Abstract
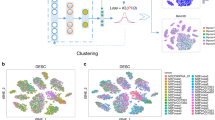
Single-cell RNA sequencing (scRNA-seq) promises to provide higher resolution of cellular differences than bulk RNA sequencing. Clustering transcriptomes profiled by scRNA-seq has been routinely conducted to reveal cell heterogeneity and diversity. However, clustering analysis of scRNA-seq data remains a statistical and computational challenge, due to the pervasive dropout events obscuring the data matrix with prevailing ‘false’ zero count observations. Here, we have developed scDeepCluster, a single-cell model-based deep embedded clustering method, which simultaneously learns feature representation and clustering via explicit modelling of scRNA-seq data generation. Based on testing extensive simulated data and real datasets from four representative single-cell sequencing platforms, scDeepCluster outperformed state-of-the-art methods under various clustering performance metrics and exhibited improved scalability, with running time increasing linearly with sample size. Its accuracy and efficiency make scDeepCluster a promising algorithm for clustering large-scale scRNA-seq data.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The scRNA-seq data that support the findings of this study are available in GitHub: https://github.com/ttgump/scDeepCluster/tree/master/scRNA-seq%20data.
Code availability
The source code, weights of trained models and the real scRNA-seq data used for experiments of scDeepCluster are available in GitHub: https://github.com/ttgump/scDeepCluster.
References
Shapiro, E., Biezuner, T. & Linnarsson, S. Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat. Rev. Genet. 14, 618–630 (2013).
Kolodziejczyk, A. A., Kim, J. K., Svensson, V., Marioni, J. C. & Teichmann, S. A. The technology and biology of single-cell RNA sequencing. Mol. Cell 58, 610–620 (2015).
MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proc. Fifth Berkeley Symposium on Mathematical Statistics and Probability Vol. 1, 281–297 (Univ. of California Press, 1967).
Bishop, C. Pattern Recognition and Machine Learning (Springer, 2006).
von Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 17, 395–416 (2007).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Han, X. et al. Mapping the mouse cell atlas by Microwell-seq. Cell 172, 1091–1107 (2018).
Angerer, P. et al. Single cells make big data: new challenges and opportunities in transcriptomics. Curr. Opin. Syst. Biol. 4, 85–91 (2017).
Xu, C. & Su, Z. Identification of cell types from single-cell transcriptomes using a novel clustering method. Bioinformatics 31, 1974–1980 (2015).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Zhang, J. M., Fan, J., Fan, H. C., Rosenfeld, D. & Tse, D. N. An interpretable framework for clustering single-cell RNA-seq datasets. BMC Bioinformatics 19, 93 (2018).
Wang, B., Zhu, J., Pierson, E., Ramazzotti, D. & Batzoglou, S. Visualization and analysis of single-cell RNA-seq data by kernel-based similarity learning. Nat. Methods 14, 414–416 (2017).
Park, S. & Zhao, H. Spectral clustering based on learning similarity matrix. Bioinformatics 34, 2069–2076 (2018).
Jianbo, S. & Malik, J. Normalized cuts and image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 22, 888–905 (2000).
Lin, P., Troup, M. & Ho, J. W. CIDR: ultrafast and accurate clustering through imputation for single-cell RNA-seq data. Genome Biol. 18, 59 (2017).
Li, W. V. & Li, J. J. An accurate and robust imputation method scImpute for single-cell RNA-seq data. Nat. Commun. 9, 997 (2018).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
Huang, M. et al. SAVER: gene expression recovery for single-cell RNA sequencing. Nat. Methods 15, 539–542 (2018).
Arisdakessian, C., Poirion, O., Yunits, B., Zhu, X. & Garmire, L. DeepImpute: an accurate, fast and scalable deep neural network method to impute single-cell RNA-seq data. Preprint at https://doi.org/10.1101/353607 (2018).
Eraslan, G., Simon, L. M., Mircea, M., Mueller, N. S. & Theis, F. J. Single-cell RNA-seq denoising using a deep count autoencoder. Nat. Commun. 10, 390 (2019).
Deng, Y., Bao, F., Dai, Q., Wu, L. & Altschuler, S. Massive single-cell RNA-seq analysis and imputation via deep learning. Preprint at https://doi.org/10.1101/315556 (2018).
Hinton, G. E. & Salakhutdinov, R. R. Reducing the dimensionality of data with neural networks. Science 313, 504–507 (2006).
Chen, J. et al. An omnibus test for differential distribution analysis of microbiome sequencing data. Bioinformatics 34, 643–651 (2018).
Bellman, R. E. Adaptive Control Processes: A Guided Tour (Princeton Univ. Press, 1961).
Hornik, K. Approximation capabilities of multilayer feedforward networks. Neural Netw. 4, 251–257 (1991).
Bengio, Y., Courville, A. & Vincent, P. Representation learning: a review and new perspectives. IEEE Trans. Pattern Anal. Mach. Intell. 35, 1798–1828 (2013).
Xie, J., Girshick, R. & Farhadi, A. Unsupervised deep embedding for clustering analysis. In Proc. 33rd International Conference on Machine Learning 478–487 (2016).
Guo, X., Gao, L., Liu, X. & Yin, J. Improved deep embedded clustering with local structure preservation. In Proc. 26th International Joint Conference on Artificial Intelligence 1753–1759 (2017).
Lin, C., Jain, S., Kim, H. & Bar-Joseph, Z. Using neural networks for reducing the dimensions of single-cell RNA-seq data. Nucleic Acids Res. 45, e156 (2017).
Ding, J., Condon, A. & Shah, S. P. Interpretable dimensionality reduction of single cell transcriptome data with deep generative models. Nat. Commun. 9, 2002 (2018).
Vincent, P., Larochelle, H., Bengio, Y. & Manzagol, P.-A. Extracting and composing robust features with denoising autoencoders. In Proc. 25th International Conference on Machine Learning 1096–1103 (2008).
Vincent, P., Larochelle, H., Lajoie, I., Bengio, Y. & Manzagol, P.-A. Stacked denoising autoencoders: learning useful representations in a deep network with a local denoising criterion. J. Mach. Learn. Res. 11, 3371–3408 (2010).
Zappia, L., Phipson, B. & Oshlack, A. Splatter: simulation of single-cell RNA sequencing data. Genome Biol. 18, 174 (2017).
Strehl, A. & Ghosh, J. Cluster ensembles—a knowledge reuse framework for combining multiple partitions. J. Mach. Learn. Res. 3, 583–617 (2003).
Hubert, L. & Arabie, P. Comparing partitions. J. Classif. 2, 193–218 (1985).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Dizaji, K. G., Herandi, A., Deng, C., Cai, W. & Huang, H. Deep clustering via joint convolutional autoencoder embedding and relative entropy minimization. In Proc. IEEE International Conference on Computer Vision 5747–5756 (IEEE, 2017).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Nair, V. & Hinton, G. E. Rectified linear units improve restricted Boltzmann machines. In Proc. 27th International Conference on Machine Learning 807–814 (Omnipress, 2010).
Maaten, L. Learning a parametric embedding by preserving local structure. In Proceedings of the Twelth International Conference on Artificial Intelligence and Statistics Vol. 5 (eds Van Dyk, D. & Welling M.) 384–391 (PMLR, 2009).
Nigam, K. & Ghani, R. Analyzing the effectiveness and applicability of co-training. In Proc. Ninth International Conference on Information and Knowledge Management Vol. 5, 86–93 (2000).
Abadi, M. et al. TensorFlow: a system for large-scale machine learning. In Proc. 12th USENIX Conference on Operating Systems Design and Implementation 265–283 (USENIX Association, 2016).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Reddi, S. J., Kale, S. & Kumar, S. On the convergence of adam and beyond. In Sixth International Conference on Learning Representations (2018).
Zeiler, M. D. ADADELTA: an adaptive learning rate method. Preprint at https://arxiv.org/abs/1212.5701 (2012).
Kingma, D. P. & Welling, M. Stochastic gradient VB and the variational auto-encoder. In Second International Conference on Learning Representations (2014).
Strehl, A. & Ghosh, J. Cluster ensembles—a knowledge reuse framework for combining multiple partitions. J. Mach. Learn. Res. 3, 583–617 (2003).
Kuhn, H. W. The Hungarian method for the assignment problem. Nav. Res. Logist. Q. 2, 83–97 (1955).
Rand, W. M. Objective criteria for the evaluation of clustering methods. J. Am. Stat. Assoc. 66, 846–850 (1971).
Author information
Authors and Affiliations
Contributions
Z.W. and Q.S. conceived and supervised the project. Z.W. led the study. T.T. designed the methods and conducted the experiments with input from J.W. T.T., J.W. and Z.W. wrote the manuscript. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Figures, table and notes
Rights and permissions
About this article
Cite this article
Tian, T., Wan, J., Song, Q. et al. Clustering single-cell RNA-seq data with a model-based deep learning approach. Nat Mach Intell 1, 191–198 (2019). https://doi.org/10.1038/s42256-019-0037-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-019-0037-0
This article is cited by
-
DANCE: a deep learning library and benchmark platform for single-cell analysis
Genome Biology (2024)
-
scCompressSA: dual-channel self-attention based deep autoencoder model for single-cell clustering by compressing gene–gene interactions
BMC Genomics (2024)
-
Classification of tropical cyclone rain patterns using convolutional autoencoder
Scientific Reports (2024)
-
Graph attention autoencoder model with dual decoder for clustering single-cell RNA sequencing data
Applied Intelligence (2024)
-
scEM: A New Ensemble Framework for Predicting Cell Type Composition Based on scRNA-Seq Data
Interdisciplinary Sciences: Computational Life Sciences (2024)