Abstract
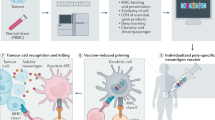
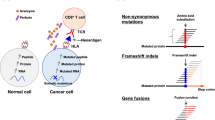
Several current immunotherapy approaches target private neoantigens derived from mutations that are unique to individual patients’ tumors. However, immunotherapeutic agents can also be developed against public neoantigens derived from recurrent mutations in cancer driver genes. The latter approaches target proteins that are indispensable for tumor growth, and each therapeutic agent can be applied to numerous patients. Here we review the opportunities and challenges involved in the identification of suitable public neoantigen targets and the development of therapeutic agents targeting them.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Data for The Cancer Genome Atlas mutation frequencies used in the analyses presented in Fig. 3 and Table 2 are available from the National Cancer Institute Genomics Data Commons (https://gdc.cancer.gov/). Data for the HLA frequencies used in the analyses presented in Tables 1 and 2 and Supplementary Table 1 are available from the Allele Frequency Net Database (http://www.allelefrequencies.net/) and National Marrow Donor Program (https://bioinformatics.bethematchclinical.org/hla-resources/haplotype-frequencies/high-resolution-hla-alleles-and-haplotypes-in-the-us-population/). Data for cancer incidence used in the analyses presented in Fig. 3 and Table 2 are available from the National Cancer Institute Surveillance, Epidemiology, and End Results Program (https://seer.cancer.gov/statfacts/html/common.html). Data for ethnicity representation in the United States used in the analyses presented in Tables 1 and 2 and Supplementary Table 1 are available from the United States Census Bureau (https://www.census.gov/quickfacts/fact/table/US/PST045219).
Change history
29 July 2021
A Correction to this paper has been published: https://doi.org/10.1038/s43018-021-00246-0
References
Schumacher, T. N., Scheper, W. & Kvistborg, P. Cancer neoantigens. Annu. Rev. Immunol. 37, 173–200 (2019).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Leko, V. & Rosenberg, S. A. Identifying and targeting human tumor antigens for T cell-based immunotherapy of solid tumors. Cancer Cell 38, 454–472 (2020).
Deniger, D. C. et al. T-cell responses to TP53 ‘hotspot’ mutations and unique neoantigens expressed by human ovarian cancers. Clin. Cancer Res. 24, 5562–5573 (2018).
Parkhurst, M. R. et al. Unique neoantigens arise from somatic mutations in patients with gastrointestinal cancers. Cancer Discov. 9, 1022–1035 (2019).
Guedan, S., Ruella, M. & June, C. H. Emerging cellular therapies for cancer. Annu. Rev. Immunol. 37, 145–171 (2019).
Reiter, J. G. et al. An analysis of genetic heterogeneity in untreated cancers. Nat. Rev. Cancer 19, 639–650 (2019).
McGranahan, N. & Swanton, C. Neoantigen quality, not quantity. Sci. Transl. Med. 11, eaax7918 (2019).
Rosenthal, R. et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 567, 479–485 (2019).
Hsiue, E. H.-C. et al. Targeting a neoantigen derived from a common TP53 mutation. Science 371, eabc8697 (2021).
Heinrich, M. C. et al. Avapritinib in advanced PDGFRA D842V-mutant gastrointestinal stromal tumour (NAVIGATOR): a multicentre, open-label, phase 1 trial. Lancet Oncol. 21, 935–946 (2020).
Hong, D. S. et al. Larotrectinib in patients with TRK fusion-positive solid tumours: a pooled analysis of three phase 1/2 clinical trials. Lancet Oncol. 21, 531–540 (2020).
Marty, R. et al. MHC-I genotype restricts the oncogenic mutational landscape. Cell 171, 1272–1283.e15 (2017).
Marty Pyke, R. et al. Evolutionary pressure against MHC class II binding cancer mutations. Cell 175, 416–428.e13 (2018).
Van den Eynden, J., Jiménez-Sánchez, A., Miller, M. L. & Larsson, E. Lack of detectable neoantigen depletion signals in the untreated cancer genome. Nat. Genet. 51, 1741–1748 (2019).
Pardoll, D. Cancer and the immune system: basic concepts and targets for intervention. Semin. Oncol. 42, 523–538 (2015).
Castle, J. C., Uduman, M., Pabla, S., Stein, R. B. & Buell, J. S. Mutation-derived neoantigens for cancer immunotherapy. Front. Immunol. 10, 1856 (2019).
Segal, N. H. et al. Epitope landscape in breast and colorectal cancer. Cancer Res. 68, 889–892 (2008).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830.e14 (2018).
Garcia-Garijo, A., Fajardo, C. A. & Gros, A. Determinants for neoantigen identification. Front. Immunol. 10, 1392 (2019).
Bassani-Sternberg, M. et al. Direct identification of clinically relevant neoepitopes presented on native human melanoma tissue by mass spectrometry. Nat. Commun. 7, 13404 (2016).
Chheda, Z. S. et al. Novel and shared neoantigen derived from histone 3 variant H3.3K27M mutation for glioma T cell therapy. J. Exp. Med. 215, 141–157 (2018).
Narayan, R. et al. Acute myeloid leukemia immunopeptidome reveals HLA presentation of mutated nucleophosmin. PLoS ONE 14, e0219547 (2019).
Wang, Q. et al. Direct detection and quantification of neoantigens. Cancer Immunol. Res. 7, 1748–1754 (2019).
Kalaora, S. et al. Combined analysis of antigen presentation and T-cell recognition reveals restricted immune responses in melanoma. Cancer Discov. 8, 1366–1375 (2018).
Arnaud, M. et al. Biotechnologies to tackle the challenge of neoantigen identification. Curr. Opin. Biotechnol. 65, 52–59 (2020).
Gerber, H.-P., Sibener, L. V., Lee, L. J. & Gee, M. H. Identification of antigenic targets. Trends Cancer 6, 299–318 (2020).
Sharkey, M. S., Lizée, G., Gonzales, M. I., Patel, S. & Topalian, S. L. CD4+ T-cell recognition of mutated B-RAF in melanoma patients harboring the V599E mutation. Cancer Res. 64, 1595–1599 (2004).
Yamamoto, T. N., Kishton, R. J. & Restifo, N. P. Developing neoantigen-targeted T cell-based treatments for solid tumors. Nat. Med. 25, 1488–1499 (2019).
Jaigirdar, A., Rosenberg, S. A. & Parkhurst, M. A high-avidity WT1-reactive T-cell receptor mediates recognition of peptide and processed antigen but not naturally occurring WT1-positive tumor cells. J. Immunother. 39, 105–116 (2016).
Tran, E. et al. Immunogenicity of somatic mutations in human gastrointestinal cancers. Science 350, 1387–1390 (2015).
Smith, K. N. et al. Persistent mutant oncogene specific T cells in two patients benefitting from anti-PD-1. J. Immunother. Cancer 7, 40 (2019).
Tran, E. et al. T-cell transfer therapy targeting mutant KRAS in cancer. N. Engl. J. Med. 375, 2255–2262 (2016).
Chen, F. et al. Neoantigen identification strategies enable personalized immunotherapy in refractory solid tumors. J. Clin. Invest. 129, 2056–2070 (2019).
Veatch, J. R. et al. Tumor-infiltrating BRAFV600E-specific CD4+ T cells correlated with complete clinical response in melanoma. J. Clin. Invest. 128, 1563–1568 (2018).
Clark, R. E. et al. Direct evidence that leukemic cells present HLA-associated immunogenic peptides derived from the BCR–ABL b3a2 fusion protein. Blood 98, 2887–2893 (2001).
Van der Lee, D. I. et al. Mutated nucleophosmin 1 as immunotherapy target in acute myeloid leukemia. J. Clin. Invest. 129, 774–785 (2019).
Biernacki, M. A. et al. CBFB–MYH11 fusion neoantigen enables T cell recognition and killing of acute myeloid leukemia. J. Clin. Invest. 130, 5127–5141 (2020).
Xie, G. et al. CAR-T cells targeting a nucleophosmin neoepitope exhibit potent specific activity in mouse models of acute myeloid leukaemia. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-020-00625-5 (2020).
Douglass, J. et al. Bispecific antibodies targeting mutant RAS neoantigens. Sci. Immunol. 6, eabd5515 (2021).
Gjertsen, M. K., Bjorheim, J., Saeterdal, I., Myklebust, J. & Gaudernack, G. Cytotoxic CD4+ and CD8+ T lymphocytes, generated by mutant p21-ras (12VAL) peptide vaccination of a patient, recognize 12VAL-dependent nested epitopes present within the vaccine peptide and kill autologous tumour cells carrying this mutation. Int. J. Cancer 72, 784–790 (1997).
Malekzadeh, P. et al. Antigen experienced T cells from peripheral blood recognize p53 neoantigens. Clin. Cancer Res. 26, 1267–1276 (2020).
Tubb, V. M. et al. Isolation of T cell receptors targeting recurrent neoantigens in hematological malignancies. J. Immunother. Cancer 6, 70 (2018).
Gros, A. et al. Recognition of human gastrointestinal cancer neoantigens by circulating PD-1+ lymphocytes. J. Clin. Invest. 129, 4992–5004 (2019).
Wang, Q. J. et al. Identification of T-cell receptors targeting KRAS-mutated human tumors. Cancer Immunol. Res. 4, 204–214 (2016).
Lo, W. et al. Immunologic recognition of a shared p53 mutated neoantigen in a patient with metastatic colorectal cancer. Cancer Immunol. Res. 7, 534–543 (2019).
Strønen, E. et al. Targeting of cancer neoantigens with donor-derived T cell receptor repertoires. Science 352, 1337–1341 (2016).
Calis, J. J. A., de Boer, R. J. & Keşmir, C. Degenerate T-cell recognition of peptides on MHC molecules creates large holes in the T-cell repertoire. PLoS Comput. Biol. 8, e1002412 (2012).
Shao, X. M. et al. High-throughput prediction of MHC class I and II neoantigens with MHCnuggets. Cancer Immunol. Res. 8, 396–408 (2020).
Bulik-Sullivan, B. et al. Deep learning using tumor HLA peptide mass spectrometry datasets improves neoantigen identification. Nat. Biotechnol. 37, 55–63 (2019).
Sarkizova, S. et al. A large peptidome dataset improves HLA class I epitope prediction across most of the human population. Nat. Biotechnol. 38, 199–209 (2020).
Abelin, J. G. et al. Defining HLA-II ligand processing and binding rules with mass spectrometry enhances cancer epitope prediction. Immunity 51, 766–779.e17 (2019).
Vita, R. et al. The immune epitope database (IEDB) 3.0. Nucleic Acids Res. 43, D405–D412 (2015).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Shao, W. et al. The SysteMHC Atlas project. Nucleic Acids Res. 46, D1237–D1247 (2018).
Hundal, J. et al. pVACtools: a computational toolkit to identify and visualize cancer neoantigens. Cancer Immunol. Res. 8, 409–420 (2020).
Wells, D. K. et al. Key parameters of tumor epitope immunogenicity revealed through a consortium approach improve neoantigen prediction. Cell 183, 818–834.e13 (2020).
Jappe, E. C. et al. Thermostability profiling of MHC-bound peptides: a new dimension in immunopeptidomics and aid for immunotherapy design. Nat. Commun. 11, 6305 (2020).
Bentzen, A. K. et al. T cell receptor fingerprinting enables in-depth characterization of the interactions governing recognition of peptide–MHC complexes. Nat. Biotechnol. 36, 1191–1196 (2018).
Zhang, S.-Q. et al. High-throughput determination of the antigen specificities of T cell receptors in single cells. Nat. Biotechnol. 36, 1156–1159 (2018).
Peng, S. et al. Sensitive detection and analysis of neoantigen-specific T cell populations from tumors and blood. Cell Rep. 28, 2728–2738.e7 (2019).
Moritz, A. et al. High-throughput peptide–MHC complex generation and kinetic screenings of TCRs with peptide-receptive HLA-A*02:01 molecules. Sci. Immunol. 4, eaav0860 (2019).
Saini, S. K. et al. Empty peptide-receptive MHC class I molecules for efficient detection of antigen-specific T cells. Sci. Immunol. 4, eaau9039 (2019).
Overall, S. A. et al. High throughput pMHC-I tetramer library production using chaperone-mediated peptide exchange. Nat. Commun. 11, 1909 (2020).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl. J. Med. 375, 819–829 (2016).
Sánchez-Paulete, A. R. et al. Antigen cross-presentation and T-cell cross-priming in cancer immunology and immunotherapy. Ann. Oncol. 28, xii44–xii55 (2017).
Marino, F. et al. Biogenesis of HLA ligand presentation in immune cells upon activation reveals changes in peptide length preference. Front. Immunol. 11, 1981 (2020).
Purcell, A. W., Ramarathinam, S. H. & Ternette, N. Mass spectrometry-based identification of MHC-bound peptides for immunopeptidomics. Nat. Protoc. 14, 1687–1707 (2019).
Klatt, M. G. et al. Solving an MHC allele-specific bias in the reported immunopeptidome. JCI Insight 5, e141264 (2020).
Cafri, G. et al. mRNA vaccine-induced neoantigen-specific T cell immunity in patients with gastrointestinal cancer. J. Clin. Invest. 130, 5976–5988 (2020).
Bocchia, M. et al. Complete molecular response in CML after p210 BCR–ABL1-derived peptide vaccination. Nat. Rev. Clin. Oncol. 7, 600–603 (2010).
Chatani, P. D. & Yang, J. C. Mutated RAS: targeting the ‘untargetable’ with T cells. Clin. Cancer Res. 26, 537–544 (2020).
Comoli, P. et al. BCR–ABL-specific T-cell therapy in Ph+ ALL patients on tyrosine-kinase inhibitors. Blood 129, 582–586 (2017).
Veatch, J. R. et al. Endogenous CD4+ T cells recognize neoantigens in lung cancer patients, including recurrent oncogenic KRAS and ERBB2 (Her2) driver mutations. Cancer Immunol. Res. 7, 910–922 (2019).
Roth, T. L. et al. Reprogramming human T cell function and specificity with non-viral genome targeting. Nature 559, 405–409 (2018).
Querques, I. et al. A highly soluble Sleeping Beauty transposase improves control of gene insertion. Nat. Biotechnol. 37, 1502–1512 (2019).
Liddy, N. et al. Monoclonal TCR-redirected tumor cell killing. Nat. Med. 18, 980–987 (2012).
Lowe, K. L. et al. Novel TCR-based biologics: mobilising T cells to warm ‘cold’ tumours. Cancer Treat. Rev. 77, 35–43 (2019).
Lu, Y.-C. et al. An efficient single-cell RNA-seq approach to identify neoantigen-specific T cell receptors. Mol. Ther. 26, 379–389 (2018).
Dijkstra, K. K. et al. Generation of tumor-reactive T cells by co-culture of peripheral blood lymphocytes and tumor organoids. Cell 174, 1586–1598.e12 (2018).
Paria, B. C. et al. Rapid identification and evaluation of neoantigen-reactive T-cell receptors from single cells. J. Immunother. https://doi.org/10.1097/CJI.0000000000000342 (2020).
Spindler, M. J. et al. Massively parallel interrogation and mining of natively paired human TCRαβ repertoires. Nat. Biotechnol. 38, 609–619 (2020).
Goebeler, M.-E. & Bargou, R. C. T cell-engaging therapies—BiTEs and beyond. Nat. Rev. Clin. Oncol. 17, 418–434 (2020).
MacKay, M. et al. The therapeutic landscape for cells engineered with chimeric antigen receptors. Nat. Biotechnol. 38, 233–244 (2020).
Skora, A. D. et al. Generation of MANAbodies specific to HLA-restricted epitopes encoded by somatically mutated genes. Proc. Natl Acad. Sci. USA 112, 9967–9972 (2015).
Miller, M. S. et al. An engineered antibody fragment targeting mutant β-catenin via major histocompatibility complex I neoantigen presentation. J. Biol. Chem. 294, 19322–19334 (2019).
Dao, T. et al. Therapeutic bispecific T-cell engager antibody targeting the intracellular oncoprotein WT1. Nat. Biotechnol. 33, 1079–1086 (2015).
Chang, A. Y. et al. A therapeutic T cell receptor mimic antibody targets tumor-associated PRAME peptide/HLA-I antigens. J. Clin. Invest. 127, 2705–2718 (2017).
Ahmed, M. et al. TCR-mimic bispecific antibodies targeting LMP2A show potent activity against EBV malignancies. JCI Insight 3, e97805 (2018).
Low, L., Goh, A., Koh, J., Lim, S. & Wang, C.-I. Targeting mutant p53-expressing tumours with a T cell receptor-like antibody specific for a wild-type antigen. Nat. Commun. 10, 5382 (2019).
Sharma, P., Harris, D. T., Stone, J. D. & Kranz, D. M. T-cell receptors engineered de novo for peptide specificity can mediate optimal T-cell activity without self cross-reactivity. Cancer Immunol. Res. 7, 2025–2035 (2019).
Riley, T. P. et al. T cell receptor cross-reactivity expanded by dramatic peptide–MHC adaptability. Nat. Chem. Biol. 14, 934–942 (2018).
Gejman, R. S. et al. Identification of the targets of T-cell receptor therapeutic agents and cells by use of a high-throughput genetic platform. Cancer Immunol. Res. 8, 672–684 (2020).
Ataie, N. et al. Structure of a TCR mimic antibody with target predicts pharmacogenetics. J. Mol. Biol. 428, 194–205 (2016).
Hellman, L. M. et al. Improving T cell receptor on-target specificity via structure-guided design. Mol. Ther. 27, 300–313 (2019).
Holland, C. J. et al. Specificity of bispecific T cell receptors and antibodies targeting peptide–HLA. J. Clin. Invest. 130, 2673–2688 (2020).
Sim, M. J. W. et al. High-affinity oligoclonal TCRs define effective adoptive T cell therapy targeting mutant KRAS-G12D. Proc. Natl Acad. Sci. USA 117, 12826–12835 (2020).
Wu, D., Gallagher, D. T., Gowthaman, R., Pierce, B. G. & Mariuzza, R. A. Structural basis for oligoclonal T cell recognition of a shared p53 cancer neoantigen. Nat. Commun. 11, 2908 (2020).
Hu, Z. et al. Personal neoantigen vaccines induce persistent memory T cell responses and epitope spreading in patients with melanoma. Nat. Med. 27, 515–525 (2021).
Blass, E. & Ott, P. A. Advances in the development of personalized neoantigen-based therapeutic cancer vaccines. Nat. Rev. Clin. Oncol. 18, 215–229 (2021).
Rosenberg, S. A., Yang, J. C. & Restifo, N. P. Cancer immunotherapy: moving beyond current vaccines. Nat. Med. 10, 909–915 (2004).
Klebanoff, C. A., Acquavella, N., Yu, Z. & Restifo, N. P. Therapeutic cancer vaccines: are we there yet? Immunol. Rev. 239, 27–44 (2011).
Vormehr, M., Türeci, Ö. & Sahin, U. Harnessing tumor mutations for truly individualized cancer vaccines. Annu. Rev. Med. 70, 395–407 (2019).
Mehta, N. K. et al. Pharmacokinetic tuning of protein–antigen fusions enhances the immunogenicity of T-cell vaccines. Nat. Biomed. Eng. 4, 636–648 (2020).
Ott, P. A. et al. A phase Ib trial of personalized neoantigen therapy plus anti-PD-1 in patients with advanced melanoma, non-small cell lung cancer, or bladder cancer. Cell 183, 347–362.e24 (2020).
Sahin, U. et al. An RNA vaccine drives immunity in checkpoint-inhibitor-treated melanoma. Nature 585, 107–112 (2020).
Romero, P. et al. The Human Vaccines Project: a roadmap for cancer vaccine development. Sci. Transl. Med. 8, 334ps9 (2016).
Türeci, Ö. et al. Challenges towards the realization of individualized cancer vaccines. Nat. Biomed. Eng. 2, 566–569 (2018).
Van Poelgeest, M. I. E. et al. Vaccination against oncoproteins of HPV16 for noninvasive vulvar/vaginal lesions: lesion clearance is related to the strength of the T-cell response. Clin. Cancer Res. 22, 2342–2350 (2016).
Schumacher, T. et al. A vaccine targeting mutant IDH1 induces antitumour immunity. Nature 512, 324–327 (2014).
Pan, J. et al. Immunoprevention of KRAS-driven lung adenocarcinoma by a multipeptide vaccine. Oncotarget 8, 82689–82699 (2017).
Morrison, A. H., Byrne, K. T. & Vonderheide, R. H. Immunotherapy and prevention of pancreatic cancer. Trends Cancer 4, 418–428 (2018).
Weber, E. W., Maus, M. V. & Mackall, C. L. The emerging landscape of immune cell therapies. Cell 181, 46–62 (2020).
Rafiq, S., Hackett, C. S. & Brentjens, R. J. Engineering strategies to overcome the current roadblocks in CAR T cell therapy. Nat. Rev. Clin. Oncol. 17, 147–167 (2020).
Depil, S., Duchateau, P., Grupp, S. A., Mufti, G. & Poirot, L. ‘Off-the-shelf’ allogeneic CAR T cells: development and challenges. Nat. Rev. Drug Discov. 19, 185–199 (2020).
Liu, E. et al. Use of CAR-transduced natural killer cells in CD19-positive lymphoid tumors. N. Engl. J. Med. 382, 545–553 (2020).
Mo, F. et al. Engineered off-the-shelf therapeutic T cells resist host immune rejection. Nat. Biotechnol. 39, 56–63 (2021).
Smith, T. T. et al. In situ programming of leukaemia-specific T cells using synthetic DNA nanocarriers. Nat. Nanotechnol. 12, 813–820 (2017).
Agarwal, S. et al. In vivo generation of CAR T cells selectively in human CD4+ lymphocytes. Mol. Ther. 28, 1783–1794 (2020).
Roybal, K. T. & Lim, W. A. Synthetic immunology: hacking immune cells to expand their therapeutic capabilities. Annu. Rev. Immunol. 35, 229–253 (2017).
Lynn, R. C. et al. c-Jun overexpression in CAR T cells induces exhaustion resistance. Nature 576, 293–300 (2019).
Yamamoto, T. N. et al. T cells genetically engineered to overcome death signaling enhance adoptive cancer immunotherapy. J. Clin. Invest. 129, 1551–1565 (2019).
Stadtmauer, E. A. et al. CRISPR-engineered T cells in patients with refractory cancer. Science 367, eaba7365 (2020).
Davenport, A. J. et al. Chimeric antigen receptor T cells form nonclassical and potent immune synapses driving rapid cytotoxicity. Proc. Natl. Acad. Sci. USA 115, E2068–E2076 (2018).
Roda-Navarro, P. & Álvarez-Vallina, L. Understanding the spatial topology of artificial immunological synapses assembled in T cell-redirecting strategies: a major issue in cancer immunotherapy. Front. Cell Dev. Biol. 7, 370 (2019).
Skokos, D. et al. A class of costimulatory CD28-bispecific antibodies that enhance the antitumor activity of CD3-bispecific antibodies. Sci. Transl. Med. 12, eaaw7888 (2020).
Salter, A. I. et al. Phosphoproteomic analysis of chimeric antigen receptor signaling reveals kinetic and quantitative differences that affect cell function. Sci. Signal. 11, eaat6753 (2018).
Ramello, M. C. et al. An immunoproteomic approach to characterize the CAR interactome and signalosome. Sci. Signal. 12, eaap9777 (2019).
Liu, Y. et al. Chimeric STAR receptors using TCR machinery mediate robust responses against solid tumors. Sci. Transl. Med. 13, eabb5191 (2021).
Harris, D. T. et al. Comparison of T cell activities mediated by human TCRs and CARs that use the same recognition domains. J. Immunol. 200, 1088–1100 (2018).
Gudipati, V. et al. Inefficient CAR-proximal signaling blunts antigen sensitivity. Nat. Immunol. 21, 848–856 (2020).
Wu, L., Wei, Q., Brzostek, J. & Gascoigne, N. R. J.Signaling from T cell receptors (TCRs) and chimeric antigen receptors (CARs) on T cells. Cell. Mol. Immunol. 17, 600–612 (2020).
Fry, T. J. et al. CD22-targeted CAR T cells induce remission in B-ALL that is naive or resistant to CD19-targeted CAR immunotherapy. Nat. Med. 24, 20–28 (2018).
Bossi, G., Buisson, S., Oates, J., Jakobsen, B. K. & Hassan, N. J. ImmTAC-redirected tumour cell killing induces and potentiates antigen cross-presentation by dendritic cells. Cancer Immunol. Immunother. 63, 437–448 (2014).
Wu, T. et al. Quantification of epitope abundance reveals the effect of direct and cross-presentation on influenza CTL responses. Nat. Commun. 10, 2846 (2019).
Huang, J. et al. A single peptide–major histocompatibility complex ligand triggers digital cytokine secretion in CD4+ T cells. Immunity 39, 846–857 (2013).
Nerreter, T. et al. Super-resolution microscopy reveals ultra-low CD19 expression on myeloma cells that triggers elimination by CD19 CAR-T. Nat. Commun. 10, 3137 (2019).
Pillai, V. et al. CAR T-cell therapy is effective for CD19-dim B-lymphoblastic leukemia but is impacted by prior blinatumomab therapy. Blood Adv. 3, 3539–3549 (2019).
Majzner, R. G. et al. Tuning the antigen density requirement for CAR T-cell activity. Cancer Discov. 10, 702–723 (2020).
Stone, J. D., Aggen, D. H., Schietinger, A., Schreiber, H. & Kranz, D. M. A sensitivity scale for targeting T cells with chimeric antigen receptors (CARs) and bispecific T-cell engagers (BiTEs). Oncoimmunology 1, 863–873 (2012).
Deng, Q. et al. Characteristics of anti-CD19 CAR T cell infusion products associated with efficacy and toxicity in patients with large B cell lymphomas. Nat. Med. 26, 1878–1887 (2020).
Nobles, C. L. et al. CD19-targeting CAR T cell immunotherapy outcomes correlate with genomic modification by vector integration. J. Clin. Invest. 130, 673–685 (2020).
Sheih, A. et al. Clonal kinetics and single-cell transcriptional profiling of CAR-T cells in patients undergoing CD19 CAR-T immunotherapy. Nat. Commun. 11, 219 (2020).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Zacharakis, N. et al. Immune recognition of somatic mutations leading to complete durable regression in metastatic breast cancer. Nat. Med. 24, 724–730 (2018).
Keskin, D. B. et al. Neoantigen vaccine generates intratumoral T cell responses in phase Ib glioblastoma trial. Nature 565, 234–239 (2019).
Hilf, N. et al. Actively personalized vaccination trial for newly diagnosed glioblastoma. Nature 565, 240–245 (2019).
Platten, M. et al. A vaccine targeting mutant IDH1 in newly diagnosed glioma. Nature https://doi.org/10.1038/s41586-021-03363-z (2021).
Linnemann, C. et al. High-throughput epitope discovery reveals frequent recognition of neo-antigens by CD4+ T cells in human melanoma. Nat. Med. 21, 81–85 (2015).
Germano, G. et al. CD4 T cell dependent rejection of beta 2 microglobulin null mismatch repair deficient tumors. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-20-0987 (2021).
Oh, D. Y. et al. Intratumoral CD4+ T cells mediate anti-tumor cytotoxicity in human bladder cancer. Cell 181, 1612–1625.e13 (2020).
Hwang, M. S. et al. Targeting loss of heterozygosity for cancer-specific immunotherapy. Proc. Natl Acad. Sci. USA 118, e2022410118 (2021).
Gonzalez-Galarza, F. F. et al. Allele Frequency Net Database (AFND) 2020 update: gold-standard data classification, open access genotype data and new query tools. Nucleic Acids Res. 48, D783–D788 (2020).
Maiers, M., Gragert, L. & Klitz, W. High-resolution HLA alleles and haplotypes in the United States population. Hum. Immunol. 68, 779–788 (2007).
Somasundaram, R. et al. Human leukocyte antigen-A2-restricted CTL responses to mutated BRAF peptides in melanoma patients. Cancer Res. 66, 3287–3293 (2006).
Malekzadeh, P. et al. Neoantigen screening identifies broad TP53 mutant immunogenicity in patients with epithelial cancers. J. Clin. Invest. 129, 1109–1114 (2019).
Linard, B. et al. A ras-mutated peptide targeted by CTL infiltrating a human melanoma lesion. J. Immunol. 168, 4802–4808 (2002).
Andersen, M. H. et al. Immunogenicity of constitutively active V599EBRaf. Cancer Res. 64, 5456–5460 (2004).
Cancer Genome Atlas Research Network et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Acknowledgements
We thank J. Cohen, M. Miller, S. Paul and K. Wright for insightful discussions, and E. Cook for assistance with Figs. 1 and 2. This work was supported by the Virginia and D. K. Ludwig Fund for Cancer Research, Lustgarten Foundation for Pancreatic Cancer Research, Commonwealth Fund, Burroughs Wellcome Career Award for Medical Scientists, Bloomberg∼Kimmel Institute for Cancer Immunotherapy, Bloomberg Philanthropies, Mark Foundation for Cancer Research, NIH Cancer Center Support Grant P30 CA006973 and National Cancer Institute grant R37 CA230400. A.H.P., B.J.M., J.D. and S.R.D. were supported by NIH T32 grant GM136577. M.F.K. was supported by NIH T32 grant AR048522.
Author information
Authors and Affiliations
Contributions
A.H.P. wrote the original draft of the manuscript. A.H.P., M.S.H., M.F.K., E.H.-C.H., J.D., S.R.D., B.J.M., C.B., D.M.P., S.B.G., N.P., K.W.K., B.V. and S.Z. reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The Johns Hopkins University has filed patent applications related to technologies described in this paper, on which A.H.P., M.S.H., E.H.-C.H., J.D., B.J.M., N.P., K.W.K., B.V., D.M.P. and S.Z. are listed as inventors: HLA-restricted epitopes encoded by somatically mutated genes (15/560,241, USPTO; 2016235251, European Patent Office); MANAbodies and Methods of Using (16/614,005, USPTO; 18802867.4, European Patent Office); MANAbodies Targeting Tumor Antigens and Methods of Using (63/059,638, USPTO; PCT/US2020/065617, World IP Organization). These applications include methods for identifying public neoantigens and the development of therapeutic agents that target these neoantigens. B.V., K.W.K. and N.P. are founders of Thrive Earlier Detection. K.W.K. and N.P. are consultants to Thrive Earlier Detection and were on its Board of Directors. B.V., K.W.K., N.P. and S.Z. own equity in Exact Sciences. B.V., K.W.K., N.P., S.Z. and D.M.P. are founders of, and serve or may serve as consultants to, ManaT Bio, and hold or may hold equity in ManaT Holdings, LLC. B.V., K.W.K., N.P. and S.Z. are founders of, hold equity in and serve as consultants to Personal Genome Diagnostics. S.Z. has a research agreement with BioMed Valley Discoveries. S.B.G. is a founder of and holds equity in AMS. K.W.K. and B.V. are consultants to Sysmex, Eisai and Cage Pharma and hold equity in Cage Pharma. B.V. is also a consultant to Catalio. K.W.K., B.V., S.Z. and N.P. are consultants to and hold equity in NeoPhore. N.P. is an advisor to and holds equity in Cage Pharma. C.B. is a consultant to DePuy Synthes and Bionaut Labs. The companies named above, as well as other companies, have licensed previously described technologies related to the work described in this paper from Johns Hopkins University. B.V., K.W.K., S.Z., N.P. and C.B. are inventors on some of these technologies. Licenses to these technologies are or will be associated with equity or royalty payments to the inventors, as well as to Johns Hopkins University. The terms of all of these arrangements are being managed by Johns Hopkins University in accordance with its conflict of interest policies. M.F.K. received personal fees from Bristol Myers Squibb and Celltrion. D.M.P. reports grant and patent royalties through his institution from Bristol Myers Squibb, a grant from Compugen, stock from Trieza Therapeutics and Dracen Pharmaceuticals and founder equity from Potenza; is a consultant for Aduro Biotech, Amgen, AstraZeneca (MedImmune/Amplimmune), Bayer, DNAtrix, Dynavax Technologies Corporation, Ervaxx, FLX Bio, Rock Springs Capital, Janssen, Merck, Tizona and Immunomic Therapeutics; is on the scientific advisory board of Five Prime Therapeutics, Catalio and WindMIL; and is on the board of directors for Dracen Pharmaceuticals.
Additional information
Peer review information Nature Cancer thanks Michal Bassani-Sternberg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Table
Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Pearlman, A.H., Hwang, M.S., Konig, M.F. et al. Targeting public neoantigens for cancer immunotherapy. Nat Cancer 2, 487–497 (2021). https://doi.org/10.1038/s43018-021-00210-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-021-00210-y
This article is cited by
-
The potential and promise for clinical application of adoptive T cell therapy in cancer
Journal of Translational Medicine (2024)
-
Cancer therapy with antibodies
Nature Reviews Cancer (2024)
-
Recent Advances and Challenges in Cancer Treatment with Car T Cell Therapy: A Novel Anti-cancer Strategy
BioNanoScience (2024)
-
Adoptive cellular immunotherapy for solid neoplasms beyond CAR-T
Molecular Cancer (2023)
-
Neoantigen-targeted TCR-engineered T cell immunotherapy: current advances and challenges
Biomarker Research (2023)