Abstract
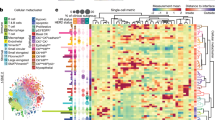
Genomic alterations shape cell phenotypes and the structure of tumor ecosystems in poorly defined ways. To investigate these relationships, we used imaging mass cytometry to quantify the expression of 37 proteins with subcellular spatial resolution in 483 tumors from the METABRIC cohort. Single-cell analysis revealed cell phenotypes spanning epithelial, stromal and immune types. Distinct combinations of cell phenotypes and cell–cell interactions were associated with genomic subtypes of breast cancer. Epithelial luminal cell phenotypes separated into those predominantly impacted by mutations and those affected by copy number aberrations. Several features of tumor ecosystems, including cellular neighborhoods, were linked to prognosis, illustrating their clinical relevance. In summary, systematic analysis of single-cell phenotypic and spatial correlates of genomic alterations in cancer revealed how genomes shape both the composition and architecture of breast tumor ecosystems and will enable greater understanding of the phenotypic impact of genomic alterations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
IMC data, including cell masks and processed single-cell data, have been deposited to the Image Data Resource (https://idr.openmicroscopy.org/) under accession code idr0076 (see https://idr.openmicroscopy.org/about/download.html). Previously published METABRIC copy number, gene expression, miRNA and targeted sequencing data that were reanalyzed here are available under accession codes EGAS00000000083, EGAS00000000122 and EGAS00001001753 at the European Genome–Phenome Archive (http://www.ebi.ac.uk/ega/). Updated METABRIC clinical data analyzed here are available as part of the supplementary information in ref. 14. All other data supporting the findings of this study are available from the corresponding authors upon reasonable request.
Code availability
In-house image preprocessing scripts are available at https://github.com/BodenmillerGroup/imctools. Other analysis code is available from the authors upon request.
References
Perou, C. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Ali, H. R. et al. Genome-driven integrated classification of breast cancer validated in over 7,500 samples. Genome Biol. 15, 431 (2014).
Ciriello, G. et al. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 45, 1127–1133 (2013).
Pereira, B. et al. The somatic mutation profiles of 2,433 breast cancers refines their genomic and transcriptomic landscapes. Nat. Commun. 7, 11479 (2016).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Wagner, J. et al. A single-cell atlas of the tumor and immune ecosystem of human breast cancer. Cell 177, 1330–1345 (2019).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Dvinge, H. et al. The shaping and functional consequences of the microRNA landscape in breast cancer. Nature 497, 378–382 (2013).
Schulz, D. et al. Simultaneous multiplexed imaging of mRNA and proteins with subcellular resolution in breast cancer tissue samples by mass cytometry. Cell Syst. 6, 25–36 (2018).
Damond, N. et al. A map of human type 1 diabetes progression by imaging mass cytometry. Cell Metab. 29, 755–768 (2019).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Haubold, C. et al. Segmenting and tracking multiple dividing targets using ilastik. Adv. Anat. Embryol. Cell Biol. 219, 199–229 (2016).
Rueda, O. M. et al. Dynamics of breast-cancer relapse reveal late-recurring ER-positive genomic subgroups. Nature 567, 399–404 (2019).
Gottfried, E. et al. Expression of CD68 in non-myeloid cell types. Scand. J. Immunol. 67, 453–463 (2008).
Costa, A. et al. Fibroblast heterogeneity and immunosuppressive environment in human breast cancer. Cancer Cell 33, 463–479 (2018).
Mitra, A. K. et al. MicroRNAs reprogram normal fibroblasts into cancer-associated fibroblasts in ovarian cancer. Cancer Discov. 2, 1100–1108 (2012).
Stingl, J. & Caldas, C. Molecular heterogeneity of breast carcinomas and the cancer stem cell hypothesis. Nat. Rev. Cancer 7, 791 (2007).
Schapiro, D. et al. histoCAT: analysis of cell phenotypes and interactions in multiplex image cytometry data. Nat. Methods 14, 873–876 (2017).
Akavia, U. D. et al. An integrated approach to uncover drivers of cancer. Cell 143, 1005–1017 (2010).
Slamon, D. J. et al. Studies of the HER-2/neu proto-oncogene in human breast and ovarian cancer. Science 244, 707–712 (1989).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Facciabene, A. et al. Tumour hypoxia promotes tolerance and angiogenesis via CCL28 and Treg cells. Nature 475, 226–230 (2011).
Dawson, S. J., Rueda, O. M., Aparicio, S. & Caldas, C. A new genome-driven integrated classification of breast cancer and its implications. EMBO J. 32, 617–628 (2013).
Cristescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362, eaar3593 (2018).
Bodenmiller, B. Multiplexed epitope-based tissue imaging for discovery and healthcare applications. Cell Syst. 2, 225–238 (2016).
Ali, H. R. et al. Association between CD8+ T-cell infiltration and breast cancer survival in 12 439 patients. Ann. Oncol. 25, 1536–1543 (2014).
Ali, H. R., Chlon, L., Pharoah, P. D., Markowetz, F. & Caldas, C. Patterns of immune infiltration in breast cancer and their clinical implications: a gene-expression-based retrospective study. PLoS Med. 13, e1002194 (2016).
Reis-Filho, J. S. & Pusztai, L. Gene expression profiling in breast cancer: classification, prognostication, and prediction. Lancet 378, 1812–1823 (2011).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Patten, D. K. et al. Enhancer mapping uncovers phenotypic heterogeneity and evolution in patients with luminal breast cancer. Nat. Med. 24, 1469–1480 (2018).
Barsoum, I. B., Koti, M., Siemens, D. R. & Graham, C. H. Mechanisms of hypoxia-mediated immune escape in cancer. Cancer Res. 74, 7185–7190 (2014).
Bristow, R. G. & Hill, R. P. Hypoxia, DNA repair and genetic instability. Nat. Rev. Cancer 8, 180–192 (2008).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
Kalluri, R. & Zeisberg, M. Fibroblasts in cancer. Nat. Rev. Cancer 6, 392–401 (2006).
Ozdemir, B. C. et al. Depletion of carcinoma-associated fibroblasts and fibrosis induces immunosuppression and accelerates pancreas cancer with reduced survival. Cancer Cell 25, 719–734 (2014).
Rhim, A. D. et al. Stromal elements act to restrain, rather than support, pancreatic ductal adenocarcinoma. Cancer Cell 25, 735–747 (2014).
Rakha, E. A. et al. Tubular carcinoma of the breast: further evidence to support its excellent prognosis. J. Clin. Oncol. 28, 99–104 (2010).
Mukherjee, A. et al. Associations between genomic stratification of breast cancer and centrally reviewed tumour pathology in the METABRIC cohort. NPJ Breast Cancer 4, 5 (2018).
Dunning, M. J., Smith, M. L., Ritchie, M. E. & Tavare, S. beadarray: R classes and methods for Illumina bead-based data. Bioinformatics 23, 2183–2184 (2007).
Kononen, J. et al. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat. Med. 4, 844–847 (1998).
Catena, R., Ozcan, A., Jacobs, A., Chevrier, S. & Bodenmiller, B. AirLab: a cloud-based platform to manage and share antibody-based single-cell research. Genome Biol. 17, 142 (2016).
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387 (2018).
Van Gassen, S. et al. FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 87, 636–645 (2015).
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Phipson, B. & Smyth, G. K. Permutation P-values should never be zero: calculating exact P-values when permutations are randomly drawn. Stat. Appl. Genet. Mol. Biol. 9, Article39 (2010).
Newson, R. B. Frequentist q-values for multiple-test procedures. Stata J. 10, 568–584 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Parker, J. S. et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 27, 1160–1167 (2009).
Yu, G. & He, Q. Y. ReactomePA: an R/Bioconductor package for Reactome pathway analysis and visualization. Mol. Biosyst.12, 477–479 (2016).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010).
Sondka, Z. et al. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nat. Rev. Cancer 18, 696–705 (2018).
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V. & Egozcue, J. J. Microbiome datasets are compositional: and this is not optional. Front. Microbiol. 8, 2224 (2017).
Blows, F. et al. Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med. 7, e1000279 (2010).
Friedman, J. H., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33 (1), 1–22 (2010).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016).
Acknowledgements
H.R.A. was supported by a Cancer Research UK (CRUK) Clinician Scientist Fellowship. We thank N. De Souza for critical review of the manuscript. The research of B.B. is funded by an SNSF R’Equip grant, an SNSF Assistant Professorship grant, an NIH grant (UC4 DK108132) and by the European Research Council (ERC) under the European Union’s Seventh Framework Program (FP/2007-2013)/ERC grant agreement 336921. The laboratories of B.B. and C.C. were supported by the CRUK IMAXT Grand Challenge for this work.
Author information
Authors and Affiliations
Consortia
Contributions
H.R.A., C.C. and B.B. conceived the study; IMC experiments were performed by H.W.J.; H.R.A. conducted data analysis with assistance from J.F.; H.R.A., C.C. and B.B. wrote the manuscript with contributions from E.D.; V.R.T.Z. built the preprocessing pipeline; H.B. and E.P. constructed the tissue microarrays; S.-F.C. and O.M.R. generated and processed genomic data; O.M.R. and E.P. collected and curated clinical data; C.C. and S.A. co-led the METABRIC consortium; C.C. and B.B. co-directed the study; and all authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.C. is a member of the External Science Panel of AstraZeneca, and his laboratory has received research grants (administered by the University of Cambridge) from Genentech, Roche, AstraZeneca and Servier. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Spatial distribution of cell types.
Representative cell masks annotated by protein expression levels (left) and by inferred cell identities separated by tumour versus stroma (right). For protein expression levels, mean counts per cell were normalised relative to all cells analysed. White scale bars (top left) represent 100 µm.
Extended Data Fig. 2 Definition of myofibroblasts and miRNA-related pathway enrichment.
a, Box-and-whisker plot of the distribution of SMA expression by stromal cell types (n = 141, 818 cells). Boxes represent the interquartile range. Lines dividing boxes indicate the median, and vertical lines represent range of expression from the 1st to 99th percentile. b, Pathway enrichment analysis of genes (hypergeometric test; adjusted for multiple comparisons) linked to miRNAs positively correlated with myofibroblast and vascular smooth muscle cell proportions (n = 371 tumours; related to Fig. 4).
Extended Data Fig. 3 Cell neighbours depend on cell phenotype.
a, Stacked bar plot depicting phenotypic composition of cell neighbours separately by each cell phenotype for all tumours. b, Stacked bar plots depicting the phenotypic composition of cells neighbouring all stromal phenotypes separately by IntClust subtype, illustrating patterns of stromal cell enrichment among IntClust subtypes (related to Fig. 4). c, Heatmap of median expression values for stromal cells based on both whole-cell and 3-pixel annular segmentation methods; rows and columns ordered by hierarchical clustering using Ward’s method.
Extended Data Fig. 4 Robustness of cell phenotype associations with tumour subtypes.
a, Example heatmap of Spearman’s correlation statistics between 100 FlowSOM groups and Phenograph clusters based on median protein expression values, to illustrate the methodology for mapping FlowSOM groups to cell phenotypes assigned using Phenograph. Solid squares indicate largest positive correlations. Rows and columns are ordered by hierarchical clustering using Ward’s method (comparison based on n = 479, 844 cells). b, Bar chart on the left shows the number of cells assigned to the Phenograph cell phenotypes. Bar chart on the right depicts frequency of FlowSOM groups mapped to each Phenograph-clustered phenotypes, arising from 100 runs of FlowSOM each generating 100 groups (10,000 groups mapped in total). This illustrates that most phenotypes were assigned by both methods and that the frequency of mapped groups was related to the number of cells represented by each phenotype. c, Bar charts comparing patterns of association (tested using linear regression) with breast cancer molecular subtypes between mapped cell phenotypes (from 100 runs of FlowSOM each generating 100 groups as in panel b) and the Phenograph-assigned phenotypes. Green and red bars distinguish between enrichment and depletion of a given cell phenotype for each molecular subtype. ‘Original’ on the x-axis indicates associations based on the Phenograph clustering methodology.
Extended Data Fig. 5 Patterns of cell phenotype and cell-cell interaction enrichment among breast cancer molecular subtypes.
a, Patterns of enrichment among metaclusters defined by combining subsets of Phenograph-assigned cell types by descriptive label (for example, fibroblasts, myofibroblasts, T cells). tSNE map on the right indicates position by cell identity (median values for 22 metaclusters computed based on n = 479, 844 cells). Coloured markers indicate significant enrichment. Depicted associations derived from linear regression (n = 390 tumours; two-sided tests; adjusted for multiple comparisons), restricted to those with an adjusted p-value < 0.05, were identified by linear models where the dark grey background is proportional to the derived point estimate, providing an indication of the relative strength of the association. b, Co-occurrence plots of cell-cell interactions identified by permutation testing and found to be significantly enriched (p-value < 0.05 after adjustment for multiple comparisons) among the molecular subtypes indicated (limited to samples that contain both cells for a given interaction; range of n between 86 and 361 tumours for depicted associations). Rows and columns correspond to cell types in the same order as labelled on the y-axis.
Extended Data Fig. 6 Distribution of cell neighbourhoods by molecular subtypes of breast cancer.
tSNE reference maps representing cell types as light grey makers (tSNE map based on median marker expression values derived from n = 479, 844 cells). Neighbourhood enrichment for each cell type within tumour molecular subtypes was determined by fitting a linear model taking mean neighbourhood values as the dependent variable and tumour molecular subtype as independent variable. Coloured markers indicate those significantly enriched within a given subtype (p-value < 0.05). Dark grey background is proportional to the point estimate from the linear model, providing an indication of the strength (size) of the association.
Extended Data Fig. 7 Associations between cell phenotypes and genome-wide copy number aberrations.
Scatter plots depicting adjusted p-values derived from linear models testing for associations between cell phenotype and genomic alterations (n = 390 tumours; two-sided; adjusted for multiple comparisons). Coloured points represent significant associations (red, gains; blue, losses). Depicted points are restricted to those associated with positive coefficients. Shown are cell phenotypes most affected by copy-number aberrations.
Extended Data Fig. 8 Relationship between genomic instability and cell type.
Box and whisker plots of the distributions of cell types as proportions versus quartiles of genomic instability. Boxes represent the interquartile range. Lines dividing boxes indicate the median, and vertical lines represent range of expression from the 1st to 99th percentile. The p-values were derived from two-sided Kruskal-Wallis tests; depicted are those cell phenotypes with a p-value < 0.05 (adjusted for multiple comparisons; n = 404 tumours).
Extended Data Fig. 9 Cell type abundance explained variance by genomic data.
Explained variances (right y-axis) for each of a series of four linear models are depicted as connected circles (n = 357 tumours). Distributions of cell type proportions per tumour (left y-axis) depicted as boxes and whiskers. Boxes represent the interquartile range. Lines dividing boxes indicate the median, and vertical lines represent range of expression from the 1st to 99th percentile.
Extended Data Fig. 10 Influence of tissue sampling on cell phenotype estimates.
Stacked bar plot depicting cell phenotype composition per tissue-microarray spot for a subset of fifteen tumours represented by at least two spots. The order of columns was determined using single-linkage hierarchical clustering. Patient IDs are on the x axis. Grey bars highlight where two tissue spots from the same tumour cluster together.
Supplementary information
Supplementary Tables
Supplementary Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ali, H.R., Jackson, H.W., Zanotelli, V.R.T. et al. Imaging mass cytometry and multiplatform genomics define the phenogenomic landscape of breast cancer. Nat Cancer 1, 163–175 (2020). https://doi.org/10.1038/s43018-020-0026-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-0026-6
This article is cited by
-
CellCharter reveals spatial cell niches associated with tissue remodeling and cell plasticity
Nature Genetics (2024)
-
Multiplex protein imaging in tumour biology
Nature Reviews Cancer (2024)
-
SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
Nature Communications (2024)
-
RGS5 maintaining vascular homeostasis is altered by the tumor microenvironment
Biology Direct (2023)
-
An end-to-end workflow for multiplexed image processing and analysis
Nature Protocols (2023)