Abstract
Hypergraphs offer an explicit formalism to describe multibody interactions in complex systems. To connect dynamics and function in systems with these higher-order interactions, network scientists have generalised random-walk models to hypergraphs and studied the multibody effects on flow-based centrality measures. Mapping the large-scale structure of those flows requires effective community detection methods applied to cogent network representations. For different hypergraph data and research questions, which combination of random-walk model and network representation is best? We define unipartite, bipartite, and multilayer network representations of hypergraph flows and explore how they and the underlying random-walk model change the number, size, depth, and overlap of identified multilevel communities. These results help researchers choose the appropriate modelling approach when mapping flows on hypergraphs.
Similar content being viewed by others
Introduction
Researchers model and map flows on networks to identify important nodes and detect significant communities1,2,3,4,5,6. From small to large system scales, random walk-based methods help to uncover the inner workings of the systems the networks represent7,8. When standard network models with dyadic relations between pairs of nodes fail to adequately represent a system’s interactions, researchers turn to higher-order models of complex systems9,10, including multilayer networks11,12,13,14 for multitype interactions, memory networks15,16,17 for multistep interactions and simplicial complexes18,19,20,21 and hypergraphs22,23,24,25 for multibody interactions.
While several methods can identify flow-based communities in multilayer11,26,27 and memory15,16,17 networks with higher-order Markov dynamics, researchers have focused on combinatorial methods to identify communities in hypergraphs28,29,30,31,32,33 and only recently begun to unravel flow-based community structures associated with random walks guided by hyperedges on hypergraphs25. However, different systems and research questions call for different random-walk and hypergraph models: random walks can be lazy, able to visit the same node multiple times in a row, or non-lazy and forced to move on. Hyperedges can have arbitrary weights, and nodes can have hyperedge-dependent weights. Because these and other models can be represented with different network types—bipartite, unipartite and multilayer—the questions multiply: How do different hypergraph random-walk models combined with different network representations change the flow dynamics at scales captured by communities?
For example, random walks on hypergraphs can model the flow of ideas in co-authorship networks. A node represents an author, and a hyperedge connects all authors of a paper. In the simplest dynamics, a random walker on a node picks a random hyperedge among those that contain the node and steps to a random node of the picked hyperedge. Then repeats. Excluding author self-links for non-lazy walks or including hyperedge weights from paper citations or using hyperedge-dependent node weights for varying author contributions are natural model variations that generate different dynamics23,24. How does the organisation of authors in nested communities from research groups to research areas change with random-walk model and representation? The many combinations of random-walk models and representations available to address specific research problems require us to ask, for different data and different questions, which model and representation is best?
To address which combination of model and representation is best for answering different questions about various hypergraph data, we derive unipartite, bipartite and multilayer network representations of hypergraph flows with identical node-visit rates for the same random-walk model. For unique node-visit rates when a representation requires directed links, we apply an unrecorded teleportation scheme robust to changes in the teleportation rate and that preserves the node-visit rates when teleportation is superfluous in undirected networks34. The information-theoretic and flow-based community detection method Infomap35 allows us to explore how different hypergraph random-walk models and network representation change the number, size, depth and overlap of identified multilevel communities. By analysing schematic and real hypergraphs, we find that the bipartite network representation requires the fewest links and enables the fastest community detection. A multilayer network representation that reinforces flows within similar layers gives the deepest modular structures with the most overlapping communities but at a high computational cost. The unipartite network representation provides a trade-off between the two, with intermediate compactness, speed, and detectable modular regularities.
Results and discussion
Modelling flows on hypergraphs
We model flows on hypergraphs with random walks, using hypergraphs with nodes V, hyperedges E with weights ω, and hyperedge-dependent node weights γ. Each hyperedge e has a weight ω(e). Each node u has a weight γe(u) for each hyperedge e incident to u, E(u) = {e ∈ E: u ∈ e}. To simplify the notation when normalising weights into probabilities, we denote node u’s total incident hyperedge weight d(u) = ∑e∈E(u)ω(e) and hyperedge e’s total node weight δ(e) = ∑u∈eγe(u)23. With these weights, a lazy random walker moves from node u at time t to node v at time t + 1 in three stages by23:
-
1.
Picking hyperedge e among node u’s hyperedges E(u) with probability \(\frac{\omega (e)}{d(u)}\).
-
2.
Picking one of the hyperedge e’s nodes v with probability \(\frac{{\gamma }_{e}(v)}{\delta (e)}\).
-
3.
Moving to node v.
Variations include non-lazy walks, which never visit the same node twice in a row with a modified second stage.
-
2.
Picking one of the hyperedge e’s nodes v ≠ u with probability \(\frac{{\gamma }_{e}(v)}{\delta (e)-{\gamma }_{e}(u)}\),
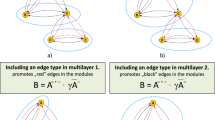
and teleporting walks, which jump to a random node at some rate to ensure that all nodes can be reached from any node in a finite number of moves, so-called ergodic walks. To model flows that tend to stay among similar hyperedges, such as among research papers with similar author lists and likely similar topics, we pick the next hyperedge based on its similarity to the previously picked hyperedge. These hyperedge-similarity walks relate to link communities to reveal pervasively overlapping modules36 and neighbourhood flow coupling to reveal intermittent communities in temporal networks37. Because hyperedge-similarity walks depend on the previously picked hyperedge, they correspond to a higher-order Markov chain model.
These hyperedge-similarity walks require multilayer networks since the other representations contain no information about the previously visited hyperedge26. For example, compare the random walker in the unipartite and multilayer schematic networks in Fig. 1b, d: once the random walker reaches node c, only the multilayer network captures that the random walker came through the hyperedge with nodes c, f and g and can use different transition rates compared with arrival through the hyperedge with nodes a, b and c. Bipartite and unipartite networks, as well as multilayer networks, can represent the other random-walk variations. Altering the random-walk process alters the node-visit rates, but a specific process has identical node-visit rates irrespective of network representation by our design.
a The schematic hypergraph with weighted hyperedges and hyperedge-dependent node weights. White circles labelled from a to j represent nodes, and large orange circles represent hyperedges incident to the nodes in each circle. Thin hyperedge borders for weight 1, medium for weight 2, and thick borders for weight 3. No node borders for node weight 1, thick borders for aggregated weights larger than 1 (Supplementary Code 1). A lazy random walk depicted with an arrow on the schematic hypergraph represented on: b a bipartite network where the unlabelled nodes represent the hyperedges, c a unipartite network and d a multilevel network with grey circles defining each layer. The colours indicate optimised module assignments, in d for hyperedge-similarity walks. The links' thicknesses are proportional to the random walk’s transition rates.
Bipartite networks offer the most direct representation of the basic three-stage random-walk process above. We represent the hyperedges with hyperedge nodes, and the three stages become a two-step walk between the nodes at the bottom and the hyperedge nodes at the top in Fig. 1b. For simplicity, we refer to them as nodes and hyperedge nodes. First a step from a node u to a hyperedge node e,
and then a step from the hyperedge node to a node v,
By starting the random walk on the nodes and taking two steps at a time, corresponding to a two-step Markov process38, hyperedge nodes are only intermediate stops with zero flow when the random walk is back on the nodes after two steps. The stationary distribution of the random walk is concentrated to the nodes. For non-lazy walks represented with bipartite networks, we use so-called state nodes35 in the hyperedge nodes. We let each incoming link to a hyperedge node connect to a state node with out-links to the hyperedge’s all nodes except the incoming link’s source node. This memory network ensures that walks are not backtracking39 (Fig. 2).
White circles with black borders represent hyperedges, and small, coloured circles within the hyperedges represent the state nodes. To prevent random walks on bipartite networks from visiting the same node at the bottom twice in a row by backtracking from the hyperedge node at the top, we use state nodes in the hyperedge nodes. Each hyperedge node requires one state node for each node in the hyperedge. The state nodes have one incoming link from its source node and outgoing links to all other nodes in the hyperedge. Colours indicate the optimised partition. The links' thicknesses are proportional to the random walks' transition rates.
To represent the random walk on a unipartite network, we project the three-stage random-walk process down to a one-step process between the nodes and describe it with the transition rate matrix
where E(u, v) = {e ∈ E: u ∈ e, v ∈ e} is the set of hyperedges incident to both nodes u and v. Each hyperedge forms a fully connected group of nodes (Fig. 1c). Unipartite networks for non-lazy walks have no self-links. The unipartite representation forms a weighted one-mode projection of the bipartite representation and requires more links with its fully connected groups of nodes.
To represent the random walk on a multilayer network, we project the three-stage random-walk process down to a one-step process on state nodes in separate layers. Each hyperedge e with weight ω(e) forms a layer α with weight ω(α). A state node uα represents u in each layer α ∈ E(u) that contains the node. All state nodes in the same layer form a fully connected set (Fig. 1d). The transition rate between state node uα in layer α and state node vβ in layer β is
Node u’s state node-visit rates in different layers sum to u’s visit rate in the unipartite and bipartite representations. With one state node per hyperedge layer that contains the node, the multilayer representation requires most nodes and links to describe the walk. But this cost from including state nodes such that all nodes have a state node for each incident hyperedge comes with benefits: the multilayer representation can describe higher-order Markov chains.
For example, to model flows that tend to stay among similar layers, we pick a hyperedge not only proportional to its weight but also proportional to its similarity to the hyperedge picked in the previous step. To include hyperedge-dependent node weight information in the similarity measure, we use one minus the Jensen–Shannon divergence between the transition rate vectors Pαv and Pβv to nodes at layers α and β as the hyperedge coupling strength,
for β ∈ E(u, v). With node u’s total incident hyperedge weight in layer α
the hyperedge-similarity walk has the transition rates
Because the transition rates at a node depend on the current layer, the random walks generate higher-order Markov dynamics that a unipartite or bipartite network representation without state nodes cannot capture.
To ensure ergodic node-visit rates, we derived an unrecorded teleportation scheme that leaves the node-visit rates unchanged when teleportation is superfluous for hypergraphs with hyperedge-independent node weights, robust to changes in the teleportation rate when teleportation is needed34 and independent of the representation (see Methods).
Mapping flows on hypergraphs
To identify flow-based communities or modules in hypergraphs, we seek to compress a modular description of random walks on the network representations. We cast the problem of finding flow-based communities in hypergraphs as a minimum-description-length problem with the map equation framework4.
The map equation measures, in bits, the optimal codelength L per step of a random walk on a network for a given node partition M with m modules. When all nodes are in the same module, the map equation is simply the Shannon entropy H of the node-visit rates \({\mathcal{P}}=\{{\pi }_{u}\}\). For the schematic example in Fig. 1 with lazy walks, the one-module codelength is
for the bipartite, unipartite, and multilayer network representations because they have the same node-visit rates. The modified hyperedge-similarity walk gives slightly different node-visit rates and codelength.
When the map equation combines within and between-module codelengths in partitions with more than one module, different representations with identical node-visit rates need no longer give the same codelength because the flows between modules can vary. For modules i = 1, …, m with
the map equation takes its general two-level form
The first term is the codelength for between-module movements, followed by the sum of codelengths for within-module movements over all modules.
When a network has modular regularities, a partition captures the modular flows when the random walker spends long times within the modules with few transitions between them. The codelength is shorter than in the one-module solution because the information required to specify a random walker’s position in a module decreases with its size. But for partitions with too many modules, the information required for describing between-module movements exceeds the gain from using small modules. The optimal partition has the shortest codelength. Its node assignment best captures the modular regularities of flows on the network.
Using the optimal three-module solution for the unipartite network representation in Fig. 1c as an example, the codelengths for the bipartite representation—with the leftmost hyperedge assigned with nodes a, b and c in Fig. 1b to match the three-module unipartite solution—and the unipartite representations are
with modules ordered from largest to smallest total flow rate. Since the node-visit rates are the same, the higher between-module flows for the bipartite representation
explain the large codelength difference. In the bipartite representation, a random walker can transition between modules even when visiting the same node multiple times in a row if an incident hyperedge belongs to a different module. Even with a zero node-visit rate that does not contribute to the codelength, a hyperedge node with nodes in multiple modules costs extra bits because its links carry flows across module boundaries. As a result, the bipartite network representation favours fewer, larger modules than the unipartite network representation.
The multilayer representation enables further compression beyond the unipartite solution because a node’s state nodes can belong to different modules. The multilayer compression gain is illustrated for the non-lazy walk on the schematic hypergraph in Fig. 1. In this example, substituting non-lazy for lazy walks does not change the optimal unipartite solution, and the map equation takes the same form as in Eq. (11), but altered node- and link-visit rates change the codelength to 2.63 bits (Table 1). Assigning node f’s two state nodes fα and fβ for its representation in the layers with nodes a, b, c and d, e, f, respectively, to modules two and three in the optimal multilayer solution changes Eq. (11) to
When modules two and three overlap in node f, less flow crosses their boundaries,
The compression gain from reduced flows between modules and within the third module is larger than the loss from adding state node fα to the second module. Overlapping modules in the multilayer hyperedge-similarity representation enable further compression because flows stay even longer within modules.
To find the optimal partitions for the different representations, we use the community-detection algorithm Infomap35. Infomap is to the map equation what the Louvain40 or the Leiden41 method is to the objective function modularity42, which favours partitions with a high internal density of links compared with a statistical null model. Infomap uses a similar search algorithm as the Leiden method but tries to find the node assignment that minimises the map equation’s codelength. Infomap can find not only shallow two-level partitions with nodes in modules, but also deeper hierarchical partitions—from top-level supermodules with multiple levels of submodules down to leaf-level modules containing the nodes—if such multilevel solutions give higher modular compression43. Infomap also finds two-level or multilevel solutions in multilayer networks26.
Using Infomap, we compare how much the different representations can compress modular flows. When mapping flows modelled by lazy and non-lazy random walks on the schematic network in Fig. 1, the optimal partitions of the bipartite networks have two communities. In contrast, the unipartite and multilayer networks have three communities and the multilayer networks with hyperedge-similarity walks have four communities (Table 1 and Fig. 3).
Darker bars represent the optimised modules in each partition, with height proportional to the flow volume of the contained nodes a to j. Streamlines connect modules that contain the same node(s). a Optimal partitions for lazy walks represented with the networks in Fig. 1b–d using the same colours. b Optimal partitions for non-lazy walks. The non-lazy bipartite representation with the same colours as in Fig. 2. h-s hyperedge-similarity.
With a state node for each hyperedge a node belongs to, the multilayer network provides Infomap with degrees of freedom that enables overlapping communities with possibly higher compression. But for this small network, only non-lazy walks give overlapping modules with 0.01 bits compression gain (Table 1). With walks that preferentially move to similar hyperedges, the optimal partitions of the multilayer hyperedge-similarity network representations for lazy and non-lazy random walks both have more overlap in four modules (Table 1 and Fig. 3). The hyperedge-similarity walks favour these overlapping modules because they stay longer within them than the regular walks.
For a given random-walk model, the representations give equivalent node-visit rates but alter the link flows, and with different link flows, the optimal partition can change. The bipartite network representation favours partitions with fewer modules than the unipartite network representation because assigning hyperedge nodes to modules implies encoding more transitions between modules. Multilayer representations, especially with walks that spend longer time among similar hyperedges, favour more overlapping modules. The random-walk model determines how much the multilayer network modules overlap. Non-lazy and hyperedge similarity walks favour overlap because they lead to longer persistence times among nodes in possibly overlapping modules.
Experiments
To illustrate how the network representation affects detected communities in real hypergraphs, we generated a collaboration hypergraph from the 734 references in Networks beyond pairwise interactions: structure and dynamics by Battiston et al.10. We modelled the referenced articles as hyperedges and their authors as nodes. Authors with multiple articles form connections between the hyperedges. We analysed the largest connected component with ∣V∣ = 361 author nodes in ∣E∣ = 220 hyperedges. The median number of authors in a hyperedge is 3, and the authors have contributed to 2.2 articles on average though most have only contributed to one.
Assuming that highly cited papers have higher influence and receive more flows23, we assigned the relative importance of references by their number of citations c in December 2020. Some references had no citations and some were highly cited. One such example is Diffusion of innovations by Everett M. Rogers, with more than 120,000 citations. To avoid disproportionally large or small hyperedge weights ω(e), we weighted the edges by the logarithm of the number of citations and added unit constants to avoid the zero citation problem,
We modelled the authors’ different contributions to articles by assigning higher weights to the first and last author23. We used the edge-dependent node weights
We assumed equal contribution for alphabetically sorted authors, and assigned all of them weight γ(v) = 1. This model ranks a co-corresponding author’s contributions lower than those of the corresponding authors.
To study how hypergraph representations and random-walk models affect the community structure, we generated bipartite, unipartite and multilayer representations for lazy and non-lazy random walks on the collaboration network. We identified nested hierarchical partitions in each network with Infomap, using 100 independent searches for each network. Infomap’s running time depends on the number of nodes, links and solution levels: the bipartite and unipartite representations finished 3–7 times faster than the multilayer representations. The non-lazy bipartite representation with many state nodes ran almost as long.
The optimised partitions for the lazy and non-lazy representations behave like the schematic example: The bipartite representations have the fewest leaf modules and highest codelengths, and the multilayer hyperedge-similarity representations have the most leaf modules and shortest codelengths, with the unipartite and the regular multilayer representations in between (Table 2). Except for the non-lazy bipartite representation with its many state nodes, the lazy representations have more leaf modules and shorter code lengths than their corresponding non-lazy representations because the lazy random walk is more confined than the non-lazy random walk.
With more nodes than in the schematic example, the solutions have more depth. The bipartite solutions have three, and the unipartite and multilayer solutions have four hierarchical levels. The unipartite and multilayer solutions also have more top modules. With non-lazy dynamics, they split the largest top module, and in the lazy dynamics, they split the two largest top modules. But the second-largest top module reunites in the hyperedge-similarity representation, with stronger connections between similar hyperedges (Fig. 4 and Supplementary Fig. 1). The unipartite and multilayer solutions are also most similar at the leaf level (Supplementary Fig. 2).
Darker bars represent the modules in each partition, with height proportional to the flow volume of the contained nodes. Streamlines connect modules that contain the same nodes. Lazy walks in a and non-lazy walks in b. Module names from the top-ranked author within each module. Colours derive from the bipartite representations' partition and differentiate author-groups that collaborate more within the group than with authors in other groups. h-s hyperedge-similarity.
In this larger example, the multilayer hyperedge-similarity representations give more overlap. The non-lazy representations result in higher average overlap because random walkers visiting a node must continue to other nodes, often in the same or a similar hyperedge layer. When random walkers from dissimilar hyperedges come together at a node, they tend to return to where they came from and favour overlapping modules. The non-lazy representations also result in higher max overlap with the same authors topping all representations (Fig. 5).
Authors in the collaboration hypergraph with the highest average effective number of assignments—the per-node module overlap measure in Eq. (25)—in the lazy and non-lazy multilayer representations (see Methods). Curves connect authors between different random-walk models. h-s hyperedge-similarity.
In line with the information-theoretic duality between finding regularities in data and compressing those data, representations that enable deeper solutions with more modules have shorter codelengths (Table 2). The lazy multilayer representation is an exception. Its optimised codelength is bound above by the lazy unipartite representation’s codelength—they have the same codelength for the same hard partition—and overlapping modules can potentially reduce the codelength. Infomap’s best codelength was instead 0.05% longer than for the lazy unipartite representation. Multilayer representations with their many state nodes and links aggravate the search problem, and Infomap could not find a better solution in 100 attempts. But the gain from overlapping modules is higher for the non-lazy multilayer representation and Infomap finds a solution with a significantly shorter codelength.
A case study on the fossil record
Palaeontologists classify major groups of marine animals archived in the fossil record into global-scale faunas that change over time44. They have used standard45 and complex network representations46 to delineate these evolutionary faunas over the past 500 million years. However, it is still unclear how such an organisation of marine animals into modules representing large-scale faunas changes with random-walk model and input network representation.
To illustrate how the network representation of the underlying paleontological data affects empirical estimates of this macroevolutionary pattern, we generated a hypergraph from genus-level fossil occurrences46 available from the Paleobiology Database47. Due to computational limitations, we restricted our analysis to fossil occurrences from the Cambrian (541 MY) to the Cretaceous (66 MY). We modelled the remained 77 geological stages in the reduced data set as hyperedges and the 13,276 fossil genera as nodes. In this hypergraph, genera occurring in multiple geological stages form connections between hyperedges. We weighted the hyperedges by dividing the number of samples where a genus occurs in a given geological stage by the total number of samples recorded at the stage, a procedure modified from ref. 48. We generated bipartite, unipartite and multilayer network representations for lazy and non-lazy random walks from the underlying palaeontology data and identified optimised partitions in the assembled networks with Infomap.
For lazy random walks, Infomap partitioned only the multilayer representations into multilevel communities, with three modules at the first hierarchical level reproducing the Cambrian, Paleozoic (with lower-level modules from Ordovician to Permian) and Mesozoic (with lower-level modules from Triassic to Cretaceous) large-scale or evolutionary faunas44,46 (Fig. 6a). Like the schematic example and the hypergraph of metabolic reaction data, the bipartite representation for the lazy random walks has the fewest leaf modules and highest codelength. The multilayer hyperedge-similarity representation has the most leaf modules, shortest codelength and highest overlap. Leaf modules in this representation can be interpreted as faunas from each geological period in the underlying data (Table 3).
Darker bars represent the modules in each partition, with height proportional to the flow volume of the contained nodes. Streamlines connect modules that contain the same nodes. Lazy walks in a and non-lazy walks in b. We show top modules when a partition lacks deeper levels and leaf modules marked with dashed lines when they exist. Module names from the geological period or era represented by the fauna assemblage. Modules belonging to the Mesozoic era in blue, Carboniferous–Permian in orange, Silurian–Devonian in red, Ordovician in green and Cambrian in purple. h-s hyperedge-similarity.
For non-lazy random walks, Infomap partitioned the bipartite representation into a multilevel solution with shorter codelength than the unipartite representation and the standard multilevel representation (Fig. 6b). The multilayer hyperedge-similarity representation also provides the most leaf modules and the highest overlap. Both multilayer representations reproduce the three large-scale or evolutionary faunas. Unlike the other representations, the multilayer hyperedge-similarity representation’s lower-level modules capture faunas from each geological period, including the Silurian.
Infomap applied to the bipartite representation of the non-lazy random walks identified similar lower-level faunas but combines Cambrian and Paleozoic into a single top module, obscuring the large-scale pattern. For lazy and non-lazy random walk models, unipartite representations fail to capture the larger-scale faunas that characterise the underlying system. Unipartite models also fail to distinguish some lower-level structures, providing a single-scale view of the system that lies between the lowest and higher levels in the multilayer solutions.
Our results suggest that representing fossil occurrence data with multilayer networks offers some advantages to quantify macroevolutionary patterns. Compared with unipartite and bipartite representations, multilayer networks enable discovering more regularities in the fossil record. Their optimised partitions provide higher compression, deeper hierarchy and a better multiscale view.
A case study on metabolic reaction data
Caenorhabditis elegans is an about 1-mm long, transparent nematode found worldwide. C. elegans is one of the most studied model organisms in molecular biology for insights about diseases’ underlying metabolic pathways49,50,51. We used the genome-scale metabolic network model called iCEL127352, which contains 1273 genes, 623 enzymes and 1985 metabolic reactions and is available at wormflux.umassmed.edu. The data include metabolic pathways such as Glu-tRNA(Gln):L-glutamine amido-ligase for Aminoacyl-tRNA biosynthesis. The corresponding reversible reaction \({\rm{ATP}}\ +\ {\rm{GLN}}-{\rm{L}}\ +\ {\rm{GLUTRNAGLN}}\ +\ {{\rm{H}}}_{2}{\rm{O}}\ \leftrightarrow \ {\rm{ADP}}\ + {\rm{GLNTRNA}}\ + {\rm{GLU}}-{\rm{L}}\ +\ {\rm{H}}\ +\ {\rm{PI}}\)with reactants on the left-hand side and products on the right-hand side requires one or more catalysing enzymes. The enzymes catalysing a reaction consist of proteins or protein complexes, which their coding genes’ Boolean logic can describe. For example, we denote the catalysing enzyme for the reaction above by C39B5.6 & Y66D12A.7 & Y41D4A.6, which corresponds to Glutamyl-tRNA(Gln) amidotransferase subunit B, Glutamyl-tRNA(Gln) amidotransferase subunit C and Glutamyl-tRNA(Gln) amidotransferase subunit A.
While standard networks with links between pairs of nodes representing reactants and products in the same reaction can provide insights about cell function3, such dyadic relations fail to capture the co-existence of multiple proteins in complexes. Instead, we use hyperedges to represent metabolic reactions and nodes to represent reactants, products and enzymes. We represent each enzymatic protein complex with genes related by Boolean ANDs by a node such that genes related by Boolean ORs form multiple nodes in the same reaction. While many other abstractions of metabolic systems are possible, this representation naturally describes protein complexes in hypergraphs. To test how different random-walk models and network representations capture functional modules of metabolites and enzymes, we generated unipartite, bipartite, and multilayer representations from the C. elegans hypergraph and identified multilevel communities with Infomap.
All hypergraph representations include modules with protein complexes otherwise overlooked in representations based on standard dyadic relationships. Again, the unipartite and multilayer representations have optimal solutions with shorter codelengths that reveal more modular regularities. The optimal solutions for the bipartite representations have fewer levels or modules (Table 4 and Fig. 7).
Modules that account for 99.9% of the flow volume are included. Darker bars represent the modules in each partition, with height proportional to the flow volume of the contained nodes. Streamlines connect modules that contain the same nodes. Lazy walks in a and non-lazy walks in b. Dashed lines surround the submodules that have the same parent module. Modules that appear together in the largest top module in the multilayer representations' partition coloured in blue. All other modules in orange. h-s hyperedge-similarity.
While the lazy and non-lazy random walk solutions are similar for several representations (Fig. 7a, b), the non-lazy walks give a deeper solution with more modules for the bipartite representation. Nevertheless, the solutions for the bipartite representations aggregate enzymes found in several metabolic processes, while the other representations include modules with enzymes representative of specific biological processes. For example, gene ontology enrichment analysis shows that Module 1:3 in the bipartite solution for non-lazy random walks includes both lipid and amino-acid metabolism. In the unipartite and multilayer representations, this module splits into distinct modules for lipid and amino-acid metabolism with more specific processes (Fig. 7b).
Only the multilayer hyperedge-similarity solutions have significant overlap (Table 4). The module overlaps constitute common metabolites such as water and NAD. Assigning these common metabolites to multiple modules compresses the data more and reveals more regularities in smaller modules. But better representing the specific biological processes come at a relatively high computational cost. Infomap takes much longer to identify overlapping modules in the multilayer networks with numerous state nodes than hard partitions in the unipartite networks. Infomap even fails to compress the multilayer network beyond the unipartite network for non-lazy random walks because the more challenging search problem offsets the tiny compression gain from overlapping modules. The unipartite representation provides a good trade-off between speed and compression, revealing more regularities than the bipartite representation much faster than the multilayer representations.
Conclusions
We have derived unipartite, bipartite, and multilayer network representations of hypergraph flows with different advantages. We used the information-theoretic and flow-based community detection method Infomap to explore how different hypergraph random-walk models and network representations change the number, size, depth and overlap of identified multilevel communities. By identifying flow-based communities both in a schematic and real hypergraphs—a small collaboration hypergraph of researchers working on networks beyond pairwise interactions, a large faunal hypergraph of sampled species across geological stages and the metabolic system of the model organism C. elegans—we found that the bipartite network representation enables the fastest community detection among the tested representations because it uses the fewest links and often has shallower solutions.
A multilayer network representation that reinforces flows within similar layers—one for each hyperedge—gave the deepest modular structures with the most module overlap. But the modular detection gain comes at a high computational cost: combining fully connected layers with other layers requires many more nodes and links than in the bipartite network representation. If the research question does not require hyperedge assignments or overlapping modules, the unipartite network representation provides a trade-off with intermediate compactness, speed and the ability to reveal modular regularities. Among the random-walk models, lazy walks typically give more modules in deeper nested structures, and non-lazy walks provide higher modular overlap. Our methods and results help researchers model and map flows on hypergraphs to study the effects of multibody interactions in complex systems.
Methods
Unrecorded teleportation
With hyperedge-independent node weights where γe(u) = γ(u) for all hyperedges e ∈ E(u), undirected weighted networks can represent the dynamics, and the stationary distribution of the random walk πu is proportional to the product of node u’s total incident hyperedge weight d(u) and weight γ(u). With normalised node-visit rates23,
For the multilayer network representation, the node-visit rates split between layers based on the node u’s incident hyperedge weight per layer state node
With hyperedge-dependent node weights γe(u), only directed weighted networks can represent the dynamics. We use random teleportation to ensure ergodic walks when deriving the node-visit rates with the power-iteration method. Unrecorded teleportation to links minimises the distortion34: in each iteration of the power-iteration method, we distribute a fraction τ = 0.15 of each node’s flow volume among all nodes proportional to their out-link weights. The remaining flow volume moves on the links proportional to their weights. In the last iteration, we move all flows on the links proportional to their weights and record all flows on links and nodes to obtain the ergodic node- and link-visit rates with unrecorded teleportation. This procedure gives equivalent visit rates as simulating a random walker that only records moves on links: with probability 1 − τ, the random walker moves to a node by following the links proportional to their weights and records the link and the target node. With probability τ, the random walker teleports without recording to the link’s source node proportional to the link weight. The normalised number of recordings of each node and link gives the visit rates.
We want teleportation applied to undirected networks—where it is unnecessary—to leave the node- and link-visit rates unchanged. We achieve this smooth teleportation by scaling the transition rates from nodes by the node-visit rates: then unrecorded teleportation proportional to the nodes’ total out-link weights followed by recorded moves on the links proportional to their weights distributes on the nodes according to the ergodic visit rates on undirected networks34. For the general case when the node weights can depend on the hyperedge, and the network may be directed, we use Eq. (18) without assuming γe(u) = γ(u) as an approximation of the node-visit rates:
for nodes and
for state nodes. With exact node-visit rates, we would obtain the stationary flow volumes on links by multiplying the transition rates by the source nodes’ visit rates. With approximate node-visit rates, instead, we obtain the link weights
for bipartite networks,
for unipartite networks, and
for multilayer networks. With unrecorded teleportation proportional to these link weights, modelling flows on hypergraphs give node-visit rates piu and link-flow rates wuv robust to changes in the teleportation rate and independent of the representation.
Module overlap metric
Modules overlap when Infomap assigns a node’s state nodes in the multilayer network representations to different modules. Measuring the overlap through the absolute number of assignments is misleading because the overlap is 2 regardless of the number of state nodes assigned to a different module than the rest. Instead, we used the effective number of assignments. If a fraction f of node u’s state nodes is assigned to the mth module in u’s module assignment set, the mth element of u’s assignment vector is \({a}_{m}^{u}=f\) and the effective number of assignments measured by the perplexity of u’s module assignments is
The effective number of assignments is one if all u’s state nodes are in one module, and it is equal to the number of assignments when the state nodes are divided evenly among u’s module assignments. We averaged over all nodes for the partition overlap.
Data availability
All data are available on GitHub (github.com/mapequation/mapping-hypergraphs). The fossil data are available on the Paleobiology Database47 (paleobiodb.org). The metabolic reaction dataset for C. elegans, iCEL127352, is available at wormflux.umassmed.edu. Furthermore, all data are available from the corresponding author upon request.
Code availability
The source code is available on GitHub (http://github.com/mapequation/mapping-hypergraphs).
Change history
28 June 2021
The original HTML version of this Article was updated shortly after publication to correct equations 12 and 15, which previously inadvertently included LaTeX \hrulefill commands.
References
Brin, S. & Page, L. The anatomy of a large-scale hypertextual web search engine. Comput. Netw. 30, 107–117 (1998).
Simonsen, I., Eriksen, K. A., Maslov, S. & Sneppen, K. Diffusion on complex networks: a way to probe their large-scale topological structures. Physica A 336, 163–173 (2004).
Guimera, R. & Amaral, L. A. N. Functional cartography of complex metabolic networks. Nature 433, 895–900 (2005).
Rosvall, M. & Bergstrom, C. T. Maps of random walks on complex networks reveal community structure. Proc. Natl. Acad. Sci. USA. 105, 1118–1123 (2008).
Delvenne, J., Yaliraki, S. & Barahona, M. Stability of graph communities across time scales. Proc. Natl. Acad. Sci. USA. 107, 12755–12760 (2010).
Mangioni, G., Jurman, G. & De Domenico, M. Multilayer flows in molecular networks identify biological modules in the human proteome. IEEE Trans. Net. Sci.Eng. 7, 411–420 (2018).
Boccaletti, S., Latora, V., Moreno, Y., Chavez, M. & Hwang, D.-U. Complex networks: structure and dynamics. Phys. Rep. 424, 175–308 (2006).
Fortunato, S. Community detection in graphs. Phys. Rep. 486, 75–174 (2010).
Lambiotte, R., Rosvall, M. & Scholtes, I. From networks to optimal higher-order models of complex systems. Nat. Phys. 15, 313–320 (2019).
Battiston, F. et al. Networks beyond pairwise interactions: structure and dynamics. Phys. Rep. 874, 1–92 (2020).
Mucha, P. J., Richardson, T., Macon, K., Porter, M. A. & Onnela, J.-P. Community structure in time-dependent, multiscale, and multiplex networks. Science 328, 876–878 (2010).
De Domenico, M. et al. Mathematical formulation of multilayer networks. Phys. Rev. X 3, 041022 (2013).
Kivelä, M. et al. Multilayer networks. J. Complex Netw. 2, 203–271 (2014).
De Domenico, M., Granell, C., Porter, M. A. & Arenas, A. The physics of spreading processes in multilayer networks. Nat. Phys. 12, 901–906 (2016).
Rosvall, M., Esquivel, A. V., Lancichinetti, A., West, J. D. & Lambiotte, R. Memory in network flows and its effects on spreading dynamics and community detection. Nat. Commun. 5, 1–13 (2014).
Scholtes, I. et al. Causality-driven slow-down and speed-up of diffusion in non-markovian temporal networks. Nat. Commun. 5, 1–9 (2014).
Xu, J., Wickramarathne, T. L. & Chawla, N. V. Representing higher-order dependencies in networks. Science Adv. 2, e1600028 (2016).
Parzanchevski, O. & Rosenthal, R. Simplicial complexes: spectrum, homology and random walks. Random Struct. Algorithms 50, 225–261 (2017).
Salnikov, V., Cassese, D. & Lambiotte, R. Simplicial complexes and complex systems. Eur. J. Phys. 40, 014001 (2018).
Iacopini, I., Petri, G., Barrat, A. & Latora, V. Simplicial models of social contagion. Nat. Commun. 10, 1–9 (2019).
Schaub, M. T., Benson, A. R., Horn, P., Lippner, G. & Jadbabaie, A. Random walks on simplicial complexes and the normalized hodge 1-laplacian. SIAM Rev. Soc. Ind. Appl. Math 62, 353–391 (2020).
Zhou, D., Huang, J. & Schölkopf, B. Learning with hypergraphs: clustering, classification, and embedding. In Advances in Neural Information Processing Systems, 1601–1608 (2007).
Chitra, U. & Raphael, B. J. Random walks on hypergraphs with edge-dependent vertex weights. In 36th International Conference on Machine Learning, ICML 2019, 2002–2011 (International Machine Learning Society (IMLS, 2019).
Carletti, T., Battiston, F., Cencetti, G. & Fanelli, D. Random walks on hypergraphs. Phys. Rev. E 101, 022308 (2020).
Carletti, T., Fanelli, D. & Lambiotte, R. Random walks and community detection in hypergraphs. J. Phys. Complex.2, 015011 (2021).
De Domenico, M., Lancichinetti, A., Arenas, A. & Rosvall, M. Identifying modular flows on multilayer networks reveals highly overlapping organization in interconnected systems. Phys. Rev. X 5, 011027 (2015).
Jeub, L. G., Mahoney, M. W., Mucha, P. J. & Porter, M. A. et al. A local perspective on community structure in multilayer networks. Netw. Sci. 5, 144–163 (2017).
Angelini, M. C., Caltagirone, F., Krzakala, F. & Zdeborová, L. Spectral detection on sparse hypergraphs. In 2015 53rd Annual Allerton Conference on Communication, Control, and Computing (Allerton), 66–73 (IEEE, 2015).
Chien, I., Lin, C.-Y. & Wang, I.-H. Community detection in hypergraphs: optimal statistical limit and efficient algorithms. In International Conference on Artificial Intelligence and Statistics, 871–879 (PMLR, 2018).
Li, P. & Milenkovic, O. Inhomogeneous hypergraph clustering with applications. In Advances in Neural Information Processing Systems (eds Guyon, I. et al.) Vol. 30 (Curran Associates, Inc., 2017). https://proceedings.neurips.cc/paper/2017/file/a50abba8132a77191791390c3eb19fe7-Paper.pdf.
Kamiński, B., Poulin, V., Prałat, P., Szufel, P. & Théberge, F. Clustering via hypergraph modularity. PloS One 14, e0224307 (2019).
Ke, Z. T., Shi, F. & Xia, D. Community detection for hypergraph networks via regularized tensor power iteration. arXiv:1909.06503 (2019).
Chodrow, P. S., Veldt, N. & Benson, A. R. Hypergraph clustering: from blockmodels to modularity. arXiv:2101.09611 (2021).
Lambiotte, R. & Rosvall, M. Ranking and clustering of nodes in networks with smart teleportation. Phys. Rev. E 85, 056107 (2012).
Edler, D. & Bohlin, L. et al. Mapping higher-order network flows in memory and multilayer networks with Infomap. Algorithms 10, 112 (2017).
Ahn, Y.-Y., Bagrow, J. P. & Lehmann, S. Link communities reveal multiscale complexity in networks. Nature 466, 761–764 (2010).
Aslak, U., Rosvall, M. & Lehmann, S. Constrained information flows in temporal networks reveal intermittent communities. Phys. Rev. E 97, 062312 (2018).
Kheirkhahzadeh, M., Lancichinetti, A. & Rosvall, M. Efficient community detection of network flows for varying markov times and bipartite networks. Phys. Rev. E 93, 032309 (2016).
Alon, N., Benjamini, I., Lubetzky, E. & Sodin, S. Non-backtracking random walks mix faster. Commun. Contemp. Math. 9, 585–603 (2007).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
Traag, V. A., Waltman, L. & Van Eck, N. J. From louvain to leiden: guaranteeing well-connected communities. Sci. Rep. 9, 1–12 (2019).
Newman, M. E. & Girvan, M. Finding and evaluating community structure in networks. Phys. Rev. E 69, 026113 (2004).
Rosvall, M. & Bergstrom, C. T. Multilevel compression of random walks on networks reveals hierarchical organization in large integrated systems. PloS One 6, e18209 (2011).
Sepkoski, J. J. A factor analytic description of the Phanerozoic marine fossil record. Paleobiology 7, 36–53 (1981).
Muscente, A. D. et al. Quantifying ecological impacts of mass extinctions with network analysis of fossil communities. Proc. Natl. Acad. Sci. USA. 115, 5217–5222 (2018).
Rojas, A., Calatayud, J., Kowalewski, M., Neuman, M. & Rosvall, M. A multiscale view of the Phanerozoic fossil record reveals the three major biotic transitions. Commun. Biol. 4, 309 (2021).
Peters, S. E. & McClennen, M. The Paleobiology Database application programming interface. Paleobiology 42, 1–7 (2016).
Rojas, A., Patarroyo, P., Mao, L., Bengtson, P. & Kowalewski, M. Global biogeography of Albian ammonoids: a network-based approach. Geology 45, 659–662 (2017).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philos. Trans. R Soc. Lond. B Biol. Sci. 314, 1–340 (1986).
Kaletta, T. & Hengartner, M. O. Finding function in novel targets: C. elegans as a model organism. Nat. Rev. Drug Discov. 5, 387–399 (2006).
Markaki, M. & Tavernarakis, N. Modeling human diseases in caenorhabditis elegans. Biotechnol. J. 5, 1261–1276 (2010).
Yilmaz, L. S. & Walhout, A. J. A caenorhabditis elegans genome-scale metabolic network model. Cell Syst. 2, 297–311 (2016).
Acknowledgements
We thank Christopher Blöcker, Leyden Fernandez, Viktor Jonsson, Michael Schaub, Jelena Smiljanić and Alexander Vergara for valuable comments that helped us improve the manuscript. A. E was supported by the Swedish Foundation for Strategic Research, Grant No. SB16-0089. A. R., D. E. and M. R. were supported by the Swedish Research Council, Grant No. 2016-00796.
The computations were enabled by resources provided by the Swedish National Infrastructure for Computing (SNIC) at High Performance Computing Center North (HPC2N), partially funded by the Swedish Research Council through grant agreement no. 2018-05973.
Funding
Open access funding provided by Umea University.
Author information
Authors and Affiliations
Contributions
A.E. and M.R. conceived the study. A.E., A.R., D.E. and M.D. performed the numerical experiments and analysed the results. A.E. and M.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Communications Physics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Eriksson, A., Edler, D., Rojas, A. et al. How choosing random-walk model and network representation matters for flow-based community detection in hypergraphs. Commun Phys 4, 133 (2021). https://doi.org/10.1038/s42005-021-00634-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-021-00634-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.