-tumor tissues from the HCC cohort (n = 50 patients, P < 0.0001, two-tailed paired t-test). Data are shown as mean ± SEM. (i) Pearson correlation analysis between lnc-CTHCC expression and YAP expression (n = 50 patients, P < 0.0001). (J) Western blot assays of YAP in HCC and paired non-tumor tissues (n = 10 paired samples). (k) IHC staining of YAP from HCC tissues with relatively high or low lnc-CTHCC expression (scale bars = 50 μm). (l) Kaplan-Meier survival curves of OS from TCGA (P = 0.0413, log-rank test). For E-F, data are shown as mean ± SEM of three independent experiments. Two-tailed paired t-test was performed. **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 8 YAP is involved in the biological function of lnc-CTHCC in HCC.
(a) The knockdown efficiencies of YAP at the RNA and protein levels in Hep3B cells. P values are as follows: P = 0.0002 (sh-1); P = 0.0001 (sh-2). (b,c) Knockdown of YAP suppressed HCC cell proliferation and metastasis in vitro. (B) Colony formation assays. P values are as follows: P = 0.0034 (sh-YAP-1); P = 0.0057 (sh-YAP-2). (C) Transwell assays. Scale bars = 50 μm. P values are as follows: P = 0.0044 (migration-sh-YAP-1); P = 0.0046 (migration-sh-YAP2); P = 0.0009(invasion-sh-YAP-1); P = 0.0013 (invasion-shYAP-2). (d-f) Knockdown of YAP suppressed lnc-CTHCC-induced increase in HCC cell proliferation and metastasis in vitro. (D) Colony formation assays. P values are as follows: P = 0.0004 (Ctrl + sh-NC and Ctrl + sh-YAP); P = 0.0065 (lnc-CTHCC + sh-NC and lnc-CTHCC + sh-YAP). (E) EdU assays. Scale bars = 50 μm. P values are as follows: P = 0.0117 (Ctrl + sh-NC and Ctrl + shYAP); P = 0.0102 (lnc-CTHCC + sh-NC and lnc-CTHCC + sh-YAP). (F) Transwell assays. Scale bars = 50 μm. P values are as follows: P = 0.0006 (Ctrl + sh-NC and Ctrl + sh-YAP); P = 0.0031 (lnc-CTHCC + sh-NC and lnc-CTHCC + sh-YAP). (g,h) The tumor growth curves and tumor weight (n = 6 mice per group, P < 0.0001). Data are shown as mean ± SEM. Two-tailed unpaired t-test was performed. (i,j) Overexpression of YAP rescued the proliferation and invasion abilities of Hep3B cells with lnc-CTHCC knockdown (scale bars = 50 μm). (I) P values are as follows: P = 0.0012 (sh-NC + NC and sh-lnc-CTHCC + NC); P = 0.0003 (sh-lnc-CTHCC + NC and sh-lnc-CTHCC + YAP). (J) P values are as follows: P = 0.0006 (sh-NC + NC and sh-lnc-CTHCC + NC); P = 0.0005 (sh-lnc-CTHCC + NC and sh-lnc-CTHCC + YAP). For A-F and I-J, data are from three biologically independent experiments and shown as mean ± SEM. Two-tailed paired t-test was performed. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 9 METTL3 promotes cell proliferation and metastasis in HCC.
(a-c) The knockdown and overexpression efficiencies of METTL3 at the RNA and protein levels in Hep3B cells. (A) P = 0.0003. (B) P = 0.0013. (C) The results are representative of three independent experiments. (d-f) Knockdown of METTL3 inhibited cell proliferation and metastasis while overexpression of METTL3 promoted cell proliferation and metastasis in Hep3B cells. (D) P values are as follows: P = 0.0009 (sh-METTL3); P = 0.0012 (Lv-METTL3). (E) Scale bars = 50 μm. P values are as follows: P < 0.0001 (sh-METTL3); P = 0.0010 (Lv-METTL3). (F) Scale bars = 50 μm. P values are as follows: P = 0.0020 (sh-METTL3); P = 0.0004 (Lv-METTL3). (g,h) Knockdown of lnc-CTHCC rescued the proliferation and invasion abilities of Hep3B cells with METTL3 overexpression. (G) P values are as follows: P = 0.0003 (NC + sh-NC and METTL3 + sh-NC); P = 0.0009 (METTL3 + sh-NC and METTL3 + sh-lnc-CTHCC). (H) Scale bars = 50 μm. P values are as follows: P = 0.0022(NC + sh-NC and METTL3 + sh-NC); P = 0.0003 (METTL3 + sh-NC and METTL3 + sh-lnc-CTHCC). For A-B and D-H, data are from three biologically independent experiments and shown as mean ± SEM. Two-tailed paired t-test was performed. **P < 0.01; ***P < 0.001; ****P < 0.0001.
Extended Data Fig. 10 Knockdown of IGF2BP1/3 inhibits cell proliferation and metastasis in HCC.
(a) The RNA levels of IGF2BP1/3 in Hep3B cells with lnc-CTHCC knockdown. (b) Western blot assays of IGF2BP1/3 in different cells with lnc-CTHCC knockdown or overexpression. The results are representative of three independent experiments. (c,d) The knockdown efficiencies of IGF2BP1/3 at the RNA and protein levels in Hep3B cells. P values are as follows: P = 0.0006 (si-IGF2BP1-1); P = 0.0006 (si-IGF2BP1-2); P = 0.0002 (si-IGF2BP3-1); P = 0.0002 (si-IGF2BP3-2). For D, the results are representative of three independent experiments. (e,f) Knockdown of IGF2BP1/3 inhibited cell proliferation and metastasis in Hep3B cells. (e) P values are as follows: P = 0.0016 (si-IGF2BP1-1); P = 0.0027 (si-IGF2BP1-2); P < 0.0001 (si-IGF2BP3-1); P < 0.0001 (si-IGF2BP3-2). (f) Scale bars = 50 μm. P values are as follows: P = 0.0012 (migration-si-IGF2BP1-1); P = 0.0003 (migration-si-IGF2BP1-2); P = 0.0009 (invasionsi-IGF2BP1-1); P < 0.0001 (invasion-si-IGF2BP1-2); P = 0.0006 (migration-si-IGF2BP3-1); P = 0.0009 (migration-si-IGF2BP3-2); P = 0.0008 (invasion-si-IGF2BP3-1); P = 0.0009 (invasion-siIGF2BP3-2). (g) Schematic diagram showing the molecular mechanisms underlying lnc-CTHCC action in hepatocellular carcinoma. For A, C and E-F, data are from three biologically independent experiments and shown as mean ± SEM. Two-tailed paired t-test was performed. **P < 0.01; ***P < 0.001; ****P < 0.0001.
Supplementary information
Supplementary Information
Supplementary Tables 1–6.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Xia, A., Yuan, W., Wang, Q. et al. The cancer-testis lncRNA lnc-CTHCC promotes hepatocellular carcinogenesis by binding hnRNP K and activating YAP1 transcription. Nat Cancer 3, 203–218 (2022). https://doi.org/10.1038/s43018-021-00315-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00315-4
This article is cited by
-
IGF2BP3 promotes glutamine metabolism of endometriosis by interacting with UCA1 to enhances the mRNA stability of GLS1
Molecular Medicine (2024)
-
The m6A modification mediated-lncRNA POU6F2-AS1 reprograms fatty acid metabolism and facilitates the growth of colorectal cancer via upregulation of FASN
Molecular Cancer (2024)
-
New horizons for the role of RNA N6-methyladenosine modification in hepatocellular carcinoma
Acta Pharmacologica Sinica (2024)
-
Bioinformatic identification reveals a m6A-binding protein, IGF2BP2, as a novel tumor-promoting gene signature in thyroid carcinoma
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
The regulation of hypoxia-related lncRNAs in hepatocellular carcinoma
Discover Oncology (2024)