Abstract
Half of the children diagnosed with neuroblastoma (NB) have high-risk disease, disproportionately contributing to overall childhood cancer-related deaths. In addition to recurrent gene mutations, there is increasing evidence supporting the role of epigenetic deregulation in disease pathogenesis. Yet, comprehensive cis-regulatory network descriptions from NB are lacking. Here, using genome-wide H3K27ac profiles across 60 NBs, covering the different clinical and molecular subtypes, we identified four major super-enhancer-driven epigenetic subtypes and their underlying master regulatory networks. Three of these subtypes recapitulated known clinical groups; namely, MYCN-amplified, MYCN non-amplified high-risk and MYCN non-amplified low-risk NBs. The fourth subtype, exhibiting mesenchymal characteristics, shared cellular identity with multipotent Schwann cell precursors, was induced by RAS activation and was enriched in relapsed disease. Notably, CCND1, an essential gene in NB, was regulated by both mesenchymal and adrenergic regulatory networks converging on distinct super-enhancer modules. Overall, this study reveals subtype-specific super-enhancer regulation in NBs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw and processed data from ChIP–seq, HiChIP, ATAC–seq and single-cell RNA-seq analyses have been deposited in the Gene Expression Omnibus under the SuperSeries number GSE136209 (containing GSE128463, GSE135756, GSE136208, GSE136279, GSE151902 and GSE158130) and at the European Genome-phenome Archive under the accession numbers EGAS00001004551 and EGAS00001004552. The epigenomic tracks and processed data are accessible for visualization at https://nbseB087.dkfz.de. The single-cell dataset for the developing mouse adrenal medulla was accessed from GSE99933. HiC data were accessed from GSE63525. Gene expression data for The Cancer Genome Atlas, Genotype-Tissue Expression Project and TARGET consortium were obtained from https://toil.xenahubs.net/download/TcgaTargetGtex_rsem_gene_tpm.gz. Transcriptomic profiles of NB from the NRC consortium were obtained from https://r2.amc.nl/. The DepMap (version DeepMap19Q2) was downloaded from https://depmap.org/. The mouse organogenesis data were from https://oncoscape.v3.sttrcancer.org/atlas.gs.washington.edu.mouse.rna/downloads. Source data are provided with this paper.
Code availability
A comprehensive Snakemake pipeline to reproduce all of the results reported in this study has been made available at https://github.com/hdsu-bioquant/project_NB_SE.
References
Furlan, A. Multipotent peripheral glial cells generate neuroendocrine cells of the adrenal medulla. Science 357, eaal3753 (2017).
Shimada, H. et al. Histopathologic prognostic factors in neuroblastic tumors: definition of subtypes of ganglioneuroblastoma and an age-linked classification of neuroblastomas. J. Natl Cancer Inst. 73, 405–416 (1984).
Matthay, K. K. et al. Neuroblastoma. Nat. Rev. Dis. Primer 2, 16078 (2016).
Maris, J. M. Recent advances in neuroblastoma. N. Engl. J. Med. 362, 2202–2211 (2010).
Janoueix-Lerosey, I. et al. Somatic and germline activating mutations of the ALK kinase receptor in neuroblastoma. Nature 455, 967–970 (2008).
Mosse, Y. P. et al. Identification of ALK as a major familial neuroblastoma predisposition gene. Nature 455, 930–935 (2008).
Schwab, M., Westermann, F., Hero, B. & Berthold, F. Neuroblastoma: biology and molecular and chromosomal pathology. Lancet Oncol. 4, 472–480 (2003).
Pugh, T. J. et al. The genetic landscape of high-risk neuroblastoma. Nat. Genet. 45, 279–284 (2013).
Ross, R. A., Spengler, B. A. & Biedler, J. L. Coordinate morphological and biochemical interconversion of human neuroblastoma cells. J. Natl Cancer Inst. 71, 741–747 (1983).
Boeva, V. et al. Heterogeneity of neuroblastoma cell identity defined by transcriptional circuitries. Nat. Genet. 49, 1408–1413 (2017).
van Groningen, T. et al. Neuroblastoma is composed of two super-enhancer-associated differentiation states. Nat. Genet. 49, 1261–1266 (2017).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Molenaar, J. J., van Sluis, P., Boon, K., Versteeg, R. & Caron, H. N. Rearrangements and increased expression of cyclin D1 (CCND1) in neuroblastoma. Genes. Chromosomes Cancer 36, 242–249 (2003).
Huang, S. et al. ZNF423 is critically required for retinoic acid-induced differentiation and is a marker of neuroblastoma outcome. Cancer Cell 15, 328–340 (2009).
Henrich, K. O. et al. CAMTA1, a 1p36 tumor suppressor candidate, inhibits growth and activates differentiation programs in neuroblastoma cells. Cancer Res. 71, 3142–3151 (2011).
Oldridge, D. A. et al. Genetic predisposition to neuroblastoma mediated by a LMO1 super-enhancer polymorphism. Nature 528, 418–421 (2015).
Lee, D. D. & Seung, H. S. Learning the parts of objects by non-negative matrix factorization. Nature 401, 788–791 (1999).
Quintero, A. et al. ShinyButchR: interactive NMF-based decomposition workflow of genome-scale datasets Biol. Methods Protoc. https://doi.org/10.1093/biomethods/bpaa0227 (2020).
Rajbhandari, P. et al. Cross-cohort analysis identifies a TEAD4 ↔ MYCN positive feedback loop as the core regulatory element of high-risk neuroblastoma. Cancer Discov. 8, 582–599 (2018).
Janoueix-Lerosey, I. et al. Overall genomic pattern is a predictor of outcome in neuroblastoma. J. Clin. Oncol. 27, 1026–1033 (2009).
ACEseq—allele specific copy number estimation from whole genome sequencing. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/210807v1.full (2017).
Ishaque, N. et al. Whole genome sequencing puts forward hypotheses on metastasis evolution and therapy in colorectal cancer. Nat. Commun. 9, 4782 (2018).
Mak, M. P. et al. A patient-derived, pan-cancer EMT signature identifies global molecular alterations and immune target enrichment following epithelial-to-mesenchymal transition. Clin. Cancer Res. 22, 609–620 (2016).
Saint-Andre, V. et al. Models of human core transcriptional regulatory circuitries. Genome Res. 26, 385–396 (2016).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Durbin, A. D. et al. Selective gene dependencies in MYCN-amplified neuroblastoma include the core transcriptional regulatory circuitry. Nat. Genet. 50, 1240–1246 (2018).
Bhasin, N., Maynard, T. M., Gallagher, P. A. & LaMantia, A.-S. Mesenchymal/epithelial regulation of retinoic acid signaling in the olfactory placode. Dev. Biol. 261, 82–98 (2003).
Li, C.-W. et al. Epithelial–mesenchymal transition induced by TNF-α requires NF-κB-mediated transcriptional upregulation of Twist1. Cancer Res. 72, 1290–1300 (2012).
Carro, M. S. et al. The transcriptional network for mesenchymal transformation of brain tumours. Nature 463, 318–325 (2010).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Ryl, T. et al. Cell-cycle position of single MYC-driven cancer cells dictates their susceptibility to a chemotherapeutic drug. Cell Syst. 5, 237–250.e8 (2017).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Biedler, J. L., Helson, L. & Spengler, B. A. Morphology and growth, tumorigenicity, and cytogenetics of human neuroblastoma cells in continuous culture. Cancer Res. 33, 2643–2652 (1973).
Ciccarone, V., Spengler, B. A., Meyers, M. B., Biedler, J. L. & Ross, R. A. Phenotypic diversification in human neuroblastoma cells: expression of distinct neural crest lineages. Cancer Res. 49, 219–225 (1989).
Anastassiou, D. et al. Human cancer cells express Slug-based epithelial–mesenchymal transition gene expression signature obtained in vivo. BMC Cancer 11, 529 (2011).
Chen, Y. A., Lu, I. L. & Tsai, J. W. Contactin-1/F3 regulates neuronal migration and morphogenesis through modulating RhoA activity. Front. Mol. Neurosci. 11, 422 (2018).
Ackermann, S. et al. A mechanistic classification of clinical phenotypes in neuroblastoma. Science 362, 1165–1170 (2018).
Eleveld, T. F. et al. Relapsed neuroblastomas show frequent RAS–MAPK pathway mutations. Nat. Genet. 47, 864–871 (2015).
Mulholland, D. J. et al. Pten loss and RAS/MAPK activation cooperate to promote EMT and metastasis initiated from prostate cancer stem/progenitor cells. Cancer Res. 72, 1878–1889 (2012).
Loboda, A. et al. A gene expression signature of RAS pathway dependence predicts response to PI3K and RAS pathway inhibitors and expands the population of RAS pathway activated tumors. BMC Med. Genomics 3, 26 (2010).
Bollag, G. et al. Loss of NF1 results in activation of the Ras signaling pathway and leads to aberrant growth in haematopoietic cells. Nat. Genet. 12, 144–148 (1996).
Voice, J. K., Klemke, R. L., Le, A. & Jackson, J. H. Four human Ras homologs differ in their abilities to activate Raf-1, induce transformation, and stimulate cell motility. J. Biol. Chem. 274, 17164–17170 (1999).
Haigis, K. M. et al. Differential effects of oncogenic K-Ras and N-Ras on proliferation, differentiation and tumor progression in the colon. Nat. Genet. 40, 600–608 (2008).
Way, G. P. et al. Machine learning detects pan-cancer ras pathway activation in The Cancer Genome Atlas. Cell Rep. 23, 172–180 (2018).
Roson-Burgo, B., Sanchez-Guijo, F., Del Cañizo, C. & De Las Rivas, J. Insights into the human mesenchymal stromal/stem cell identity through integrative transcriptomic profiling. BMC Genomics 17, 944 (2016).
Ambros, I. M. et al. Role of ploidy, chromosome 1p, and Schwann cells in the maturation of neuroblastoma. N. Engl. J. Med. 334, 1505–1511 (1996).
Shimada, H. et al. Terminology and morphologic criteria of neuroblastic tumors. Cancer 86, 349–363 (1999).
Dagogo-Jack, I. & Shaw, A. T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 15, 81–94 (2018).
Schramm, A. et al. Mutational dynamics between primary and relapse neuroblastomas. Nat. Genet. 47, 872–877 (2015).
Schmitt, M. & Pawlita, M. High-throughput detection and multiplex identification of cell contaminations. Nucleic Acids Res. 37, e119 (2009).
Hauser, S. et al. Isolation of novel multipotent neural crest-derived stem cells from adult human inferior turbinate. Stem Cells Dev. 21, 742–756 (2012).
Keshelava, N., Seeger, R. C., Groshen, S. & Reynolds, C. P. Drug resistance patterns of human neuroblastoma cell lines derived from patients at different phases of therapy. Cancer Res. 58, 5396–5405 (1998).
Blecher-Gonen, R. et al. High-throughput chromatin immunoprecipitation for genome-wide mapping of in vivo protein–DNA interactions and epigenomic states. Nat. Protoc. 8, 539–554 (2013).
Peifer, M. et al. Telomerase activation by genomic rearrangements in high-risk neuroblastoma. Nature 526, 700–704 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Landt, S. G. et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 38, 675–678 (2020).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Phanstiel, D. H., Boyle, A. P., Heidari, N. & Snyder, M. P. Mango: a bias-correcting ChIA-PET analysis pipeline. Bioinformatics 31, 3092–3098 (2015).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Sherwood, R. I. et al. Discovery of directional and nondirectional pioneer transcription factors by modeling DNase profile magnitude and shape. Nat. Biotechnol. 32, 171–178 (2014).
Lachmann, A., Giorgi, F. M., Lopez, G. & Califano, A. ARACNe-AP: gene network reverse engineering through adaptive partitioning inference of mutual information. Bioinformatics 32, 2233–2235 (2016).
Margolin, A. A. et al. ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinformatics 7, S7 (2006).
Alvarez, M. J. et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat. Genet. 48, 838–847 (2016).
Henrich, K. O. et al. Integrative genome-scale analysis identifies epigenetic mechanisms of transcriptional deregulation in unfavorable neuroblastomas. Cancer Res. 76, 5523–5537 (2016).
Zhang, W. et al. Comparison of RNA-seq and microarray-based models for clinical endpoint prediction. Genome Biol. 16, 133 (2015).
Worst, B. C. et al. Next-generation personalised medicine for high-risk paediatric cancer patients—the INFORM pilot study. Eur. J. Cancer 65, 91–101 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
DeLuca, D. S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Wu, D. & Smyth, G. K. Camera: a competitive gene set test accounting for inter-gene correlation. Nucleic Acids Res. 40, e133 (2012).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Acknowledgements
We thank the patients and their parents for making the tumor specimens that were analyzed in this study available. We also thank the German Neuroblastoma Biobank for providing these samples. This work was supported by the e:Med initiative (SYSMED-NB; grant number 01ZX1307D to F.W.), BMBF PRECiSe grant 021L0076B (to C.H. and A.K.S.), the German Cancer Consortium (DKTK) Joint Funding Program, ERACoSysMed grant OPTIMIZE-NB (to F.W.), European Union grant number 259348 (to F.W.), the German Cancer Research Center (DKFZ) Intramural Program for Interaction Projects (NCT3.0 ENHANCE to F.W., A.K.S., A.Q. and C.H.) and the DKFZ Heidelberg Center for Personalized Oncology (HIPO) and National Center for Tumor Diseases (NCT) Precision Oncology Program (F.W. and N. I.). M.G. was supported by doctoral fellowships of the German–Israeli Helmholtz Research School in Cancer Biology. We thank J. Kreth for technical assistance. We also acknowledge the DKFZ Genomics and Proteomics Core Facility for helping us to obtain high-quality data for the sequencing analyses.
Author information
Authors and Affiliations
Contributions
The study was conceived, designed and interpreted by M.G., A.K.S., D. Dreidax, A.Q., C.H. and F.W. The ChIP–seq, ATAC–seq and HiChIP experiments and analyses were performed by M.G., Y.-G.P. and D. Dreidax. All of the computational analyses, data processing and data analyses were done by A.K.S., A.Q. and C.H. The RNA-seq experiments were conducted by S.J. The cell viability studies and functional assays were conducted and analyzed by M.G., S.K., E.W., J.M. and E.A. Regulatable cell models were established and flow cytometry assays were performed by J.v.N. Technical assistance and bioinformatics support were provided by J.K., N.I., D. Doncevic., P.S and U.H.T. Assistance with the primary and relapsed tumor RNA expression data analyses was provided by K.-O.H., D.T.W.J. and S.M.P. Reagents, materials and analysis tools were contributed by S.J., E.A., R.V. and T.G.P.G. The manuscript was prepared by M.G., A.K.S., A.Q., D. Dreidax., C.H. and F.W., with support from K.-O.H. and S.M.P. All authors approved the current version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Super enhancers in neuroblastoma.
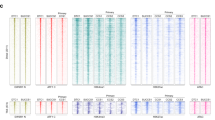
a, Quality parameters for the H3K27ac ChIP-seq analysis in neuroblastoma tumors (n=60) and cell lines (n=25), including (from top to bottom) - number of SEs per sample; number of core regulatory circuits (CRCs) identified in each sample; number of aligned reads in millions; number of peaks called by MACS2; fraction of reads in MACS2 peaks (FRiP); normalized strand correlation (NSC), relative strand correlation (RSC) and PCR bottleneck coefficient (PBC) as defined by ENCODE55; sample clinical annotations. b, Pairwise Spearman correlation and the corresponding p values between all of the QC parameters shown in (a) and signature activity scores measured in tumors (top panel) and cell lines (bottom panel). c, Saturation analysis of the super enhancers identified in neuroblastoma showing that we identify about two-thirds (120Mb) of SE genomic region of the expected total of 200Mb. We estimate that about 1000 samples will be required to achieve the power to identify almost all SEs in neuroblastoma (d, e) The percentage of NB tumor-specific SEs recovered from SEs identified from other tissue and cells, including SE identified in previous studies from neuroblastoma cell lines10,11 (expanded view of Fig. 1b) (f) Distribution of the expression of genes (n=1476) predicted to be regulated by the NB-specific SE across a panel of normal human tissues (n=7832, GTEx; green) and tumor samples (n=9843, TCGA and TARGET; red), including an independent NB cohort (n=162, TARGET). The box plot shows the first and third quartiles (top and bottom of the box), the median (line within the box), and the values at +/- 1.5 times the interquartile range of the first and third quartile (vertical lines).
Extended Data Fig. 2 Subtypes of neuroblastoma based on super enhancers activity.
a, Schematic representation of the Non-Negative Matrix Factorization (NMF). Any non-negative matrix, for instance, a gene expression matrix with rows as genes and columns as samples can be factorized into a matrix W and H which contains the contribution (activity) of each gene and sample toward each of the identified signatures respectively. b, Hierarchical clustering of the NB tumor (n=60) and cell lines (n=25) based on the H3K27ac signal over the NB SEs (n=1973 SEs). c, Combined NMF analysis of the NB tumor (n=60) and cell lines (n=25) based on the H3K27ac signal at the NB SEs. d), River-plot representation of the stability of the signatures extracted from the H3K27ac SE signal in tumors (left) and cell lines (right) (n=60 NB tumors and n=25 NB cell lines). The vertical axis represents the different factorization ranks (which equals to the number of signatures extracted, k=2 to 6) and the ribbons indicate the similarity of the signatures defined for different factorization ranks. For example, in cell lines, the MES signature (right-most ribbon) is stable for all factorization ranks. In tumors, the MES signature appears for factorization ranks k>2. e, MES signature activity scores in tumors and cell lines. In cell lines we observe two distinct population of cells with very high or low MES activity.
Extended Data Fig. 3 Robustness and specificity of the super enhancer derived subtypes of neuroblastomas.
a, Comparison of the subtype specifics signature activity scores defined from ChIP-seq data (signatureSE, x-axis) and RNA-seq data (signatureSE-Exp, y-axis) (n=47 NB samples) (b) Hierarchical clustering of the 579 NB bulk RNA-seq samples using the expression of the SE target genes (left, bottom) or the most variable genes (right, bottom). The annotation bars above the heatmap indicate clinical parameters (MYCN amplification / Stage / Age / Risk / Relapse) and the activity score to the four NMF signatures, as well as the tumor classification based on the highest activity score (NMF signature) (c) NMF analysis of the 579 NB bulk RNA-seq samples based on the expression of the 1476 most variable genes (equal to the number of unique SE target genes, see Fig. 2d). d, Enrichment analyses of the signature specific genes among the terms classified as cell migration and EMT (red circles in Fig. 1c), based on the NMF analysis of the 1476 most variable genes. P-values are computed using a one-sided Fisher’s exact test.
Extended Data Fig. 4 Validation of neuroblastoma super enhancer subtypes in two independent cohorts (NRC and TARGET).
a, NMF analysis based on the expression of the SE target genes in NB tumors from the NRC dataset (n=283 tumor samples, n=972 target genes) and (b) from the TARGET dataset (n=162 tumor samples, n=1428 target genes). c, d, Scatter plots indicating the correlation of the signature specific gene activity between NB tumor and (c) NRC dataset (n=972 target genes) or (d) the TARGET dataset (n=1428 target genes). Each dot represents a gene, and the x/y coordinates represent the contribution (or weight) of the gene to the corresponding signature. Pearson’s correlation values are indicated. e,f, Association analysis of known clinical and molecular variables to the (e) NRC dataset NMF signatures (n=5 signatures, n=10 clinical covariates) and to (f) TARGET dataset NMF signatures (n=5 signatures, n=20 clinical covariates). Arrows indicate the signatures that are most correlated to our MESSE-Exp signature. Unadjusted one-sided hypothesis Wilcoxon signed-rank test P-values are shown. g, Comparison of median RAS signature gene expression in samples assigned to NMF signatures in the NRC dataset (n=68 Sig1, n=49 Sig2, n=42 Sig3, n=35 Sig4, n=89 Sig5, NB samples from the NRC cohort) or (h) the TARGET dataset (n=20 Sig1, n=10 Sig2, n=57 Sig3, n=34 Sig4, n=41 Sig5, NB samples from the TARGET cohort). Two-sided t-test P-values computed between indicated groups are shown. The MES signature could be recovered (NRC-Sig4 and TARGET-Sig1/Sig2) from both independent datasets (NRC and TARGET).
Extended Data Fig. 5 Clinical and molecular features of the neuroblastoma super enhancer subtypes.
(a) Cox proportional hazards regression model showing the association of various clinical covariates and the epigenetic signatures of neuroblastoma with overall survival. Error bars represent the 95% confidence intervals and the indicated p values are from log rank test. Sample size for every group is indicated on the figure. (b) Comparison of signature activity scores and tumor purity as estimated using ACEseq21,22 method in the ChIP-seq (top) and RNA-seq (bottom) defined signatures. The Pearson correlation coefficient is indicated for each plot. The activity scores are discretized into high (red) and low (cyan) groups using kmeans (k=2) and the vertical line denotes 50% tumor purity. For all signatures in both cohort, majority of high (red) score samples also have high tumor purity. (n=44 NB samples from the ChIP-seq cohort (top), and n=124 NB samples from the RNA-seq cohort (top) for which purity scores were available) (c, d) Comparison of the activity scores for the MNA-LR and MNA-HR signatures, derived from SE signal (c), or SE target gene expression (d). (e) H3K27ac signal of signature specific SE (each box panel, top) and the target gene expression of the corresponding SEs (each box panel, bottom) across samples specific to each of the four subtypes. P values are computed using a two-sided Wilcoxon test.
Extended Data Fig. 6 Core regulatory circuits in neuroblastomas.
a, Example of a CRC comprising of SMAD3, MYC and FOSL2 in the SH-EP cell line (MES). ChIP-seq of H3K27ac (upper track) and ATAC-seq (middle track) at super enhancers. Footprints of the respective TFs are indicated as dots below (lower track). b, Distribution of the CRC TFs across primary tumors and cell lines (n=60 NB tumors and n=25 NB cell lines). Samples in the columns are ordered by the subtypes and CRC TFs in the rows are according to their frequency of occurrence in the subtypes (based on the row order of Fig. 3b). Clustering of TFs according to their frequency of occurrence in CRCs of tumors only (c) and cell lines only d, based on the respective SE-defined subgroups. The TFs are labelled according to the module definition of Fig. 3b.
Extended Data Fig. 7 The regulatory landscape in neuroblastoma.
a, Overview of the neuroblastoma (NB) transcription factor (TF) regulome construction and estimation of TF activity score in each of the four signatures identified from NB tumors (expanded view of Fig. 3a). The heatmap depicts the clustering based on TF activity scores across all samples. (b) The distribution of TF activity scores of TFs (n=24) annotated in the mesenchymal module (Fig. 3b) among the different subtypes. P-values are computed using a two-sided Wilcoxon test. c, Difference in chromatin accessibility (using ATAC-seq) (t-statistics) in footprints of CRC-TFs for mesenchymal (MES) SK-N-AS cell line versus adrenergic (ADRN) KELLY cell line colored by differential MES vs. ADRN TF activity (d) Fraction of chromatin interactions (using Hi-ChIP) involving mesenchymal-specific (purple, in SK-NAS) and adrenergic (green, in CLB-GA) super enhancers and enhancers. Regions were classified as MES and ADRN based on a differential binding analysis of 14 MES samples (6 cell lines and 8 tumors) against 71 ADRN samples (19 cell lines and 52 tumors). e, Distribution of the TF activity score of TFs selected for knockdown experiments (n=32 selected TF, blue) or not (n=33 non selected TF, yellow), showing that there is no bias in the TF selection. P-values are computed using a two-sided Wilcoxon test. f, Contingency table showing the number of ADRN/MES TFs (columns) vs. the number of TFs selected or not for knockdown experiments (rows). Pvalues are computed using a two-sided chi-square test which shows that there is no bias in the selection of validated TFs. The box plots in (b,e) shows the first and third quartiles (top and bottom of the box), the median (line within the box), and the values at +/- 1.5 times the interquartile range of the first and third quartile (vertical lines). Individual TFs are shown as dots.
Extended Data Fig. 8 Super enhancer controlled regulators and targets in neuroblastoma.
a–c, Using the data from the Cancer Dependency Map project30,77 - (a) Knockdown CERES score of CCND1 across multiple cancer types (n=23 cancer types from n=549 cancer cell lines). Lower values represent higher sensitivity of the cell to CCND1 knockdown. b, Comparison of CCND1 expression between neuroblastoma (n=25) and other cancer cell-lines (n=1175). P-values are computed using a two-sided Wilcoxon test. c, Expression of CCND1 in cancer cell lines (n=26 cancer types from n =1176 cancer cells). The box plots (a-c) shows the first and third quartiles (top and bottom of the box), the median (line within the box), and the values at +/- 1.5 times the interquartile range of the first and third quartile (vertical lines). Individual TFs are shown as dots. d, Footprint count of MES and ADRN CRC TFs located inside SE regions of NB-essential genes described in Fig. 13a in MES SK-N-AS, MYCN non-amplified ADRN CLB-GA and MYCN-amplified ADRN KELLY cells. e, Scatter plot showing, for each TF, the MES TF activity (x-axis) against the enrichment of its target genes in the genes differentially expressed between relapse and primary tumors (data from n=3 matched primary/relapse NB tumor pairs for n=1240 TFs) (y-axis, -log10(FDR)). Expanded view of Fig. 6b, including all transcription factors. Only the master transcription factors are labelled.
Extended Data Fig. 9 Molecular features of the mesenchymal subtype.
a, b, Scatter plots showing, for each of the (n=579 NB tumors) the Spearman correlation (rho) between the median expression of the (a) RAS signature genes or (b) JUN/FOS target genes (x-axis) and the activity score of the samples according to the four signatures derived from the expression matrix (MYCNSE-Exp; MESSE-Exp; MNA-LRSE-Exp; MNA-HRSE-Exp) (y-axis). Samples with mutations of RAS-pathway genes are highlighted with red crosses in (a). c, Scatter plot, similar to (a), using an independent RAS signature gene set, displaying a similar positive Spearman correlation (shown for n=579 NB tumors), (d) Spearman correlation (rho) of n=1000 random genesets of size equal to the RAS signature geneset with MES exposure, showing the significance of the observed correlation in (c). The permutation pvalue was computed as 1 + number of random correlation values greater or equal than the observed correlation value divided by the number of random sampling performed (e) Spearman correlation of the RAS activity scores computed using two independent RAS genesets, namely the Loboda et al. geneset (x-axis, used in panel a) and the Way et al. geneset. Samples (n=579) are colored according to their exposure to the MES signature; mutated samples are indicated along with the name of the mutated gene. (f) Venn diagram showing the intersection between the two RAS genesets used in this analysis. g, h, i, Spearman correlation of the expression of TNFRSF12A (CD266) used as a mesenchymal marker in Fig. 7e with (g) VIM, (h) FN1 and (i) the MESSE-Exp activity score across all NB samples (n=579) (j) The Spearman’s correlation distribution of TNFRSF12A with an independent mesenchymal gene signature (red) and 1000 random genesets of the same size (grey) (k) Expression of the MES marker gene TNFRSF12A (CD266) in NB cell lines (n=23) from the Cancer Dependency Map project and (l) in NB patient samples separated by high Mesenchymal (MES) and low Mesenchymal (ADRN) signature activity. The box plot shows the first and third quartiles (top and bottom of the box), the median (line within the box), and the values at +/- 1.5 times the interquartile range of the first and third quartile (vertical lines). Values beyond this range are shown as dots. P-values are computed using a two-sided Wilcoxon test. m, Scatter plot of gene detected and total RNA count in single cells from HRAS induced and control. The cells are colored by the estimated mitochondrial gene expression.
Extended Data Fig. 10 Reanalysis of all major results using only the high purity tumor samples.
Comparison of In-house SEs to In-house SEs derived from high purity samples. a, Percentage of NB tumor-specific SEs (in-house SEs) recovered from SEs identified in previous studies from neuroblastoma cell lines, and from high purity NB tumor-specific SEs (high purity in-house SEs). b, Pearson’s correlation of sample activity scores derived from the complete ChIP-seq cohort NMF (n=41 out of 60 NB samples) (rows) to the high purity (columns) ChIP-seq cohort NMF (n=41 high purity NB samples). Likely, due to a smaller number of samples, we now obtain five stable signatures in the high purity cohort instead of four (full cohort) from the NMF analysis (see columns), two of which uniquely correspond to the MYCN signature, while the rest have a one-to-one correspondence with the MES, MNA-HR and MNA-LR signatures from the full cohort (rows). c, Pearson’s correlation of sample activity scores derived from the complete RNA-seq cohort NMF (n=111 out of 579 NB samples) (rows) to the high purity (columns) RNA-seq cohort NMF (n = 111 high purity NB samples). In (b,c), Pearson’s correlation values and two-sided P values are shown. Shaded bands represent 95% confidence intervals of linear regression slopes. d, UMAP projection of mouse adrenal medulla cells at E12.5; colors indicate the inferred cell type based on marker genes (same as Fig. 5a). e, Projection of high purity NB tumor transcriptomic profiles (n=111 NB samples) (colored dots) onto the landscape shown in (d) defined by the mouse adrenal medulla cells (grey dots). The tumor samples are colored according to (d) and the quantification of the most frequent neighboring single cells is shown in the inset bar plot. f, Proportion of high purity MES samples among samples mapping to cell types from the mouse adrenal medulla (n=17 out of 93 high purity NB samples assigned to the Chromaffin and SCPs clusters, no MES samples were assigned to the Bridge, Dividing Cells mitotic, and Sympathoblasts clusters). g, h, Correlation between Mesenchymal activity scores and RAS activity scores using the published signatures from Loboda et.al (g) and Way et. al (h) using only the samples with tumor purity >50%. Spearman’s rank correlation values and two-sided P-values are shown.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Tables
Supplementary Tables 1–10.
Source data
Source Data Fig. 1
Statistical and numerical source data
Source Data Fig. 2
Statistical and numerical source data
Source Data Fig. 3
Statistical and numerical source data
Source Data Fig. 4
Statistical and numerical source data
Source Data Fig. 5
Statistical and numerical source data
Source Data Fig. 6
Statistical and numerical source data
Source Data Fig. 7
Statistical and numerical source data
Source Data Fig. 7
Uncropped western blot
Source Data Extended Data Fig. 1
Statistical and numerical source data
Source Data Extended Data Fig. 2
Statistical and numerical source data
Source Data Extended Data Fig. 3
Statistical and numerical source data
Source Data Extended Data Fig. 4
Statistical and numerical source data
Source Data Extended Data Fig. 5
Statistical and numerical source data
Source Data Extended Data Fig. 6
Statistical and numerical source data
Source Data Extended Data Fig. 7
Statistical and numerical source data
Source Data Extended Data Fig. 8
Statistical and numerical source data
Source Data Extended Data Fig. 9
Statistical and numerical source data
Source Data Extended Data Fig. 10
Statistical and numerical source data
Rights and permissions
About this article
Cite this article
Gartlgruber, M., Sharma, A.K., Quintero, A. et al. Super enhancers define regulatory subtypes and cell identity in neuroblastoma. Nat Cancer 2, 114–128 (2021). https://doi.org/10.1038/s43018-020-00145-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00145-w
This article is cited by
-
Identification of MYCN non-amplified neuroblastoma subgroups points towards molecular signatures for precision prognosis and therapy stratification
British Journal of Cancer (2024)
-
Etiology of super-enhancer reprogramming and activation in cancer
Epigenetics & Chromatin (2023)
-
Development and validation of a 21-gene prognostic signature in neuroblastoma
Scientific Reports (2023)
-
Identification of a novel eighteen-gene signature of recurrent metastasis neuroblastoma
Journal of Molecular Medicine (2023)
-
SOX11 regulates SWI/SNF complex components as member of the adrenergic neuroblastoma core regulatory circuitry
Nature Communications (2023)