Abstract
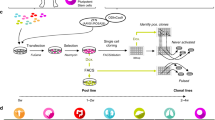
Patient-derived xenografts (PDXs) are high-fidelity in vivo tumor models that accurately reflect many key aspects of human cancer. In contrast to cancer cell lines or genetically engineered mouse models, the utility of PDXs has been limited by the inability to perform targeted genome editing of these tumors. To address this limitation, we have developed methods for CRISPR-Cas9 editing of PDXs using a tightly regulated, inducible Cas9 vector that does not require in vitro culture for selection of transduced cells. We demonstrate the utility of this platform in PDXs to analyze genetic dependencies by targeted gene disruption and to analyze mechanisms of acquired drug resistance by site-specific gene editing, using templated homology-directed repair. This flexible system has broad application to other explant models and substantially augments the utility of PDXs as genetically programmable models of human cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Unmodified gel images for Figs. 1–3 and 5 are provided as Source Data files (Unmodified_Gels_Fig1 - Unmodified_Gels_Fig4). Numerical source data for Main Figs. 1–6 and Extended Data Figs. 1–4 are provided as Source Data files (SourceData_Fig1 – SourceData_Fig6 and SourceData_ExtendedData_Fig1 – SourceData_ExtendedData_Fig4). All other data supporting the findings of this study are available from the corresponding authors upon reasonable request.
Code availability
The computer code that supports the findings of this study is available from the corresponding author upon request.
References
Townsend, E. C. et al. The public repository of xenografts enables discovery and randomized phase II-like trials in mice. Cancer Cell 29, 574–586 (2016).
Krepler, C. et al. A comprehensive patient-derived xenograft collection representing the heterogeneity of melanoma. Cell Rep. 21, 1953–1967 (2017).
Bruna, A. et al. A biobank of breast cancer explants with preserved intra-tumor heterogeneity to screen anticancer compounds. Cell 167, 260–274 (2016).
Drapkin, B. J. et al. Genomic and functional fidelity of small cell lung cancer patient-derived xenografts. Cancer Discovery 8, 600–615 (2018).
Daniel, V. C. et al. A primary xenograft model of small-cell lung cancer reveals irreversible changes in gene expression imposed by culture in vitro. Cancer Res. 69, 3364–3373 (2009).
Poirier, J. T. et al. DNA methylation in small cell lung cancer defines distinct disease subtypes and correlates with high expression of EZH2. Oncogene 34, 5869–5878 (2015).
Guo, S. et al. Molecular pathology of patient tumors, patient-derived Xenografts, and cancer cell lines. Cancer Res. 76, 4619–4626 (2016).
Puca, L. et al. Patient-derived organoids to model rare prostate cancer phenotypes. Nat. Commun. 9, 2404 (2018).
Beshiri, M. L. et al. A PDX/organoid biobank of advanced prostate cancers captures genomic and phenotypic heterogeneity for disease modeling and therapeutic screening. Clin. Cancer Res. 24, 4332–4345 (2018).
Gao, H. et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat. Med. 21, 1318–1325 (2015).
Gardner, E. E. et al. Chemosensitive relapse in small cell lung cancer proceeds through an EZH2-SLFN11 axis. Cancer Cell 31, 286–299 (2017).
Lallo, A., Schenk, M. W., Frese, K. K., Blackhall, F. & Dive, C. Circulating tumor cells and CDX models as a tool for preclinical drug development. Transl. Lung Cancer Res. 6, 397–408 (2017).
Sánchez-Rivera, F. J. & Jacks, T. Applications of the CRISPR-Cas9 system in cancer biology. Nat. Rev. Cancer 15, 387–395 (2015).
Ventura, A. & Dow, L. E. Modeling cancer in the CRISPR Era. Annu. Rev. Cancer Biol. 2, 111–131 (2018).
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017).
Tschaharganeh, D. F., Lowe, S. W., Garippa, R. J. & Livshits, G. Using CRISPR-Cas to study gene function and model disease in vivo. FEBS J. 283, 3194–3203 (2016).
Siolas, D. & Hannon, G. J. Patient-derived tumor xenografts: transforming clinical samples into mouse models. Cancer Res. 73, 5315–5319 (2013).
Meca-Cortés, O. et al. CRISPR-Cas9-mediated knock-in application in cell therapy: a non-viral procedure for Bystander treatment of glioma in mice. Mol. Ther. Nucleic Acids 8, 395–403 (2017).
Ablain, J., Durand, E. M., Yang, S., Zhou, Y. & Zon, L. I. A CRISPR-Cas9 vector system for tissue-specific gene disruption in zebrafish. Dev. Cell 32, 756–764 (2015).
Kabadi, A. M., Ousterout, D. G., Hilton, I. B. & Gersbach, C. A. Multiplex CRISPR-Cas9-based genome engineering from a single lentiviral vector. Nucleic Acids Res. 42, e147 (2014).
Dow, L. E. et al. Conditional reverse Tet-transactivator mouse strains for the efficient induction of TRE-regulated transgenes in mice. PLoS ONE 9, e95236–11 (2014).
Dow, L. E. et al. Inducible in vivo genome editing with CRISPR-Cas9. Nat. Biotechnol. 33, 390–394 (2015).
Gangopadhyay, S. A. et al. Precision control of CRISPR-Cas9 using small molecules and light. Biochemistry 58, 234–244 (2019).
González, F. et al. An iCRISPR platform for rapid, multiplexable, and inducible genome editing in human pluripotent stem cells. Stem Cell 15, 215–226 (2014).
Wu, M. et al. Conditional gene knockout and reconstitution in human iPSCs with an inducible Cas9 system. Stem Cell Res. 29, 6–14 (2018).
Cao, J. et al. An easy and efficient inducible CRISPR/Cas9 platform with improved specificity for multiple gene targeting. Nucleic Acids Res. 60, gkw660–10 (2016).
Verma, N. et al. TET proteins safeguard bivalent promoters from de novo methylation in human embryonic stem cells. Nat. Genet. 50, 83–95 (2017).
Gaines, P. & Wojchowski, D. M. pIRES-CD4t, a dicistronic expression vector for MACS- or FACS-based selection of transfected cells. Biotechniques 26, 683–688 (1999).
Bonini, C. et al. HSV-TK gene transfer into donor lymphocytes for control of allogeneic graft-versus-leukemia. Science 276, 1719–1724 (1997).
Wang, X. et al. A transgene-encoded cell surface polypeptide for selection, in vivo tracking, and ablation of engineered cells. Blood 118, 1255–1263 (2011).
Hirenallur-Shanthappa, D. K., Ramírez, J. A. & Iritani, B. M. in Immunodeficient Mice: The Backbone of Patient-Derived Tumor Xenograft Models. Patient Derived Tumor Xenograft Models 57–73, Ch. 5 (Elsevier, 2016). https://doi.org/10.1016/B978-0-12-804010-2.00005-9
Koretzky, G. A. Multiple roles of CD4 and CD8 in T cell activation. J. Immunol. 185, 2643–2644 (2010).
Loew, R., Heinz, N., Hampf, M., Bujard, H. & Gossen, M. Improved Tet-responsive promoters with minimized background expression. BMC Biotechnol. 10, 81 (2010).
Zhou, X., Vink, M., Klaver, B., Berkhout, B. & Das, A. T. Optimization of the Tet-On system for regulated gene expression through viral evolution. Gene Ther. 13, 1382–1390 (2006).
Kim, S., Bae, T., Hwang, J. & Kim, J.-S. Rescue of high-specificity Cas9 variants using sgRNAs with matched 5′ nucleotides. Genome Biol. 18, 218 (2017).
Zafra, M. P. et al. Optimized base editors enable efficient editing in cells, organoids and mice. Nat. Biotechnol. 36, 888–893 (2018).
Heinz, N. et al. Retroviral and transposon-based Tet-regulated all-in-one vectors with reduced background expression and improved dynamic range. Hum. Gene Ther. 22, 166–176 (2011).
Kumar, M., Keller, B., Makalou, N. & Sutton, R. E. Systematic determination of the packaging limit of lentiviral vectors. Hum. Gene Ther. 12, 1893–1905 (2001).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Helling, B. et al. A specific CD4 epitope bound by tregalizumab mediates activation of regulatory T cells by a unique signaling pathway. Immunol. Cell Biol. 93, 396–405 (2015).
Cawthorne, C., Swindell, R., Stratford, I. J., Dive, C. & Welman, A. Comparison of doxycycline delivery methods for Tet-inducible gene expression in a subcutaneous xenograft model. J. Biomol. Tech. 18, 120–123 (2007).
Mao, Z., Bozzella, M., Seluanov, A. & Gorbunova, V. Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair 7, 1765–1771 (2008).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Dow, L. E. et al. A pipeline for the generation of shRNA transgenic mice. Nat. Protoc. 7, 374–393 (2012).
Beard, C., Hochedlinger, K., Plath, K., Wutz, A. & Jaenisch, R. Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis 44, 23–28 (2006).
Charles, J. P. et al. Monitoring the dynamics of clonal tumour evolution in vivo using secreted luciferases. Nat. Commun. 5, 3981 (2014).
van Rijn, S. et al. Functional multiplex reporter assay using tagged Gaussia luciferase. Sci. Rep. 3, 1046 (2013).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Wroblewska, A. et al. Protein barcodes enable high-dimensional single-cell CRISPR screens. Cell. 175, 1141–1155 (2018).
Poirier, J. T. CRISPR libraries and screening. Prog. Mol. Biol. Transl. Sci. 152, 69–82 (2017).
Winters, I. P. et al. Multiplexed in vivo homology-directed repair and tumor barcoding enables parallel quantification of Kras variant oncogenicity. Nat. Commun. 8, 2053 (2017).
Kleinstiver, B. P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Casini, A. et al. A highly specific SpCas9 variant is identified by in vivo screening in yeast. Nature 36, 265–271 (2018).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Sastry, L., Johnson, T., Hobson, M. J., Smucker, B. & Cornetta, K. Titering lentiviral vectors: comparison of DNA, RNA and marker expression methods. Gene Ther. 9, 1155–1162 (2002).
Zhang, B. et al. The significance of controlled conditions in lentiviral vector titration and in the use of multiplicity of infection (MOI) for predicting gene transfer events. Genet. Vaccines Ther. 2, 6 (2004).
Wang, M. et al. Humanized mice in studying efficacy and mechanisms of PD-1-targeted cancer immunotherapy. FASEB J. 32, 1537–1549 (2018).
Gray, J. T. & Zolotukhin, S. Design and construction of functional AAV vectors. Methods Mol. Biol. 807, 25–46 (2011).
Campeau, E. et al. A versatile viral system for expression and depletion of proteins in mammalian cells. PLoS ONE 4, e6529 (2009).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Cossarizza, A. et al. Guidelines for the use of flow cytometry and cell sorting in immunological studies. Eur. J. Immunol. 47, 1584–1797 (2017).
Acknowledgements
We thank the MSKCC Flow Cytometry Core for their technical assistance, members of the Antitumor Assessment Core Facility for assistance with in vivo experiments and all members of the Rudin laboratory for critical comments. We thank the CCIB at Massachusetts General Hospital for the use of the CCIB DNA Core Facility and the Boston Children’s Hospital Viral Core (Core grant 5P30EY012196, NEI). This work was supported by National Institutes of Health U01 CA199215, U24 CA213274, P01 CA129243, P30 CA008748 and R01 CA197936 (C.M.R. and J.T.P.).
Author information
Authors and Affiliations
Contributions
C.H.H. performed most of the experiments described here and drafted the manuscript. E.A.C. performed key in vitro assays including editing of GFP in A549GFP cells and of EGFR in PC9 cells. E.dS. directs the xenograft core facility in which the EGFR gene editing in vivo under drug selection was performed. N.S.S. and A.Q.V. provided technical assistance. G.H. provided statistical analysis. C.M.R. and J.T.P. supervised the project and helped draft the manuscript. All authors contributed to editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.M.R. has consulted regarding oncology drug development with AbbVie, Amgen, Ascentage, AstraZeneca, BMS, Celgene, Daiichi Sankyo, Genentech/Roche, Ipsen, Loxo and PharmaMar and is on the scientific advisory boards of Elucida, Bridge and Harpoon. All other authors have no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
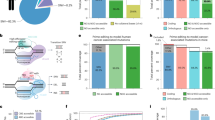
Extended Data Fig. 1 Truncated CD4T is a size-efficient selectable marker for flow cytometry and immunomagnetic selection.
a, Domain structure of wildtype human CD4 and truncated constructs CD4ΔD2-4;ΔIC and CD4ΔD3-4;ΔIC (hereafter CD4T), where the indicated deletions are replaced by flexible linkers. Heatmap depicts flow cytometry staining intensity of commercially available α-CD4 antibodies, which target the indicated extracellular domains of CD4, to the indicated CD4 constructs. S, signal peptide; TM, transmembrane domain; IC, intracellular domain; MFI, mean fluorescence intensity b, Flow cytometry analysis of A549 cells expressing CD4T and enriched by fluorescence activated cell sorting (FACS). Data is representative of three independent experiments with similar results. c, Immunomagnetic enrichment of A549 cells expressing CD4T, with the indicated enrichment factors (above). Mean percent CD4 positive is displayed for n = 3 independent cell culture replicates. Data is representative of two independent experiments with similar results. Error bars are SD. d, α-CD4 staining strategy with domain D1 and D3 targeting antibodies to differentiate CD4T and full-length CD4 using flow cytometry. A mixture of GFP-negative cells expressing full length CD4 and GFP-positive cells expressing CD4T were stained with the indicated α-CD4 antibody clones. Double positive cells were exclusively GFP-negative (expressing full length CD4) and cells single positive for the domain D1 targeting α-CD4 antibody were exclusively GFP-positive (expressing CD4T). Data is representative of n = 3 independent cell culture replicates from a single experiment. Data for experiments in panels b-d are available as source data (SourceData_ExtendedData_Fig1).
Extended Data Fig. 2 Competition assay utilizing sgTrack vectors effectively determines fitness effects of gene disruption.
a, Competition assay of sgRPA1-2 in A549 with no Cas9, constitutive Cas9 expression from lentiCas9-Blast, or dox-inducible Cas9 expression from pSpCTRE (left, middle and right panels, respectively). Competition assays were performed with (solid) or without (dashed) dox. Mean fluorescence expression is displayed for n = 3 independent cell culture replicates from a single experiment. Error bars are SD, with SD < plotting character not drawn. b, Formulas for fitness score and log ratio (left). Comparison of fitness score and log ratio analysis methods (right). Mean fitness score or mean log ratio are displayed for n = 3 independent cell culture replicates from a single experiment. Error bars are SD, with SD < plotting character not drawn. c, In vitro competition assays in A549 with sgRPA1-1, sgKRAS-1, and sgKRAS-2 (left, middle and right panels, respectively). Log ratio calculations and line assignments are as described in panel b. Mean log ratio is displayed for n = 3 independent cell culture replicates from a single experiment. Error bars are SD, with SD < plotting character not drawn. Data for experiments in panels a-c are available as source data (SourceData_ExtendedData_Fig2).
Extended Data Fig. 3 CD4T induction in SpCTRE PDXs and gating strategy for in vivo competition assay analysis.
a, Representative flow cytometry analysis of CD4T induction for the competition assay with sgRPA1-1 in MSK-LX369. Data is representative of n = 5 mice. b, Flow cytometry analysis of sgRPA1-1 (mCherry) and sgNTC (GFP) in tumors from panel a. Cells are first gated on CD4T positive or CD4T induced populations as shown in panel a. Data is representative of n = 5 mice. c, Log ratio of MSK-LX369 competition assays with the indicated sgRNAs for control or dox-treated mice gated on either CD4T positive or CD4T induced populations. Mean log ratio is displayed for n = 5 mice. Error bars are SD, with SD < plotting character not drawn. A two-sided Wilcoxon rank sum test was used to determine statistical significance. d, Percent of CD4T induced cells from dox-treated mice for competition assays with the indicated sgRNAs in the SpCTRE PDXs MSK-LX369, JHU-LX55a, and MSK-LX29 (left, middle, and right panels, respectively). Mean percent CD4 positive cells is displayed for n = 4 or n = 5 mice as indicated. Error bars are SD. e, Mean time to sacrifice in days for parental and SpCTRE PDXs for n = 2 to n = 46 mice. Error bars are SD, with SD < plotting character not drawn. Data for experiments in panels a-e are available as source data (SourceData_ExtendedData_Fig3).
Extended Data Fig. 4 Evaluation of acquired osimertinib resistance driven by rAAV-mediated homology-directed repair in PC9 cells in vitro.
a, Number of PC9 cells under the indicated conditions for n = 2 independent cell culture replicates from a single experiment. Error bars are SD, with SD < plotting character not drawn. b, Osimertinib dose response curves of PC9 cells from the rAAV, osimertinib or control, DMSO treatment arms from panel a. Mean relative luminescence is displayed for n = 3 independent cell culture replicates from a single experiment. Error bars are SD, with SD < plotting character not drawn. c, d, Sequencing analysis of PC9 cells from the rAAV, osimertinib group at (c) t = 7 and (d) t = 35 days for n = 2 biologically independent replicates from panel a. Data for experiments in panels a-d are available as source data (SourceData_ExtendedData_Fig4).
Supplementary information
Supplementary Tables
Supplementary Tables S1–S3
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Rights and permissions
About this article
Cite this article
Hulton, C.H., Costa, E.A., Shah, N.S. et al. Direct genome editing of patient-derived xenografts using CRISPR-Cas9 enables rapid in vivo functional genomics. Nat Cancer 1, 359–369 (2020). https://doi.org/10.1038/s43018-020-0040-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-0040-8
This article is cited by
-
TWIST1 is a critical downstream target of the HGF/MET pathway and is required for MET driven acquired resistance in oncogene driven lung cancer
Oncogene (2024)
-
Patient-derived xenograft models in cancer therapy: technologies and applications
Signal Transduction and Targeted Therapy (2023)
-
Dissecting metastasis using preclinical models and methods
Nature Reviews Cancer (2023)
-
Combined proteomics and CRISPR‒Cas9 screens in PDX identify ADAM10 as essential for leukemia in vivo
Molecular Cancer (2023)
-
CRISPR in cancer biology and therapy
Nature Reviews Cancer (2022)