Abstract
Artificial intelligence (AI) researchers and radiologists have recently reported AI systems that accurately detect COVID-19 in chest radiographs. However, the robustness of these systems remains unclear. Using state-of-the-art techniques in explainable AI, we demonstrate that recent deep learning systems to detect COVID-19 from chest radiographs rely on confounding factors rather than medical pathology, creating an alarming situation in which the systems appear accurate, but fail when tested in new hospitals. We observe that the approach to obtain training data for these AI systems introduces a nearly ideal scenario for AI to learn these spurious ‘shortcuts’. Because this approach to data collection has also been used to obtain training data for the detection of COVID-19 in computed tomography scans and for medical imaging tasks related to other diseases, our study reveals a far-reaching problem in medical-imaging AI. In addition, we show that evaluation of a model on external data is insufficient to ensure AI systems rely on medically relevant pathology, because the undesired ‘shortcuts’ learned by AI systems may not impair performance in new hospitals. These findings demonstrate that explainable AI should be seen as a prerequisite to clinical deployment of machine-learning healthcare models.
Similar content being viewed by others
Main
The prospect of applying artificial neural networks to the detection of COVID-19 in chest radiographs has generated interest from machine learning (ML) researchers and radiologists alike, given its potential to (1) help guide management in resource-limited settings that lack sufficient numbers of the gold-standard polymerase chain reaction with reverse transcription (RT-PCR) assay and (2) clarify cases of suspected false negatives from the RT-PCR assay1,2. Although numerous recent publications and preprints report machine learning models with high performance at this task3,4,5,6,7,8, the trustworthiness of these models needs to be evaluated rigorously before deployment in a clinical setting9.
Our findings in this study support the troubling possibility that these models fail to learn the true underlying pathology reflecting the presence of COVID-19 and instead leverage spurious associations between the presence or absence of COVID-19 and radiographic features that reflect variations in image acquisition, that is, ‘shortcuts’10. Although such spurious associations may arise in any dataset, we have observed that many recent ML models for radiographic detection of COVID-19 were trained using data with the potential for near worst-case confounding. These datasets are composed of an exclusively COVID-19-negative source and a COVID-19-positive source, such that any systematic differences between the sources correlate perfectly with COVID-19 status3,4,5,6,7,8. Similar combinations of data sources, where the source label correlates with disease status, have also been used to train AI systems for the detection of COVID-19 in computed tomography scans11 (although the non-public nature of the data precludes experimental verification of the extent of shortcut learning in this setting) and for other medical imaging tasks12,13, implying that our findings have broad implications for the field of medical machine learning.
In this Article, we evaluate the trustworthiness of recent deep learning models for COVID-19 detection from chest radiographs. After training deep convolutional neural networks14,15 (‘Datasets and preprocessing’ section and Supplementary Fig. 1) in the manner of these previous publications3,4,5,6,7,8, we evaluate their performance in new hospital systems. We then interrogate the extent to which these models rely on confounds by identifying the most important image features using state-of-the-art explainable artificial intelligence (AI) techniques, including both saliency maps and generative adversarial networks (GANs)16,17,18,19. These enquiries reveal how seemingly high-performance AI systems may derive the majority of their performance from the exploitation of undesired shortcuts, highlighting the need to verify that AI systems rely on the desired signals. Finally, we evaluate several methods to alleviate the problem of shortcut learning in this setting, demonstrating the importance of improved data quality for the creation of robust and useful models.
Results
Overview of the experimental approach
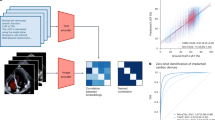
Before examining our main results, we first outline our experimental approach (Fig. 1a). To begin, we reviewed the literature to examine the datasets and models used for the detection of COVID-19 from chest radiographs, with attention focused on studies with the potential for ‘worst-case confounding’. After choosing representative networks, we built two datasets: one that reproduces the data used in previous studies and a second that enables external validation on new hospitals. In a first experiment, we evaluated models that were trained on one dataset using test images from the other dataset, under the expectation that a model that relies on valid medical pathology—which should not change between datasets—should maintain high performance. We then probed deeper into specific shortcuts that these models leverage, using techniques from explainable AI.
a, A neural network model is trained to detect COVID-19 using radiographs from either of two datasets, and then evaluated on both datasets to learn how performance may drop in deployment (that is, a generalization gap). Intepretability methods are then applied to infer what the model learned and which features were important for its decisions. Whereas dataset I draws radiographs from multiple hospital systems as well as cropped images from publication figures, dataset II draws radiographs from multiple hospitals from a single regional hospital system. b, Characteristics of the datasets used in this study. c, Model evaluation scheme (top) and corresponding receiver operating characteristic (ROC) curves (bottom), which show the performance of our neural network models evaluated on both an internal test set (new, held-out examples from the same data source as the training radiographs) and an external test set (radiographs from a new hospital system). Inset numbers indicate areas under the ROC curves, where a larger area corresponds to higher performance (area under the curve (AUC), mean ± standard deviation). The difference between internal and external test set performance is the generalization gap.
In a ‘model-centric’ approach, which focuses on the specific portions of the radiographs that contribute most to the predictions of our models in particular, we built saliency maps using expected gradients19. In essence, this approach attributes importance to each pixel of a radiograph based on the gradients of our models, while avoiding issues such as saturation or an arbitrary choice of baseline. We complemented this model-centric approach with a data-centric approach, focusing on the key aspects of the data that could be used to distinguish COVID-19-positive and COVID-19-negative cases. Specifically, we applied generative adversarial networks (CycleGANs17) to transform COVID-19-positive radiographs to appear COVID-19-negative and vice versa, in the sense that key image features are transformed, such that a network can no longer discriminate between the real images of a given pathology label and the transformed images from the opposite class18. Rather than use our classifier networks to perform this discrimination task, we instead trained new discriminator networks simultaneously with generator networks that transformed the images, such that this experiment focused on key aspects of our data, rather than our classifiers in particular.
To further validate these findings, we went on to perform ‘region-swapping’ experiments in which we swapped out portions of radiographs that our explainable AI approaches identified as important, with the expectation that changes to truly important regions would have a large impact on our classifiers’ outputs. We concluded by evaluating approaches to mitigate shortcut learning from the perspectives of both generalization performance and model explainability.
Literature review of model and dataset construction
In our investigation, we aimed to determine the extent to which shortcut learning affects AI systems for COVID-19 detection in chest radiographs, which is complicated by the diversity of these systems. We therefore trained a series of 10 models with various architectures, including state-of-the-art networks that were tailor-made for the detection of COVID-19 in chest radiographs4,6,20 and multiple ‘off-the-shelf’, general-purpose architectures14,15,21,22. For our primary models, we chose a network based on the DenseNet-121 architecture14, which we judged faithfully replicated the modelling choices of recent high-performance models for COVID-19 classification, while also following established best practices for classification of pathologies from chest radiographs using deep learning. Alongside these primary models, we also investigate multiple secondary models to help probe the generality of our findings and the extent to which they apply to AI systems found in the wild. These secondary models include the COVID-Net network, which was custom-designed for the detection of COVID-19 via a machine-based architecture search4, the DarkCovidNet model, which was modified from a standard DarkNet-19 model for the purpose of COVID-19 detection6, and the CV19-Net model20, which was built by ensembling 20 DenseNet-121 networks and motivates our primary model, which uses the same architecture without ensembling, given that ensembling did not provide performance gains but substantially increases the computational complexity (‘Evaluation of models on new hospital systems’ section).
To train and evaluate these models, we created two datasets (Fig. 1a and Supplementary Table 1). Dataset I consisted of COVID-19-positive radiographs from the GitHub-COVID repository23, which aggregates radiographs from publication figures and other online sources with different geographic origins. We supplemented these with COVID-19-negative radiographs from the ChestX-ray14 repository of the National Institutes of Health (NIH)24, which originate from a single hospital in the United States.
Dataset I is similar to the datasets used for training in recent publications on AI for COVID-19 detection3,4,5,6,7,8. Specifically, four of these publications3,5,6,7 combine the GitHub-COVID repository with either the NIH repository24 or the similar Radiological Society of North America pneumonia dataset25, which was derived from the NIH repository. Two others4,8 similarly combine these repositories, but then supplement them with additional COVID-19-positive images from other online repositories, many of which have since been added to the GitHub-COVID repository. Given the continually evolving nature of many of these repositories, the precise set of images used in each study remains unclear and additional uncertainty is introduced by the dearth of documentation on the source of some images or the validity of their labels (for example, in the ActualMed and Fig. 1 databases at https://github.com/agchung/Actualmed-COVID-chestxray-dataset and https://github.com/agchung/Figure1-COVID-chestxray-dataset). This uncertainty notwithstanding, our core observation is that numerous well-cited studies build their datasets by gathering COVID-19-positive radiographs from various sources, as exemplified most thoroughly by the GitHub-COVID repository (in which the image sources and labelling method are clearly documented), and then combine these with COVID-19-negative radiographs originating from the NIH repository, so that we judge that our dataset I fairly represents the key aspects of the data used in these earlier works. Other publications20,26,27,28 generally use non-public data, precluding our ability to audit their models, and do not share this issue of strong correlation between data source labels and COVID-19 status. However, based on our review of the literature, we find this issue in an alarming proportion of the publications, including many of the most high-profile studies4,5,6.
Unlike the datasets used in recent publications, which collected COVID-19-positive and -negative images from disparate sources, dataset II corresponds to a seemingly more ideal case where both COVID-19-positive and -negative images were drawn from similar sources. This dataset, which comprises the PadChest and BIMCV-COVID-19+ repositories (Fig. 1a,b), consisted of radiographs from a single region and published by a shared research team, although BIMCV-COVID-19+ represents a greater diversity of hospitals than PadChest, and the repositories were acquired over different time periods29,30.
Evaluation of models on new hospital systems
After training on dataset I, we evaluated our models for reliance on confounding factors by comparing the predictive performance on an internal test set (new, held-out radiographs from dataset I) to performance on external radiographs from dataset II. Although our models attain high performance on internal test data, half of the models’ predictive performance is lost when testing on dataset II (Fig. 1c, left). This performance drop (the generalization gap) suggests that these models rely on source-specific confounds in the radiographs, as we would expect models that use genuine markers of pathology to generalize well10. This finding held true for all nine additional architectures we examined, including those that were custom-tailored in recent studies for the detection of COVID-19 in radiographs (Supplementary Figs. 2 and 3).
Although we initially expected that a dataset built from radiographs drawn from a single region would be less likely to contain spurious correlations that enable ML models to take shortcuts, we found that models trained on dataset II also exhibit high performance on internal test data and low performance on external test data (Fig. 1c, right and Supplementary Fig. 2). Thus, dataset-level confounding may be a serious issue, even in datasets derived from more similar sources such as hospitals from a single region, contrary to the conclusions of contemporary work31. These findings argue for routine reporting of metadata on potential patient, hospital system and preprocessing confounds. By illuminating the construction of radiographic datasets in greater detail, these data will make it easier for domain experts to identify likely sources of confounding. Additionally, these metadata enable the construction of models that explicitly control for confounds, providing a route to AI systems that generalize well even in the context of confounded training data32,33,34. By contrast, we note that a popular set of approaches to improve generalization performance, known as ‘unsupervised domain adaptation’, are precluded by the presence of worst-case confounding because these methods rely on learning models invariant to data-source labels, which will be perfectly correlated with the pathology labels35.
Alternative hypotheses do not explain poor generalization
To verify the hypothesis that exploitation of dataset-specific confounding leads to poor generalization performance, we investigated alternative explanations for the generalization gap. Previous publications have suggested that more complex models—that is, those with higher capacity—may be particularly prone to learning confounds36, so we evaluated the generalization performance of simpler models, including a logistic regression and a simple convolutional neural network architecture, but found that the generalization gap did not improve (Supplementary Fig. 3). This result further supports the broad applicability of our findings, because the generalization gap was present regardless of network architecture, aligning with a previous study that showed that radiograph classification performance is robust to neural network architecture37. Similarly, we found that replacing the multilabel classification scheme of our original models with a simpler single-label classification scheme (‘Model architecture and training procedure’ section) did not improve generalization performance.
In addition to the choice of model architecture, an alternative explanation for poor generalization performance is that, rather than the model learning a spurious correlation that does not generalize, the model learns a genuine relationship between a radiograph’s appearance and its COVID-19 label that still does not generalize. One such scenario is that the COVID-19 detection task differs between training and test time, which may occur in our datasets given that most of the images in the GitHub-COVID dataset were cropped from scientific publications and thus are perhaps more likely to show radiographic evidence of COVID-19, while labels in the BIMCV dataset are derived solely from RT-PCR or serology and therefore may or may not feature radiographic evidence of COVID-19. However, when we modified the label scheme of BIMCV-COVID-19+ such that radiographs are only labelled positive if a radiologist noted evidence of COVID-19, the generalization gap persisted (Supplementary Fig. 4), suggesting that such a concept shift between training and test time does not explain the performance difference, leaving the use of spurious correlations as the best explanation38.
Explainable AI identifies spurious confounders
We further interrogated the trained AI models using saliency maps16,39,40, which highlight the regions of each radiograph that contribute most to the models’ prediction (Supplementary Note 1 and Supplementary Fig. 5), to determine specific confounds exploited by deep convolutional networks for COVID-19 detection. Although our saliency maps sometimes highlight the lung fields as important (Fig. 2a), which suggests that our model may take into account genuine COVID-19 pathology, concerningly, the saliency maps also highlight regions outside the lung fields that may represent confounds. The saliency maps frequently highlight laterality markers that originate during the radiograph acquisition process (Fig. 2a and Supplementary Fig. 6), which differ in style between the COVID-19-negative and COVID-19-positive datasets, and similarly highlight arrows and other annotations that are uniquely found in the publication-sourced radiographs of the GitHub-COVID data source23 (Supplementary Fig. 7), which aligns with a previous study finding that ML models can learn to detect pneumonia based on spurious differences in text on radiographs41. Our saliency maps also indicate that the image edges, the diaphragm and the cardiac silhouette are important for our models’ predictions of a patient’s COVID-19 status, although these regions are not among those routinely used by radiologists to assess for COVID-1942 and instead probably reflect dataset-level differences in patient positioning and radiographic projection, that is, the anterior–posterior (AP) view versus posterior–anterior (PA) view34. Reliance on such confounds, which do not consistently correlate with COVID-19 status in outside datasets, helps explain the previously observed poor generalization performance.
a, Saliency maps for our neural network models indicating the regions of each radiograph with the greatest influence on the models’ prediction. Top: in a COVID-19-negative radiograph, in addition to the highlighting in the lung fields (open arrow), the saliency maps also emphasize laterality tokens (filled arrow). Middle: in a COVID-19-positive radiograph, the most intensely highlighted regions of the image are the bottom corners (arrows), outside of the lung fields. Bottom: in a COVID-19-positive radiograph, the only highlighted region is the diaphragm (arrow). The colour bar indicates saliency map pixel importances by percentile. b, Radiographs and their corresponding transformations by a GAN, illustrating systematic differences that enable neural networks to differentiate between COVID-19-positive and -negative radiographs. COVID-19-negative images are transformed by the GAN to appear as if they were COVID-19-positive, and vice versa. Comparison of images before and after transformation with a GAN visualizes important image features for COVID-19 prediction. Blue boxes indicate alterations to the opacity of the lung fields, which may represent the network’s attention to genuine COVID-19 pathology. Red solid boxes indicate altered laterality markers, and red dashed boxes indicate altered radiopacity at the image borders, both of which may spuriously correlate with a patient’s COVID-19 status in the training data. Figure adapted with permission from ref. 52, H. Winther et al. (a, bottom; b, bottom row); and ref. 53, Springer Nature Ltd (b, top row).
To further investigate what features could be used by an ML model to differentiate between the COVID-19-positive and COVID-19-negative datasets, we trained GANs to transform COVID-19-negative radiographs to resemble COVID-19-positive radiographs and vice versa. This technique should capture a broader range of features than saliency maps, as the GANs are optimized to identify all possible features that differentiate the datasets. Consistent with our knowledge of how radiologists detect evidence of COVID-19 in chest radiographs, the GAN increases the radiopacity or radiolucency of the lung fields bilaterally to respectively add or remove evidence of COVID-19, indicating that neural network models are capable of learning genuine markers of COVID-19 (Fig. 2b, blue boxes and Supplementary Figs. 8 and 9). However, the generative networks frequently add or remove laterality markers and annotations (Fig. 2b, solid red boxes), reinforcing our observation from saliency maps that these spurious confounds also enable ML models to differentiate the COVID-19-positive and COVID-19-negative radiographs. The generative networks additionally alter the radiopacity of image borders (Fig. 2b, dashed red boxes), supporting our previous assertion that systematic, dataset-level differences in patient positioning and radiographic projection provide an undesirable shortcut for ML models to detect COVID-19. Given this strong evidence that ML models can leverage spurious confounds to detect COVID-19, we also investigated the extent to which our classifiers, in particular, relied on the features altered by the GAN. We found that images transformed by the GANs were reliably predicted by the classifiers to be the transformed class rather than the original class (Supplementary Fig. 10), demonstrating that the majority of features used by our classifiers were altered by the GAN; that is, the features identified by the GAN are approximately a superset of those used by the classifiers. Thus, the image transformations from the GANs enable us to see hypothetical versions of the same radiographs that would have caused our classifiers to predict the opposite COVID-19 status.
Experimental validation of factors identified by interpretability methods
We next aimed to experimentally validate the importance of spurious confounds to our models by manually modifying key features (Fig. 3a,b). We first swapped laterality markers from a COVID-19-positive and COVID-19-negative image, and found that introduction of a laterality marker more common in COVID-19-positive images increased the models’ predicted odds that the patient had COVID-19, while the converse also held. As a control, we compared to randomly swapped image patches of the same size and found that the change in model output from swapping laterality markers is significantly greater than expected by random (Fig. 3a), indicating that laterality markers are key features leveraged by our models to determine a patient’s COVID-19 status. Although these markers vary consistently between the datasets (Fig. 4 and Supplementary Figs. 7–9), these markers would not reliably indicate COVID-19 status in more general settings. We similarly investigated the shoulder region of radiographs, which was often highlighted as an important feature in our saliency maps (Supplementary Fig. 7), and found that moving the clavicle region of a radiograph to the top border of the radiograph increased the models’ predicted odds that the patient has COVID-19 (Fig. 3b and Supplementary Fig. 11), suggesting that the models leverage the consistent but medically irrelevant difference in patient positioning between the COVID-19-negative and COVID-19-positive data sources. To verify whether these findings held on a population basis, we sampled a random subset of the radiographs and repeated our experiments involving the swapping of laterality markers and movement of the shoulder region (Supplementary Fig. 12), which confirmed that our models indeed leverage these shortcuts throughout the dataset.
a, Left: text markers on radiographs are highlighted by saliency maps as important for COVID-19 prediction. The exchange of laterality markers between a pair of COVID-19-positive and COVID-19-negative images significantly shifts the output when compared to swapping random patches of the same size: Δ positive image (log odds) = −5.63 (empirical P = 9.99 × 10−4 based on Monte Carlo substitution of random image patches, n = 1,000); Δ negative image (log odds) = 13.85 (P = 5.00 × 10−3, n = 1,000) (‘Experimental validation of feature attributions’ and ‘Statistics’ sections). Grey dots in the distribution plots (right) correspond to the change in model output after swapping random image patches, which were used as a negative control. Red dots correspond to the change in model output for the radiographs with swapped laterality markers. b, The positioning of patient shoulders may impact COVID-19 prediction. Saliency maps highlight the shoulder region as important predictors of COVID-19 positivity after (but not before) this region is moved to the top of the image (left). This patch increased model output significantly more than random patches of the same size moved to the same corners (Δ model output = 6.57, empirical P = 5.00 × 10−3, n = 1,000). Grey dots in the distribution plot (right) correspond to radiographs with randomly selected patches. The red dot corresponds to the radiograph with the shoulder regions moved.
Shortcuts have a variable effect on generalization
Importantly, some shortcuts will impair generalization performance, but other shortcuts will not. While the large generalization gap is explained well by shortcut learning, a portion of the remaining external test set performance may still be due to shortcuts that happen to generalize for our datasets. Both types of shortcut are undesirable, because even those that generalize between our datasets may not consistently generalize to other settings, and the use of clinical rather than strictly radiological information extracted from these radiographs may be redundant, depending on the clinical workflow.
To analyse which shortcuts may contribute to poor generalization, we considered clinical metadata (Supplementary Table 1) and average images from each repository (Fig. 4). Among the shortcuts that do not generalize are the textual markers, which were clearly identified by our explainability approaches as important for prediction of COVID-19 but appear differently in the COVID-19-negative and COVID-19-positive images from each repository (Fig. 4). In addition, the radiographic projection, which may contribute to (but does not completely explain) the importance of the image edges and shoulder position, does not generalize between the datasets (Fig. 1b, ‘% AP images’ row) and therefore may contribute to poor generalization performance.
Among the shortcuts that do generalize (at least between our datasets) are aspects of patient positioning that do not result from the radiographic projection. These aspects of patient positioning also probably contribute to the previously observed importance of image edges and shoulder position, and they maintain a consistent relationship with COVID-19-negative and COVID-19-positive radiographs in each dataset (Fig. 4), despite the inconsistent relationship of the radiographic projection with COVID-19 status. An additional factor that may generalize well is patient sex, because, within both datasets, a higher proportion of males were COVID-19-positive (Supplementary Table 1). Taken together with our observation that half of our models’ performance is attributable to confounds that do not generalize well, we conclude that only a minority of our models’ performance is attributable to monitoring for genuine COVID-19 pathology.
Given that radiographic projection and patient sex are diffusely represented in radiographs and therefore less clearly pointed out by our explainability approaches, we also validated whether our models could leverage these factors as shortcuts. We reasoned that, for a model to be able to leverage these concepts as shortcuts, the same model (when retrained) must be able to predict these concepts well. Indeed, our models accurately predict both the radiographic projection and patient sex for both internal and external test data (Fig. 5), which supports that these concepts are easily learned and available to be leveraged as shortcuts. Considering that these concepts are easily learned and are also predictive of COVID-19 status (that is, they are correlated with COVID-19 in our datasets), we judge that our networks probably incorporate this information to predict COVID-19 status.
Improved data mitigate shortcut learning
Given this strong evidence that neural networks leverage dataset-level differences as shortcuts for COVID-19 status, we enquired to what extent this issue might be mitigated. Although an initial hypothesis may be that the choice of neural network architecture determines the propensity for shortcut learning, all architectures that we examined displayed similar evidence for shortcut learning, as quantified by the generalization performance (Supplementary Fig. 2). Although our tests hinted that data augmentation may help alleviate shortcut learning, the effect was small and not statistically significant (Supplementary Fig. 2b; external test set ROC-AUC of 0.76 ± 0.04 versus 0.79 ± 0.03 before and after data augmentation, respectively, when trained on dataset I, P = 0.22, U = 6 based on a Mann–Whitney U-test; external test set ROC-AUC of 0.70 ± 0.05 versus 0.69 ± 0.05 before and after data augmentation, respectively, when trained on dataset II, P = 1.00, U = 13 using Mann–Whitney U-test).
In principle, an attractive solution to mitigate shortcut learning is to remove the image factors that the models leverage as shortcuts. However, in practice, it is difficult to remove all such image factors. As a simple test case, we enquired whether removing textual markers by cropping to the centre 75% of each radiograph would reduce shortcut learning and thus improve generalization performance. After retraining our models on these cropped radiographs, we found that such cropping does not improve generalization performance (Supplementary Fig. 13), which naively may suggest that these textual markers do not contribute to shortcut learning. However, considering the consistent identification of this factor by saliency maps, the CycleGANs and manual image modifications (Figs. 2a,b and 3a), a more likely explanation is that a multitude of redundant shortcuts exist, such that a model may shift its attention toward other shortcuts in the absence of a particular shortcut. Conjecturally, such image attributes could include the size of the lung fields relative to the image, the positioning of the scapular shadows, the size of the cardiac silhouette, image intensities or textural features that enable inference of the data source.
Perhaps a more reliable solution to remove the image factors that enable shortcut learning is to simply collect data that is less confounded. To test this hypothesis, we created a third dataset (dataset III) to represent a nearly optimal case, where the COVID-19-positive and -negative cases were taken from the BIMCV-COVID-19+ repository and its paired BIMCV-COVID-19− repository (https://bimcv.cipf.es/bimcv-projects/bimcv-covid19/), respectively, which were collected from the same hospitals over the same time period (Supplementary Fig. 14). If this near-optimal dataset solved the ‘shortcut problem’, then we would expect that models trained on these data may (1) attain higher performance on an external test set, because bona fide pathology should transfer between datasets while shortcuts may or may not, and (2) exhibit a lower generalization gap, in the sense that performance on an internal test set would not as drastically misrepresent the true performance, as measured on external data. We trained models to detect COVID-19 in dataset III and then tested these models on external data from dataset I, and compared these results to models that were trained on dataset II and tested on dataset I. Despite that dataset III contains ~1/20th the images of dataset II, it attains significantly higher performance on external data (Fig. 6), and exhibits little generalization gap (Supplementary Fig. 15), suggesting that collection of less confounded data indeed alleviates the issue of shortcut learning. Furthermore, saliency maps for the model trained on dataset III tend to attribute more importance to the lung fields, where COVID-19 pathology would be expected, than to potentially confounding regions, as compared to the equivalent saliency maps generated for the model trained on dataset II (Supplementary Fig. 16), although the saliency maps still show some attention toward shortcuts. Taken together, these findings argue for careful collection of data so as to minimize the potential for shortcut learning, with continued caution that improved data collection may only partially solve the problem.
a, To evaluate whether improved data collection mitigates shortcut learning, we train classifiers on dataset II and dataset III, then test both on the same external data (dataset I). b, Evaluation of generalization performance as measured by ROC curves. Inset values indicate area under the ROC curve (AUC, mean ± standard deviation, n = 5). The AUC of models trained on dataset III is significantly greater than the AUC of models trained on dataset II (P = 0.016 based on a two-tailed Mann–Whitney U-test, corresponding U = −2.4).
Discussion
ML models that were built and trained in the manner of recent studies generalize poorly and owe the majority of their performance to the learning of shortcuts. This undesired behaviour is due partially to the synthesis of training data from separate datasets of COVID-19-negative and COVID-19-positive images, which introduces near worst-case confounding and thus abundant opportunity for models to learn these shortcuts. Importantly, because undesirable ‘shortcuts’ may be consistently detected in both internal and external domains, our results warn that external test set validation alone may be insufficient to detect poorly behaved models.
Previous studies also audited AI systems for the detection of COVID-19 in radiographs, with mixed success at identification of shortcuts. In a simple yet clever approach, one study found that models retain high performance when examining only the borders of radiographs, such that genuine COVID-19 pathology was removed from the images31. This study concurs with our findings but comments primarily on the possibility of this issue rather than its occurrence in the wild, though it is nonetheless alarming. The study that introduces the COVID-Net model also audits its model, using a saliency map approach known as ‘GSInquire’, but, in contrast, does not identify evidence of shortcut learning in a set of three published images4. Given the similarity of that study’s training data to our own dataset I and the large generalization gap that we observe with the same architecture, we suspect that shortcut learning probably did occur, and it remains unclear whether auditing decisions about additional radiographs beyond the three presented would have revealed evidence of shortcut learning or if the GSInquire approach, which is not available through a public-facing repository, fails to identify the shortcuts. A number of other studies that involve datasets with severe confounding between pathology and image source3,5,6,7,8 similarly audit their models using saliency map approaches (most prominently, the Grad-CAM approach43) and report findings on one to three radiographs, without noting evidence of shortcut learning. Based on this pattern, we recommend that researchers examine and report results from explainable AI or saliency map approaches on a population level, employing a sampling-based approach as necessary, and to remain sceptical of high performances in the absence of external validation. Moreover, we find that population-level audits using saliency maps are highly labour-intensive to perform in a rigorous manner and may depend on domain knowledge, which motivates future approaches for explainable AI in medical imaging that simplify population-level analysis.
Our findings support common-sense solutions to alleviate shortcut learning in AI systems for radiographic COVID-19 detection, including (1) improved collection of training data, that is, data in which radiographs are collected and processed in a way matching the target population of a future AI system and (2) improved choice of the prediction task to involve more clinically relevant labels, such as a numeric quantification of the radiographic evidence for COVID-1927,44. However, we demonstrate that shortcut learning may occur even in a more ideal data collection scenario, highlighting the importance of explainable AI and principled external validation. Although AI promises eventual benefits to radiologists and their patients, our findings demonstrate the need for continued caution in the development and adoption of these algorithms9.
Methods
Model architecture and training procedure
For our primary neural network, we used a convolutional neural network with the DenseNet-121 architecture to predict the presence versus absence of COVID-1914. This architecture has not only been used in a variety of recent models for COVID-19 classification4,5, but has also been used for the diagnosis of non-COVID pneumonia34,39, as well as for more general radiographic classification45.
Following the approach in recent COVID-19 models4,5, we first pre-trained the model on ImageNet, a large database of natural images46. Forcing models to first learn general image features should also serve as an inductive bias to prevent overfitting on domain-specific features34. After ImageNet pre-training, the final 1,000-node classification layer of the trained ImageNet model was removed and replaced by a 15-node layer, corresponding to the 14 pathologies recorded in the ChestX-ray14 dataset plus an additional node corresponding to COVID-19 pathology. Only the prediction for COVID-19 was used for evaluating the model, but we followed previous works that showed simultaneous learning of multiple tasks was useful for achieving the highest predictive performance39. To obtain a consistent label scheme, labels in the GitHub-COVID, PadChest and BIMCV-COVID-19+ repositories were mapped to the 14 ChestX-ray14 categories.
The model was optimized end to end using mini-batch stochastic gradient descent with a batch size of 16, momentum parameter of 0.9, weight decay of 10−4 and learning rate of 0.01, which was decreased by a factor of 10 every five epochs. We chose a binary cross-entropy loss as the optimization criterion. To prevent overfitting, we monitored the area under the ROC curve (AUC) for COVID-19 classification on a held-out validation set, and chose the epoch with the highest validation AUC as the final model. All models were trained for 30 epochs, which was long enough for all models to reach a maximum in the validation AUC. All models were trained using the PyTorch software library47, version 1.4, on NVIDIA RTX 2080 TI graphics processing units and required ~5 h of training time per replicate.
We also examined three architectures that were designed in previous publications specifically for the task of COVID-19 detection, with the hypothesis that these specialized architectures may better learn genuine COVID-19 pathology and generalize better to external data. These architectures were CV19-Net20, DarkCovidNet6 and COVID-Net4. We trained these models on datasets I and II, following the image preprocessing procedures, data augmentation pipelines and optimization schemes used in the original publications (we note that although dataset I is analogous to the original datasets used to train DarkCovidNet and COVID-Net, CV19-Net was trained on data that are not publicly available). For both CV19-Net and DarkCovidNet, the base architectures were downloaded from the torchvision library47, then modified to match the descriptions in each respective paper. The COVID-Net network was adapted from an open-source, PyTorch implementation (by Ilias Papastratis; https://github.com/iliasprc/COVIDNet). For the CV19-Net paper, the data augmentation pipeline was altered to match the pipeline in the original paper: when loading images, each radiograph is additionally randomly flipped with probability 0.5 then rotated between −30° and 30°. To disentangle performance differences due to the ensembling present in the CV19-Net architecture from performance differences due to the change in data augmentation, we also trained a single DenseNet-121 model with the same data augmentation steps as CV19-Net. In the case of CV19-Net and DarkCovidNet, we maintained the same multilabel classification task (that is, the 14 ChestX-ray14 labels plus a label for COVID-19) to facilitate optimal comparison between architectures. In the case of the COVID-Net architecture, due to problems with vanishing and exploding gradients when using the full multilabel classification task, we reduced our full label set to only the three labels used in the COVID-Net paper (COVID-19 Pneumonia, Non-COVID Pneumonia, No Pneumonia). We also trained additional, popular architectures that were not tailored specifically for COVID-19 detection, including MobileNetv221 and ResNeXt-5022. These networks were again modified from the ImageNet-pretrained base models in the torchvision library47. We trained these architectures using the same preprocessing scheme and optimization parameters as for our DenseNet-121 models, again replacing the standard, 1,000-label classification layers with an analogous layer for our 15 labels.
To test the hypothesis that lower-capacity models may not learn spurious correlations36, we also trained two lower-capacity models. The first, an AlexNet model15, was trained in the same manner as the DenseNet-121, with the weights randomly initialized rather than pretrained on ImageNet. The second was a logistic regression with ‘deep features’: because individual pixels do not have stable semantic meaning over different samples in the dataset, we first extract a set of 1,024 higher-level features using the feature embedding (that is, the activations of the penultimate layer) of a DenseNet-121 trained on ImageNet and then fit a logistic regression to these fixed features. This procedure is accomplished by training the DenseNet-121 architecture with the weights of its feature embedding subnetwork frozen. The AlexNet and logistic regression were optimized using the same training parameters as the full DenseNet-121 model specified above. The fact that lower-capacity models did not generalize better in our setting may be due to the fact that Sagawa et al. focus on a reweighted training scheme36, while our models were trained to minimize empirical risk to replicate the training schemes used by recent COVID-19 detection models (see above).
Datasets and preprocessing
To train and evaluate our models, we combined images from five large open-access repositories of chest radiographs into three datasets (Fig. 1a and Supplementary Table 1). The first, which we refer to as dataset I, was designed to replicate the datasets used to develop and evaluate the most popular COVID-19 diagnostic models4. In this dataset, we collected COVID-19-negative images from the NIH ChestX-ray14 repository, representing 112,120 radiographs from 30,805 patients from the NIH Clinical Center24. We collected COVID-19-positive images from the GitHub-COVID repository23 (commit ID 9b9c2d5; https://github.com/ieee8023/covid-chestxray-dataset/commit/9b9c2d5), representing 408 radiographs from 262 patients, where the data were originally collected from figures in scientific publications and assorted web sources of COVID-19-positive cases.
The second dataset, which we refer to as dataset II, was designed to represent a more ideal case in terms of domain confounding—both COVID-19-positive and COVID-19-negative images were acquired from hospitals from a common region and were published by a shared research team. We collected COVID-19-negative images from the PadChest repository, representing 96,270 radiographs from 63,939 patients from a hospital in Valencia, Spain29. The COVID-19-positive images in our dataset were taken from the BIMCV-COVID-19+ dataset (version 1), which represents 1,596 images from 1,015 patients (after exclusions), from the same regional hospital system in Valencia, Spain30. We note that while PadChest and BIMCV-COVID-19+ originate from the same region, potential for confounding remains, because (1) PadChest was collected from a single hospital whereas BIMCV-COVID-19+ was collected from multiple hospitals and (2) the repositories were collected over different time periods, over which image acquisition techniques may have changed.
The third dataset, referred to as dataset III, was designed to represent the most ideal case in terms of domain confounding. Unlike dataset II, the COVID-19-positive and COVID-19-negative images were collected not only from the same region, but also from the same hospitals and over the same time period. Like dataset II, the COVID-19-positive images were collected from the BIMCV-COVID-19+ repository. The COVID-19-negative images were taken from the corresponding BIMCV-COVID-19− repository, which includes 3,086 images from 2,327 patients (after exclusions).
Following the recommendations by Cohen et al.48, we filtered radiographs from the online repositories to include only PA and upright AP radiographs. Lateral radiographs, AP supine radiographs, radiographs with unknown projections and computed tomography scans were excluded from the datasets. Images with absent radiographic windowing information, which was necessary to display radiographs from the BIMCV-COVID-19+ and BIMCV-COVID-19− repositories, were also excluded.
We partitioned each repository into training, validation and test folds, ensuring that all radiographs of any given patient belong to a single fold. Because the ChestX-ray14 dataset specifies a ‘test’ partition, we used these radiographs as part of our dataset I test fold. Of the remaining portion, 5% were reserved as a validation fold, while the rest were used directly for training. In the PadChest, BIMCV-COVID-19+ and BIMCV-COVID-19− repositories, we reserved 5% of the radiographs for testing and 5% of the remaining radiographs for validation. Owing to the smaller size of the GitHub-COVID repository, we reserved 10% of the radiographs for testing and 10% of the remaining radiographs for validation. With the exception of the ChestX-ray14 test fold, which was held fixed as explained above, the folds were drawn at random for each model replicate.
Model interpretability using saliency maps
To generate saliency maps, which enable interpretation of machine learning models by assigning importance values to each pixel of an input image, we apply a state-of-the-art approach known as ‘Expected Gradients’19. Broadly, this approach captures the notion of ‘importance’ by tracking how each pixel of an image impacts the output of the model when contrasted with a set of non-informative baseline examples, where the impact is measured by accumulating the model’s gradients (a mathematical measure of a model’s sensitivity to small changes in a feature) as the image is interpolated from the baseline example to the image of interest. Formally, the Expected Gradients attribution ϕ for an input sample x and input feature i is defined as
where D represents a background distribution from which reference samples \(x^{\prime}\) are drawn, f represents the model, and the parameter α enables interpolation between the baseline x′ and the input sample x. This method is an extension of the popular saliency map approach ‘Integrated Gradients’, which is the special case of Expected Gradients in which there is only a single reference sample.
For our application, Expected Gradients improves over Integrated Gradients in terms of the accuracy of its saliency maps19 and the inclusion of multiple reference samples, which avoids the choice of a single reference that may be arbitrary but nonetheless impactful upon the resultant saliency maps49. Finally, path-based approaches like Expected Gradients and Integrated Gradients are preferable to other methods for generating saliency maps because they are theoretically principled: these methods are provably guaranteed to attribute importance to important pixels and guaranteed not to attribute importance to unimportant pixels (Supplementary Note 1) 16.
As the background distribution D for Expected Gradients, we used the COVID-19-negative images from the training dataset for each model we explain. Intuitively, we are explaining how the output of our model for our input image x differs on average from the output of the model for images in the training data D. We demonstrate that Expected Gradients is not overly sensitive to choice of D by comparing the saliency maps for several radiographs with a background distribution of images from the training data to attributions for those same radiographs with a background distribution of images from the external dataset, and find that the resultant attributions are similar (Supplementary Fig. 17).
Data interpretability using CycleGAN
To attain visual explanations of the differences between COVID-19-positive and COVID-19-negative images in each dataset, we aimed to understand which characteristics of the chest radiograph would have to change to make a COVID-19-negative image appear to be a COVID-19-positive image, and vice versa. Formally, let \({\mathcal{X}}\) be a domain of COVID-19-negative images and let \({\mathcal{Y}}\) be a domain of COVID-19-positive images. Our goal is to learn a mapping \(G:{\mathcal{X}}\mapsto {\mathcal{Y}}\) that takes a COVID-19-negative chest radiograph, \(X\in {\mathcal{X}}\), and transforms it so that it is indistinguishable from COVID-19-positive chest radiographs. We also aim to learn the inverse transformation, \(F:{\mathcal{Y}}\mapsto {\mathcal{X}}\).
Because GANs have previously been shown to be effective for the interpretation of neural networks, we learn these two transformations using the CycleGAN approach17,18. The mappings G and F are learned by two neural networks, which are optimized in conjunction with two discriminator networks \({D}_{{\mathcal{Y}}}\) and \({D}_{{\mathcal{X}}}\). These networks are optimized to minimize a series of losses. The first, referred to as the adversarial loss, encourages the mapping functions G and F to match the distribution of generated images from each source domain to the true data distribution of each target domain:
where pdata(X) and pdata(Y) represent the data distributions for each domain. In addition to the adversarial loss, the networks are also trained to enforce cycle consistency, meaning that F(G(X)) = X. This is desirable, because it enforces a similarity between the original and transformed images. The loss here is
The full loss that is optimized then is simply the sum of these three losses:
To understand which image features are important in distinguishing the domains \({\mathcal{X}}\) and \({\mathcal{Y}}\), we transform a COVID-19-negative radiograph \(X\in {\mathcal{X}}\) or a COVID-19-positive radiograph \(Y\in {\mathcal{Y}}\) using the learned generator networks G or F to map the image to the opposite domain. We then compare which image features are changed in the transformation.
Our CycleGAN networks were implemented in Python 3.7 using the PyTorch software library and an open-source implementation of the CycleGAN approach (by Aitor Ruano; https://github.com/aitorzip/PyTorch-CycleGAN). To attain comparable training time, the networks were trained for 3,000 epochs (dataset I) or 1,000 epochs (dataset II). Each network required approximately one week of training time on an NVIDIA RTX 2080 graphics processing unit.
Experimental validation of feature attributions
We experimentally validated our findings from saliency maps and GANs by modifying important radiographic features. To detect whether the higher-level features that our saliency maps highlight are major contributors to the model’s classification, we used methods inspired by a behavioural testing approach50. For example, saliency maps highlight dataset-specific laterality markers and text within the images. If these text markers are indeed important, then moving a marker from a COVID-19-positive image to a COVID-19-negative image should increase the predicted log odds of COVID-19. For a pair of COVID-19-positive and COVID-19-negative images, we swap the text markers and measure the change in the output for each image. To assess the significance of the change in the model’s output at the level of each individual image, we generate empirical P values by comparing to a null distribution generated by swapping 1,000 random patches of each image of the same dimensions as the text markers (Fig. 3a). We conduct a similar experiment to validate whether the shoulder regions frequently highlighted in the saliency maps have a significant impact on the model’s decisions. We observe that the shoulder region of COVID-19-positive images tends to appear at the upper image border, while the shoulder region of COVID-19-negative images appears slightly lower. Furthermore, the saliency maps highlight the clavicles and shoulders of the COVID-19-positive images, but not of the COVID-19-negative images. We hypothesized that the model was looking for the presence of shoulders in the upper corners of the image. To test our hypothesis, we moved the clavicles and shoulders of a COVID-19-negative image to the top corners of the radiograph and measured the change in model output (Fig. 3b). We tested for statistical significance at the level of individual images by generating empirical P values. Our distribution was generated by randomly sampling and replacing 1,000 patches of the same size as the shoulder region, following the same procedure as described for the laterality markers.
To verify the significance of these regions for our models at a population level, we repeated the procedure described in the paragraph above for a sample of randomly selected radiographs from the datasets (Supplementary Fig. 12). For the dataset-specific laterality markers (Supplementary Fig. 12, left), we randomly sampled 10 COVID-19-negative images with laterality or other text markers and 10 COVID-19-positive images with laterality or other text markers. To test for the significance of the text markers across the datasets, we used a Wilcoxon signed rank test to compare the distribution of the magnitudes of changes in model output after swapping the text markers to the distribution of the magnitudes of the average changes in model output after swapping 1,000 random patches of the same size (P = 8.86 × 10−5, Siegel’s T statistic = 0.0). For the positioning of the shoulder regions (Supplementary Fig. 12, right), we randomly sampled 20 COVID-19-negative images. We then used a Wilcoxon signed rank test to compare the distribution of changes in model output after moving the clavicles and shoulder regions to the top of the image with the distribution of the average changes in model output after moving 1,000 random patches of the same size (P = 8.86 × 10−5, Siegel’s T statistic = 0.0).
Statistics
In our experiments involving manual modification of radiographs (Fig. 3a,b and Supplementary Fig. 11), we computed empirical P values by first generating the distribution of the change in the model output (in log odds space) for a set of random, non-specific modifications as described in each caption. The P value was then calculated as (r + 1)/(n + 1) where r is the number of non-specific modifications that produced a greater increase in model output (greater magnitude decrease in Fig. 3a, top row) and n is the total number of non-specific modifications51.
To compare the generalization performance of models (for example, Fig. 6), we performed a two-tailed Mann–Whitney U-test, given that the ROC-AUC values are bounded by 0 and 1 and therefore unlikely to be normally distributed.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this Article.
Data availability
All radiographs are compiled from publicly available data repositories. The ChestX-ray14 repository is available at https://nihcc.app.box.com/v/ChestXray-NIHCC. The GitHub-COVID dataset is available at https://github.com/ieee8023/covid-chestxray-dataset. The PadChest repository is available at https://bimcv.cipf.es/bimcv-projects/padchest/. The BIMCV-COVID19 repositories are available at https://bimcv.cipf.es/bimcv-projects/bimcv-covid19/.
Code availability
All of the code necessary to reproduce our experimental findings can be found at https://github.com/suinleelab/cxr_covid (archived at https://doi.org/10.5281/zenodo.4623792).
References
Mossa-Basha, M. et al. Policies and guidelines for COVID-19 preparedness: experiences from the University of Washington. Radiology https://doi.org/10.1148/radiol.2020201326 (2020).
Kundu, S., Elhalawani, H., Gichoya, J. W. & Kahn, C. E.Jr How might AI and chest imaging help unravel COVID-19’s mysteries? Radiol. Artificial Intell 2, 3 (2020).
Ghoshal, B. & Tucker, A. Estimating uncertainty and interpretability in deep learning for coronavirus (COVID-19) detection. Preprint at https://arxiv.org/pdf/2003.10769.pdf (2020).
Wang, L., Lin, Z. Q. & Wong, A. COVID-Net: a tailored deep convolutional neural network design for detection of COVID-19 cases from chest X-ray images. Sci. Rep. 10, 19549 (2020).
Hemdan, E. E.-D., Shouman, M. A. & Karar, M. E. COVIDX-Net: a framework of deep learning classifiers to diagnose COVID-19 in X-ray images. Preprint at https://arxiv.org/pdf/2003.11055.pdf (2020).
Ozturk, T. et al. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 121, 103792 (2020).
Brunese, L., Mercaldo, F., Reginelli, A. & Santone, A. Explainable deep learning for pulmonary disease and coronavirus COVID-19 detection from X-rays. Comput. Methods Programs Biomed. 196, 105608 (2020).
Karim, M. et al. DeepCOVIDExplainer: explainable COVID-19 predictions based on chest X-ray images. Preprint at https://arxiv.org/pdf/2004.04582.pdf (2020).
Laghi, A. Cautions about radiologic diagnosis of COVID-19 infection driven by artificial intelligence. Lancet Digit. Health 2, e225 (2020).
Geirhos, R. et al. Shortcut learning in deep neural networks. Nat. Mach. Intell. 2, 665–673 (2020).
Harmon, S. A. et al. Artificial intelligence for the detection of COVID-19 pneumonia on chest CT using multinational datasets. Nat. Commun. 11, 4080 (2020).
Lakhani, P. & Sundaram, B. Deep learning at chest radiography: automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology 284, 574–582 (2017).
Al-Masni, M. A., Kim, D.-H. & Kim, T.-S. Multiple skin lesions diagnostics via integrated deep convolutional networks for segmentation and classification. Comput. Methods Programs Biomed. 190, 105351 (2020).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 4700–4708 (IEEE, 2017).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. In Proc. 25th International Conference on Neural Information Processing Systems 1097–1105 (ACM, 2012).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. In Proc. 34th International Conference on Machine Learning Vol. 70, 3319–3328 (PMLR, 2017).
Zhu, J.-Y., Park, T., Isola, P. & Efros, A. A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proc. 2017 IEEE International Conference on Computer Vision 2242–2251 (IEEE, 2017).
Singla, S., Pollack, B., Chen, J. & Batmanghelich, K. Explanation by progressive exaggeration. In International Conference on Learning Representations (2019).
Erion, G., Janizek, J. D., Sturmfels, P., Lundberg, S. M. & Lee, S.-I. Improving performance of deep learning models with axiomatic attribution priors and expected gradients. Nat. Mach. Intell. https://doi.org/10.1038/s42256-021-00343-w (2021).
Zhang, R. et al. Diagnosis of COVID-19 pneumonia using chest radiography: value of artificial intelligence. Radiology https://doi.org/10.1148/radiol.2020202944 (2020).
Sandler, M., Howard, A., Zhu, M., Zhmoginov, A. & Chen, L.-C. MobileNetV2: inverted residuals and linear bottlenecks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 4510–4520 (IEEE, 2018).
Xie, S., Girshick, R., Dollár, P., Tu, Z. & He, K. Aggregated residual transformations for deep neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 5987–5995 (IEEE, 2017).
Cohen, J. P., Morrison, P. & Dao, L. COVID-19 image data collection. GitHub https://github.com/ieee8023/covid-chestxray-dataset
Wang, X. et al. ChestX-ray8: hospital-scale chest X-ray database and benchmarks on weakly-supervised classification and localization of common thorax diseases. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 2097–2106 (IEEE, 2017).
Radiological Society of North America. RSNA pneumonia detection challenge. kaggle https://www.kaggle.com/c/rsna-pneumonia-detection-challenge/data
Wehbe, R. M. et al. DeepCOVID-XR: an artificial intelligence algorithm to detect COVID-19 on chest radiographs trained and tested on a large US clinical dataset. Radiology https://doi.org/10.1148/radiol.2020203511 (2020).
Li, M. D. et al. Automated assessment and tracking of COVID-19 pulmonary disease severity on chest radiographs using convolutional Siamese neural networks. Radiol. Artif. Intell. 2, e200079 (2020).
Murphy, K. et al. COVID-19 on chest radiographs: a multireader evaluation of an artificial intelligence system. Radiology 296, E166–E172 (2020).
Bustos, A., Pertusa, A., Salinas, J.-M. & de la Iglesia-Vayá, M. PadChest: a large chest X-ray image dataset with multi-label annotated reports. Med. Image Anal. 66, 101797 (2020).
Vayá, M. d. l. I. et al. BIMCV COVID-19+: a large annotated dataset of RX and CT images from COVID-19 patients. Preprint at https://arxiv.org/pdf/2006.01174.pdf (2020).
Maguolo, G. & Nanni, L. A critic evaluation of methods for COVID-19 automatic detection from X-ray images. Inform. Fusion 76, 1–7 (2021).
Castro, D. C., Walker, I. & Glocker, B. Causality matters in medical imaging. Nat. Commun. 11, 3673 (2020).
Richens, J. G., Lee, C. M. & Johri, S. Improving the accuracy of medical diagnosis with causal machine learning. Nat. Commun. 11, 3923 (2020).
Janizek, J. D., Erion, G., DeGrave, A. J. & Lee, S.-I. An adversarial approach for the robust classification of pneumonia from chest radiographs. In Proc. ACM Conference on Health, Inference and Learning 69–79 (ACM, 2020).
Ganin, Y. et al. Domain-adversarial training of neural networks. J. Mach. Learn. Res. 17, 2096–2030 (2016).
Sagawa, S., Raghunathan, A., Koh, P. W. & Liang, P. An investigation of why overparameterization exacerbates spurious correlations. In Proc. 37th International Conference on Machine Learning (ICML) Vol. 119, 8346–8356 (PMLR, 2020).
Bressem, K. K. et al. Comparing different deep learning architectures for classification of chest radiographs. Sci. Rep. 10, 13590 (2020).
Quionero-Candela, J., Sugiyama, M., Schwaighofer, A. & Lawrence, N. D. Dataset Shift in Machine Learning (MIT Press, 2009).
Rajpurkar, P. et al. CheXNet: radiologist-level pneumonia detection on chest X-rays with deep learning. Preprint at https://arxiv.org/pdf/1711.05225.pdf (2017).
Mitani, A. et al. Detection of anaemia from retinal fundus images via deep learning. Nat. Biomed. Eng. 4, 18–27 (2020).
Zech, J. R. et al. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: a cross-sectional study. PLoS Med. 15, e1002683 (2018).
Ng, M.-Y. et al. Imaging profile of the COVID-19 infection: radiologic findings and literature review. Radiol. Cardiothorac. Imaging 2, e200034 (2020).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. Int. J. Comput. Vision 128, 336–359 (2020).
Wong, H. Y. F. et al. Frequency and distribution of chest radiographic findings in COVID-19 positive patients. Radiology https://doi.org/10.1148/radiol.2020201160 (2020).
Gale, W., Oakden-Rayner, L., Carneiro, G., Bradley, A. P. & Palmer, L. J. Detecting hip fractures with radiologist-level performance using deep neural networks. Preprint at https://arxiv.org/pdf/1711.06504.pdf (2017).
Deng, J. et al. ImageNet: a large-scale hierarchical image database. In Proc. 2009 IEEE Conference on Computer Vision and Pattern Recognition 248–255 (IEEE, 2009).
Paszke, A. et al. PyTorch: an imperative style, high-performance deep learning library. In Advances in Neural Information Processing Systems 32 (eds Wallach, H. et al.) 8024–8035 (Curran Associates, 2019); http://papers.neurips.cc/paper/9015-pytorch-an-imperative-style-high-performance-deep-learning-library.pdf
Cohen, J. P. et al. 2020. COVID-19 image data collection: prospective predictions are the future. GitHub https://github.com/ieee8023/covid-chestxray-dataset
Sturmfels, P., Lundberg, S. & Lee, S.-I. Visualizing the impact of feature attribution baselines. Distill 5, e22 (2020).
Ribeiro, M. T., Wu, T., Guestrin, C. & Singh, S. Beyond accuracy: behavioral testing of NLP models with checklist. In Proc. 58th Annual Meeting of the Association for Computational Linguistics 4902–4912 (Association for Computational Linguistics, 2020); https://www.aclweb.org/anthology/2020.acl-main.442
North, B. V., Curtis, D. & Sham, P. C. A note on the calculation of empirical P values from Monte Carlo procedures. Am. J. Human Genet. 71, 439–441 (2002).
Winther, H. et al. COVID-19 image repository. figshare https://doi.org/10.6084/m9.figshare.12275009
Jin, Y.-H. et al. A rapid advice guideline for the diagnosis and treatment of 2019 novel coronavirus (2019-nCoV) infected pneumonia (standard version). Mil. Med. Res. 7, 4 (2020).
Acknowledgements
This work was funded by the National Science Foundation (CAREER DBI-1552309 to S.-I.L.) and the National Institutes of Health (R35 GM 128638 and R01 AG061132 to S.-I.L.). We thank H. Chen and G. Erion for providing feedback while the manuscript was being written. We thank A. Bustos for clarifying the characteristics of the PadChest and BIMCV-COVID-19+ datasets. We also thank D. Janizek for insight into the interpretation of COVID-19 on chest radiographs.
Author information
Authors and Affiliations
Contributions
J.D.J. conceived the study. A.J.D. and J.D.J. prepared datasets, designed experiments and wrote software. S.-I.L. supervised the study. A.J.D., J.D.J. and S.-I.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Machine Intelligence thanks Kayhan Batmanghelich and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Note 1 and Figs. 1–17.
Rights and permissions
About this article
Cite this article
DeGrave, A.J., Janizek, J.D. & Lee, SI. AI for radiographic COVID-19 detection selects shortcuts over signal. Nat Mach Intell 3, 610–619 (2021). https://doi.org/10.1038/s42256-021-00338-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-021-00338-7
This article is cited by
-
Generalizable disease detection using model ensemble on chest X-ray images
Scientific Reports (2024)
-
Analysis of large-language model versus human performance for genetics questions
European Journal of Human Genetics (2024)
-
Modelling dataset bias in machine-learned theories of economic decision-making
Nature Human Behaviour (2024)
-
Towards trustworthy seizure onset detection using workflow notes
npj Digital Medicine (2024)
-
Improving deep neural network generalization and robustness to background bias via layer-wise relevance propagation optimization
Nature Communications (2024)