Abstract
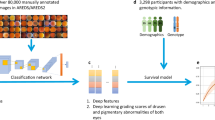
Both genetic and environmental factors influence the etiology of age-related macular degeneration (AMD), a leading cause of blindness. AMD severity is primarily measured by images of the fundus of the retina and recently developed machine learning methods can successfully predict AMD progression using image data. However, none of these methods have used both genetic and image data for predicting AMD progression. Here we used both genotypes and fundus images to predict whether an eye had progressed to late AMD with a modified deep convolutional neural network. In total, we used 31,262 fundus images and 52 AMD-associated genetic variants from 1,351 subjects from the Age-Related Eye Disease Study, which provided disease severity phenotypes and fundus images available at baseline and follow-up visits over a period of 12 years. Our results showed that fundus images coupled with genotypes could predict late AMD progression with an averaged area-under-the-curve value of 0.85 (95% confidence interval 0.83–0.86). The results using fundus images alone showed an averaged area under the receiver operating characteristic curve value of 0.81 (95% confidence interval 0.80–0.83). We implemented our model in a cloud-based application for individual risk assessment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All the phenotype data and fundus images of AREDS participants required are available from dbGap (accession phs000001.v3.p1). The genotype data on AREDS subjects have been reported earlier9 and are available from dbGap (accession phs001039.v1.p1). The UK Biobank test dataset was obtained from UK Biobank (application number 43252).
Code availability
The prediction models with Python implementation and a detailed tutorial are available at https://github.com/QiYanPitt/AMDprogressCNN and a web-based graphical user interface is also available at http://www.pitt.edu/~qiy17/amdprediction.html.
References
Stark, K. et al. The German AugUR study: study protocol of a prospective study to investigate chronic diseases in the elderly. BMC Geriatr. 15, 130 (2015).
Congdon, N. et al. Causes and prevalence of visual impairment among adults in the United States. Arch. Ophthalmol. 122, 477–485 (2004).
Swaroop, A., Chew, E. Y., Rickman, C. B. & Abecasis, G. R. Unraveling a multifactorial late-onset disease: from genetic susceptibility to disease mechanisms for age-related macular degeneration. Annu. Rev. Genom. Hum. Genet. 10, 19–43 (2009).
Fritsche, L. G. et al. Age-related macular degeneration: genetics and biology coming together. Annu. Rev. Genom. Hum. Genet. 15, 151–171 (2014).
Ratnapriya, R. & Chew, E. Y. Age-related macular degeneration-clinical review and genetics update. Clin. Genet. 84, 160–166 (2013).
Grassmann, F., Fauser, S. & Weber, B. H. The genetics of age-related macular degeneration (AMD)–novel targets for designing treatment options? Eur. J. Pharm. Biopharm. 95, 194–202 (2015).
Holz, F. G. et al. Progression of geographic atrophy and impact of fundus autofluorescence patterns in age-related macular degeneration. Am. J. Ophthalmol. 143, 463–472 (2007).
Fritsche, L. G. et al. Seven new loci associated with age-related macular degeneration. Nat. Genet. 45, 439 (2013).
Fritsche, L. G. et al. A large genome-wide association study of age-related macular degeneration highlights contributions of rare and common variants. Nat. Genet. 48, 134–143 (2016).
Yan, Q. et al. Genome-wide analysis of disease progression in age-related macular degeneration. Hum. Mol. Genet. 27, 929–940 (2018).
Chen, W. et al. Genetic variants near TIMP3 and high-density lipoprotein-associated loci influence susceptibility to age-related macular degeneration. Proc. Natl Acad. Sci. USA 107, 7401–7406 (2010).
The Age-Related Eye Disease Study Research Group. The Age-Related Eye Disease Study (AREDS): design implications. AREDS Report No. 1. Control Clin. Trials 20, 573–600 (1999).
Abramoff, M. D. et al. Improved automated detection of diabetic retinopathy on a publicly available dataset through integration of deep learning. Invest. Ophthalmol. Vis. Sci. 57, 5200–5206 (2016).
Gulshan, V. et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. J. Am. Med. Assoc. 316, 2402–2410 (2016).
Kermany, D. S. et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell 172, 1122–1131 (2018).
Devalla, S. K. et al. A deep learning approach to digitally stain optical coherence tomography images of the optic nerve head. Invest. Ophthalmol. Vis. Sci. 59, 63–74 (2018).
Grassmann, F. et al. A deep learning algorithm for prediction of age-related eye disease study severity scale for age-related macular degeneration from color fundus photography. Ophthalmology 125, 1410–1420 (2018).
Burlina, P. M. et al. Automated grading of age-related macular degeneration from color fundus images using deep convolutional neural networks. JAMA Ophthalmol. 135, 1170–1176 (2017).
Poplin, R. et al. Prediction of cardiovascular risk factors from retinal fundus photographs via deep learning. Nat. Biomed. Eng. 2, 158–164 (2018).
Peng, Y. et al. DeepSeeNet: a deep learning model for automated classification of patient-based age-related macular degeneration severity from color fundus photographs. Ophthalmology 126, 565–575 (2019).
Abramoff, M. D., Garvin, M. K. & Sonka, M. Retinal imaging and image analysis. IEEE Rev. Biomed. Eng. 3, 169–208 (2010).
Yousefi, S. et al. Learning from data: recognizing glaucomatous defect patterns and detecting progression from visual field measurements. IEEE Trans. Biomed. Eng. 61, 2112–2124 (2014).
Chaudhuri, S., Chatterjee, S., Katz, N., Nelson, M. & Goldbaum, M. Detection of blood vessels in retinal images using two-dimensional matched filters. IEEE Trans. Med. Imaging 8, 263–269 (1989).
Burlina, P. M. et al. Use of deep learning for detailed severity characterization and estimation of 5-year risk among patients with age-related macular degeneration. JAMA Ophthalmol. 136, 1359–1366 (2018).
Youden, W. J. Index for rating diagnostic tests. Cancer 3, 32–35 (1950).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Schmidt-Erfurth, U. et al. Prediction of individual disease conversion in early AMD using artificial intelligence. Invest. Ophthalmol. Vis. Sci. 59, 3199–3208 (2018).
Hao, J., Kim, Y., Mallavarapu, T., Oh, J. H. & Kang, M. Cox-PASNet: pathway-based sparse deep neural network for survival analysis. In 2018 IEEE Int. Conf. on Bioinformatics and Biomedicine (BIBM) 381–386 (IEEE, 2018).
The Age-Related Eye Disease Study Research Group. The Age-Related Eye Disease Study system for classifying age-related macular degeneration from stereoscopic color fundus photographs. AREDS Report No. 6. Am. J. Ophthalmol. 132, 668–681 (2001).
Davis, M. D. et al. The Age-Related Eye Disease Study severity scale for age-related macular degeneration. AREDS Report No. 17. Arch. Ophthalmol. 123, 1484–1498 (2005).
Swaroop, A., Branham, K. E., Chen, W. & Abecasis, G. Genetic susceptibility to age-related macular degeneration: a paradigm for dissecting complex disease traits. Hum. Mol. Genet. 16, R174–R182 (2007).
Ferris, F. L. 3rd et al. Clinical classification of age-related macular degeneration. Ophthalmology 120, 844–851 (2013).
Angermueller, C., Parnamaa, T., Parts, L. & Stegle, O. Deep learning for computational biology. Mol. Syst. Biol. 12, 878 (2016).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Szegedy, C., Vanhoucke V., Ioffe S., Shlens J. & Wojna Z. Rethinking the inception architecture for computer vision. In Proc. IEEE Conf. on Computer Vision and Pattern Recognition (IEEE, 2016).
Christopher, M. et al. Performance of deep learning architectures and transfer learning for detecting glaucomatous optic neuropathy in fundus photographs. Sci. Rep. 8, 16685 (2018).
Russakovsky, O. et al. ImageNet large scale visual recognition challenge. Int. J. Comp. Sci. 115, 211–252 (2015).
Ding, Y. et al. Bivariate analysis of age-related macular degeneration progression using genetic risk scores. Genetics 206, 119–133 (2017).
Chollet, F. Deep learning for humans: keras. GitHub https://github.com/keras-team/keras (2015).
Graf, E., Schmoor, C., Sauerbrei, W. & Schumacher, M. Assessment and comparison of prognostic classification schemes for survival data. Stat. Med. 18, 2529–2545 (1999).
Kotikalapudi, R. et al. Neural network visualization toolkit for keras. GitHub https://github.com/raghakot/keras-vis (2017).
Ribeiro, M. T., Singh, S. & Guestrin, C. “Why should I trust you?”: explaining the predictions of any classifier. In Proc. 22nd ACM SIGKDD Int. Conf. on Knowledge Discovery and Data Mining 1135–1144 (ACM, 2016).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44, 837–845 (1988).
Acknowledgements
H.H. was partially supported by NSF IIS 1852606 and IIS 1837956. A.S. was supported by NEI Intramural Research Program grant number ZIAEY000546. The independent validation has been conducted using the UK Biobank Resource under Application Number 43252.
Author information
Authors and Affiliations
Contributions
Q.Y., Y.D. and W.C. conceived the project and supervised it. Q.Y. performed data processing and analysis. Q.Y. and H.X. designed the study. Q.Y., A.S. and E.Y.C. interpreted the data. Q.Y. and D.E.W. wrote the paper and Y.D., W.C., H.H., A.S. and E.Y.C. commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11 and Tables 1–3
Rights and permissions
About this article
Cite this article
Yan, Q., Weeks, D.E., Xin, H. et al. Deep-learning-based prediction of late age-related macular degeneration progression. Nat Mach Intell 2, 141–150 (2020). https://doi.org/10.1038/s42256-020-0154-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-020-0154-9
This article is cited by
-
Deep learning predicts prevalent and incident Parkinson’s disease from UK Biobank fundus imaging
Scientific Reports (2024)
-
Automated diagnosis of plus disease in retinopathy of prematurity using quantification of vessels characteristics
Scientific Reports (2024)
-
Automatic diagnosis of keratitis using object localization combined with cost-sensitive deep attention convolutional neural network
Journal of Big Data (2023)
-
Developing and validating a multivariable prediction model which predicts progression of intermediate to late age-related macular degeneration—the PINNACLE trial protocol
Eye (2023)
-
Ocular disease detection systems based on fundus images: a survey
Multimedia Tools and Applications (2023)