Abstract
The arrangement of network nodes in hyperbolic spaces has become a widely studied problem, motivated by numerous results suggesting the existence of hidden metric spaces behind the structure of complex networks. Although several methods have already been developed for the hyperbolic embedding of undirected networks, approaches able to deal with directed networks are still in their infancy. Here, we present a framework based on the dimension reduction of proximity matrices reflecting the network topology, coupled with a general conversion method transforming Euclidean node coordinates into hyperbolic ones even for directed networks. While proposing a measure of proximity based on the shortest path length, we also incorporate an earlier Euclidean embedding method in our pipeline, demonstrating the widespread applicability of our Euclidean-hyperbolic conversion. Besides, we introduce a dimension reduction technique that maps the nodes directly into the hyperbolic space of any number of dimensions with the aim of reproducing a distance matrix measured on the given (un)directed network. According to various commonly used quality scores, our methods are capable of producing high-quality embeddings for several real networks.
Similar content being viewed by others
Introduction
Networks offer an intuitive and general approach to the study of complex systems that has become extremely widespread in the recent decades1,2,3. The staggering amount of research in this direction has shown that the statistics of the underlying graph structure can highlight previously unseen properties in systems ranging from interactions within the cell up to the level of the entire human society1,2,3,4,5. The most well-known features that seem to be more or less universal across the majority of the complex networks are the small-world property6,7, the high clustering coefficient8, the scale-free degree distribution9,10 and a well-pronounced community structure11,12,13.
Grasping the above properties all at once with a simple network model is a challenging task for which hyperbolic approaches offer an intuitive framework. The basic idea of hyperbolic network models is to place the nodes in the hyperbolic space and connect them with a probability decaying as the function of the hyperbolic distance14,15,16,17,18,19,20,21,22. Remarkably, the networks generated in this way are usually small-world, highly clustered and scale-free14,15, and according to recent results they can easily display a strong community structure as well18,20,23,24,25,26,27. In parallel with revealing the notable properties of hyperbolic models, several studies suggested the existence of hidden geometric spaces behind the structure of real networks as well, ranging from protein interaction networks28,29 through brain networks30,31 to the Internet32,33,34,35,36 or the world trade network37, leading to important discoveries about the self-similarity38 and the navigability of networks32,39,40.
These advancements opened a further frontier in the research focusing on the relationship between hyperbolic spaces and complex networks centred on the problem of hyperbolic embedding, where the task is to find an optimal arrangement of the network nodes in the hyperbolic space for a given network structure that we inputted33. A natural idea in this respect is likelihood optimisation16,41, where a loss function is formulated (and minimised) based on the assumption that the input network was generated by a given hyperbolic network model. A prominent method following this idea is HyperMap16, working with a generalised version of the popularity-similarity optimisation (PSO) model15 called the E-PSO model. Another possibility is the application of dimension reduction techniques to matrices that represent the network topology, such as in the Laplacian-based Network Embedding (LaBNE) technique42 (relying on the Laplacian matrix of the graph to be embedded) and the family of coalescent embeddings43 (building on different matrices of distances measured along the graph after pre-weighting), where the dimension reduction yields a Euclidean embedding, the radial coordinates of which are converted then to hyperbolic ones in accordance with the PSO model, or in the hydra (hyperbolic distance recovery and approximation) method44, where the dimension reduction yields node positions in the hyperboloid model of the hyperbolic space that are finally converted to an embedding in the Poincaré ball representation. Dimension reduction and the optimisation of the angular node coordinates with respect to a given hyperbolic network model can be also combined. Such a combination was applied for the Laplacian-based embedding45 with the E-PSO model46 and the so-called \({{\mathbb{S}}}^{1}/{{\mathbb{H}}}^{2}\) model47, and also for a coalescent embedding43 that was coupled with a local likelihood optimisation according to the E-PSO model17. A further alternative approach for embedding networks into hyperbolic spaces is offered by artificial neural networks, whose objective is to learn a low-dimensional representation of the input network48,49,50,51. Although these methods are more difficult to interpret and their setup is usually more complicated compared to the previous approaches, they can also allow the inclusion of additional node (or link) features such as attributes, annotations, text, etc. in the learning process.
Even though the aforementioned methods achieved notable success and have been shown to provide high-quality embeddings for a number of different networks, almost all of them lack a very important capability: to take into account the link directions when dealing with directed network input. In general, directed connections can indicate asymmetric relations between the nodes (e.g., the dominant-subordinate relations in hierarchical networks52,53, the consumer-producer relations in food webs, etc.) or may signal some sort of flow over the links. Consequently, nodes with mainly incoming links may have a very different function in the system compared to nodes with mainly outgoing links or nodes having a balanced amount of in- and out-neighbours, and the directionality may play an important role also on the level of communities54. In this light, it seems that ignoring link directions during the preparation of an embedding can lead to a considerable amount of information loss. The only embedding methods41,51 that can take into account the directed nature of a network and use hyperbolic geometry either creates two-dimensional hyperbolic embeddings with a likelihood optimisation technique based on a directed \({{\mathbb{S}}}^{1}/{{\mathbb{H}}}^{2}\) model, or assigns to each network node a Gaussian distribution with a mean vector given in the hyperboloid model of the hyperbolic space, where the parameters of the representation of the nodes are learned using a neural network, and the asymmetry of the relations between the nodes can manifest itself in the Kullback-Leibler divergence between the Euclidean mapping of the corresponding distributions.
Motivated by the above, here we propose a general, albeit also simple framework for embedding directed networks into hyperbolic spaces of any number of dimensions, representing the topological distances and the connection probabilities through hyperbolic distances. Due to the possibly different functions of the sources and the targets in directed systems, our approach assigns separate source and target positions to each one of the network nodes, allowing large flexibility in how the directed nature of the input may affect the obtained embedding. This means that in the two-dimensional case, the output of our method can be visualised on a pair of disks (one of which contains the nodes at their source coordinates and the other at their target positions), where the links always point from the “source disk” to the “target disk”.
In order to keep the approach model-independent, the calculation of the node positions is based on a dimension reduction of a matrix encapsulating the distance relations in the network. The result of the dimension reduction of a proximity matrix can be already treated as a Euclidean embedding of the network. To obtain the hyperbolic coordinates from the Euclidean node arrangement, we introduce a transformation designed to preserve the attractivity of a given radial position from the point of view of link creation. With the help of this transformation, we can incorporate the output of several directed Euclidean embedding methods for gaining a hyperbolic layout of the studied network. Along this line, in the present work, we also apply the Euclidean HOPE (High-Order Proximity preserved Embedding) algorithm55, and transform its output in the same manner as the results of the here-proposed Euclidean embeddings. Finally, inspired by the undirected hyperbolic embedding method hydra (hyperbolic distance recovery and approximation)44, we also introduce a directed embedding approach that yields hyperbolic coordinates based on the dimension reduction of a Lorentz product matrix calculated from node-node distances measured along the inputted network, providing a hyperbolic layout without embedding the network first into the Euclidean space.
We test all the proposed methods both on synthetic and real networks. We examine the mapping accuracy56, which is a measure of embedding quality characterising the correlation between the shortest path lengths and the pairwise geometric relations of the nodes. We also evaluate the performance of the embeddings in graph reconstruction problems, where the task is to distinguish the connected node pairs of the embedded network from the unconnected ones according to geometric measures associated with the node pairs. Lastly, the embeddings are also compared to each other based on their navigability via greedy routing, which corresponds to a simple navigation protocol where we always try to proceed towards the destination node based only on the spatial position of the current neighbours.
Results
In this section, we first outline the studied embedding framework and describe the quality functions used for characterising the performance of the different methods. This is followed by the results obtained for a couple of directed real networks.
The studied embedding algorithms
In this paper, we consider embeddings of directed networks, which—due to the possible different roles of the same node as a source or as a target of links—result in two distinct sets of coordinates (i.e., source and target coordinates). In Fig. 1, we provide a concise flowchart of the considered embedding methods, the full detailed description of which is given in Supplementary Note 1. Note that all the studied methods are deterministic, yielding always the same node arrangement for a given network.
The left side of the figure traces the algorithmic steps for creating a hyperbolic embedding with the High-Order Proximity preserved Embedding (HOPE), our TRansformation of EXponential shortest Path lengths to EuclideaN measures (TREXPEN) and their variants by converting the Euclidean node arrangement obtained from them to a hyperbolic one with our model-independent conversion (MIC). The right side of the figure shows the algorithmic steps of our method named TRansformation of EXponential shortest Path lengths to hyperbolIC measures (TREXPIC), which embeds networks directly in the hyperbolic space. The embedding parameters are written in red: the parameters α and q adjust how the elements of the reduced matrices depend on the distances measured along the graph to be embedded, d denotes the number of dimensions of the embedding space, ζ (usually set to 1) tunes the curvature of the hyperbolic space, and C (usually set to 2) controls the extent of the graph in the hyperbolic space when using MIC.
Embedding into the hyperbolic space through the conversion of a Euclidean node arrangement
The three main steps of the algorithms described by the left side of the flowchart in Fig. 1 can be summarised in the following way:
-
1.
Preparation of a proximity matrix P based on the network topology.
-
2.
Decomposition of this matrix for performing dimension reduction and obtaining a Euclidean embedding, i.e. a lower-dimensional representation in the Euclidean space.
-
3.
Model-independent conversion (MIC) of the Euclidean source and target coordinates into position vectors in the native representation of the hyperbolic space.
The proximity matrix can be defined in multiple alternative ways, and since steps (2) and (3) are always the same, we name the different methods based on the choice of P. In the High-Order Proximity preserved Embedding (HOPE)55, the applied proximity matrix is the Katz matrix, where the intuitive meaning of a matrix element is that it corresponds to the weighted sum of the paths between the corresponding pair of nodes, where longer paths are more or less suppressed with the help of the adjustable parameter α. As an alternative for embedding via the Katz matrix, we introduce the method TRansformation of EXponential shortest Path lengths to EuclideaN measures, abbreviated as TREXPEN, where the proximity matrix P is composed of exponential shortest path lengths in the form of
where SPLs→t denotes the shortest path length from node s to node t, and 0 < q is a decay parameter similar in nature to the α parameter of the Katz matrix. Note that for node pairs s and t where t is unreachable from s, the above matrix element Pst and also the element of the Katz matrix becomes zero, which enable us to embed weakly connected components, not only strongly connected parts of directed networks.
The usage of a proximity matrix (where large values indicate small distances or large similarities) has the advantage compared to distance matrices that it yields such Euclidean embeddings in which smaller topological distances can be associated primarily with larger inner products of the position vectors instead of smaller Euclidean distances, providing the possibility to effortlessly separate the contribution of the radial and the angular node coordinates in the geometric relations. However, when equating only non-negative proximity values with Euclidean inner products, the angular range of the node coordinates becomes restricted. Therefore, we also consider a centred version of the proximity matrices by shifting the mean of the matrix elements to zero, which is expected to broaden the angular range of the node coordinates. We shall refer to the embeddings where the mean of the proximities is set to zero before the matrix decomposition as HOPE-S and as TREXPEN-S (where the suffix “-S” refers to the shifting of the elements of P). Another alternative considered here is that we return to the original (non-shifted) proximity matrices, but discard the first and use from the second to the d + 1th dimension for creating a d-dimensional embedding. The rationale behind this approach is that when embedding the network, we are interested in the positions of the nodes relative to each other, whereas the first component in the dimension reduction usually contains information mainly about the point cloud as a whole, relative to the origin. We shall refer to the embedding methods relying on the second to d + 1th dimensions as HOPE-R and as TREXPEN-R (where the suffix “-R” refers to the removal of the first dimension). These circular Euclidean node arrangements in which the high connection probabilities are represented with high inner products can serve as a good candidate for a Euclidean-hyperbolic conversion that maps the high Euclidean inner products to small hyperbolic distances.
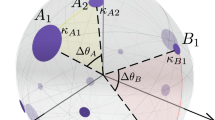
In our hyperbolic embedding methods, we used the native representation of the hyperbolic space14, which is commonly used both in hyperbolic network models15,21,22,26 and hyperbolic embeddings16,17,43,47. This representation visualises the d-dimensional hyperbolic space of curvature K = − ζ2 < 0 in the Euclidean space as a d-dimensional ball of infinite radius (to which we refer as the native ball), in which the radial coordinate of a point (i.e., its Euclidean distance measured from the centre of the ball) is equal to the hyperbolic distance between the point and the centre of the ball, and the Euclidean angle formed by two hyperbolic lines is equal to its hyperbolic value. The hyperbolic distance is measured along a hyperbolic line, which is either an arc going through the points in question and intersecting the ball’s boundary perpendicularly or—if the ball centre falls on the Euclidean line that connects the examined points—the corresponding diameter of the ball. According to the commonly applied approximating form of the hyperbolic distance14 given by \({x}_{s\to t}\,\,\approx {r}_{s}^{{{{{{{{\rm{source}}}}}}}}}+{r}_{t}^{{{{{{{{\rm{target}}}}}}}}}+\frac{2}{\zeta }\cdot \ln \left(\frac{{\theta }_{s\to t}}{2}\right)\), a smaller hyperbolic distance xs→t between the source position of node s and the target position of node t—the indicator of a higher connection probability—can originate from small radial coordinates \({r}_{s}^{{{{{{{{\rm{source}}}}}}}}}\) and \({r}_{t}^{{{{{{{{\rm{target}}}}}}}}}\) and/or a small angular distance θs→t. Another intuitive consequence of the above distance formula is that nodes with low radial coordinates are more attractive since their hyperbolic distance can become small in a larger angular region compared to nodes with large radial coordinates.
On the other hand, the Euclidean embedding methods we consider provide layouts where node pairs with high proximity values (and presumably, also high connection probabilities) obtain position vectors yielding a high inner product value. As the inner product between the source position of node s and the target position of node t is simply \({r}_{s}^{{{{{{{{\rm{source}}}}}}}}}\cdot {r}_{t}^{{{{{{{{\rm{target}}}}}}}}}\cdot \cos ({\theta }_{s\to t})\), high connection probability in the Euclidean space can originate from large radial coordinates and/or small angular distances.
Since small angular distance is favourable from the point of view of both a large Euclidean inner product and a small hyperbolic distance, we transfer the angular coordinates from the Euclidean space without modification to the hyperbolic ball, similarly to the practice in several previous embedding algorithms from the literature42,43,47. However, the situation is more complex in terms of the radial coordinates, since a high inner product requires large radial coordinates in the Euclidean space, whereas a low hyperbolic distance favours small radial coordinates in the hyperbolic ball. Nevertheless, relying on the expectation that Euclidean and the hyperbolic radial arrangements of the same network should represent the same attractivity relations, we can presume that if the radial positions of the embedding from both geometries are converted to the same space, then the node arrangements that are formed in the common space must be consistent with each other. More precisely, we assume that the node arrangements obtained in the common space from the Euclidean and the hyperbolic radial coordinates reflect the same radial attractivity of any node compared to the highest one.
We use the linearly expanding half-line as the pass-through between the polynomially expanding Euclidean and the exponentially expanding hyperbolic spaces. For this, we take the well-known formulas for the spherical volume, and define the coordinate on the half-line of Euclidean and hyperbolic radial values to be equal to the volume of a sphere with the radius equal to the original radial coordinate in the given metric space, resulting in
Then, our assumption about the reconcilability of the node coordinates calculated on the half-line from the Euclidean and the hyperbolic radial coordinates can be formalised for any node i as
where we have also taken into account that the attractivity of the nodes increases in the Euclidean and decreases in the hyperbolic space with the radial coordinate (and that the radially most attractive node is at the maximal radial coordinate \({r}_{{{{{{{{\rm{Euc,max}}}}}}}}}\) in the Euclidean space, and at the minimal radial coordinate \({r}_{{{{{{{{\rm{hyp}}}}}}}},\min }\) in the hyperbolic space).
By fixing the maximal radius in the hyperbolic space, we can use Eqs. (2)–(4) for calculating the hyperbolic radial coordinate of the nodes based on their Euclidean radial coordinate. Our suggestion for the largest possible radial coordinate in the hyperbolic ball is \({r}_{{{{{{{{\rm{hyp}}}}}}}},\max }=\frac{C}{\zeta }\cdot \ln (N)\), where C is a constant. With this choice, the hyperbolic volume scales as \({V}_{d}^{{{{{{{{\rm{hyp}}}}}}}}} \sim {N}^{C\cdot (d-1)}\) with the number of nodes N, and at C = 2 we obtain the same volume as we would have in a network generated by the PSO model15,22. Based on that, the radial coordinate in the hyperbolic ball can be expressed as
where further details of the calculation are given in Sect. S1.5 of Supplementary Note 1. Besides, Sect. S2.3 of Supplementary Note 2 demonstrates that MIC, our model-independent Euclidean-hyperbolic conversion of the radial coordinates can outperform the widely used17,42,43,46 PSO-based transformation even on such hyperbolic networks that were generated by the PSO model.
As an illustration of MIC, in Fig. 2 we show two-dimensional embeddings of an undirected E-PSO network16,17 that was generated from N = 1000 number of nodes, setting the average degree to \(\bar{k}\approx 2\cdot (m+L)=2\cdot (3+2)=10\) (where one can interpret m as the number of external links that emerge in each time step and L as the net number of added and removed internal links per time step), the popularity fading parameter to β = 0.8 (corresponding to the decay exponent γ = 1 + 1/β = 2.25 of the degree distribution \({{{{{{{\mathcal{P}}}}}}}}(k) \sim {k}^{-\gamma }\)), and the temperature T = 0 (resulting in an average clustering coefficient of \(\bar{c}=0.806\)). During the network generation, the nodes appeared one by one with increasing radial coordinate and connected to a given number of hyperbolically closest ones of the previously appeared nodes. Aiming at connections of small hyperbolic distances basically means that the new nodes tended to connect to nodes of small radial coordinates and/or small angular distance from them. In our Euclidean embeddings that represent small topological distances as large inner products, the early-appearing nodes that collected the highest number of links during the network formation become placed in the outermost positions, as the radial attractivity of the nodes increases outwards in this case. However, when transforming these layouts into hyperbolic ones, the largest hubs are transferred back to the innermost positions that possess the highest radial attractivity from the point of view of the minimisation of the hyperbolic distances. Besides, both our Euclidean and hyperbolic embeddings seem to preserve the angular arrangement of the nodes, reflecting the common preference of both geometries towards the relatively small angular distances of the connected pairs.
Here we use embeddings of an undirected network that was created by a generalised version of the popularity-similarity optimisation model. The node degrees are indicated by the node sizes: nodes with more connections are depicted by larger markers. The nodes are coloured in each layout according to the angular coordinates originally assigned in the hyperbolic plane of curvature K = − ζ2 = − 1 by the E-PSO model. The depicted HOPE-R embeddings were created using α = 5.97 ⋅ 10−3, while the TREXPEN-R layouts were obtained at q = 3.89. We used C = 2 and ζ = 1 for all the hyperbolic embeddings. We also show the result of carrying out the optional step of shifting the centre of mass (COM) of the Euclidean node arrangement before converting it to a hyperbolic one.
Embedding directly into the hyperbolic space with TREXPIC
The above-discussed hyperbolic embedding methods rely on the implicit assumption that the Euclidean embedding obtained in the first stages of the algorithms is able to capture the most important features of the network structure. This dependence on the Euclidean methods can be avoided by embedding directly into the hyperbolic space, as it was done e.g. in the hydra approach44 on undirected networks. In order to provide also such an algorithm that follows this alternative path, we propose the method TRansformation of EXponential shortest Path lengths to hyperbolIC measures, abbreviated as TREXPIC in the following.
As it was utilised in the hydra method44, the Lorentz product defined between two position vectors as \(\underline{y}\circ \underline{z}={y}_{1}{z}_{1}-({y}_{2}{z}_{2}+{y}_{3}{z}_{3}+\cdots +{y}_{d+1}{z}_{d+1})\) enables the calculation of the hyperbolic distance in the hyperboloid representation of the d-dimensional hyperbolic space via the formula \(x(\underline{y},\underline{z})=\frac{1}{\zeta }\cdot {{{{{{{\rm{acosh}}}}}}}}(\underline{y}\circ \underline{z})\). Thus, if we construct a distance matrix D between the nodes where the matrix element Dst estimates the hyperbolic distance from node s to node t, then using the formula \({L}_{st}=\cosh (\zeta \cdot {D}_{st})\) we obtain a matrix containing the estimated pairwise Lorentz products. Here, we suggest using
where q > 0 is an adjustable parameter that controls how fast our distance measure increases towards the larger shortest path lengths. The advantage of this choice compared to using simply the shortest paths themselves as in the hydra approach44 is that it makes all the matrix elements finite even in weakly connected components.
Based on the matrix of Lorentz products, we created low-dimensional hyperbolic embeddings in the hyperboloid model with the help of dimension reduction. For this, we used singular value decomposition (SVD) as opposed to hydra, which performs eigendecomposition. Then, using a mapping between the hyperboloid model and the native representation of the hyperbolic space, we obtained a layout in the native ball that is comparable with the output of the previous embedding methods.
Directed embedding into two-dimensional spaces
As a first illustration of the results that can be obtained from our framework, in Fig. 3 we show the embeddings of synthetic directed networks generated by the stochastic block model (SBM)57,58 (using the Python function ‘stochastic_block_model’ available in the ‘NetworkX’ package at https://networkx.org/documentation/stable/reference/generated/networkx.generators.community.stochastic_block_model.html) in both Euclidean and hyperbolic spaces in the case of setting the number of dimensions to d = 2, allowing the display of the achieved layouts in a simple manner. In the top half of the figure (Fig. 3a–f) we show the results for a graph with an apparent community structure (where the diagonal elements of the connection probability matrix of the blocks are larger), while in the bottom half of the figure (Fig. 3g–l) the embedded network has an “anti-community” structure (where the off-diagonal connection probabilities are larger). According to these layouts, the considered embedding methods were able to correctly separate the different blocks and provide an angular arrangement that reflects the most important features of the network structure in an easy-to-observe manner. Further layouts of the SBM networks are displayed in Sect. S3.1 of Supplementary Note 3.
a An assortative block matrix used for generating the input network for the embeddings displayed in panels b–f, where the embedding method is named in the panel title and the colouring of the nodes indicates their block membership according to panel a. The source and the target coordinates are shown in separate planes (where a single node can appear both as a source and as a target of links). As a consequence, the links of the network appear as lines between the “source plane” and the “target plane”. g A disassortative block matrix used for generating the input network for the embeddings shown in panels h–l. The colouring of the nodes in panels h–l reflects their block membership according to panel g.
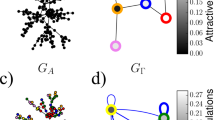
Next, in Fig. 4 we present embeddings of the network of political weblogs59 (downloaded from http://konect.cc/networks/dimacs10-polblogs/), for which several quantitative results are provided in the next section, in both the Euclidean and the hyperbolic plane. As it can be seen here, the nodes of different attributes tend to become grouped into different angular regions in the embeddings. More examples of the automatic separation of the ground-truth communities of real networks are provided in Sect. S3.2 of Supplementary Note 3.
The coordinates obtained for the source and the target representations of the nodes are displayed in separate planes for better visibility. Thus, the links always point from a node on the “source plane” to a node on the “target plane”. The node colours indicate the political leaning of the corresponding weblog. Larger node sizes in the “source plane” and in the “target plane” correspond to larger out- and in-degrees, respectively. Panels a–c correspond to embeddings based on Katz proximity, while our exponential proximity was used in the case of panels d–f. Panels a and d show the non-circular Euclidean embeddings created by HOPE and TREXPEN, respectively. Panels b and e depict the circular Euclidean layouts yielded by the HOPE-R and the TREXPEN-R variants, while panels c and f present the hyperbolic embeddings obtained from these using our Euclidean-hyperbolic conversion method MIC.
Performance of HOPE, TREXPEN, their several variants and TREXPIC on real directed networks
We tested the proposed embedding methods on the following directed real networks:
-
A subnetwork of N = 505 number of nodes and E = 2081 number of edges extracted from Wikipedia’s norm network of 201560, where Wikipedia pages are connected to each other with directed edges that correspond to hyperlinks. We created the subgraph by omitting all nodes for which the highest value of the topic distribution does not reach 80%, i.e. we kept only the pages for which the topic was not too uncertain.
-
The transcriptional regulation network61 (downloaded from https://www.weizmann.ac.il/mcb/UriAlon/download/collection-complex-networks) of the yeast Saccharomyces cerevisiae, describing E = 1063 number of interactions between N = 662 number of regulatory proteins and genes. The links point from the regulating objects toward the regulated ones. The mode of regulation was considered to be the same in each case, i.e. we did not differentiate between activators and repressors.
-
A network59 (downloaded from http://konect.cc/networks/dimacs10-polblogs/) of E = 19,021 hyperlinks among N = 1222 number of U.S. political weblogs from before the 2004 presidential election. The blogs are characterised by their political leaning, forming 2 groups: left/liberal and right/conservative.
-
A word association network62 (downloaded from http://w3.usf.edu/FreeAssociation/) of N = 4865 number of nodes and E = 41,964 number of links that point from the cue words toward the associated words.
Note that we carried out the same analysis as below for four additional directed real networks in Supplementary Note 5, and in Supplementary Note 7 we also show some results regarding the embeddings of two undirected real networks, confirming that the methods proposed here are able to compete with previous, well-known dimension reduction techniques. In addition, in Supplementary Note 6, we show the significance of the directedness of the links in the examined directed real networks by comparing their directed embeddings to the embeddings of their undirected counterpart.
Since a node with zero out- and in-degree does not have any role neither as a source nor as a target, it cannot be represented in the embedding (will not have neither a source nor a target position). Therefore, we only embedded the largest weakly connected component (WCC) of each graph—the above-listed N and E values refer to these. Throughout this section, we discarded the link weights given in some of the datasets and assigned the weight 1 to each edge. To learn about how our embeddings treat real link weights, see Supplementary Note 8.
In the following subsections, we evaluate the embedding performance on the above-listed four directed networks in three aspects: we examine mapping accuracy, graph reconstruction and greedy routing. The detailed description of the applied measures is provided in the Methods section. During the measurements, we took into consideration all the possible node pairs in each task for the two smaller graphs (namely the network of Wikipedia pages and the yeast transcription network), but—because of the high computational intensity—accomplished the evaluation of the embedding performance only on sampled sets of node pairs in the case of the two larger graphs (i.e. the network of political blogs and the word association network). The details of the applied sampling procedures are given in the Methods section.
We always tested HOPE-S, HOPE-R, TREXPEN-S and TREXPEN-R both with and without shifting the centre of mass (COM) of the node positions to the origin, but depicted here only the results of the better option. Note that shifting all the nodes by the same vector does not change the pairwise (Euclidean or hyperbolic) distances, but modifies the pairwise inner products of the nodes in a Euclidean embedding, and also changes the hyperbolic node arrangement that can be obtained from that via MIC. The difference between the quality scores achieved with or without shifting the COM is demonstrated by Supplementary Note 4 and Supplementary Note 7: usually the Euclidean embeddings are hindered by the displacement of the COM, whereas MIC—and the hyperbolic embeddings resulting from it—can benefit from the balancing of the Euclidean node arrangement.
In every task, the tested number of dimensions were \(d=2,3,4,8,\ldots ,{2}^{n}\le \frac{N}{10}\), \(n\in {\mathbb{{Z}^{+}}}\) for all the embedding methods, where the condition d ≤ N/10 is intended to ensure a considerable dimension reduction. Note that while the embeddings obtained in high-dimensional spaces may be able to capture more information precisely, relatively high importance can be attributed also to the d = 2 and the d = 3 settings that are the only ones yielding directly visualisable node arrangements.
In HOPE and its variants, we tested 15 number of α values that we sampled from the interval \(\left[\frac{1}{200\cdot {\rho }_{{{{{{{{\rm{spectral}}}}}}}}}({{{{{{{\boldsymbol{A}}}}}}}})},\frac{1}{{\rho }_{{{{{{{{\rm{spectral}}}}}}}}}({{{{{{{\boldsymbol{A}}}}}}}})}\right]\) for each network (see Sect. S1.2 of Supplementary Note 1), where ρspectral(A) is the spectral radius of the adjacency matrix A. In the case of TREXPEN and its variants, we always tested 15 number of q values sampled from the interval \([-\ln (0.9)/{{{{{{{{\rm{SPL}}}}}}}}}_{\max },-\ln (1{0}^{-50})/{{{{{{{{\rm{SPL}}}}}}}}}_{\max }]\) (see Sect. S1.3 of Supplementary Note 1), where \({{{{{{{{\rm{SPL}}}}}}}}}_{\max }\) is the largest finite shortest path length occurring in the given network. For TREXPIC, we tested 15 number of q values from the interval \([\ln (1.0/0.9999)\cdot {{{{{{{{\rm{SPL}}}}}}}}}_{\max },\ln (10)\cdot {{{{{{{{\rm{SPL}}}}}}}}}_{\max }]\) for each network (see Sect. S1.6 of Supplementary Note 1). The suitability of these parameter intervals is demonstrated by Supplementary Note 4, where we show through the example of the Wikipedia network that the performance of the examined methods typically reaches a maximum within these ranges and declines at the boundaries. It is important to emphasize that we did not try to find the exact optimum of the embedding parameters, meaning that slight variances between the different embedding methods have to be treated with caution since these may simply be a consequence of the imperfection of the parameter settings and the method that seems to be worse may prevail over the other at a better parameter setting.
The curvature K = − ζ2 of the hyperbolic space was set to − 1 for all the hyperbolic embeddings—the role of the curvature is discussed in Sect. S4.1 of Supplementary Note 4. And lastly, we always used C = 2 in MIC, which choice is supported by Supplementary Note 4.
Mapping accuracy
A simple measure of the embedding quality is provided by the mapping accuracy56, defined as the Spearman’s correlation coefficient (that we calculated with the Python function ‘spearmanr’ available in the ‘scipy.stats’ package at https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.spearmanr.html) between the shortest path lengths and given geometric measures of the node pairs in an embedded network. In this study, the examined geometric measures were the Euclidean distance and the additive inverse of the inner product in the case of the Euclidean embeddings, and the hyperbolic distance for the hyperbolic node arrangements. In all cases, we considered the quality of those embeddings to be better, which yielded higher positive values of the correlation coefficient, meaning that we expected all the investigated methods to minimise the distances and/or maximise the inner products between the positions of the nodes that are close to each other according to the network topology.
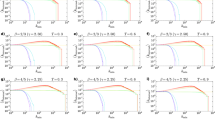
In Fig. 5, we show the mapping accuracy on the four test networks, i.e. the network of Wikipedia pages, the transcription network, the network of political blogs and the word association network. As expected, TREXPEN, its variants and TREXPIC yield higher correlations between the shortest path lengths and the geometric measures compared to HOPE and its variants in most of the cases since HOPE considers all the paths between the nodes to a certain extent, not only the shortest ones. The best overall results were produced by Euclidean embeddings, but the hyperbolic methods do not fall behind much and, in the meantime, typically prevail over the Euclidean node arrangements when considering the distances between the nodes instead of the inner products.
Each panel refers to a real network named in the title of the panel. For the networks in panels a and b, we measured the mapping accuracy examining each node pair connected by at least one directed path, whereas for the larger networks in panels c and d, the mapping accuracy was measured on five samples of 500,000 node pairs connected by at least one directed path. In the case of the larger networks, we always considered the average of the performances over the five samples and depicted the corresponding standard deviations with (usually very small) grey error bars. The colours indicate the used geometric measure, as listed in the common legend at the bottom of the figure. We plotted only the best results in each panel, obtained with the parameter setting that yielded the highest values of the mapping accuracy. Note that the 0 values denote that the given methods have not achieved any positive value. The bars were created considering all the tested number of dimensions, whereas the horizontal lines show the best two-dimensional performances achieved among all the embedding methods.
Graph reconstruction
To quantify the ability of the node arrangements provided by our embedding methods to reflect the topology of the inputted networks, we accomplished graph reconstruction trials aiming at the differentiation between the connected and the unconnected node pairs of the examined networks based on pairwise geometric measures. For this, we embedded the whole largest WCC for each one of the studied networks, and ranked the source-target node pairs according to the Euclidean distance, the inner product or the hyperbolic distance between them, assuming simply that smaller distances and/or higher inner products refer to higher proximities along the graph, and thus, larger connection probabilities.
As a baseline, we measured the graph reconstruction performance of some local methods that, contrary to the embeddings, do not use the whole graph to give an estimation of the connection probability of a given node pair. We associated higher connection probabilities with higher numbers of common neighbours63, higher node degrees (preferential attachment64) and higher values of 3 directed variations of the originally undirected resource allocation index65—for details, see the Methods section. In our figures, we always indicate for each quality measure only the best result obtained among these (altogether 5) tested local methods.
We evaluated the graph reconstruction performance with 3 measures: Prec ∈ [0, 1] denotes the precision obtained when treating the number of links \({{{{{{{\mathcal{E}}}}}}}}\) that have to be reconstructed as a known input (i.e. the proportion of the actual links among the first \({{{{{{{\mathcal{E}}}}}}}}\) node pairs in the order assigned by the given connection probability measure), the area under the precision-recall (PR) curve AUPR ∈ (0, 1] and the area under the receiver operating characteristic (ROC) curve AUROC ∈ [0, 1]. All of these are increasing functions of the graph reconstruction performance. For more details, see the Methods section.
Figure 6 presents the embedding quality with respect to the graph reconstruction task of the examined four networks: Fig. 6a–c refer to the subgraph of Wikipedia’s norm network, Fig. 6d–f depict the results obtained for the transcriptional regulation network, Fig. 6g–i deal with the network of U.S. political weblogs, while Fig. 6j–l show the values achieved in the case of the word association network. The usage of Katz proximities (in HOPE and its variants) and the exponential proximities (in TREXPEN and its variants) or distances (in TREXPIC) both seem to be expedient in this task. While generally the inner product in the Euclidean embeddings seems to be the best proxy for the connection probability, in the network of political blogs, with regard to the area under the PR curve (Fig. 6h) the best method in the two-dimensional case is a hyperbolic one. Furthermore, when focusing on the distance-based representations of the network topology, the hyperbolic embeddings clearly outperform the Euclidean ones that often even struggle to surpass the performance of the local methods.
For the networks in panels a–f, the task was to reconstruct all the links (Esampled = E), whereas for the network of political blogs in panels g–j and for the word association network in panels j–l, due to the large network size, the task was to reconstruct five samples of Esampled = 5000 and Esampled = 500 number of links, respectively. In the case of the larger networks, we always considered the average of the quality scores over the five samples and depicted the corresponding standard deviations with (usually very small) grey error bars. Each row of panels refers to a real network indicated in the row title, while the different columns show the different quality measures that we studied, given by the precision obtained when reconstructing the first Esampled most probable links (1st column), the area under the precision-recall (PR) curve (2nd column), and the area under the ROC curve (3rd column). The colours indicate the applied geometric measure, as listed in the common legend at the bottom of the figure. Using the bars, we plotted only the best results regarding all the performance measures, considering all the tested number of dimensions. The horizontal lines in colour show the best two-dimensional performances achieved among all the embedding methods, whereas the grey horizontal lines correspond to the baselines provided by the random predictor and the best local method.
Greedy routing
The navigability of an embedded network can be measured via the greedy routing32,66,67, corresponding to the process when a walker tries to reach a given destination node from a starting node, always knowing only the position of the end of the links that spring from the current node compared to the position of the destination node. In our hyperbolic embeddings, we minimised in each step among the current neighbours their hyperbolic distance from the position of the destination node occupied as a target of links, while in Euclidean embeddings we tested both the minimisation of the Euclidean distance and the maximisation of the inner product. An embedded network is considered to be more navigable if its greedy routing score43GR-score ∈ [0, 1] is higher, expressing a larger success rate in reaching the destination node and/or a smaller hop-length of the successful greedy routes.
In Fig. 7, we depict the achieved greedy routing scores with the corresponding success rates and average hop-lengths for the examined starting node-destination node pairs in the studied four real networks. For all of these networks, the best GR-scores are achieved in the hyperbolic space; however, the distance-based routing performed in the Euclidean space is usually also effective. The inner product generally does not seem to be well usable for navigating on networks in the Euclidean space. Besides, in this task HOPE and its variants clearly fall behind the methods that we introduced here building on exponential proximities or distances instead of Katz proximities.
For the networks in panels a–f, the task was to perform greedy routing between each node pair connected by at least one directed path, whereas for the larger networks in panels g–l, the task was to perform greedy routing in five samples of 500,000 node pairs connected by at least one directed path. In the case of the larger networks, we always considered the average of the quality scores over the five samples and depicted the corresponding standard deviations with (usually very small) grey error bars. The colours indicate the used geometric measure as listed in the common legend at the bottom of the figure. We plotted in each panel for each method only the result of the parameter setting that turned out to be the best according to the GR-score. The bars were created considering all the tested number of dimensions, whereas the horizontal lines show the best two-dimensional average performances achieved among all the embedding methods. Each row of panels refers to a real network named in the row title, and the different columns correspond to different quality measures: the 1st column shows the greedy routing score (the higher the better), the 2nd column corresponds to the success rate of greedy routing (the higher the better), and the 3rd column depicts the average hop-length of the successful greedy paths (the smaller the better), where the grey bars indicate the average of the hop-length of the shortest paths connecting the node pairs for which the greedy routing was successful.
Discussion
We introduced a general framework based on the dimension reduction of proximity matrices for embedding directed networks into Euclidean and hyperbolic spaces of any number of dimensions. A key feature of our Euclidean embedding method TREXPEN is that it assigns both a source and a target position vector to each network node when aiming to capture the asymmetry of the connections in directed input graphs. The proximity matrix used in TREXPEN considers only the length of the shortest paths contrary to calculating all path lengths as it is done in the well-known HOPE algorithm55, and according to our experiments, this may be suitable for obtaining higher quality embeddings. This was especially striking in the case of the greedy routing score, where the usage of our exponential proximities instead of Katz proximities55 was proven to be strongly advantageous. In addition, our exponential proximity measure can be applied without any difficulty also on weighted networks, as it is described in Supplementary Note 8.
We also proposed a model-independent conversion between Euclidean and hyperbolic embeddings that does not assign any specific hyperbolic network model as the origin of the network to be embedded. The suggested transformation is based on the assumption that high connection probabilities are represented by large inner products in a circular Euclidean node arrangement on the one hand, and by low hyperbolic distances in the corresponding hyperbolic layout on the other hand. According to the results, with the help of this transformation both the output of our method TREXPEN and that of HOPE (with some minor modification) can be converted into directed hyperbolic embeddings of high quality. In addition, inspired by the hydra method44 proposed for undirected networks, we also developed the TREXPIC algorithm that can arrange directed networks in the hyperbolic space in a straightforward manner, without the need of creating a Euclidean embedding as an intermediate step.
The embedding techniques developed in this paper are all based on dimension reduction, hence providing an efficient and also model-independent approach for achieving an optimal representation of directed networks in both Euclidean and hyperbolic spaces. In two dimensions, the obtained hyperbolic layouts seemed to be more pleasant to the human eye compared to their Euclidean counterparts. This is due to the fact that the large number of radially unattractive nodes are placed in the outer regions of the hyperbolic disk, whereas they are gathered around the origin on the Euclidean plane. Meanwhile, the radial arrangements provided by TREXPIC did not seem to be so informative visually due to the relatively small differences between the radial coordinates, even though the measured quality scores were competitive with that of the proposed conversion-based hyperbolic algorithms. Treating the number of dimensions of the embedding space as a free parameter, all of our methods can utilize the benefits of the increased number of dimensions (noting, however, that the number of dimensions was still significantly lower compared to the system size in our experiments). We demonstrated the excellent usability of HOPE, TREXPEN, their variants and TREXPIC for different tasks via experiments carried out on real networks of several disciplines, including e.g. networks between webpages, word associations, and a transcriptional regulation network.
It is worth emphasizing that in our measurements regarding the mapping accuracy, the graph reconstruction performance and the navigability, the hyperbolic distance was the only geometric measure using which relatively good quality scores have been achieved in all of the different tasks. Among the examined three measures, the Euclidean distance performed the worst in mapping accuracy and especially in graph reconstruction, where it was often outperformed even by the simple local methods that we tested, while the results obtained using the Euclidean inner product lagged behind both that of the Euclidean and the hyperbolic distances in greedy routing. These findings clearly justify the competitiveness of the hyperbolic embeddings. In recent years, several studies examined the emergent properties of random networks of different geometries14,68,69 and the indicators of different hidden geometries behind networks70,71. In this work, we did not pursue to reveal how certain network properties are connected to the type and the dimension of the geometrical space underlying the networks; however, our embedding framework may contribute to further investigations on this topic by enabling the placement of real networks in different geometrical spaces of any number of dimensions.
Methods
This section provides the exact definition of the measures and methods used for evaluating the embedding performance. Note that none of the examined quality indicators assumes any specific model as the generator of the embedded network, i.e., all the applied evaluation processes are model independent, just like our embedding methods. For the details and the explanations regarding the studied embedding algorithms, see Supplementary Note 1.
Mapping accuracy
To evaluate the performance of the embedding methods in expressing the distance relations measured along the graph by means of geometric measures, we calculated a mapping accuracy measure ACCm ∈ [−1, +1] also used for undirected networks56. It was defined as the Spearman’s correlation coefficient (that we calculated with the Python function ‘spearmanr’ available in the ‘scipy.stats’ package at https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.spearmanr.html) between the shortest path lengths of a network and the pairwise distances between the network nodes in the embedding space—either Euclidean or hyperbolic. However, in the case of the Euclidean embeddings, the Euclidean distance was not the only geometric measure that was examined, but the correlation of the shortest path lengths with the inner products was also calculated.
Naturally, in directed networks we took into account the directedness of the paths and compared the hop-length of the shortest path from node s to node t to the distance or the inner product measured between the source position vector of node s and the target position vector of node t. We always discarded those s − t node pairs in our calculations, for which the examined graph does not contain any connecting paths, i.e. between which the shortest path length is infinity, and also disregarded the pairing of each node with itself (characterised by a shortest path length of 0) since the location of the target representation of a node compared to its own source position does not influence the quality of the embedding in itself, but only via the relations of the node’s two representations with the other nodes. Besides, to reduce the computational cost, in networks having >500,000 number of start-destination node pairs that could be used for the evaluation of the mapping accuracy, we estimated this quality measure based on 5 random samples of 500,000 proper node pairs. Note that when all the proper node pairs of a network are considered, then the calculation of the mapping accuracy is deterministic, and thus, there is no need for the repetition of its computation.
Evaluation of the embedding performance in graph reconstruction
We examined how precisely the embedding methods can represent the presence and the absence of the pairwise connections of an inputted network via the graph reconstruction task, similarly to previous studies in the literature55,72. Here the question is whether the connected and the unconnected node pairs can be distinguished based on pairwise measures that are derived with full knowledge of the network topology and can be interpreted as a proxy of the connection probability. Regarding the embedding techniques, this means that we embedded the whole largest WCC of a network in the Euclidean or the hyperbolic space, arranged the node pairs in the increasing order of the Euclidean distance, the additive inverse of the inner product or the hyperbolic distance and compared the set of node pairs appearing at the beginning of the order (i.e. below a given threshold of the applied geometric measure) to the list of links in the network. Besides the embeddings, we also tested local methods in graph reconstruction, where the decreasing order of the connection probability is estimated by the decreasing order of such measures that depend solely on the immediate neighbourhood of the two nodes in question. The assumptions of the applied local methods were the following:
-
Common neighbours: In undirected networks, the larger number of common neighbours of two nodes are often associated with a larger connection probability63. In directed networks, we assumed that the larger the number of paths of hop-length 2 from node s to node t, the higher the probability of the link from node s to node t.
-
Preferential attachment: In undirected networks, a simple proximity measure is given by the product of the node degrees in the examined node pair64. In the directed case, we applied this concept as the following: the larger the product of the out-degree of node s and the in-degree of node t (considering also the link s → t since we deal with graph reconstruction and not link prediction), the higher the probability of the link from node s to node t.
-
Resource allocation index: The resource allocation index RAI applies one of the simplest ways for reducing the contribution of the common neighbours of high degrees to the connection probability and assigning more weight to the common neighbours of low degrees, which provide more specific connections between the examined two nodes. For undirected networks, the resource allocation index is defined65 as
$${{{{{{{\rm{RAI}}}}}}}}(i,j)=\mathop{\sum}\limits_{c\in {{{{{{{\rm{CN}}}}}}}}(i,j)}\frac{1}{{k}_{c}},$$(7)where \({{{{{{{\rm{CN}}}}}}}}(i,j)\) denotes the set of the common neighbours of the examined two nodes i and j, and kc stands for the degree of the common neighbour c. Larger values of RAI are presumed to indicate larger connection probabilities. For directed networks, we identified the set of common neighbours \({{{{{{{\rm{CN}}}}}}}}(s,t)\) for the ordered node pair s, t as the nodes that are reachable from node s in one step and from which node t is reachable in one step, and tested 3 versions of RAI(s, t), in which we substituted kc in Eq. (7) with either the out-degree, the in-degree, or the total degree of the common neighbour c.
In every case, the order between node pairs that have the same value of the given measure of connection probability was set randomly.
In the smaller networks, we considered all the possible node pairs in the graph reconstruction task with the exception of the pairing of each node with itself (since self-loops are disregarded by the embeddings) and those node pairs in which the out-degree of the source node or the in-degree of the target node is 0 (since to a node with 0 out- or in-degree no position is assigned by the embedding methods as source or target, respectively). In those larger graphs where the total number of the proper source-target pairs exceeds 500,000, we applied a random sampling of the connected and the unconnected node pairs. To obtain such samples that well represent the total dataset, it is important to set the ratio between the number of sampled links and the total number of sampled node pairs equal to the ratio between the total number of links and the total number of proper node pairs in the network73,74. In order to keep the computational cost within reasonable limits, we set the number of links Esampled in each sample low enough to ensure that the total size of the sample (i.e. the sum of the number of links and the corresponding number of unconnected node pairs) remains under 500,000. When measuring the embedding quality on such samples, we always repeated the sampling and the reconstruction of the given links 5 times. However, since—at proper settings of the embedding parameters—it is very rare that the same value of the given geometric measure (i.e. the same connection probability) becomes assigned to more than one node pair yielding an indefinite ordering between them, and therefore, the graph reconstruction itself is rather deterministic, we did not repeat the evaluation of the graph reconstruction performance in those cases where all the proper node pairs were considered.
We characterised the embedding performance in graph reconstruction with the following three measures (that can be also used for evaluating link prediction accuracy75), each of which is an increasing function of the embedding quality:
-
The precision at \({{{{{{{\mathcal{E}}}}}}}}\) number of node pairs labelled as connected, i.e. \({{{{{{{\rm{Prec@}}}}}}}}{{{{{{{\mathcal{E}}}}}}}}\in [0,1]\) is defined as the proportion of the actual links among the \({{{{{{{\mathcal{E}}}}}}}}\) number of guesses corresponding to the first \({{{{{{{\mathcal{E}}}}}}}}\) node pairs in the decreasing order of the given measure of the connection probability. In our measurements, we always set \({{{{{{{\mathcal{E}}}}}}}}\) to the number of links to be reconstructed—that is, to the total number of links E in the smaller WCCs and to the number of sampled links Esampled in the case of the larger networks—and denoted the corresponding precision by Prec. For a random predictor, Prec was calculated for each network as the ratio between the number of actual links and all the node pairs in the examined set.
-
The precision-recall (PR) curve depicts the proportion of the actual links among all the node pairs that become labelled as connected (i.e. the precision) as a function of the proportion of the links that are successfully identified among all the links that have to be restored (i.e. the recall or true positive rate), where moving between the different points of the curve corresponds to changing the threshold value of the given connection probability measure or, in other words, shifting the point in the node pair order that separates the node pairs that we label as connected from those that we label as unconnected. (We computed the precision-recall pairs for different probability thresholds with the Python function ‘precision_recall_curve’ available in the ‘sklearn.metrics’ package at https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_recall_curve.html.) To give an overall description of the performances obtained at the different thresholds, we calculated AUPR ∈ (0, 1] (with the Python function ‘auc’ available in the ‘sklearn.metrics’ package at https://scikit-learn.org/stable/modules/generated/sklearn.metrics.auc.html) that is the area under the PR curve76. In the case of a random predictor, the precision-recall curve is a horizontal line at the precision value given by the ratio between the number of actual links and all the node pairs in the examined set, yielding an AUPR equal to this constant precision value.
-
The receiver operating characteristic (ROC) curve presents the proportion of the links that are successfully identified among all the links that have to be restored (i.e. the recall or true positive rate) as a function of the proportion of the actually unconnected node pairs that become labelled as connected (i.e. the false positive rate) obtained using different threshold values of the given measure associated with the connection probability. (We computed the receiver operating characteristic curve with the Python function ‘roc_curve’ available in the ‘sklearn.metrics’ package at https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_curve.html.) To summarize this curve in a single number, we calculated AUROC ∈ [0, 1] (with the Python function ‘auc’ available in the ‘sklearn.metrics’ package at https://scikit-learn.org/stable/modules/generated/sklearn.metrics.auc.html) that is the area under the ROC curve, corresponding to the probability that a randomly chosen connected node pair gets ranked over a randomly chosen unconnected node pair in the order of the examined connection probability measure77,78. For a random predictor, the ROC curve is a straight line between the points (0, 0) and (1, 1) with AUROC = 0.5.
Evaluation of the embedding performance in greedy routing
To characterise the navigability of the embedded networks, similarly to several other studies16,17,43,56, we examined the efficiency of the greedy routing32,66,67 on them. The aim of greedy routing is to walk along the network’s edges from a starting node s to a destination node t using the possible least number of steps, leaning solely on local information, namely the geometric distance of the current neighbours from the destination.
In our measurements, we adopted a rather general stepping rule, where the greedy router being at node i always moves along that outgoing link of node i that points toward the neighbour having a target position of the smallest geometric measure in relation to the target position of the destination node among all the current neighbours. The examined geometric measures for which the local minimisation was performed were the Euclidean distance or the additive inverse of the inner product in the Euclidean embeddings, and the hyperbolic distance in the hyperbolic cases. Returning to a node that has already been visited in the current walk indicates that the walk between the given pair of starting and destination nodes can not be accomplished in a greedy way. Thus, two simple measures of the greedy routing’s quality are the average hop-length of the successful greedy routes (that reached the destination and have not stopped at any other node) and the fraction of successful greedy walks. Besides, we also measured the greedy routing score43 (GR-score ∈ [0, 1], the higher the better), which we define for directed networks as
where \({\ell }_{s\to t}^{{{{{{{{\rm{(SP)}}}}}}}}}\) stands for the shortest path length from node s to another node t—which is infinity if there is no path in the graph leading from s to t –, and \({\ell }_{s\to t}^{{{{{{{{\rm{(GR)}}}}}}}}}\) denotes the greedy routing hop-length between the same pair of starting and destination nodes—which is set to infinity if the routing fails to reach node t from node s. To allow the investigation of weakly connected networks where not all the nodes are reachable from every node, we always took into account only those starting node-destination node pairs that are connected by at least one path in the graph, i.e., for which \({\ell }_{s\to t}^{{{{{{{{\rm{(SP)}}}}}}}}}\) is finite, and thus, the greedy routing is at least theoretically possible. Therefore, the total number Npaths of the examined start-destination pairs can be <N ⋅ (N − 1), and the summations in Eq. (8) go over only the nodes that function as a source of links in the network, i.e. the nodes of non-zero out-degree (contained by the set S) and the destinations to which leads at least one directed path from node s (contained by the set Ts for a given starting node s, not including node s).
For large networks, it is not feasible to take into consideration each possible node pair, but using a large enough random sample of the node pairs, the performance of an embedding in greedy routing can still be well estimated. In this study, we maximised the number of start-destination node pairs for which the greedy routing was attempted at 500,000 for each network, meaning that in those networks where the total number of node pairs connected by at least one path of finite length was larger than this limit, we randomly sampled 500,000 number of such node pairs and performed the greedy routing only between the selected starting and destination nodes. For those networks where thus not all the possible node pairs were examined, we repeated the node pair sampling and the greedy routing 5 times. Otherwise, since—at proper settings of the embedding parameters—it is very rare that two or more neighbouring nodes have the exact same geometric relation with the destination and the greedy router has to choose randomly between them, and therefore, the greedy routing itself is rather deterministic, we carried out greedy routing only once for all the proper node pairs of a network.
Data availability
All data generated during the current study are available from the corresponding author upon request. The subnetwork extracted from Wikipedia’s norm network of 201560 is available at https://github.com/BianKov/TREXPEN_TREXPIC/tree/main/embeddingDirectedNetworks/wikipedia. The yeast transcription network61 is available at https://www.weizmann.ac.il/mcb/UriAlon/download/collection-complex-networks. The network of political blogs59 is available at http://konect.cc/networks/dimacs10-polblogs/. The word association network62 is available at http://w3.usf.edu/FreeAssociation/.
Code availability
The code used for embedding undirected/directed, unweighted/weighted networks using HOPE, TREXPEN, their variants and TREXPIC is available at https://github.com/BianKov/TREXPEN_TREXPIC.
References
Albert, R. & Barabási, A.-L. Statistical mechanics of complex networks. Rev. Mod. Phys. 74, 47–97 (2002).
Mendes, J. F. F. & Dorogovtsev, S. N. Evolution of Networks: From Biological Nets to the Internet and WWW (Oxford Univ. Press, Oxford, 2003).
Newman, M. E. J., Barabási, A.-L. & Watts, D. J. (eds.) The Structure and Dynamics of Networks (Princeton University Press, Princeton and Oxford, 2006).
Holme, P. & Saramäki, J. (eds.) Temporal Networks (Springer, Berlin, 2013).
Barrat, A., Barthelemy, M. & Vespignani, A. Dynamical Processes on Complex Networks (Cambridge University Press, Cambridge, 2008).
Milgram, S. The small world problem. Psychol. Today 2, 60–67 (1967).
Kochen, M. (ed.) The Small World (Ablex, Norwood (N.J.), 1989).
Watts, D. J. & Strogatz, S. H. Collective dynamics of ’small-world’ networks. Nature 393, 440–442 (1998).
Faloutsos, M., Faloutsos, P. & Faloutsos, C. On power-law relationships of the internet topology. Comput. Commun. Rev. 29, 251–262 (1999).
Barabási, A.-L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Fortunato, S. Community detection in graphs. Phys. Rep. 486, 75–174 (2010).
Fortunato, S. & Hric, D. Community detection in networks: a user guide. Phys. Rep. 659, 1–44 (2016).
Cherifi, H., Palla, G., Szymanski, B. & Lu, X. On community structure in complex networks: challenges and opportunities. Appl. Netw. Sci. 4, 117 (2019).
Krioukov, D., Papadopoulos, F., Kitsak, M., Vahdat, A. & Boguñá, M. Hyperbolic geometry of complex networks. Phys. Rev. E 82, 036106 (2010).
Papadopoulos, F., Kitsak, M., Serrano, M. Á., Boguñá, M. & Krioukov, D. Popularity versus similarity in growing networks. Nature 489, 537 EP (2012).
Papadopoulos, F., Psomas, C. & Krioukov, D. Network mapping by replaying hyperbolic growth. IEEE/ACM Trans. Netw. 23, 198–211 (2015).
Kovács, B. & Palla, G. Optimisation of the coalescent hyperbolic embedding of complex networks. Sci. Rep. 11, 8350 (2021).
Zuev, K., Boguñá, M., Bianconi, G. & Krioukov, D. Emergence of soft communities from geometric preferential attachment. Sci. Rep. 5, 9421 (2015).
Muscoloni, A. & Cannistraci, C. V. A nonuniform popularity-similarity optimization (npso) model to efficiently generate realistic complex networks with communities. N. J. Phys. 20, 052002 (2018).
García-Pérez, G., Serrano, M. & Boguñá, M. Soft communities in similarity space. J. Stat. Phys. 173, 775–782 (2017).
Yang, W. & Rideout, D. High dimensional hyperbolic geometry of complex networks. Mathematics https://doi.org/10.3390/math8111861 (2020).
Kovács, B., Balogh, S. G. & Palla, G. Generalised popularity-similarity optimisation model for growing hyperbolic networks beyond two dimensions. Sci. Rep. 12, 968 (2022).
Wang, Z., Li, Q., Xiong, W., Jin, F. & Wu, Y. Fast community detection based on sector edge aggregation metric model in hyperbolic space. Phys. A: Stat. Mech. Appl. 452, 178–191 (2016).
Wang, Z., Li, Q., Jin, F., Xiong, W. & Wu, Y. Hyperbolic mapping of complex networks based on community information. Phys. A: Stat. Mech. Appl. 455, 104–119 (2016).
Kovács, B. & Palla, G. The inherent community structure of hyperbolic networks. Sci. Rep. 11, 16050 (2021).
Muscoloni, A. & Cannistraci, C. V. A nonuniform popularity-similarity optimization (npso) model to efficiently generate realistic complex networks with communities. N. J. Phys. 202, 052002 (2018).
Balogh, S. G., Kovács, B. & Palla, G. Maximally modular structure of growing hyperbolic networks. arXiv https://doi.org/10.48550/arXiv.2206.08773 (2022).
Higham, D. J., Rašajski, M. & Pržulj, N. Fitting a geometric graph to a protein-protein interaction network. Bioinformatics 24, 1093–1099 (2008).
Kuchaiev, O., Rašajski, M., Higham, D. J. & Pržulj, N. Geometric de-noising of protein-protein interaction networks. PLoS Comput. Biol. 5, 1–10 (2009).
Cannistraci, C., Alanis-Lobato, G. & Ravasi, T. From link-prediction in brain connectomes and protein interactomes to the local-community-paradigm in complex networks. Sci. Rep. 3, 1613 (2013).
Tadić, B., Andjelković, M. & S̃uvakov, M. Origin of hyperbolicity in brain-to-brain coordination networks. Front. Phys. 6, 7 (2018).
Boguñá, M., Krioukov, D. & Claffy, K. Navigability of complex networks. Nat. Phys. 5, 74–80 (2009).
Boguñá, M., Papadopoulos, F. & Krioukov, D. Sustaining the internet with hyperbolic mapping. Nat. Commun. 1, 62 (2010).
Jonckheere, E., Lou, M., Bonahon, F. & Baryshnikov, Y. Euclidean versus hyperbolic congestion in idealized versus experimental networks. Internet Math. 7, 1–27, https://doi.org/10.1080/15427951.2010.554320 (2011).
Bianconi, G. Interdisciplinary and physics challenges of network theory. EPL (Europhys. Lett.) 111, 56001 (2015).
Chepoi, V., Dragan, F. F. & Vaxès, Y. Core congestion is inherent in hyperbolic networks. In Proc. 28th Annual ACM-SIAM Symposium on Discrete Algorithms (ed. Klein, P. N.) 2264–2279 (SIAM, 2017).
García-Pérez, G., Boguñá, M., Allard, A. & Serrano, M. Á. The hidden hyperbolic geometry of international trade: World trade atlas 1870–2013. Sci. Rep. 6, 33441 (2016).
Serrano, M. A., Krioukov, D. & Boguñá, M. Self-similarity of complex networks and hidden metric spaces. Phys. Rev. Lett. 100, 078701 (2008).
Gulyás, A., Bíró, J., Kőrösi, A., Rétvári, G. & Krioukov, D. Navigable networks as nash equilibria of navigation games. Nat. Commun. 6, 7651 (2015).
Muscoloni, A. & Cannistraci, C. V. Geometrical congruence and efficient greedy navigability of complex networks. arXiv https://doi.org/10.48550/arXiv.2005.13255 (2020).
Shen, D., Wu, Z., Di, Z. & Fan, Y. An asymmetric popularity-similarity optimization method for embedding directed networks into hyperbolic space. Complexity 2020, 8372928 (2020).
Alanis-Lobato, G., Mier, P. & Andrade-Navarro, M. Efficient embedding of complex networks to hyperbolic space via their laplacian. Sci. Rep. 6, 301082 (2016).
Muscoloni, A., Thomas, J. M., Ciucci, S., Bianconi, G. & Cannistraci, C. V. Machine learning meets complex networks via coalescent embedding in the hyperbolic space. Nat. Commun. 8, 1615 (2017).
Keller-Ressel, M. & Nargang, S. Hydra: a method for strain-minimizing hyperbolic embedding of network- and distance-based data. J. Complex Networks https://doi.org/10.1093/comnet/cnaa002 (2020).
Belkin, M. & Niyogi, P. Advances in Neural Information Processing Systems Vol. 14 (MIT Press, 2001).
Alanis-Lobato, G., Mier, P. & Andrade-Navarro, M. A. Manifold learning and maximum likelihood estimation for hyperbolic network embedding. Appl. Netw. Sci. 1, 10 (2016).
García-Pérez, G., Allard, A., Serrano, M. Á. & Boguñá, M. Mercator: uncovering faithful hyperbolic embeddings of complex networks. N. J. Phys. 21, 123033 (2019).
Chamberlain, B. P., Clough, J. & Deisenroth, M. P. Neural embeddings of graphs in hyperbolic space. arXiv https://doi.org/10.48550/arXiv.1705.10359 (2017).
Chami, I., Ying, Z., Ré, C. & Leskovec, J. Advances in Neural Information Processing Systems Vol. 32 (Curran Associates, Inc., 2019).
McDonald, D. & He, S. Heat: Hyperbolic embedding of attributed networks. In Intelligent Data Engineering and Automated Learning—IDEAL 2020: 21st International Conference, Guimaraes, Portugal, November 4–6, 2020, Proceedings, Part I, 28–40 (Springer-Verlag, Berlin, Heidelberg, 2020).
McDonald, D. & He, S. Hyperbolic embedding of attributed and directed networks. In IEEE Transactions on Knowledge and Data Engineering 1–12 (IEEE, 2022).
Palla, G., Tibély, G., Mones, E., Pollner, P. & Vicsek, T. Hierarchical networks of scientific journals. Palgrave Commun. 1, 15016 (2015).
Palla, G. et al. Hierarchy and control of ageing-related methylation networks. PLoS Comput. Biol. 17, e1009327 (2021).
Palla, G., Farkas, I. J., Pollner, P., Derényi, I. & Vicsek, T. Directed network modules. N. J. Phys. 9, 186 (2007).
Ou, M., Cui, P., Pei, J., Zhang, Z. & Zhu, W. Asymmetric transitivity preserving graph embedding. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, KDD ’16, 1105–1114 (Association for Computing Machinery, New York, 2016).
Zhang, Y.-J., Yang, K.-C. & Radicchi, F. Systematic comparison of graph embedding methods in practical tasks. Phys. Rev. E 104, 044315 (2021).
Holland, P. W., Laskey, K. B. & Leinhardt, S. Stochastic blockmodels: first steps. Soc. Netw. 5, 109–137 (1983).
Wang, Y. J. & Wong, G. Y. Stochastic blockmodels for directed graphs. J. Am. Stat. Assoc. 82, 8–19 (1987).
Adamic, L. A. & Glance, N. The political blogosphere and the 2004 U.S. election: divided they blog. In Proc. 3rd International Workshop on Link Discovery, LinkKDD ’05, 36–43 (Association for Computing Machinery, New York, NY, USA, 2005).
Heaberlin, B. & DeDeo, S. The Evolution of Wikipedia’s Norm Network. https://www.mdpi.com/1999-5903/8/2/14 (2016).
Costanzo, M. C. et al. YPD, PombePD and WormPD: model organism volumes of the BioKnowledge library, an integrated resource for protein information. Nucleic Acids Res. 29, 75–79 (2001).
Nelson, D. L., McEvoy, C. L. & Schreiber, T. A. The university of south florida free association, rhyme, and word fragment norms. Behav. Res. Methods Instrum. Comput. 36, 402–407 (2004).
Liben-Nowell, D. & Kleinberg, J. The link prediction problem for social networks. In Proc. Twelfth International Conference on Information and Knowledge Management, CIKM ’03, 556–559 (Association for Computing Machinery, USA, 2003).
Huang, Z., Li, X. & Chen, H. Link prediction approach to collaborative filtering. In Proc. 5th ACM/IEEE-CS Joint Conference on Digital Libraries, JCDL ’05, 141–142 (Association for Computing Machinery, New York, NY, USA, 2005).
Zhou, T., Lü, L. & Zhang, Y.-C. Predicting missing links via local information. Eur. Phys. J. B 71, 623–630 (2009).
Kleinberg, J. Navigation in a small world. Nature 406, 845 (2000).
Muscoloni, A. & Cannistraci, C. V. Navigability evaluation of complex networks by greedy routing efficiency. Proc. Natl Acad. Sci. USA 116, 1468–1469 (2019).
Krioukov, D. Clustering implies geometry in networks. Phys. Rev. Lett. 116, 208302 (2016).
Valdivia, E. A. Random geometric graphs on euclidean balls. arXiv https://arxiv.org/abs/2010.13734 (2020).
Kennedy, W. S., Saniee, I. & Narayan, O. In 2016 IEEE International Conference on Big Data 3344–3351 (Big Data, 2016).
Litvak, N., Michielan, R. & Stegehuis, C. Detecting hyperbolic geometry in networks: why triangles are not enough. arXiv https://arxiv.org/abs/2206.01553 (2022).
Goyal, P. & Ferrara, E. Graph embedding techniques, applications, and performance: a survey. Knowl.-Based Syst. 151, 78–94 (2018).
Yang, Y., Lichtenwalter, R. N. & Chawla, N. V. Evaluating link prediction methods. Knowl. Inf. Syst. 45, 751–782 (2015).
Sinha, A., Cazabet, R. & Vaudaine, R. Systematic biases in link prediction: comparing heuristic and graph embedding based methods. In Complex Networks and Their Applications VII (eds Aiello, L. M. et al.) 81–93 (Springer International Publishing, Cham, 2019).
Kitsak, M., Voitalov, I. & Krioukov, D. Link prediction with hyperbolic geometry. Phys. Rev. Res. 2, 043113 (2020).
Boyd, K., Eng, K. H. & Page, C. D. In Machine Learning and Knowledge Discovery in Databases (eds Blockeel, H., Kersting, K., Nijssen, S. & Železný, F.) 451–466 (Springer Berlin Heidelberg, Berlin, Heidelberg, 2013).
Hanley, J. A. & McNeil, B. J. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143, 29–36 (1982).
Clauset, A., Moore, C. & Newman, M. E. J. Hierarchical structure and the prediction of missing links in networks. Nature 453, 98–101 (2008).
Acknowledgements
This project has received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement no. 101021607 and was partially supported by the National Research, Development and Innovation Office of Hungary under grant no. K128780 and grant no. RRF-2.3.1-21-2022-00006, Datadriven Health Division of Health Security NL.
Funding
Open access funding provided by Eötvös Loránd University.
Author information
Authors and Affiliations
Contributions
B.K. and G.P. developed the concept of the study, B.K. implemented and tested the embedding methods, B.K. pre-processed the network data, performed the analyses and prepared the figures, B.K. and G.P. wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Physics thanks Emil Saucan and the other, anonymous, reviewer for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kovács, B., Palla, G. Model-independent embedding of directed networks into Euclidean and hyperbolic spaces. Commun Phys 6, 28 (2023). https://doi.org/10.1038/s42005-023-01143-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-023-01143-x
This article is cited by
-
Geometric description of clustering in directed networks
Nature Physics (2024)
-
Greedy routing optimisation in hyperbolic networks
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.