Abstract
Protein–protein interaction networks have been used to investigate the influence of SARS-CoV-2 viral proteins on the function of human cells, laying out a deeper understanding of COVID–19 and providing ground for applications, such as drug repurposing. Characterizing molecular (dis)similarities between SARS-CoV-2 and other viral agents allows one to exploit existing information about the alteration of key biological processes due to known viruses for predicting the potential effects of this new virus. Here, we compare the novel coronavirus network against 92 known viruses, from the perspective of statistical physics and computational biology. We show that regulatory spreading patterns, physical features and enriched biological pathways in targeted proteins lead, overall, to meaningful clusters of viruses which, across scales, provide complementary perspectives to better characterize SARS-CoV-2 and its effects on humans. Our results indicate that the virus responsible for COVID–19 exhibits expected similarities, such as to Influenza A and Human Respiratory Syncytial viruses, and unexpected ones with different infection types and from distant viral families, like HIV1 and Human Herpes virus. Taken together, our findings indicate that COVID–19 is a systemic disease with potential effects on the function of multiple organs and human body sub-systems.
Similar content being viewed by others
Introduction
The COVID-19 pandemic, with global impact on multiple crucial aspects of human life, is still a public health threat in most areas of the world. Despite the ongoing investigations aiming to find a viable cure, our knowledge of the nature of the disease is still limited, especially regarding the similarities and differences it has with other viral infections. On the one hand, SARS-CoV-2 shows high genetic similarity to SARS-CoV1—the virus causing 2003 coronavirus outbreak—and its infection shares a number of symptoms with some other respiratory diseases, such as flu caused by Influenza virus. On the other hand, drugs usually used to treat different infection types, like AIDS caused by Human Immunodeficiency Virus (HIV), are under investigation to treat COVID-192,3,4, suggesting potentially unexplored parallel between the function of other viruses and SARS-CoV-2. Characterizing these (dis)similarities can result in a deeper understanding of the novel coronavirus and facilitate the search for reliable treatments.
With the rise of network medicine5,6,7,8,9,10, methods developed for complex networks analysis have been widely adopted to efficiently investigate the interdependence among genes, proteins, biological processes, diseases, and drugs11. Especially, protein–protein interactions (PPI)12 play an essential role in every cellular process and, therefore, PPI network analysis has been extensively used to predict protein function and understand signal transduction pathways in normal or altered conditions. The human PPI networks can include direct (physical) and indirect (functional) interactions, identified through a wide range of experimental and computational techniques.
Additionally, since PPIs are potential drug targets, a better understanding of the interactomes is also essential in drug development. In fact, interactomes are characterized by topological modules bridged by a small number of cross-module PPI13, organized into modular hierarchies14 essential for efficient information exchange15,16 and, consequently, for the system function.
Moreover, PPI network analysis has been used for characterizing the interactions between viral and human proteins in case of SARS-CoV-217,18,19, providing insights into the structure and function of the virus20 and identifying, for instance, drug repurposing strategies21,22,23,24. Very recently, the molecular analysis unraveled the potential reason behind the fact that SARS-CoV-2 infections lead to diverse outcomes for COVID-19, the disease being more severe and lethal preferentially for males and for older patients rather than children and young adults25,26,27.
A comprehensive comparison of SARS-CoV-2 against other viruses has the potential to unravel hidden (dis)similarities with the effects of existing and well-known viral agents, opening the opportunity to network-based applications which complement the more standard ones. However, such a systematic analysis is still missing or limited to a few viruses biologically similar to SARS-CoV-2: recently, the comparative analysis against other zoonotic coronaviruses causing Severe Acute Respiratory Syndrome (SARS) in 2002 and Middle East Respiratory Syndrome in 2012, revealed the existing of pan-viral disease mechanisms28.
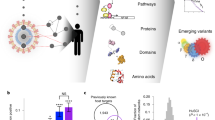
Here, we use statistical physics and techniques from computational biology to analyze pan-viral patterns of 93 viruses, including SARS-CoV-2. We consider the virus–human PPI as an interdependent system with two parts, human PPI network targeted by viral proteins. We carry out a multiscale analysis of virus–host interactomes to highlight how viral interactions impact and perturb the PPI network. In Fig. 1 we illustrate, schematically, the multiscale nature of this work, and the features we extract from the interactomes. To discover pan-viral patterns, we feed advanced machine-learning techniques with the output of physics and biology analyses in order to cluster together viruses with similar physical, biological, or biophysical features. Our findings indicate that SARS-CoV-2 groups with a distinct number of pathogens depending on the physical scale and on the biological information used, providing complementary perspective on its functional effects on organs and human sub-systems. For instance, we find proximity with pathogens such as Human Respiratory Syncytial virus while being very close to other clusters including HIV1 and Herpesvirus, suggesting that COVID-19 exhibits properties typical of systemic diseases. The results of these analyses confirmed the peculiar similarity found between SARS-CoV-2 and viruses from distant families. By integrating all the results obtained from each analysis, we reached a final clustering for viruses which accounts, simultaneously, for biological and physical features from micro to macro scales. Our finding shed light on the unexplored aspects of SARS-CoV-2 from the perspective of statistical physics of complex networks. The presented framework opens the doors for further theoretical developments aiming to characterize structure and dynamics of virus–host interactions, as well as grounds for further experimental investigation and potentially novel clinical treatments, since one can exploit knowledge about existing drug-target interactions related to known viral agents to perform network-based prediction of drug candidates for SARS-CoV-2 from viruses exhibiting similar properties from a statistical physics and biological point of view, thus complementing existing and more biologically only approaches.
Schematic illustration of virus–host interactions across scales, where viral proteins attack human protein targets and the corresponding effects are investigated with distinct techniques from statistical physics of complex networks. Addition of viral components δG to the human protein–protein interaction (PPI) network G generates the virus–human interactome \(G^{\prime}\). Micro: percolation analysis evaluates static structural properties (\(S(G^{\prime} )\) size of the giant connected component) and robustness of the interactome under removal of proteins. Here, the underlying biological hypothesis is that viral proteins might inhibit the usual function of human targets, and we map this activity into the removal of protein from the system. We also test another less invasive hypothesis: the viral proteins interact with the human targets while altering, and not just inhibiting, their functions: the resulting perturbations are propagated (dashed lines mimicking the propagation) and we analyze the system response36,79. Meso: in this case, the underlying hypothesis is that viral proteins alter the function of the human interactome at the mesoscopic level, i.e., interfering with the functional organization in modules (green shaded areas) typical of biomolecular systems13,14,80. This interference is mapped into the isolation of the target proteins, and the modular and hierarchical re-organization of the interactome is detected according to two popular methods for community and hierarchy detection39,40. Macro: viral interactions δG perturb macroscopic properties of the interactome which are captured by the analysis of the network density matrix46,47 von Neumann entropy, Massieu function (ϕ(β, G)) and energy functions at temporal scale β.
Results
Here, we use data regarding the viral proteins and their interactions with human proteins for 93 viruses (see “Methods”). To obtain the virus–human interactomes, we link the data to the BIOSTR Human PPI network (19,945 nodes and 737,668 edges)29,30 built from data fusion of two comprehensive public repositories (see “Methods” and Fig. 2). We also refer to Supplementary Note 1 and Supplementary Figs. 1–4, for summarizing statistics about viruses size, targeted human proteins, and viral families.
BIOSTR human PPI (protein–protein interactions) used in this study, is obtained from data fusion of two comprehensive public repositories, namely STRING and BIOGRID (see the text for details). The network consists of N = 19,945 proteins linked by ∣E∣ = 737,668 edges, and the largest connected component (99.8% nodes, 99.6% edges) is shown (a). Proteins targeted by viruses are highlighted in two ways. On the one hand, markers of distinct size identify targeted proteins: bigger the marker larger the number of times a protein is targeted by viruses in our dataset. On the other hand, distinct colored markers of constant size encode distinct viruses (93 in total, including SARS-CoV-2) and the same color scheme is used to show the contribution of each virus to one of the most frequently targeted proteins (b), TP53, as an example. See Supplementary Figs. 1–3 for more information on the contribution of viruses in targeting a number of other human proteins.
Mapping biology into mathematical models
To allow the analysis from the perspective of statistical physics of complex networks, we first need to map the biology of our problem into mathematical assumptions that can be used operationally. On the one hand, viral proteins try to coopt cellular processes, from protein translation to nuclear transport, through a complex web of PPI. On the other hand, the response of human cells consists in initiating transcriptional programs which activate the adaptive immune system innate and anti-viral countermeasures to control and mitigate virus’ replication. However, DNA and RNA viruses behave differently: the first ones target proteins to alter either human cellular processes or metabolic processes—or both simultaneously—while the second ones tend to target proteins involved into RNA processing, intracellular transport and localization within the cell, preferentially31.
It is worth remarking that our hypotheses in this work do not correspond to a difference between DNA and RNA viruses but, instead, they are intended to provide an operational framework to support the choice of the analytical techniques used in this study. Here, we will consider the following mapping, regardless of the type of virus, whether RNA or DNA, to allow for a consistent comparison of results across all families of viruses considered in this work:
-
(1)
Type-I: the interaction between a viral protein and a human target is assumed to inhibit the function of the latter, destroying its existing interactions with the human interactome. This approach induces a specific change in the function of sub-system the target belongs to and, potentially, in the function of the whole interactome.
-
(2)
Type-II: the interaction between a viral protein and a human target is assumed to perturb the function of the latter, propagating such a perturbation systemically according to some specific biomolecular dynamics.
Note that more sophisticated approaches are also possible: for instance, one can randomly rewire a fraction—or the whole set—of the interactions involving the target protein, thus preserving the overall network connectivity while only altering the functionality of the system. While the Type-I approach inhibits a target, the Type-II also encodes the activation of novel interactions: however, in this second case, the results might depend on the way rewiring is performed—e.g., within or across functional modules—and, to avoid the dependence of our results from the methodology used for rewiring, we prefer to keep the lowest possible number of assumptions and degrees of freedom to employ only Type-I and Type-II approaches.
Percolation of the interactomes and perturbation propagation: microscopic analyses
In this section we introduce two analyses performed on virus–host interactomes at the microscopic scale to detect virus (dis)similarities. A complete discussion of methods and results is presented in the Supplementary Notes 5 and 6. On the one hand, we investigate percolation processes, that, in the past, have been proved useful to shed light on several aspects of protein-related networks, such as in the identification of functional clusters32 and protein complexes33, the verification of the quality of functional annotations34 or identification of critical properties35. These successful applications motivate us to investigate percolation properties of virus–host interactomes. However, it turns out that percolation does not offer valuable insights when it comes at differentiating the topological response of our set of viruses under protein removals (see Supplementary Fig. 11), because the interactomes are too similar between each other. On the other hand, we take a dynamical approach and consider a regulatory dynamic process evolving on top of the reconstructed interactome with the aim of assessing differences between viral agents in the way they impact this system, by means of a dynamic perturbation in its steady state36,37. We employ recent definitions of correlation functions36,38 to quantify the system response. We find that while this approach returns interesting insight regarding the amount of perturbation distributed by single targeted proteins (see Supplementary Fig. 12), there is need for more analyses to bring a comprehensive picture. Therefore, these types of microscopic analyses do not allow us to achieve our goal and we devote the rest of the article to investigate alternative approaches to differentiate between these so topologically similar interactomes.
Functional organization in modules and hierarchy: mesoscale analysis
In this section we analyze how the modular and hierarchical organization of the human interactome changes in response to perturbations caused by viral agents, to shed light on the impact on the functional organization of human proteins and their interactions. Here, the underlying assumption is that the viral proteins alter the functional role of their targets in such a way that they impact on the overall function of the system: operatively, this alteration is mapped into the isolation of protein targets from the network. This method alters the modular structure and the hierarchical organization, leading to a change in the number of functional modules and the hierarchical structure of protein groups. We quantify this change by measuring the number of modules obtained through multiscale modularity maximization based on the Louvain method39 and through the Bayesian inference of a hierarchical degree-corrected stochastic block model (DCSBM)40. The hierarchical structure is probed by extending iteratively the analysis on the network of community nodes, where each module is treated as a supernode of a higher level network. These properties are measured for both the un-targeted human PPI and the targeted virus–human PPI network, the relative change being quantified in the number of modules and in the modularity, captured by ΔModules and ΔModularity, respectively (see Fig. 3). The Louvain method suggests that viral interactions tend to increase the number of modules, decrease the modularity and reduce the number of levels in the hierarchy, indicating a decentralization of functions and a large-scale change in how information is exchanged, respectively. According to our resutls, SARS-CoV-2 exhibits a non-negligible positive change in modularity, like HPV type 16, Influenza A, and Bunyavirus. When analyzed from the perspective of Bayesian inference, we find a larger number of modules on average with respect to Louvain and an opposite trend: after viral interactions, modularity increases in most of the cases. Overall, a few viruses do not alter the hierarchical organization of the human interactome, the trend being a reduction in the number of levels, indicating that information exchange across units might be less efficient15,16. We also compare the new partitioning of functional modules of the targeted interactomes to the un-targeted groups of proteins, via normalized mutual information41 and Variation of Information (see Supplementary Fig. 7 in Supplementary Note 3). Some of the largest variations are detected for human coronaviruses in the Coronaviridae family. SARS-CoV and Coronavirus-229E, despite not having shared targeted human proteins with SARS-CoV-2, and impacting a sensibly lower number of proteins, show a variation in the new modular structure comparable to SARS-CoV-2. A result confirmed by both Louvain and DCSBM community detection methods.
Different methods to detect the mesoscale functional organization of human PPI (protein–protein interactions) highlight the impact of viral interactions on the human interactome in terms of variations in the number of modules, modularity, and hierarchical levels. Axis with virus labels is shared between all panels. a The method based on multiscale modularity maximization (Louvain) shows that viral interactions can significantly disrupt modules, leading to a higher number of smaller modules, while reducing modularity in most of the cases. The relative impact is relevant for SARS-CoV-2 and Influenza A. b The degree-corrected stochastic block model (DCSBM) shows a more heterogeneous pattern, where isolation of targeted proteins can lead to the merging of communities and more modular structure, or vice versa. c The number of levels (N Levels) of the hierarchical structure of targeted interactomes is presented, compared to the un-targeted reference structure (human PPI, dashed red line). Virus–host interactions often lead to a shallower hierarchical arrangement of communities.
Analysis of macroscopic properties: spectral information
In this section, we use statistical physics of complex networks to analyze the macroscopic features of virus–human PPI networks. A variety of methods have been introduced to analyze the information content of complex networks42,43. Since networks can be viewed as collections of entangled entities, a density matrix can be used to describe their state as in quantum statistical mechanics. While some choices of the density matrix have been shown to be unphysical44,45, Gibbsian-like density matrices have been successfully used to define spectral entropy46,47 and estimate the information content of empirical complex networks at multiple scales, with applications ranging from transportation systems48 to the human microbiome46 and the human brain49. In fact, it has been shown that such density matrices describe the short to long range interactions between the nodes, and their Von Neumann entropy encodes the diversity of information dynamics within the structures16. The goal of this section is to study and compare the effect of viral components on the state of information dynamics in the human protein–protein network.
The density matrix can be defined in terms of the combinatorial Laplacian matrix L = D − A, where D is defined as Dij = kiδij, where δij = 1 if i = j and otherwise δij = 0, and \({k}_{i}=\mathop{\sum}\limits_{j}{A}_{ij}\) denotes the degree of ith node. The Laplacian matrix governs the diffusion dynamics on top of the network and is involved in the linear stability analysis of many complex dynamics, such as synchronization. Here we use the Gibbs state given by:
which is defined in terms of the propagator of a diffusion process on top of the network, where β encodes the temporal scale for signal propagation, normalized by the partition function \({\mathcal{Z}}(\beta ,G)=\,\text{Tr}\,\left({e}^{-\beta {\bf{L}}}\right)\), which has an elegant physical meaning in terms of dynamical trapping for diffusive flows48. Consequently, the counterpart of Massieu function—also known as free entropy—in statistical physics can be defined for networks as:
Note that a low value of the Massieu function indicates high information flow between the nodes. The von Neumann entropy can be directly derived from the Massieu function by:
encoding the information content of graph G. In the following, we use the above quantities to compare the interactomes corresponding to different virus–host interactomes. In fact, as the number of viral nodes is much smaller than the number of human proteins, we model each virus–human interdependent system \(G^{\prime}\) as a perturbation of the large human PPI network G (see Fig. 4).
a The BIOSTR human interactome G is targeted by viral proteins, considered as perturbations δG, to build the virus–human interactome \(G^{\prime}\). Here, the SARS-CoV-2 interactome is shown, while excluding the 10% human proteins with the highest degree, for clarity. The interdependence is reflected in the macroscopic functions of the network, perturbing the thermodynamic-like features, at different temporal scales for signal propagation characterized by β. The perturbations of the macroscopic properties of the human protein–protein interaction network (b), reflected in the von Neumann entropy \(\delta {\mathcal{S}}(\beta ,G^{\prime} )\)and the Massieu function \(\delta {\boldsymbol{\phi }}(\beta ,G^{\prime} )\) are shown, where each dot corresponds to a specific virus and the color of the dotts shows their clustering (see the text). More specifically, the von Neumann entropy and Massieu function perturbations caused by each virus, are used as features defining a two-dimensional space, k-means algorithm is used to cluster the viruses at different scales corresponding to different temporal scales β ≈ 1, 3, 5. The trajectories (c) indicate that the perturbation caused by each virus change with β, leading to possibly different clustering at different temporal scales. The color of each trajectory in b is set by the clustering plot given at β ≈ 3 (b).
After considering the viral perturbations due to each virus, the von Neumann entropy and Massieu function of the human PPI network change slightly, as follows:
-
\(\delta {\mathcal{S}}(\beta ,G^{\prime} )={\mathcal{S}}(\beta ,G^{\prime} )-{\mathcal{S}}(\beta ,G)\)
-
\(\delta {\boldsymbol{\phi }}(\beta ,G^{\prime} )={\boldsymbol{\phi }}(\beta ,G^{\prime} )-{\boldsymbol{\phi }}(\beta ,G)\)
In our analysis of the perturbations, the temporal scale β is used as a resolution parameter tuned to characterize the node–node interactions, from short to long range16.
Based on the magnitude of perturbations, caused by the viral components, and using k-means algorithm, a widely adopted clustering technique, we group the viruses together (see Fig. 4)—i.e., the perturbations in Von Neumann entropy and Massieu function shape our two-dimensional feature space and the number of clusters has been calculated using the elbow method at each temporal scale β = 1, 3, 5. A more advanced clustering and the full description of the cluster members at different characteristic propagation time scales is presented later in the text.
Gene ontology and pathways enrichment analysis
To understand if these findings were biologically relevant, we have further performed a clustering analysis on the 93 viruses based on the human proteins they interact with (Supplementary Data 5). We consider a human protein as a “shared target” if it was reported to bind both to a SARS-CoV-2 protein and another virus’ protein, according to the PPI data retrieved from http://viruses.string-db.org (Supplementary Table 1). Out of the 332 human proteins directly targeted by SARS-CoV-2, only 18 of them were found to be also targeted by other viruses, among which Herpes viruses, HPV type 16, Reovirus or Encephalomyocarditis virus (Supplementary Table 1). Figure 5a shows that SARS-CoV-2 does not indeed cluster with any other virus on the basis of shared protein interactors (Supplementary Fig. 2).
Viruses are clustered based on their (a) shared human protein direct targets (first range interactors) and (b) the enriched Reactome pathways containing these proteins. Binary distance has been used to perform the hierarchical clustering in all subplots. Red boxes highlight the cluster containing SARS-CoV-2. Enrichment analyses were performed with the R package clusterProfiler50 with a cutoff of p value <0.005. Viruses for which we could not find any enriched Reactome pathway do not appear on the enrichment heatmaps. Complete results of these enrichment analyses are available as Supplementary material (Supplementary Data 1 and 2).
We then extended our clustering analysis to biological pathways and processes in which these targeted proteins are involved. The R package clusterProfiler50 allows to perform enrichment analysis of gene clusters and was used to identify statistically enriched Reactome pathways51 and Gene Ontology terms52 potentially targeted by the viruses although through multiple different proteins. Considering enriched Reactome pathways, SARS-CoV-2 was shown to have the highest similarity with Bunyavirus and Reovirus (Fig. 5b). The same clustering analysis on Biological Processes as defined by the Gene Ontology database showed that SARS-CoV-2 clusters with Rotavirus C, another virus of the Reoviridae family (Supplementary Fig. 5).
These two methods to assess virus similarities (based either on their targeted proteins, or on their relative enriched pathways among these proteins) are complementary. Although Bunyavirus does not share any human protein target with SARS-CoV-2 (Supplementary Table 1), it is still found to be the most similar to SARS-CoV-2 based on their shared targeted biological pathways (which are mostly related to mitotic checkpoint controls, see Supplementary Table 2).
To investigate whether SARS-CoV-2 would cluster with other viruses at a higher distance, we extended the clustering analysis to the human proteins located one node further of the proteins directly targeted by viruses (referring to them as second-order interactors, Supplementary Data 6). Figure 6a shows that based on the similarity of these second-order interactors, SARS-CoV-2 clusters with more viruses including Hepatitis B and C, HIV-1, Influenza A, Herpesvirus 1/2/8, Varicella, Cytomegalovirus, HPV16, Epstein-Barr and Bunyavirus. Based on enriched pathways from first-order and second-order targets, SARS-CoV-2 clusters with viruses of Bluetongue, West Nile, Cucumber mosaic, Bunyavirus, Reovirus, Rotavirus C, Newcastle disease, Vesicular stomatitis Indiana, Measles, and Myxoma (Supplementary Fig. 6). Gene Ontology Biological Processes-based clustering using first- and second-order targets shows an association of SARS-CoV-2 with more viruses as well, including Human SARS coronavirus, Bunyavirus, HPV16/18, HIV-1/2, African swine fever, Simian virus 40, Avian infectious bronchitis, Influenza A, Herpesvirus 1/2/8, Hepatitis B/C, cytomegalovirus, and Epstein-Barr virus (Fig. 6b). These latter clusters based on enrichments including second-order viral interactors highlight non-trivial functional similarity between viruses of different families, possibly retrieved with the statistical physics approaches mentioned previously, and in agreement with the results described in the last section. Full investigation of these (dis)similarities require further experimental investigations and is beyond the scope of this work.
Viruses are clustered based on their (a) shared human protein targets including direct and secondary interactors and (b) the enriched biological processes (Gene Ontology) containing these direct and secondary targets. Binary distance has been used to perform the hierarchical clustering in all subplots. Red boxes highlight the cluster containing SARS-CoV-2. Enrichment analyses were performed with the R package clusterProfiler50 with a cutoff of p value <0.005. Viruses for which we could not find any enriched GO biological process do not appear on the enrichment heatmaps. Complete results of these enrichment analyses are available as Supplementary Data 3 and 4.
Clusters of viruses
In previous sections, we analyzed the effect of viruses on the human interactome, across different scales. Each analysis, coupled with embedding techniques and clustering algorithms, can be used to investigate the (dis)similarities of viruses from a specific point of view. Here, we use the UMAP dimensionality reduction technique—a machine-learning technique exploiting the hidden geometry of the data—and HDBSCAN—a hierarchical method exploiting spatial density and accounting for the presence of noise—clustering algorithm to groups together the viruses according to their biological and physical effects. We combine the result of different analyses as features to perform the UMAP embedding, to provide an integrated view of virus clusters, identified via HDBSCAN algorithm (for more information and a detailed list of features used, see Supplementary Note 4 and Supplementary Figs. 8–10). In this section, we present the clustering according to three analyses, one based on physical methods, another based on biological and the last one based on their combination (see Fig. 7).
Clustering of viruses according to combination of the features obtained from the mesoscale (including modularity and depth of hierarchy), macroscale (including Massieu and Von Neumann values calculated at temporal scale β = 3), and microscale analyses (including cumulative perturbation, see Supplementary Note 6) and biological analyses (including Gene Ontology GO2, Pathways enrichment PW2, and protein interactors INT2 for first- and second-order shared interactors). To map the multidimensional feature space into a 2d space, we use the UMAP dimensionality reduction technique and find the clusters by means of the HDBSCAN algorithm. In all panels, viruses are shown as dots where their colors indicates their membership in clusters and their size is proportional to the reliability of their assignment to that cluster. In each panel, the labels are added to the viruses that cluster with SARS-CoV-2 and located at the intersection of the dashed lines. a Features from mesoscale organization coupled with spectral entropy and Massieu function perturbations. b Embedding using gene ontology and pathways enrichment analyses. c Features from micro-, meso-, and macroscale analyses are combined with biological analyses GO2, PW2, and INT2.
More specifically, when the mesoscale organization is combined with the results obtained from the spectral entropy and Massieu function (β = 3), SARS-CoV-2 is clustered with Influenza A (Puerto Rico), Human Herpesvirus, Human Parovirus B19, and Mrine Minute virus. Instead, combining GO and Pathways enrichment analyses for second-range interactions, the novel coronavirus exhibits more similarity to Influenza A (Puerto Rico), HIV-1, Epstein-Barr virus, and Vaccina virus. Finally, combining all the mentioned features with microscopic analysis of perturbation propagation and the analysis of second interactors comparison, we find Human Herpesvirus, Epstein-Barr virus, Varicella Zoster virus, Hepatitis C virus in the same cluster with SARS-CoV-2. In the discussion, we report on the clustering results according to each analysis and, also, elaborate on the similarities between the results obtained from physical and biological approaches and their integration.
Discussion
Our knowledge of COVID-19 is still far from being complete. To enhance our understanding of properties of the virus responsible for this emerging disease, one possibility is to compare, at a molecular level, the effects of its interactions with the human interactome against the effects of well-known viral agents. By measuring such effects from multiple analysis, one can use the results to cluster together viruses in order to learn about potential hidden pan-viral relationships. However, comparing COVID-19 against other viral infections is still a challenge, since various approaches can be adopted to characterize and categorize the complex nature of viruses and their impact on human cells.
In this study, we used an approach based on statistical physics to analyze virus–human PPI outlining 93 different viral infections. Our findings suggest that microscopic analyses such as percolation and perturbation propagation are not sensitive to the differentiating features of networks, due to the similarity of interactomes and the high level of details which is a characteristic of microscale analysis (see “Methods”).
Thus, we investigated the effect of virual components on the mesoscale organization of human protein–protein interactome. We used the UMAP dimensionality reduction technique with the HDBSCAN clustering algorithm to find the viruses exhibiting the highest similarity to SARS-CoV-2 in the way they affect the functional modularity, including Influenza A (Puerto Rico) and Marine Minute virus. While this analysis provides mesoscopic details about the impact of viruses on the human proteins, it is not sufficient to identify and compare the global effects of viruses. Therefore, to complement the mesoscale analysis, we used thermodynamic-like quantities—such as the von Neumann entropy and the Massieu function—to quantify the effect of viruses on human interactome, across multiple scales determined by the resolution parameter β. We used the HDBSCAN algorithm again and find SARS-CoV-2 showing similarity to Human Respiratory Syncytial virus at small scales, while at larger scales where the interplay between the topology of virus–host interaction and information flow dynamics becomes more relevant, Measles virus is found in their cluster. It is also worth pointing out that in the geometric space determined by UMAP, the cluster containing SARS-CoV-2 is very close to other clusters including viruses such as SARS-CoV, Human Herpesvirus, and HIV-1, suggesting that SARS-CoV-2 exhibits physical and biological features which makes it similar to viruses well known for their systemic effects, rather than for localized ones. Our findings suggest unexplored relationships between SARS-CoV-2, Herpesvirus, and HIV-1, motivating further theoretical and experimental investigations.
Furthermore, our biological pathways enrichment analysis highlighted that SARS-CoV-2 might impact specific pathways also targeted by other viruses, from different families, although their human protein targets were found to be different, in the strict sense.
In fact, we included the biological analysis based on enrichment with gene ontology and biological pathways, considering only first direct interactors, and then second-order interactors. Concerning the direct interactors, although the approach solely based on protein similarity did not allow to highlight any relevant cluster, 18 human proteins were found to be targeted both by SARS-CoV-2 and by other viruses. Surprisingly, no other members from the coronavirus family were found to share human targets with SARS-CoV-2. However, when using pathway enrichment analysis, we observed that SARS-CoV-2 clustered with Bunyavirus (La Crosse encephalitis) and Reovirus. It is worth noting that La Crosse encephalitis virus can cause inflammation of the brain and its symptoms include nausea, headache, vomiting (in milder cases) and seizures, coma, paralysis, and permanent brain damage (in severe cases)53,54. Additionally, ribavirin has been shown to be effective against La Crosse encephalitis virus both in vitro and in infected patients55,56. Several clinical trials using the same drug to treat COVID-19 are also ongoing57,58,59. Reoviruses can affect the gastrointestinal system (such as rotaviruses) and the respiratory tract. Although they are mostly non-pathogenic in humans, a strain of bat origin has been found to be associated with an acute respiratory disease in humans60. When the second-range targets were included in the clustering analysis, SARS-CoV-2 was observed to share secondary targets, and thus clustered with a wider range of viruses, including viruses responsible for skin and eye infections (Varicella, Cytomegalovirus), or attacking the hepatic (Hepatitis B/C), immune (HIV-1/2), respiratory (Influenza, Epstein-Barr), neurological system (Bunyavirus), or more systemic-infectious viruses (Herpes). This apparent similarity with such diverse viruses may help explain the wide variety of symptoms and organs involved with SARS-CoV-2 infection and COVID-19.
We reach similar conclusions based on both physical and biological approaches, providing evidence for the systemic effect of the novel coronavirus. Noticeably, even when all the considered approaches are combined to reach an integrated view of the virus clusters, we observe the same similarity between SARS-CoV-2 and viruses such as Herpes.
It is worth mentioning that the SARS-CoV-2 outbreak is very recent and its PPI is not yet available on the STRING repository. Therefore, for this particular virus, we relied on a study published in Nature17, in April. We acknowledge the possibility that our results might be affected by the limitations of the currently available data sets.
Overall, our framework opens the doors for further analyses of viral agents from the perspective of combining statistical physics and computational biology, highlighting the sensitivity of macroscopic functions, such as spectral entropy, to small variations across interaction networks and, more specifically, virus–host interactomes. Even though other analyses, such as the perturbation propagation patterns, lack the same sensitivity, according to our results it provides microscopic details about the interactions between viral and human proteins that complement the macroscopic view, together enhancing our understanding of the novel SARS-CoV-2 from a new perspective, which can provide a mathematical ground for the exploration of further clinical treatments and biological understanding.
The most likely application in this direction is drug repurposing, i.e., the identification of new roles of an existing drug to discover previously unknown therapies for untreated diseases. Usually, drugs are combined together to trigger their most direct effects, i.e., at the first-order neighborhood of their targets: however, this approach does not account for potential interference at a systemic level, and databases of empirically discovered side effects have to be taken maintained to be interrogated61,62. Conversely, network-based drug repurposing has the potential to capture those systemic effects, reducing side effects63, an application already being explored for SARS-CoV-2 combining biological information with AI-based techniques21,22,23,24. Our findings complement the ongoing efforts, providing information on similarities between SARS-CoV-2 and other viruses that can be exploited as an additional layer of information for network-based drug repositioning.
Finally, we would like to comment on a more speculative, but extremely fascinating, connection between our findings and latest evidence on the impact of COVID-19 on immune response. On the one hand, in the recent years the study of the human virome64—a part of the microbiome—enhanced our knowledge of its relationships with systemic inflammation, immunophenotype, and disease susceptibility, to mention a few. Usually, the human immune system monitors and co-exists with the virome: however, deviations from this equilibrium condition happening, for instance, when immunity is hampered because of a pathogen like SARS-CoV-2, can lead to the proliferation of other viruses which are successfully suppressed in normal circumstances. This perturbation of the immune system state might lead, as a consequence, to bacterial and viral co-infections, as confirmed by meta-analysis of host pathways in SARS-CoV-2 and its potential copathogens65. It is tempting to consider viruses clustered with SARS-CoV-2 as natural candidates for such co-infections. Intriguingly, the recent literature on this topic is in agreement with this possibility, for instance in the case of the Influenza A66, Epstein-Barr67, HIV68 as well as other respiratory69,70 viruses, such as respiratory syncytial virus and adenovirus.
On the other hand, it is known that some viruses are able to module the development of autoimmune diseases71 through distinct mechanisms, such as molecular mimicry and bystander activation72. SARS-CoV-2 might be in this class of viruses (see73 and refs. therein) and the recent finding for a pathological role for exoproteome-directed autoantibodies in COVID-1974.
Taken together, such an experimental evidence calls for further analysis to gain deeper insights about the physical and biological features of SARS-CoV-2.
Methods
Overview of the dataset
The human interactome used in this study combines PPI from two of the largest repository publicly available to date, namely STRING v10.512—publicly available at https://string-db.org/cgi/download.pl—and BIOGRID v3.5.18275,76—publicly available at https://downloads.thebiogrid.org/BioGRID/Release-Archive/BIOGRID-3.5.182/). For a consistent analysis, all protein names and aliases have been standardized to follow the common nomenclature of official symbols of NCBI gene database (ftp://ftp.ncbi.nlm.nih.gov/gene/DATA/GENE_INFO/Mammalia/ (accessed: 28/03/2020)77). In the following we will refer to this comprehensive network, in standardized format, as BIOSTR.
The virus–host interactions for 93 viruses are collected from the STRING database—publicly available at http://viruses.string-db.org/. We consider interactions of any type as long as their confidence (score) is equal or larger than 0.7. For each virus, we record the targeted human proteins and build a virus–host interactome by merging this information with BIOSTR. While BIOSTR contains 19,945 proteins, the number of human proteins in each human-virus interactome is 19,929, as we excluded the disconnected components. Also, our analyses are focused only on the human interactome and virus–human interactions, discarding the virus–virus interactions.
It is worth noting that to build the COVID-19 virus–host interactions, a different procedure had to be used. In fact, since the SARS-CoV-2 is too novel we could not find its PPI in the STRING repository and we have considered, instead, the targets experimentally observed in Gordon et al.17, consisting of 332 human proteins. The remainder of the procedure used to build the virus–host PPI is the same as before.
Figure 2 shows a visualization of the human interactome where proteins targeted by viruses are highlighted. It is worth noting that viruses target a certain number of proteins which have interesting functions in the interactome. In fact, based on our dataset, TP53 (Tumor Protein p53, NCBI Gene ID: 7157) is the most targeted node: it is responsible for inducing changes in metabolism, DNA repair, apoptosis and cell cycle arrest, and its mutations are associated with several human cancers. Other relevant targets (see Fig. 2) include GK (Glycerol Kinase, NCBI Gene ID: 2710), an important enzyme contributing to regulate metabolism and glycerol uptake, and its mutations are associated with glycerol kinase deficiency; TBP (TATA-box Binding Protein, NCBI Gene ID: 6908), which composes the transcription factor IID, which coordinates the activities of more than 70 polypeptides to initiate the transcription by RNA polymerase II; TLR4 (Toll Like Receptor 4, NCBI Gene ID: 7099), relevant for recognizing pathogens and activating innate immunity; STAT2 (Signal Transducer and Activator of Transcription 2, NCBI Gene ID: 6773), acting as a transcription activator within the cell nucleus: it is likely that it contributes to block interferon-alpha response by adenovirus; PTGS2 (Prostaglandin-endoperoxide Synthase 2, NCBI Gene ID: 5743), a key enzyme involved in the process of prostaglandin biosynthesis; IFIH1 (Interferon Induced with Helicase C domain 1, NCBI Gene ID: 64135), encoding MDA5, an intracellular sensor of viral RNA responsible for triggering the innate immune response: it is fundamental for activating the process of pro-inflammatory response that includes interferons, for this reason it is targeted by several virus families which are able to hinder the innate immune response by evading its specific interferon response.
Gene ontology, reactome pathway, and clustering analysis
The compareCluster function in clusterProfiler R package was used to perform the Reactome pathway enrichment analysis on viral target proteins with a p value cutoff of 0.005. The parameters “enrichPathway” and “enrichGO” with ontology “BP” were used to retrieve enriched Reactome pathways and biological processes from Gene Ontology, respectively. They are based on hypergeometric distribution to calculate enrichment test for GO terms and Reactome pathways, determining whether some protein sets within the same Reactome pathway or defined by particular GO terms are more represented than expected randomly. Enrichment analysis output results were binarized and clustering was performed using pheatmap R package with binary distance and complete method.
Data availability
Data available in figshare repository78.
Code availability
The code to perform the analysis will be available upon request.
References
Andersen, K. G., Rambaut, A., Lipkin, W. I., Holmes, E. C. & Garry, R. F. The proximal origin of SARS-CoV-2. Nat. Med. 26, 450–452 (2020).
Young, B. E. et al. Epidemiologic features and clinical course of patients infected with SARS-CoV-2 in singapore. JAMA 323, 1488 (2020).
Cao, B. et al. A trial of lopinavir–ritonavir in adults hospitalized with severe covid-19. N. Engl. J. Med. 382, 1787–1799 (2020).
Choy, K.-T. et al. Remdesivir, lopinavir, emetine, and homoharringtonine inhibit SARS-CoV-2 replication in vitro. Antivir. Res. 178, 104786 (2020).
Barabási, A.-L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12, 56–68 (2011).
Ivanov, P. C., Liu, K. K. & Bartsch, R. P. Focus on the emerging new fields of network physiology and network medicine. N. J. Phys. 18, 100201 (2016).
Zhou, X., Menche, J., Barabási, A.-L. & Sharma, A. Human symptoms–disease network. Nat. Commun. 5, 1–10 (2014).
Silverman, E. K. & Loscalzo, J. Network medicine approaches to the genetics of complex diseases. Discov. Med. 14, 143 (2012).
Goh, K.-I. et al. The human disease network. Proc. Natl Acad. Sci. USA 104, 8685–8690 (2007).
Halu, A., De Domenico, M., Arenas, A. & Sharma, A. The multiplex network of human diseases. NPJ Syst. Biol. Appl. 5, 1–12 (2019).
Sonawane, A. R., Weiss, S. T., Glass, K. & Sharma, A. Network medicine in the age of biomedical big data. Front. Genet. 10 https://doi.org/10.3389/fgene.2019.00294 (2019).
Szklarczyk, D. et al. String v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Valente, A. X. & Cusick, M. E. Yeast protein interactome topology provides framework for coordinated-functionality. Nucleic Acids Res. 34, 2812–2819 (2006).
Ryan, C. J. et al. Hierarchical modularity and the evolution of genetic interactomes across species. Mol. Cell 46, 691–704 (2012).
Lynn, C. W., Papadopoulos, L., Kahn, A. E. & Bassett, D. S. Human information processing in complex networks. Nat. Phys. 16, 965–973 (2020).
Ghavasieh, A., Nicolini, C. & De Domenico, M. Statistical physics of complex information dynamics. Phys. Rev. E 102, 052304 (2020).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Cui, H. et al. Structural genomics and interactomics of 2019 Wuhan novel coronavirus, 2019-ncov, indicate evolutionary conserved functional regions of viral proteins. bioRxiv https://doi.org/10.1101/2020.02.10.942136 (2020).
Vandelli, A., Monti, M., Milanetti, E., Ponti, R. D. & Tartaglia, G. G. Structural analysis of sars-cov-2 and prediction of the human interactome. https://arxiv.org/abs/2003.13655v4 (2020).
Estrada, E. Fractional diffusion on the human proteome as an alternative to the multi-organ damage of SARS-CoV-2. Chaos: Interdiscip. J. Nonlinear Sci. 30, 081104 (2020).
Zhou, Y. et al. Network-based drug repurposing for novel coronavirus 2019-ncov/sars-cov-2. Cell Discov. 6, 1–18 (2020).
Sadegh, S. et al. Exploring the SARS-CoV-2 virus-host-drug interactome for drug repurposing. Nat. Commun. 11 https://doi.org/10.1038/s41467-020-17189-2 (2020).
Gysi, D. M. et al. Network medicine framework for identifying drug repurposing opportunities for covid-19. Preprint at https://arxiv.org/abs/2004.07229 (2020).
Ray, S., Lall, S., Mukhopadhyay, A., Bandyopadhyay, S. & Schönhuth, A. Predicting potential drug targets and repurposable drugs for covid-19 via a deep generative model for graphs. Preprint at https://arxiv.org/pdf/2007.02338 (2020).
Bastard, P. et al. Autoantibodies against type i ifns in patients with life-threatening covid-19. Science. 370, eabd4585 (2020).
Zhang, Q. et al. Inborn errors of type i ifn immunity in patients with life-threatening covid-19. Science. 370, eabd4570 (2020).
Meffre, E. & Iwasaki, A. Interferon deficiency can lead to severe COVID. Nature https://doi.org/10.1038/d41586-020-03070-1 (2020).
Gordon, D. E. et al. Comparative host-coronavirus protein interaction networks reveal pan-viral disease mechanisms. Science https://science.sciencemag.org/content/early/2020/10/14/science.abe9403.full.pdf (2020).
Verstraete, N. et al. CovMulNet19, integrating proteins, diseases, drugs, and symptoms: a network medicine approach to COVID-19. Netw. Syst. Med. 3, 130–141 (2020).
Verstraete, N. et al. Covmulnet19.zip. figshare https://figshare.com/articles/CovMulNet19_zip/12563192/2 (2020).
Durmuş, S. & Ülgen, K. Ö. Comparative interactomics for virus–human protein–protein interactions: DNA viruses versus RNA viruses. FEBS Open Bio. 7, 96–107 (2017).
Zhang, S., Ning, X. & Zhang, X.-S. Identification of functional modules in a ppi network by clique percolation clustering. Computational Biol. Chem. 30, 445–451 (2006).
Wang, J., Liu, B., Li, M. & Pan, Y. Identifying protein complexes from interaction networks based on clique percolation and distance restriction. BMC Genomics 11, S10 (2010).
Gilks, W. R., Audit, B., de Angelis, D., Tsoka, S. & Ouzounis, C. A. Percolation of annotation errors through hierarchically structured protein sequence databases. Math. Biosci. 193, 223–234 (2005).
Kim, J., Krapivsky, P., Kahng, B. & Redner, S. Infinite-order percolation and giant fluctuations in a protein interaction network. Phys. Rev. E 66, 055101 (2002).
Barzel, B. & Barabási, A.-L. Universality in network dynamics. Nat. Phys. 9, 673–681 (2013).
Maslov, S. & Ispolatov, I. Propagation of large concentration changes in reversible protein-binding networks. Proc. Natl Acad. Sci. USA 104, 13655–13660 (2007).
Barzel, B. & Biham, O. Quantifying the connectivity of a network: the network correlation function method. Phys. Rev. E. 80 https://doi.org/10.1103/physreve.80.046104 (2009).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech.: Theory Exp. 2008, P10008 (2008).
Peixoto, T. P. Hierarchical block structures and high-resolution model selection in large networks. Phys. Rev. X 4, 011047 (2014).
Danon, L., Díaz-Guilera, A., Duch, J. & Arenas, A. Comparing community structure identification. J. Stat. Mech.: Theory Exp. 2005, P09008 (2005).
Cimini, G. et al. The statistical physics of real-world networks. Nat. Rev. Phys. 1, 58–71 (2019).
Radicchi, F., Krioukov, D., Hartle, H. & Bianconi, G. Classical information theory of networks. J. Phys.: Complex. 1, 025001 (2020).
Passerini, F. & Severini, S. The Von Neumann entropy of networks. SSRN Electron. J. https://doi.org/10.2139/ssrn.1382662 (2008).
De Domenico, M., Nicosia, V., Arenas, A. & Latora, V. Structural reducibility of multilayer networks. Nat. Commun. 6, 1–9 (2015).
De Domenico, M. & Biamonte, J. Spectral entropies as information-theoretic tools for complex network comparison. Phys. Rev. X 6, 041062 (2016).
Biamonte, J., Faccin, M. & De Domenico, M. Complex networks from classical to quantum. Commun. Phys. 2 https://doi.org/10.1038/s42005-019-0152-6 (2019).
Ghavasieh, A. & De Domenico, M. Enhancing transport properties in interconnected systems without altering their structure. Phys. Rev. Res. 2, 013155 (2020).
Nicolini, C., Forcellini, G., Minati, L. & Bifone, A. Scale-resolved analysis of brain functional connectivity networks with spectral entropy. NeuroImage 211, 116603 (2020).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterprofiler: an r package for comparing biological themes among gene clusters. Omics: A J. Integr. Biol. 16, 284–287 (2012).
Fabregat, A. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 46, D649–D655 (2018).
Consortium, G. O. The gene ontology (go) database and informatics resource. Nucleic Acids Res. 32, D258–D261 (2004).
McJunkin, J. E. et al. La crosse encephalitis in children. N. Engl. J. Med. 344, 801–807 (2001).
Jones, T. F. et al. Newly recognized focus of la crosse encephalitis in tennessee. Clin. Infect. Dis. 28, 93–97 (1999).
Cassidy, L. F. & Patterson, J. L. Mechanism of la crosse virus inhibition by ribavirin. Antimicrobial Agents Chemother. 33, 2009–2011 (1989).
McJunkin, J. E. et al. Safety and pharmacokinetics of ribavirin for the treatment of la crosse encephalitis. Pediatr. Infect. Dis. J. 30, 860–865 (2011).
Hung, I. F.-N. et al. Triple combination of interferon beta-1b, lopinavir–ritonavir, and ribavirin in the treatment of patients admitted to hospital with covid-19: an open-label, randomised, phase 2 trial. Lancet 395, 1695–1704 (2020).
Khalili, J. S., Zhu, H., Mak, N. S. A., Yan, Y. & Zhu, Y. Novel coronavirus treatment with ribavirin: groundwork for an evaluation concerning covid-19. J. Med. Virol. 92, 740–746 (2020).
Tong, S. et al. Ribavirin therapy for severe covid-19: a retrospective cohort study. Int. J. Antimicrobial Agents 56, 106114 (2020).
Chua, K. B. et al. A previously unknown reovirus of bat origin is associated with an acute respiratory disease in humans. Proc. Natl Acad. Sci. USA 104, 11424–11429 (2007).
Kuhn, M., Campillos, M., Letunic, I., Jensen, L. J. & Bork, P. A side effect resource to capture phenotypic effects of drugs. Mol. Syst. Biol. 6, 343 (2010).
Kuhn, M., Letunic, I., Jensen, L. J. & Bork, P. The sider database of drugs and side effects. Nucleic Acids Res. 44, D1075–D1079 (2016).
Jarada, T. N., Rokne, J. G. & Alhajj, R. A review of computational drug repositioning: strategies, approaches, opportunities, challenges, and directions. J. Cheminformatics 12, 1–23 (2020).
Virgin, H. W. The virome in mammalian physiology and disease. Cell 157, 142–150 (2014).
Vavougios, G. D. Overlapping host pathways between sars-cov-2 and its potential copathogens: an in silico analysis. Infect. Genet. Evolution 86, 104602 (2020).
Kondo, Y., Miyazaki, S., Yamashita, R. & Ikeda, T. Coinfection with sars-cov-2 and influenza a virus. BMJ Case Rep. CP 13, e236812 (2020).
García-Martínez, F. J., Moreno-Artero, E. & Jahnke, S. Sars-cov-2 and ebv coinfection. Med. Clin. 155, 319–320 (2020).
Lai, C.-C., Wang, C.-Y. & Hsueh, P.-R. Co-infections among patients with covid-19: the need for combination therapy with non-anti-sars-cov-2 agents? J. Microbio. Immunol. Infect. 53, 505–512 (2020).
Ma, L. et al. Coinfection of sars-cov-2 and other respiratory pathogens. Infect. Drug Resistance 13, 3045 (2020).
Burrel, S. et al. Co-infection of sars-cov-2 with other respiratory viruses and performance of lower respiratory tract samples for the diagnosis of covid-19. Int. J. Infect. Dis. 102, 10–13 (2020).
Fujinami, R. S. Viruses and autoimmune disease–two sides of the same coin? TRENDS Microbiol. 9, 377–381 (2001).
Fujinami, R. S., von Herrath, M. G., Christen, U. & Whitton, J. L. Molecular mimicry, bystander activation, or viral persistence: infections and autoimmune disease. Clin. Microbiol. Rev. 19, 80–94 (2006).
Galeotti, C. & Bayry, J. Autoimmune and inflammatory diseases following covid-19. Nat. Rev. Rheumatol. 16, 413–414 (2020).
Wang, E. Y. et al. Diverse functional autoantibodies in patients with covid-19. medRxiv https://www.medrxiv.org/content/early/2020/12/12/2020.12.10.20247205.1.full.pdf (2020).
Stark, C. et al. Biogrid: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006).
Oughtred, R. et al. The biogrid interaction database: 2019 update. Nucleic Acids Res. 47, D529–D541 (2019).
Murphy, M. et al. Gene help: integrated access to genes of genomes in the reference sequence collection. In Gene Help [Internet] (Bethesda (MD), National Center for Biotechnology Information (US), 2019). Available from: https://doi.org/https://www.ncbi.nlm.nih.gov/books/NBK3841/.
Ghavasieh, A., Bontorin, S., Artime, O., Verstraete, N. & De domenico, M. Panvirus93interactomes. figshare https://figshare.com/articles/dataset/PanVirus93Interactomes/14103311/1. (2021)
Hens, C., Harush, U., Haber, S., Cohen, R. & Barzel, B. Spatiotemporal signal propagation in complex networks. Nat. Phys. 15, 403–412 (2019).
Guimera, R. & Amaral, L. A. N. Functional cartography of complex metabolic networks. Nature 433, 895–900 (2005).
Acknowledgements
The authors thank Vera Pancaldi for useful discussions.
Author information
Authors and Affiliations
Contributions
A.G., O.A., N.V., and S.B. performed numerical experiments and data analysis. M.D.D. conceived and designed the study. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ghavasieh, A., Bontorin, S., Artime, O. et al. Multiscale statistical physics of the pan-viral interactome unravels the systemic nature of SARS-CoV-2 infections. Commun Phys 4, 83 (2021). https://doi.org/10.1038/s42005-021-00582-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-021-00582-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.