Abstract
Despite progress in immunotherapy, identifying patients that respond has remained a challenge. Through analysis of whole-exome and targeted sequence data from 5,449 tumors, we found a significant correlation between tumor mutation burden (TMB) and tumor purity, suggesting that low tumor purity tumors are likely to have inaccurate TMB estimates. We developed a new method to estimate a corrected TMB (cTMB) that was adjusted for tumor purity and more accurately predicted outcome to immune checkpoint blockade (ICB). To identify improved predictive markers together with cTMB, we performed whole-exome sequencing for 104 lung tumors treated with ICB. Through comprehensive analyses of sequence and structural alterations, we discovered a significant enrichment in activating mutations in receptor tyrosine kinase (RTK) genes in nonresponding tumors in three immunotherapy treated cohorts. An integrated multivariable model incorporating cTMB, RTK mutations, smoking-related mutational signature and human leukocyte antigen status provided an improved predictor of response to immunotherapy that was independently validated.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Whole-exome sequencing data that support the findings of this study have been deposited in the database of Genotypes and Phenotypes (dbGaP) and the European Genome-phenome Archive under accession codes phs001940.v1.p1 and EGAS00001003892, respectively (cohort 1). Source data for the TCGA tumor samples were retrieved from http://cancergenome.nih.gov. WES-derived somatic mutation calls from the TCGA PanCancer Atlas MC3 project were retrieved from the NCI Genomic Data Commons46 (https://gdc.cancer.gov/about-data/publications/mc3-2017). Previously published genomic data, reanalyzed here, were obtained from dbGaP under accession code phs000980.v1.p1 (cohort 2) and from the CBioPortal for Cancer Genomics2,6 (https://www.cbioportal.org/study/summary?id=nsclc_pd1_msk_2018 and https://www.cbioportal.org/study/summary?id=tmb_mskcc_2018). All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Somatic mutations were identified using the VariantDx software44. Missense mutations were evaluated for their potential as cancer drivers by CHASMplus (v.1.0.0)45 with consequence prediction using CRAVAT (v.4.3)47. HLA genotyping was performed using OptiType (v.1.2)48. MHC class I prediction of neoantigens used netMHCpan49 and netCTLpan50. Mutational signatures were quantified using the deconstructSigs R package (v.1.8.0). Somatic copy number profiles were estimated using CNV-kit54, FACETS (v.0.5.0)56 and as previously described13. POLYSOLVER was applied to detect somatic mutations in class I HLA genes33. LOHHLA was applied to determine loss of HLA genes32. Further statistical analyses were performed using the SPSS software program (v.25.0.0) and R v.3.2 and higher.
References
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Rizvi, H. et al. Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J. Clinical Oncol. 36, 633–641 (2018).
Hellmann, M. D. et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. New Eng. J. Med 378, 2093–2104 (2018).
Hellmann, M. D. et al. Genomic features of response to combination immunotherapy in patients with advanced non-small-cell lung cancer. Cancer Cell 33, 843–852 (2018).
Yarchoan, M., Hopkins, A. & Jaffee, E. M. Tumor mutational burden and response rate to PD-1 inhibition. New Eng. J. Med. 377, 2500–2501 (2017).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Raphael, B. J., Dobson, J. R., Oesper, L. & Vandin, F. Identifying driver mutations in sequenced cancer genomes: computational approaches to enable precision medicine. Genome Med. 6, 5 (2014).
Shi, W. et al. Reliability of whole-exome sequencing for assessing intratumor genetic heterogeneity. Cell Rep. 25, 1446–1457 (2018).
Viray, H. et al. A prospective, multi-institutional diagnostic trial to determine pathologist accuracy in estimation of percentage of malignant cells. Arch. Pathol. Lab. Med. 137, 1545–1549 (2013).
Gettinger, S. et al. Impaired HLA class I antigen processing and presentation as a mechanism of acquired resistance to immune checkpoint inhibitors in lung cancer. Cancer Disc. 7, 1420–1435 (2017).
Chowell, D. et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science 359, 582–587 (2018).
Wood, D. E. et al. A machine learning approach for somatic mutation discovery. Science Transl. Med. 10, eaar7939 (2018).
Anagnostou, V. et al. Evolution of neoantigen landscape during immune checkpoint blockade in non-small cell lung cancer. Cancer Disc. 7, 264–276 (2017).
Miao, D. et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat. Genet. 50, 1271–1281 (2018).
Forde, P. M. et al. Neoadjuvant PD-1 blockade in resectable lung cancer. New Engl. J. Med 378, 1976–1986 (2018).
Krause, D. S. & Van Etten, R. A. Tyrosine kinases as targets for cancer therapy. New Engl. J. Med. 353, 172–187 (2005).
Liu, C. et al. BRAF inhibition increases tumor infiltration by T cells and enhances the antitumor activity of adoptive immunotherapy in mice. Clinical Cancer Res. 19, 393–403 (2013).
Shen, J. et al. ARID1A deficiency promotes mutability and potentiates therapeutic antitumor immunity unleashed by immune checkpoint blockade. Nat. Medicine 24, 556–562 (2018).
Peng, W. et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Disc. 6, 202–216 (2016).
Kato, S. et al. Hyperprogressors after immunotherapy: analysis of genomic alterations associated with accelerated growth rate. Clinical Cancer Res. 23, 4242–4250 (2017).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. New Engl. J. Med. 13, 2016 (2016).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Gao, J. et al. Loss of IFN-gamma pathway genes in tumor cells as a mechanism of resistance to anti-CTLA-4 therapy. Cell 167, 397–404 (2016).
Knijnenburg, T. A. et al. Genomic and molecular landscape of DNA damage repair deficiency across the Cancer Genome Atlas. Cell Rep. 23, 239–254 (2018).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic beta-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Van Allen, E. M. et al. Genomic correlates of response to CTLA4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Bassani-Sternberg, M. et al. Direct identification of clinically relevant neoepitopes presented on native human melanoma tissue by mass spectrometry. Nature Commun. 7, 13404 (2016).
Luksza, M. et al. A neoantigen fitness model predicts tumour response to checkpoint blockade immunotherapy. Nature 551, 517–520 (2017).
Smith, K. N. et al. Persistent mutant oncogene-specific T cells in two patients benefitting from anti-PD-1. J. Immunother. Cancer 7, 40 (2019).
McGranahan, N. et al. Allele-specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271 (2017).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
Shin, D. S. et al. Primary resistance to PD-1 blockade mediated by JAK1/2 mutations. Cancer Disc. 7, 188–201 (2016).
Koyama, S. et al. Adaptive resistance to therapeutic PD-1 blockade is associated with upregulation of alternative immune checkpoints. Nature Commun. 7, 10501 (2016).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Cristescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362, eaar3593 (2018).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Sabari, J. K. et al. PD-L1 expression, tumor mutational burden, and response to immunotherapy in patients with MET exon 14 altered lung cancers. Ann. Oncol. 29, 2085–2091 (2018).
Hu-Lieskovan, S. et al. Improved antitumor activity of immunotherapy with BRAF and MEK inhibitors in BRAF(V600E) melanoma. Science Trans. Med. 7, 279ra241 (2015).
Roh, W. et al. Integrated molecular analysis of tumor biopsies on sequential CTLA-4 and PD-1 blockade reveals markers of response and resistance. Science Trans. Med. 9, eaah3560 (2017).
Anagnostou, V. et al. Dynamics of tumor and immune responses during immune checkpoint blockade in non-small cell lung cancer. Cancer Res. 79, 1214–1225 (2019).
Anagnostou, V. et al. Immuno-oncology trial endpoints: capturing clinically meaningful activity. Clinical Cancer Res. 23, 4959–4969 (2017).
Jones, S. et al. Personalized genomic analyses for cancer mutation discovery and interpretation. Science Trans. Med. 7, 283ra253 (2015).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Ellrott, K. et al. Scalable open science approach for mutation calling of tumor exomes using multiple genomic pipelines. Cell Syst. 6, 271–281 (2018).
Masica, D. L. et al. CRAVAT 4: cancer-related analysis of variants toolkit. Cancer Res. 77, e35–e38 (2017).
Szolek, A. et al. OptiType: precision HLA typing from next-generation sequencing data. Bioinformatics 30, 3310–3316 (2014).
Nielsen, M. & Andreatta, M. NetMHCpan-3.0; improved prediction of binding to MHC class I molecules integrating information from multiple receptor and peptide length datasets. Genome Med. 8, 33 (2016).
Stranzl, T., Larsen, M. V., Lundegaard, C. & Nielsen, M. NetCTLpan: pan-specific MHC class I pathway epitope predictions. Immunogenetics 62, 357–368 (2010).
Kim, Y., Sidney, J., Pinilla, C., Sette, A. & Peters, B. Derivation of an amino acid similarity matrix for peptide: MHC binding and its application as a Bayesian prior. BMC Bioinformatics 10, 394 (2009).
Balasubramanian, S. et al. Using ALoFT to determine the impact of putative loss-of-function variants in protein-coding genes. Nature Commun. 8, 382 (2017).
Rosenthal, R. deconstructSigs: identifies signatures present in a tumor sample. R version 1.8.0 https://CRAN.R-project.org/package=deconstructSigs (2016).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput. Biol. 12, e1004873 (2016).
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Shannon, C. E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Aran, D., Sirota, M. & Butte, A. J. Systematic pan-cancer analysis of tumour purity. Nature Commun. 6, 8971 (2015).
Niknafs, N., Beleva-Guthrie, V., Naiman, D. Q. & Karchin, R. Subclonal hierarchy inference from somatic mutations: automatic reconstruction of cancer evolutionary trees from multi-region next generation sequencing. PLoS Comput. Biol. 11, e1004416 (2015).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830 e814 (2018).
Sidney, J., Peters, B., Frahm, N., Brander, C. & Sette, A. HLA class I supertypes: a revised and updated classification. BMC Immunol. 9, 1 (2008).
Carlson, C. S. et al. Using synthetic templates to design an unbiased multiplex PCR assay. Nature Commun. 4, 2680 (2013).
Acknowledgements
We thank members of our laboratories for critical review of the manuscript. This work was supported in part by US National Institutes of Health grants (nos. CA121113 to V.E.V. and V.A., CA006973 to D.M.P. and V.E.V. and CA180950 to V.E.V.), the Commonwealth Foundation (to V.E.V. and S.B.B.), the Bloomberg-Kimmel Institute for Cancer Immunotherapy (to V.A., P.M.F., J.R.B., D.M.P. and V.E.V.), the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (to V.E.V. and S.B.B.), the Eastern Cooperative Oncology Group—American College of Radiology Imaging Network (to V.A.), MacMillan Foundation (to V.A.), the V Foundation (to V.A., V.E.V. and S.B.B.), Swim Across America (to V.A.), the William R. Brody Faculty Scholarship (to R.K.), the SU2C-ACS Lung Cancer Dream Team (to P.M.F. and E.G.), PRIME Oncology (to J.N.), the SU2C DCS International Translational Cancer Research Dream Team Grant (no. SU2C-AACR-DT1415 to V.E.V), the SU2C-LUNGevity-American Lung Association Lung Cancer Interception Dream Team (to J.B. and V.E.V.), the VARI-SU2C Epigenetics Dream Team Grant (to S.B.B.), the Allegheny Health Network—Johns Hopkins Research Fund (to V.A. and V.E.V.), the LUNGevity Foundation (to V.A. and P.M.F.), the Jim Toth Foundation (to J.B. and S.B.B.), the Rising Tide Foundation (to J.B. and S.B.B.) and the Mark Foundation for Cancer Research (to V.E.V.). Stand Up To Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research.
Author information
Authors and Affiliations
Contributions
V.Anagnostou and V.E.V. conceived and designed the study. V.Anagnostou, K.Ma., J.N., K.H., K.Mo., F.L., M.L., S.R., J.E.R., K.N.S., V.Adleff, K.R., Z.B., L.R., B.L., J.F., C.L.H., D.S.E., C.G., F.V., P.I., A.S.B., E.G., M.V.B., J.R.B. and P.M.F. acquired data. V.Anagnostou, N.N., D.C.B, J.R.W., Q.K.L., R.K., D.M.P., S.B.B., J.R.B., R.B.S., P.M.F. and V.E.V. analyzed and interpreted data. All authors have approved the submitted version and are accountable for their contributions.
Corresponding authors
Ethics declarations
Competing interests
V.A. receives research funding from Bristol-Meyer Squibb. P.M.F. receives research funding for clinical trials from Bristol-Myers Squibb, AstraZeneca/MedImmune, Kyowa, Corvus and Novartis and is a consultant/advisory board member for AbbVie, AstraZeneca, Bristol-Meyers Squibb, Boehringer Ingelheim, Celgene, EMD Serono, Inivata, Janssen, Lilly, Merck, and Novartis. J.R.W. is founder and owner of Resphera Biosciences, and is a consultant for Personal Genome Diagnostics. J.N. receives research funding from Merck, AstraZeneca/MedImmune, Kyowa and Calithera and is a consultant/advisory board member of Bristol-Myers Squibb, AstraZeneca/MedImmune and Takeda. C.L.H. is a consultant/advisory board member for AbbVie, Bristol-Myers Squibb and Genentech, receives research funding from Merrimack, GlaxoSmithKline, AbbVie, Bristol-Myers Squibb, and GlaxoSmithKline. B.L. is a consultant/advisory board member of AstraZeneca, Celgene, Genentech Rouche, Eli Lilly and Takeda and receives research funding from Celgene and Boehringer Ingleheim. J.R.B. receives research grants from Bristol-Myers Squibb, AstraZeneca/MedImmune and Merck and is a consultant/advisory board member for Bristol-Myers Squibb, AstraZeneca/MedImmune and Merck. M.V.B. is a paid consultant for Cepheid Inc. J.I. owns LumaCyte, LLC. Stock. D.M.P. receives research support from Bristol-Myers Squibb, Compugen and Potenza Therapeutics and is a consultant for Aduro Biotech, Amgen, AstraZeneca/Medimmune, Bayer, Compugen, DNAtrix, Five Prime, GlaxoSmithKline, ImmuneXcite, Jounce Therapeutics, Neximmune, Pfizer, Rock Springs Capital, Sanofi and Vesuvius/Tizona. V.E.V. is a founder of Personal Genome Diagnostics, a member of its Scientific Advisory Board and Board of Directors, and owns Personal Genome Diagnostics stock, which are subject to certain restrictions under university policy. V.E.V. is an advisor to Takeda Pharmaceuticals. Within the last 5 years, V.E.V. has been an advisor to Daiichi Sankyo, Janssen Diagnostics and Ignyta. The terms of these arrangements are managed by Johns Hopkins University in accordance with its conflict of interest policies.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Correlations between tumor purity and tumor mutation burden estimates from whole exome and targeted next-generation sequencing.
(a-h) Paired tumor-normal whole exome sequencing data from 3,788 TCGA samples from 8 tumor types (BLCA, N=405 tumors, BRCA, N=778 tumors, COAD, N=287 tumors, HNSCC, N=504 tumors, KIRC, N=367 tumors, LUAD, N=508 tumors, LUSC, N=477 tumors, SKCM, N=462 tumors) for which immune targeted approaches have shown promising clinical benefit were analyzed and revealed a positive correlation between obsTMB and tumor purity in bladder cancer (Pearson’s R=0.083, p=0.095), lung adenocarcinoma (Pearson’s R=0.092, p=0.038), and lung squamous cell carcinoma (Pearson’s R=0.089, p=0.051). A loess model was fitted to the mutation sequence data for each tumor type. For colon adenocarcinoma, tumor samples with a hypermutator genotype are shown in gray and were excluded from the model fit. Pearson’s correlation coefficient (R) was used to assess correlations between continuous variables and p values are based on two-sided testing. (i) TMB scores derived from targeted sequencing and tumor purity assessments were retrieved from a published cohort of 1,661 tumors treated with immune checkpoint blockade (Samstein et al., Nat Genet, 2019) and non-parametric correlations were evaluated by the Spearman’s rho statistic. A significant correlation between obsTMB and tumor purity was identified for NSCLC (N=254 tumors, Spearman rho=0.29, p=2.4x10-6), bladder cancer (N=148 tumors, Spearman rho=0.18, p=0.03), esophagogastric cancer (N=103 tumors, Spearman rho=0.19, p=0.05) and head and neck cancer (N=100 tumors, Spearman rho=0.18, p=0.07). The purple solid line indicates a linear regression fit and the purple shaded area represents the estimated 95% confidence interval of the probability density of TMB at each level of purity.
Extended Data Fig. 2 Tumor purity correlates with TMB estimates in the immunotherapy treated cohorts.
The correlations of tumor purity with tumor mutational burden and clinical response in 957 TCGA NSCLC samples and in immunotherapy cohorts 1 and 2 are shown in a-f. (a-b) TCGA lung adenocarcinomas-LUAD (N=508 tumors; a) and lung squamous cell carcinomas-LUSC (N=477 tumors; b) with a higher degree of normal contamination had a significantly lower TMB compared to tumors with a tumor purity > 50% (Mann-Whitney U test p=0.06 and p=1.3e-7 for LUAD and LUSC respectively). (c-d) In cohorts 1 and 2, the correlation between obsTMB and tumor purity was particularly pronounced for tumor purity less than 30% (Mann Whitney U test p=0.008 and p=0.08 for overall comparisons of TMB across tumor purity tiers for cohort 1 (N=87 tumors; c) and cohort 2 respectively (N=30 tumors; d). (e-f) Tumor purity was associated with clinical benefit from ICB when mutation-based purity was used, which is most likely attributed to the contribution of TMB in the mutation-based purity calculation; however no difference in tumor purity was found between responding and non-responding tumors when copy-number based tumor purity (N=74 tumors) and adjusted tumor purity (N=87 tumors) was used in cohort 1 (Mann Whitney U test p=0.14 and Mann Whitney U test p=0.22 respectively; e) and cohort 2 (Mann Whitney U test p=0.72 and Mann Whitney U test p=0.6 respectively; f). Seventy four and twenty one tumor samples from cohort 1 and 2 respectively had evaluable copy-number based tumor purity (Methods). The vertical line in each whisker plot signifies the median and the ends the upper and lower quartile, while the whiskers extend 1.5 times the interquartile range from the hinges. P values are based on two-sided testing.
Extended Data Fig. 3 Effect of TMB correction for the TCGA and immunotherapy-treated cohorts.
(a) Corrected TMB was replotted with tumor purity for TCGA samples to visualize whether our approach generated overcorrected TMB values on the lower end of tumor purity (N=3,788 tumors, BLCA N=405 tumors, BRCA N=778 tumors, COAD N=287 tumors, HNSCC N=504 tumors, KIRC N=367 tumors, LUAD N=508 tumors, LUSC N=477 tumors, SKCM N=462 tumors). We did not observe such phenomenon for all tumor types evaluated, suggesting that our correction factors did not generate high TMB-low tumor purity outliers. Additionally, we found that after using our TMB correction approach, there was now no longer a positive correlation between cTMB and tumor purity (BLCA Pearson R 0.023 p=0.64, BRCA R=-0.013 p=0.71, COAD R=0.019 p=0.0.77, HNSCC R=0.031 p=0.48, KIRC R=0.015 p=0.78, LUAD R=0.071 p=0.11, LUSC R=0.069 p=0.13, SKCM R=0.048 p=0.31). Blue line indicates a linear fit. The Pearson correlation coefficient (R) was used to assess correlations between continuous variables and p values are based on two-sided testing. (b) Effect of TMB correction for tumor purity in cohort 1. Distribution of observed (black circles) and corrected TMB for patients in cohort 1 (N=89 patients) are shown for each tumor purity tier. Corrected TMB values are denoted by purple circles for tumor purity 0.1-0.25 and green circles for tumor purity >0.25, error bars represent 95% confidence intervals. cTMB values are capped at 1000. The second tertile of cTMB was used as a cut-off point to classify tumors as high vs low TMB categories in order to avoid bias related to multiple cutoff point selection. After correcting for tumor purity in cohort 1, ten patients were reclassified in TMB categories. As would be expected with lower tumor purity, there were four patients with tumor purity <20% that switched class from the low observed TMB to the high corrected TMB category but there was also one patient with tumor purity >20% that switched from low observed TMB to high corrected TMB category and also five patients with higher tumor purity (range 46–82%) that switched class from the high observed TMB to the low corrected TMB category. Switching between both TMB categories would be expected as our approach generates corrected TMB estimates that better capture the true TMB distribution and that in turn would affect the percentile threshold that defines high TMB more accurately. NA; radiographic response non evaluable.
Extended Data Fig. 4 Impact of molecular smoking signature on outcome for immune checkpoint blockade and association with TMB.
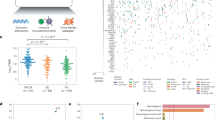
(a-b) In silico dilution experiments of single base substitutions to evaluate the power to accurately determine contribution of a dominant mutation signature. Mutation signature analyses were performed on whole exome data from 985 NSCLC tumors (N=508 lung adenocarcinomas and N=477 squamous cell carcinomas) obtained through TCGA. Seventy-six NSCLC tumors (N=64 lung adenocarcinomas and N=12 squamous cell carcinomas) had a tumor mutation load >250 and a molecular smoking signature >75% and were further selected for an in silico dilution series. (a) Mutation counts were diluted from maximum count to a minimum of 5 using random resampling (3 re-sampling replicates per dilution level), to evaluate consistency and divergence in the predicted presence of a smoking signature. Connected points in (a) indicate mean values per case per dilution series. On average, 20 mutations were sufficient to predict the presence of a smoking signature at a 50% level. (b) Mutational load below 20 mutations lead to a 30% difference from the original contribution of the C>A transversion rich signature value and therefore represents a threshold beyond which, there is a significant deviation from accurately determining a dominant mutation signature. Canter values indicate the mean values per dilution level and error bars indicate standard error of the mean. (c) Patients with a molecular smoking mutational signature derived durable benefit from immune checkpoint blockade (N=80, log rank p=0.031). TMB did not fully explain the association between molecular smoking signature and ICB response. (d) The number of clonal mutations was plotted against the percent contribution of the molecular smoking signature for each tumor (N=74 tumors in total, N=33 responding tumors and N=41 non-responding tumors). Each dot represents a tumor and each tumor is color coded by response, with responding tumors showed in orange and non-responding tumors showed in blue. While there is a correlation between the number of clonal mutations and a dominant molecular smoking signature, there are tumors with a low clonal TMB (dotted red line indicates a TMB threshold used to classify TMB in high/low categories) but a dominant molecular smoking signature which derive benefit from immune checkpoint blockade. The median point estimate and 95% confidence intervals for survival were estimated by the Kaplan–Meier method and survival curves were compared by using the nonparametric log rank test. Log rank p values are based on two-sided testing.
Extended Data Fig. 5 Impact of RTK mutations on outcome and genomic drivers associated with response to immune checkpoint blockade in cohort 2.
(a-b) Activating RTK mutations identified a group of patients with dismal prognosis in both cohort 1 (N=89 patients, log rank p=0.005) and cohort 2 (N=34 patients, log rank p=0.009). (c) In cohort 2 (N=31), responding tumors had a higher total and clonal TMB compared to non-responding tumors (Mann Whitney U test p=0.006 and p=0.004 respectively). Progression-free survival, histology and tumor purity are shown as separate panels. Patients with a clinical response had a higher contribution of the molecular smoking signature (N=15 patients, Mann Whitney U test p=0.054). There were no differences in tumor aneuploidy between responders and non-responders (N=74 patients, Mann Whitney U test p=0.72). A significant enrichment in RTK activating mutations, including point mutations and amplifications in EGFR, amplifications in ERBB2 and MET exon 14 skipping, was found in non-responding tumors (N=89 total patients, chi-squared p=0.056). (d) A third cohort of NSCLC patients treated with ICB was obtained from CBioportal (N=240 patients) as described in detail in the Methods section; for this cohort, sequence and copy number alterations, as well as outcome information were publicly available. Patients with activating RTK mutations in EGFR, ERBB2, MET, FGFR1 and IGF1R had a significantly shorter progression-free survival (log rank p=0.035). We defined response as durable clinical benefit if complete, partial response or stable disease was achieved with a duration >6months. The median point estimate and 95% confidence intervals for survival were estimated by the Kaplan–Meier method and survival curves were compared by using the nonparametric log rank test. CNV; copy number variation. cTMB; corrected TMB, RTK; receptor tyrosine kinase, Hist; histotype, TP; tumor purity, SS; smoking signature, AI; allelic imbalance. Dots indicate hotspot mutations, and × denotes loss of heterozygosity of the wild type allele. P values are based on two-sided testing.
Extended Data Fig. 6 Copy number analyses for NSCLC tumors in cohort 1.
Given that deletions in IFN-ɣ genes have been described as a potential mechanism of intrinsic resistance to immunotherapy, we investigated whether there is an enrichment in IFN-ɣ related gene copy number variation in non-responding tumors. (a) A cluster of IFN-ɣ related genes (IFNE, IFNA1, IFNA2, IFNA4, IFNA5, IFNA6, IFNA8, IFNA14, IFNA21, IFNW1 and IFNB1) is located on chromosome 9 (p21.3), in close proximity to the CDKN2A locus. (b) The locus that contains both the IFN-ɣ related genes and CDKN2A was frequently found to be deleted; an example of such homozygous deletion is shown for case CGLU262. The vertical axes denote the relative copy ratio (log2 scale), and the integer copy number levels assigned to genomic bins (circles) and segments. Purple and green boxes mark the coordinates of IFN gene cluster and CDKN2A, respectively. (c) The frequency of homozygous deletions in IFNE, IFNA1, IFNA2, IFNA4, IFNA5, IFNA6, IFNA8, IFNA14, IFNA21, IFNW1 and IFNB1 was similar in responding and non-responding tumors and more importantly, these deletions co-occurred with CDKN2A loss in 86% of the cases, whereas CDKN2A deletions also occurred independently. Given that, CDKN2A and the group of IFN-ɣ pathway genes lie on chromosome 9p within a span of 917 Kb, IFN-ɣ deletions may be passengers in the setting of a driver CDKN2A deletion. Seventy five tumors had evaluable copy number estimates (Methods). (d) A genome-wide analysis of copy number profiles revealed genomic regions with copy number gains and losses and was used to determine the extent of tumor aneuploidy. The relative copy ratio (LogR) values quantifying the abundance of each genomic region compared to the genome average (ploidy) are shown after correction for tumor purity in responding (N=33 tumors) and non-responding tumors (N=41 tumors). Red and blue shades indicate copy gains and losses, respectively, whereas white marks copy neutral regions. There was no significant difference in the degree of aneuploidy assessed by the fraction of genome with allelic imbalance between the two groups (Mann Whitney U test p=0.367, FDR-corrected p=0.65). We defined response as durable clinical benefit (DCB) if complete, partial response or stable disease was achieved with a duration >6months. P values are based on two-sided testing. CAN; copy number aberration, NDB; non durable clinical benefit.
Extended Data Fig. 7 Pathway enrichment analysis for DNA damage repair genes and the WNT pathway and distribution of hotspot mutations and associated potentially immunogenic MANAs in cohort 1.
(a) We investigated co-occurrence of mutations in DNA damage repair genes involved in base excision repair (DDR-BER), DNA damage sensoring (DS), the Fanconi anemia pathway (FA), homologous recombination (DDR-HR), mismatch repair (DDR-MMR), nucleotide excision repair (DDR-NER), non-homologous end joining (DDR-NHEJ) and translesion DNA synthesis (DDR-TLS). Mutations were characterized by consequence (missense, frameshift, nonsense, splice site, in-frame) and recurrence (hotspots) and loss of the wild type allele was considered in case of truncating mutations (biallellic inactivation). A similar analysis was performed for genes involved in the WNT pathway. A high TMB tumor with biallellic inactivation of MLH1 and a tumor with a gain-of-function beta-catenin hotspot mutation were identified among responders (N=41 patients) and non-responders (N=46 patients) respectively, with no additional significant differences in genomic alterations in the DDR and WNT pathways between responders and non-responders. We defined response as durable clinical benefit (DCB) if complete, partial response or stable disease was achieved with a duration >6months. (b) The number of mutations with at least one fit MANA (determined as neopeptides with a predicted MHC affinity < 50nM for which the wild type peptides has a predicted MHC affinity of > 1000nM) in each tumor, divided by clonality and hotspot status is shown in the top distribution graph. Clinical response and overall survival are shown in the middle panel. Clonal hotspot frameshifts and in-frame insertions and deletions in ANTRX2, TP53, EGFR, ASXL1, NOTCH2, ZFP36L2, FAM171B, SLC35F5, CD93 and SLAMF1 generated fit MANAs shown in the lower panel. There was no difference in the fraction of clonal fit MANAs between responding (N= 33 tumors) and non-responding (N=41 tumors, Mann Whitney U test p=0.65) tumors. We defined response as durable clinical benefit (DCB) if complete, partial response or stable disease was achieved with a duration >6months. NDB: No durable benefit.
Extended Data Fig. 8 HLA genetic variation and TMB correlations.
(a) We investigated whether there is an enrichment for chromosome 6p which contains the HLA class I loci for 3,767 tumors from TCGA (BLCA N=89 tumors, BRCA N=737 tumors, COAD N=348 tumors, GBM N=483 tumors, HNSCC N=269 tumors, KIRC N=370 tumors, LUAD N=287 tumors, LUSC N=257 tumors, OV N=456 tumors, READ N=146 tumors, UCEC N=325 tumors). LOH events in NSCLC were compared to the background arm-level allelic imbalance of the same tumor type and across tumor types. Chromosome 6p losses were not more frequent compared to other chromosomal arm level deletions (on the contrary the degree of chromosome 6p LOH was lower compared to other chromosomal arms deletions in lung tumors, N=544 tumors, Fisher’s exact p=0.037). In contrast, when chromosome 6p LOH events were compared between lung tumors and 9 tumor types (BLCA, BRCA, COAD, GBM, HNSCC, KIRC, OV, READ and UCEC), we found that LOH events involving chromosome 6p that contains the HLA class I loci are more frequent in lung cancer (NSCLC N=533, other tumors N= 3223, 17.3% vs. 8.2%, Fisher’s exact p=6.8e-10), without any evidence for positive selection of these events in advanced stage disease. Analysis of variance (ANOVA) was applied to assess the correlation of germline homozygosity in HLA class I genes with tumor mutation burden in 6 tumor types (BLCA, BRCA, COAD, HNSCC, KIRC, LUAD and LUSC, total N=3,601). (b) Germline HLA zygosity was not correlated with TMB for the vast majority of tumors examined with the exception of bladder cancer (N=396 tumors, ANOVA p=0.02). (c) Germline and tumor HLA class I status was combined to determine the number of unique HLA class I alleles in each tumor. The number of unique HLA class I alleles appeared to correlate with TMB such that tumors with a higher number of unique HLA class I alleles in the tumor harbored a lower non-synonymous mutation load for BLCA (N=86 tumors, ANOVA p=0.02) and HNSCC (N=244, ANOVA p=0.07). When tumors heterozygous for all three HLA class I loci (6 HLA class I alleles) where compared to tumors that were homozygous in all three HLA class I loci (3 HLA class I alleles), a lower TMB was noted in the tumors with the more intact antigen presentation capacity (N=86 tumors, Wilcoxon p=0.05 for BLCA, N=693 tumors p=0.09 for BRCA, N=244 tumors p=0.01 for HNSCC, N=284 tumors p=0.01 for LUAD). (d-g) HLA class I distribution by supertype and association with TMB. Individual HLA-I alleles were classified into discrete supertypes, based upon similar peptide-anchor-binding specificities. (d) TMB did not differ among different HLA-A supertypes (N=89 patients, Kruskal-Wallis p=0.45). (e) Similarly, there were no differences in TMB among different HLA-B supertypes (N=89 patients, Kruskal-Wallis p=0.45. (f) HLA-A supertype distribution for cases in cohort 1. (g) HLA-B supertype distribution for cases in cohort 1. Center values of the box plots in (d) and (e) indicate median values and error bars denote 95% confidence intervals. P values are based on two-sided testing.
Extended Data Fig. 9 Impact of HLA genetic variation on survival.
(a-d) Association of HLA class I haplotypes and outcome. There was no association between HLA class I haplotypes and overall survival (N=89 patients, log rank p=0.175; a). The same observations held true for HLA-B haplotypes (N=89 patients, log rank p=0.95; b). (c) There was a trend towards longer overall survival for TMB high tumors with maximal germline HLA class I heterozygosity compared to tumors with low TMB and maximal germline HLA class I homozygosity (N=31 patients, log rank p=0.096). (d) Maximal germline heterozygosity vs. homozygosity with more than one HLA loci was not associated with outcome (N=89 patients, log rank p=0.95). (e) Cases with maximal germline HLA class I heterozygosity were found to have a less clonal TCR repertoire (N=60 patients, Kruskal-Wallis p=0.01). Center values of the box plots in (e) indicate median values and error bars denote 95% confidence intervals. P values are based on two-sided testing.
Extended Data Fig. 10 Correlations among genomic and immune features and multivariable model for prediction of outcome to immune checkpoint blockade, excluding patients on chemo-immunotherapy and tumors collected at the time of resistance in cohort 1.
(a) Relationships among different features were assessed by the Spearman’s rho statistic for cohort 1 (N=89 patients) and p values were corrected for multiple comparisons. Four clusters of parameters were identified, related to RTK mutations, HLA genetic variation, tumor aneuploidy and TMB. The color of each dot refers to the Spearman rho coefficient value (darkest blue being 1 and darkest red being -1) and the size of each dot is proportional to the strength of the correlation. Three stars indicate an FDR adjusted p value of <0.05, two stars indicate an FDR adjusted p value of <0.1 and one star denotes an FDR adjusted p value of <0.2. (b-c) Corrected TMB, RTK mutations, molecular smoking signature and HLA germline variation were combined in a multivariable Cox proportional hazards regression model and a risk score was calculated for each case based on the weighted contribution of each parameter. Among 84 patients in total, patients with a higher risk score (N=28 patients) had a significantly shorter overall survival in cohort 1 (13 months vs. 38 months, HR=3.16, 95% CI: 1.68-5.91, log rank p=0.0002; b). The model was trained in cohort 1, fixed and applied in cohort 2 revealing a significantly shorter progression-free survival for high risk patients in cohort 2 (3 months vs. 8 months, HR=2.73, 95% CI 1.15-6.45, log rank p=0.017; c). The second tertile of the risk score was used to classify patients in high risk (top 33.3%, N=28 patients for cohort 1 and N=11 patients for cohort 2) and low risk (bottom 66.6%, N=56 patients for cohort 1 and N=23 patients for cohort 2) groups. The median point estimate and 95% confidence intervals for survival were estimated by the Kaplan–Meier method and survival curves were compared by using the nonparametric log rank test. Log rank p values are based on two-sided testing.
Supplementary information
Source data
Source Data Fig. 1
Source data for Fig. 1.
Source Data Fig. 2
Source data for Fig. 2.
Source Data Fig. 3
Source data for Fig. 3.
Source Data Fig. 4
Source data for Fig. 4.
Source Data Fig. 5
Source data for Fig. 5.
Source Data Fig. 6
Source data for Fig. 6.
Source Data Extended Data Fig. 2
Source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9
Source data for Extended Data Fig. 9.
Source Data Extended Data Fig. 10
Source data for Extended Data Fig. 10.
Rights and permissions
About this article
Cite this article
Anagnostou, V., Niknafs, N., Marrone, K. et al. Multimodal genomic features predict outcome of immune checkpoint blockade in non-small-cell lung cancer. Nat Cancer 1, 99–111 (2020). https://doi.org/10.1038/s43018-019-0008-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-019-0008-8
This article is cited by
-
Unveiling the mechanisms and challenges of cancer drug resistance
Cell Communication and Signaling (2024)
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
Exploiting temporal aspects of cancer immunotherapy
Nature Reviews Cancer (2024)
-
Assessment of human leukocyte antigen-based neoantigen presentation to determine pan-cancer response to immunotherapy
Nature Communications (2024)
-
Tumour mutational burden: clinical utility, challenges and emerging improvements
Nature Reviews Clinical Oncology (2024)