Abstract
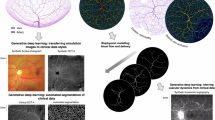
Vascular disease is one of the leading causes of death and threatens human health worldwide. Imaging examination of vascular pathology with reduced invasiveness is challenging due to the intrinsic vasculature complexity and non-uniform scattering from bio-tissues. Here, we report VasNet, a vasculature-aware unsupervised learning algorithm that augments pathovascular recognition from small sets of unlabelled fluorescence and digital subtraction angiography images. VasNet adopts a multi-scale fusion strategy with a domain adversarial neural network loss function that induces biased pattern reconstruction by strengthening features relevant to the retinal vasculature reference while weakening irrelevant features. VasNet delivers the outputs ‘Structure + X’ (where X refers to multi-dimensional features such as blood flows, the distinguishment of blood dilation and its suspicious counterparts, and the dependence of new pattern emergence on disease progression). Therefore, explainable imaging output from VasNet and other algorithm extensions holds the promise to augment medical diagnosis, as it improves performance while reducing the cost of human expertise, equipment and time consumption.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
A pretrained model and all experimental images appearing in the Article and the Supplementary Information are publicly available at https://doi.org/10.6084/m9.figshare.11986962. Other detailed data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Our PyTorch implementation of VasNet, training data and test samples are available from GitHub (https://github.com/mjiUST/VasNet) or https://doi.org/10.5281/zenodo.3762820.
Change history
08 January 2021
A Correction to this paper has been published: https://doi.org/10.1038/s42256-021-00292-4
References
Tortora, G. J. & Derrickson, B. H. Principles of Anatomy and Physiology (Wiley, 2018).
Field, T. S. & Hill, M. D. Cerebral venous thrombosis: we should ask the right questions to get better answers. Stroke 50, 1598–1604 (2019).
Portegies, M., Koudstaal, P. & Ikram, M. Cerebrovascular disease. Handbook Clin. Neurol. 138, 239–261 (2016).
Hu, X., De Silva, T. M., Chen, J. & Faraci, F. M. Cerebral vascular disease and neurovascular injury in ischemic stroke. Circ. Res. 120, 449–471 (2017).
Brown, R. D. Jr & Broderick, J. P. Unruptured intracranial aneurysms: epidemiology, natural history, management options and familial screening. Lancet Neurol. 13, 393–404 (2014).
Kisler, K., Nelson, A. R., Montagne, A. & Zlokovic, B. V. Cerebral blood flow regulation and neurovascular dysfunction in Alzheimer disease. Nat. Rev. Neurosci. 18, 419–434 (2017).
Sweeney, M. D., Kisler, K., Montagne, A., Toga, A. W. & Zlokovic, B. V. The role of brain vasculature in neurodegenerative disorders. Nat. Neurosci. 21, 1318–1331 (2018).
Franks, I. Gut microbes might promote intestinal angiogenesis. Nat. Rev. Gastroenterol. Hepatol. 10, 3 (2013).
Hanauer, S. B. Update on the etiology, pathogenesis and diagnosis of ulcerative colitis. Nat. Clin. Pract. Gastroenterol. Hepatol. 1, 26–31 (2004).
Torres, J., Mehandru, S., Colombel, J.-F. & Peyrin-Biroulet, L. Crohn’s disease. Lancet 389, 1741–1755 (2017).
Lopera, J. E. Embolization in trauma: principles and techniques. Semin. Intervent. Radiol. 27, 014–028 (2010).
Burke, C. T. & Mauro, M. A. Bronchial artery embolization. Semin. Intervent. Radiol. 21, 43–48 (2004).
Rilling, W. S. & Chen, G. W. Preoperative embolization. Semin. Intervent. Radiol. 21, 3–9 (2004).
Hong, G., Antaris, A. L. & Dai, H. Near-infrared fluorophores for biomedical imaging. Nat. Biomed. Eng. 1, 0010 (2017).
Wan, H. et al. A bright organic NIR-II nanofluorophore for three-dimensional imaging into biological tissues. Nat. Commun. 9, 1171 (2018).
Hong, G. et al. Through-skull fluorescence imaging of the brain in a new near-infrared window. Nat. Photon. 8, 723–730 (2014).
Martínez-Corral, I. et al. In vivo imaging of lymphatic vessels in development, wound healing, inflammation and tumor metastasis. Proc. Natl Acad. Sci. USA 109, 6223–6228 (2012).
Jia, Y. et al. Quantitative optical coherence tomography angiography of vascular abnormalities in the living human eye. Proc. Natl Acad. Sci. USA 112, E2395–E2402 (2015).
Zhang, W. et al. High-resolution, in vivo multimodal photoacoustic microscopy, optical coherence tomography and fluorescence microscopy imaging of rabbit retinal neovascularization. Light Sci. Appl. 7, 103 (2018).
Sakadžić, S. & Wang, L. V. High-resolution ultrasound-modulated optical tomography in biological tissues. Opt. Lett. 29, 2770–2772 (2004).
Yao, J., Maslov, K. I., Shi, Y., Taber, L. A. & Wang, L. V. In vivo photoacoustic imaging of transverse blood flow by using Doppler broadening of bandwidth. Opt. Lett. 35, 1419–1421 (2010).
Huang, C.-H. et al. High-resolution structural and functional assessments of cerebral microvasculature using 3D Gas ΔR2*-mMRA. PLoS ONE 8, e78186 (2013).
Wang, L. V. & Yao, J. A practical guide to photoacoustic tomography in the life sciences. Nat. Methods 13, 627–638 (2016).
Nasiriavanaki, M. et al. High-resolution photoacoustic tomography of resting-state functional connectivity in the mouse brain. Proc. Natl Acad. Sci. USA 111, 21–26 (2014).
Meijering, E. H., Niessen, W. J. & Viegever, M. Retrospective motion correction in digital subtraction angiography: a review. IEEE Trans. Med. Imaging 18, 2–21 (1999).
Jeans, W. The development and use of digital subtraction angiography. Br. J. Radiol. 63, 161–168 (1990).
Christiansen, E. M. et al. In silico labeling: predicting fluorescent labels in unlabeled images. Cell 173, 792–803 (2018).
Wang, Y. et al. Accurate quantification of astrocyte and neurotransmitter fluorescence dynamics for single-cell and population-level physiology. Nat. Neurosci. 22, 1936–1944 (2019).
Rivenson, Y. et al. Virtual histological staining of unlabelled tissue-autofluorescence images via deep learning. Nat. Biomed. Eng. 3, 466–477 (2019).
Kermany, D. S. et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell 172, 1122–1131 (2018).
Cole, J. H. et al. Predicting brain age with deep learning from raw imaging data results in a reliable and heritable biomarker. NeuroImage 163, 115–124 (2017).
Li, S., Deng, M., Lee, J., Sinha, A. & Barbastathis, G. Imaging through glass diffusers using densely connected convolutional networks. Optica 5, 803–813 (2018).
Li, Y., Xue, Y. & Tian, L. Deep speckle correlation: a deep learning approach toward scalable imaging through scattering media. Optica 5, 1181–1190 (2018).
Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 1, 206–215 (2019).
Lee, H. et al. An explainable deep-learning algorithm for the detection of acute intracranial haemorrhage from small datasets. Nat. Biomed. Eng. 3, 173–182 (2019).
Mahapatra, D., Bozorgtabar, B., Hewavitharanage, S. & Garnavi, R. Image super resolution using generative adversarial networks and local saliency maps for retinal image analysis. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention 382–390 (Springer, 2017).
Son, J., Park, S. J. & Jung, K.-H. Retinal vessel segmentation in fundoscopic images with generative adversarial networks. Preprint at https://arxiv.org/abs/1706.09318 (2017).
Zhu, J.-Y., Park, T., Isola, P. & Efros, A. A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proceedings of the IEEE International Conference on Computer Vision 2223–2232 (IEEE, 2017).
Murray, C. D. The physiological principle of minimum work: I. The vascular system and the cost of blood volume. Proc. Natl Acad. Sci. USA 12, 207–214 (1926).
West, G. B., Brown, J. H. & Enquist, B. J. A general model for the origin of allometric scaling laws in biology. Science 276, 122–126 (1997).
Ganin, Y. et al. Domain-adversarial training of neural networks. J. Mach. Learn. Res. 17, 2096–2030 (2016).
Dey, N. et al. Richardson–Lucy algorithm with total variation regularization for 3D confocal microscope deconvolution. Microsc. Res. Tech. 69, 260–266 (2006).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Lu, B., Chen, J.-C. & Chellappa, R. Unsupervised domain-specific deblurring via disentangled representations. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 10225–10234 (IEEE, 2019).
Dice, L. R. Measures of the amount of ecologic association between species. Ecology 26, 297–302 (1945).
Gegundez-Arias, M. E., Aquino, A., Bravo, J. M. & Marin, D. A function for quality evaluation of retinal vessel segmentations. IEEE Trans. Med. Imaging 31, 231–239 (2012).
Laroui, H. et al. Dextran sodium sulfate (DSS) induces colitis in mice by forming nano-lipocomplexes with medium-chain-length fatty acids in the colon. PLoS ONE 7, e32084 (2012).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay‐Kumar, M. Dextran sulfate sodium (DSS)‐induced colitis in mice. Curr. Protoc. Immunol. 104, 15–25 (2014).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
O’Donnell, T. F. Jr, Rasmussen, J. C. & Sevick-Muraca, E. M. New diagnostic modalities in the evaluation of lymphedema. J. Vasc. Surg, Venous Lymphat. Disord. 5, 261–273 (2017).
Greives, M. R., Aldrich, M. B., Sevick-Muraca, E. M. & Rasmussen, J. C. Near-infrared fluorescence lymphatic imaging of a toddler with congenital lymphedema. Pediatrics 139, e20154456 (2017).
Tzeng, E., Hoffman, J., Darrell, T. & Saenko, K. Simultaneous deep transfer across domains and tasks. In Proceedings of the IEEE International Conference on Computer Vision 4068–4076 (IEEE, 2015).
Fraz, M. M. et al. Blood vessel segmentation methodologies in retinal images—a survey. Comput. Methods Programs Biomed. 108, 407–433 (2012).
Staal, J., Abràmoff, M. D., Niemeijer, M., Viergever, M. A. & Van Ginneken, B. Ridge-based vessel segmentation in color images of the retina. IEEE Trans. Med. Imaging 23, 501–509 (2004).
Watts, L. T., Zheng, W., Garling, R. J., Frohlich, V. C. & Lechleiter, J. D. Rose Bengal photothrombosis by confocal optical imaging in vivo: a model of single vessel stroke. J. Visualized Exp. 23, e52794 (2015).
Acknowledgements
We thank H. Fan for insightful suggestions regarding the optics experiments, X. Yuan for camera troubleshooting, J. Cai for fruitful discussions about data processing in the bowel vasculature section and H. Zhao for drawing illustrations. The work was supported by the National Natural Science Foundation of China (grants 61722209, 61860206003, 61971255 and 81872368), funds from the Shenzhen Science and Technology Innovation Committee (grants KQJSCX20180327143623167 and JCYJ20180508152130899) and a fund from Shenzhen Development and Reform Commission Subject Construction Project [2017]143.
Author information
Authors and Affiliations
Contributions
L.F., S.M., Q.D. and L.H. conceived this project. S.M., L.F., Y.W., S.J. and M.J. designed the experiments. Y.W. and S.J. performed animal experiments. M.J. innovated and implemented the algorithm. M.J., X.W., Y.W. and J.F. tested the algorithm and performed the data analysis. F.D. contributed the X-ray and DSA data under ethical approval from the PLA General Hospital. Q.D. oversaw the experimental design and progress. S.M., L.F., M.J., Y.W. and J.W. prepared the manuscript. All authors reviewed the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14, Discussion and Table 1.
Supplementary Video 1
In vivo fluorescence imaging of cerebral vasculature of a healthy mouse at a 25 Hz frame acquisition rate. As ICG is injected, VasNet recovers the dynamic changes of vascular structure of a healthy cerebrum.
Supplementary Video 2
In vivo fluorescence imaging of cerebral vasculature of a mouse with thrombosis model at a 25 Hz frame acquisition rate. As ICG is injected, VasNet recovers the dynamic changes of vascular structure of a diseased cerebrum.
Supplementary Video 3
Augmenting diagnosis of bleeding from dynamic DSA vascular images acquired from human. VasNet outputs image with vasculature reconstructed. The bleeding features are augmented along with the dynamics of dilation.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Y., Ji, M., Jiang, S. et al. Augmenting vascular disease diagnosis by vasculature-aware unsupervised learning. Nat Mach Intell 2, 337–346 (2020). https://doi.org/10.1038/s42256-020-0188-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-020-0188-z