Abstract
The past two decades have seen significant successes in our understanding of networked systems, from the mapping of real-world networks to the establishment of generative models recovering their observed macroscopic patterns. These advances, however, are restricted to pairwise interactions and provide limited insight into higher-order structures. Such multi-component interactions can only be grasped through simplicial complexes, which have recently found applications in social, technological, and biological contexts. Here we introduce a model to grow simplicial complexes of order two, i.e., nodes, links, and triangles, that can be straightforwardly extended to structures containing hyperedges of larger order. Specifically, through a combination of preferential and/or nonpreferential attachment mechanisms, the model constructs networks with a scale-free degree distribution and an either bounded or scale-free generalized degree distribution. We arrive at a highly general scheme with analytical control of the scaling exponents to construct ensembles of synthetic complexes displaying desired statistical properties.
Similar content being viewed by others
Introduction
All beauty, richness, and harmony in the emergent dynamics of a complex system largely depend on the specific way in which its elementary components interact. The last 20 years have seen the birth and development of the multidisciplinary field of network science, wherein a variety of systems in physics, biology, social sciences, and engineering has been modeled by networks of coupled units, in the attempt to unveil the mechanisms underneath the observed systems’ functionality1,2,3,4,5,6.
But the fundamental limit of such a representation is that networks capture only pairwise interactions, whereas the function of many real-world systems not only involves dyadic connections, but rather is the outcome of collective actions at the level of groups of nodes7. For instance, in ecological systems, three or more species compete for food or territory8. Similar multicomponent interactions appear in functional9,10,11,12,13 and structural14 brain networks, protein interaction networks15, semantic networks16, multiauthors scientific collaborations17, offline and online social networks18,19, trigenic interactions in gene regulatory networks20,21, and spreading of social behavior due to multiple, simultaneous, social interactions22.
Simplicial complexes (SCs), being structures formed by simplices of different dimensions (nodes, links, triangles, tetrahedra, etc.), can effectively map the relationships between any number of components. Originally introduced over two decades ago23, SCs are becoming increasingly relevant, thanks to the enhanced resolution of current datasets and the recent advances in data analysis techniques24,25. As real data are being accumulated, we encounter a theoretical challenge: how to synthesize SCs that faithfully reproduce the observed structural features. Significant progresses were made in extending to SCs static graph models, such as random graphs26,27,28, configuration models29,30,31, or activity-driven models32. The fact is however that in many circumstances (such as in the case of scientists collaborating with and citing each other), the network is the result of a growing process, and therefore another approach to model SCs is that of out-of-equilibrium models. Most of the models falling within this category have a geometrical interpretation33 and are variations of the so-called “network geometry with flavor” (NGF) model34,35,36,37,38, which aims at providing a theoretical basis to characterize the underlying geometry of complex networks39 or to explore hidden geometries in complex materials by aggregating SCs as fundamental building blocks40. While the original NGF model adds one34 or more37 maximal simplices of fixed dimension at each time step of the growth process, Fountoulakis et al.41 introduced even the option to remove an existing simplex when a new one is added.
In this paper, we discuss the NGF model proposed by Courtney and Bianconi37 that is able to grow SCs of order two, i.e., structures made of nodes, links and triangles, making use of preferential and nonpreferential rules, and we generalize this model by proposing a model that makes a combination of the two mechanisms. The resulting SCs are characterized by two distributions, the classic degree distribution P(k), capturing the fraction of nodes with degree k, and the generalized degree distribution P(kℓ), where kℓ characterizes the number of triangles supported by each link l = (i, j). We show that our generative model always yields a power-law scaling in P(k), recovering the ubiquitously observed scale-free property42,43, and, at the same time, it allows full control over P(kℓ), i.e., bounded or scale-free with any desired scaling exponent. Indeed, P(kℓ) has been shown to play a crucial role in the emergence of collective behavior, such as synchronization44. We will furthermore show that our methods can be straightforwardly extended to grow structures containing hyperedges of size larger than two.
Results and discussion
SCs growth models
Let us, for the time being and unless otherwise specified, concentrate on NGFs models of SCs of order two proposed by Courtney and Bianconi37. The purpose is to grow a network of N nodes featuring a transitivity coefficient T = 145,46, thus implying that each link is part of a connected triplet of nodes (a triad) which forms a triangle. This is done by starting at t = 0 with an elementary network seed, consisting in an initial clique (an all-to-all connected network) of size N0 ≪ N. Then, at each successive times t = 1, 2, 3, ..., N − N0, a new node is added to the graph. The added node selects mtri already existing links, and forms connections with the 2mtri nodes located at the ends of such links, thus generating mtri new triangles (when mtri > 1, a further condition is enforced that the mtri selected links are not pairwise adjacent, to avoid multiple links from the added node to single existing nodes in the graph).
Such a procedure can be conducted with or without adopting a preferential attachment rule. In the latter case, the probability that the node added at time t selects the specific link (i,j) to form its connections is taken to be \({P}_{ij}(t)=\frac{1}{{N}_{0}\left({N}_{0}-1\right)/2+2{m}_{{\rm{tri}}}(t-1)}\), i.e., the mtri links are randomly selected among all those which already exist in the graph at time t − 1 with equal probability (and therefore without applying a preferential rule). In the former case, instead, one has that \({P}_{ij}(t)=\frac{{k}_{ij}(t-1)}{{\sum }_{i,j}{k}_{ij}(t-1)}\), which implies that the larger is the number of triangles a given edge (i,j) is part of at time t − 1 the larger the probability for that edge of being selected to form a new triangle with the added node. These models are the NGF models proposed by Courtney and Bianconi37 for d = 2, s = 0 (uniform attachment) and d = 2, s = 1 (preferential attachment).
Figure 1a, c reports the degree distribution P(k) of the networks (of size N = 10,000 nodes) generated by the two methods. One immediately sees that in both cases the graphs feature a clear power-law scaling, i.e., the scale-freeness which is indeed characterizing the vast majority of real-world networks1,2,3,4,5,6,42,43. In Fig. 1b, d, we report, instead, a visualization of a typical synthesized network with N = 200 and mtri = 1.
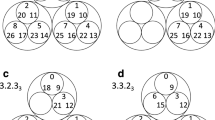
a, b The model without preferential attachment and c, d the model with preferential attachment. a, c Log–log plot of the node degree distribution P(k) of the resulting networks for different values of the number of triangles mtri added at each step of the growth process (see color code in the legend). Data are obtained as an ensemble average over 100 different realizations of a network with size N = 10,000 nodes. Dotted, dashed, and dash-dotted lines in a and c correspond to the analytical predictions given by Eqs. (3) and (9). b, d Schematic visualization of generated networks with N = 200 and mtri = 1. The size of the nodes correlates with their influence in the network in terms of their eigenvector centrality55, such that a node with large size implies a high eigenvector centrality, and therefore the node is connected to many nodes also of large size with high eigencentrality. The width of each link ℓ = (i,j) is proportional to the square root of the link degree kℓ, the number of triangles the link is adjacent to, and the color of the links encodes the supported number of triangles kℓ as reported in the bars at the right of both panels.
We also measure the generalized degree kℓ. Given a graph of N nodes and its adjacency matrix A (the N × N matrix with entries aij = 1 if nodes i and j are connected by a link, and aij = 0 otherwise), the generalized degree kij of the link between node i and j is the (i, j)-entry of the matrix A ⊘ A2 (with the symbol ⊘ standing for the Hadamard product), that is, \({k}_{ij}={(A)}_{ij}\oslash {({A}^{2})}_{ij}\), which accounts for the number of triangles in which the link ij participates. Looking at Fig. 1, one can immediately see that while the two generated networks are essentially heterogeneous in the node degree, the properties of kℓ appear instead to be very different. In particular, the nonpreferential case leads to a much more restricted range of kℓ values as compared to that generated by the preferential rule (see the two color bars at the right of the panels). Moreover, the structure obtained with nonpreferential attachment is homogeneous in terms of kℓ (almost all the six colors, each one representing a given value of kℓ, are visible). On the opposite, the preferential rule generates an SC with a high heterogeneity in kℓ: almost all links are red (they have the lowest as possible value of kℓ), and only one link (the black one) displays a value of kℓ equal to the maximum in the distribution.
These latter features are more quantitatively visible in Fig. 2, which reports the distribution P(kℓ) of the generalized degree kℓ for the two cases. Comparing Fig. 2a, b, one immediately realizes that while P(kℓ) is exponentially decaying when the growth is realized with nonpreferential attachment (Fig. 2a), its scaling is a clear power law in the presence of preferential attachment. A relevant conclusion at this stage is that the two cases are imprinting a completely different topology in the triangular structures (reflected by completely different scaling properties in the distribution of the generalized degree kℓ).
The distribution P(kℓ) vs. the generalized—link—degree kℓ (number of triangles a link ℓ is adjacent to) obtained by growing a network of size N = 10,000 with the nonpreferential (a) and with the preferential (b) attachment models. The data refer to an ensemble average over 100 different realizations of the growth process. Notice that a is in log–linear scale, whereas b is in log–log scale. Legends in both panels report the color code for the number of triangles mtri added at each step of the growth process. The dashed line in a is used for exponential solution given by Eq. (6), while the dashed line in b is used for the power-law solution given by Eq. (12).
Nonpreferential model (NGF37 with d = 2, s = 0)
We now furnish a full analytic treatment, and provide the rigorous expressions for the distributions P(k) and P(kℓ). We start with the case of nonpreferential attachment, and we call N(k,t) the number of nodes with degree k at time t. Its rate equation reads
where \(\frac{dN(k,t)}{dt}\equiv N(k,t+1)-N(k,t)\), and δ is the Kronecker delta function.
For N(t) ≡ ∑kN(k,t) ≃ t, one seeks a solution of the form N(k,t) = tP(k), where P(k) is assumed to be time independent. Since the total number of edges is ~2mtrit, one has ∑kkN(k,t) ≃ 4mtrit, and the rate equation for P(k) becomes
Such a latter equation is the same as Eq. (6) introduced by Boccaletti et al.47, and its solution (for k ≥ 2mtri) is
which perfectly fits the data (see the dotted, dashed-dotted, and dashed black lines in Fig. 1a). A similar, exponential, degree distribution has also been reported in refs. 36,39,48.
Then, one can consider Ne(kℓ,t) as the number of edges participating in kℓ triangles at time t. Its rate equation is
where Ne(t) ≃ 2mtrit and the unwritten terms account for the formation of triangles from two or more linked edges. For the distribution of the generalized degree kℓ, one has that P(kℓ) = Ne(kℓ,t)/Ne(t), and the recursive equation is
admitting the following solution:
The exponential function (6) is reported as a dashed line in Fig. 2a, and one can see that the fit with numerical simulations is rather good, especially for the case mtri = 1.
Preferential model (NGF37 with d = 2, s = 1)
The analytic treatment of the preferential attachment case is far more complicated, as it implies the demonstration of a couple of theorems and the extensive use of a known lemma. We limit here ourselves to furnish the main results without reporting all the (sometimes cumbersome) formal mathematical steps, whereas the interested reader can find the full details within the “Methods” section. Calling again N(k,t) the number of vertices with degree k at time t, its recurrence relation (see details in “Methods”) can be written as
for k > 2mtri, and
for k = 2mtri.
In order to obtain an expression for P(k), one supposes that \(\frac{N(k,t+1)}{t+1}=\frac{N(k,t)}{t}\) for large t. Therefore, one gets \(P(2{m}_{{\rm{tri}}})=\frac{3}{3+2{m}_{{\rm{tri}}}}\) and \(P(k)=P(k-2)\frac{k/2+1}{k/2+1.5}\), which ultimately gives \(P(k)=\frac{3}{3+2{m}_{{\rm{tri}}}}\mathop{\prod }\nolimits_{l = {m}_{{\rm{tri}}}+1}^{k/2}\frac{l-1}{l+1.5}\) or alternatively
where Γ is here the gamma function. The power-law scaling predicted by Eq. (9) fits remarkably well the numerical data (see the dotted, dashed-dotted, and dashed black lines in Fig. 1c) and was also measured in refs. 36,39.
As for P(kℓ), one calls again Ne(kℓ,t) the number of edges participating in kℓ triangles at time t. The recurrence relation for Ne(kℓ,t) (see details in “Methods”) is
for kℓ > 1, and
for kℓ = 1. Imposing that \(\frac{{N}_{{\rm{e}}}({k}_{{\rm{\ell }}},t+1)}{t+1}=\frac{{N}_{{\rm{e}}}({k}_{{\rm{\ell }}},t)}{t}\) for large t, one gets an equation for P(kℓ) which reads as \(P({k}_{{\rm{\ell }}})=P({k}_{{\rm{\ell }}}-1)\frac{{k}_{{\rm{\ell }}}-1}{{k}_{{\rm{\ell }}}+3}\), with \(P(1)=\frac{3}{4}\). The solution is
The power-law function (12) is reported as a dashed line in Fig. 2b, and one can observe that the fit is, once again, extremely good.
Mixed model
Finally, our study can be extended and generalized to a mixed model, through which it is possible to effectively imprint any desired power-law scaling in the triangular structure of the network. Contrary to the previous two cases, our mixed model cannot be encompassed within the framework of the NGF36. In particular, we consider the case in which the probability that the node added at time t selects the specific link (i,j) to form its connections is
for some constants A and B. Here, B is nonnegative, Nℓ(t) ~ 2mtrit is the number of links at moment t and ∑ijkij(t − 1) = 3mtrit is the sum of the generalized degrees of all edges. Notice that A = 0 and \(B=\frac{2}{3}\) (\(A=\frac{1}{2}\) and B = 0) recovers the preferential (nonpreferential) case discussed above.
From the constraint that the sum of all probabilities must be equal to 1, it follows that A and B must obey \(2A+\frac{3}{2}B=1\), so that A = 1/2 − 3/4B. Furthermore, Pij(t) must be nonnegative for all (i,j) (also those for which kij = 1), and this gives the following bounds for A and B: \(-1\le A\le \frac{1}{2}\) and 0 ≤ B ≤ 2. Under these conditions, one can analytically demonstrate (see “Methods” for details) that for strictly positive B values, the resulting networks display scale-free distributions \(P({k}) \sim {k}^{-{\gamma _{\ell}}}\) and \(P({k}_{{{\ell }}}) \sim {k}_{{\rm{\ell }}}^{-{\gamma _ {\ell}}}\) with exponents given by
and
For B = 0, one has instead (see “Methods” for full details) \(P({k}_{{\rm{\ell }}})=\frac{2}{{3}^{{k}_{{\rm{\ell }}}}}\). Equation (14) implies that γ values are between 2 (B = 2) and 3 (B = 0), whereas γℓ is equal to 2 for B = 2, and tends to infinity as B tends to 0 according to Eq. (15). On its turn, this means that choosing B between 1 and 2, γ and γℓ can be preselected ad libitum between 2 and 3, i.e., the imprinted structures of links and triangles feature well defined mean values of the degrees, but unbounded fluctuations as the system grows in size.
In Fig. 3, we report P(k) and P(kℓ) for three distinct values of B. It is seen that the fit between analytic predictions and numerically generated data is always remarkably good. Moreover, the figure demonstrates that our method constitutes a highly general scheme by means of which one can construct, in a fully flexible way, ensembles of synthetic complexes displaying any desired statistical properties [from the condition of Fig. 3a, d featuring a super scale-freeness—where even the mean degrees diverge in the thermodynamic limit—to any milder condition which characterizes in fact many networks from the real world].
Node degree probability distributions P(k) (a–c) and link degree probability distributions P(kℓ) (d–f) obtained by growing networks of size N = 50,000 with the mixed model, at three different values of the parameter B introduced in Eq. (13). The data (blue lines) refer to an average over 100 different realizations of the growth process. Dotted lines report, for comparison, the scaling exponents γ and γℓ predicted by Eqs. (14) and (15), respectively.
Extension to d uniform hypergraphs
The mixed model for SCs can be straightforwardly extended to grow uniform d hypergraphs. For simplicity, let us start by setting N0 = d + 1 nodes in a ring, such that, for d = 2 we have a triangle (2-hyperlink), for d = 3 a square (3-hyperlink), for d = 4 a pentagon (4-hyperlink), and so on. At each time step, a structure of d − 1 new nodes forming an open ring with d − 2 edges (see the sketches in Fig. 4a–c) are added to the network, in order to close a d hyperedge with an existing link. Once again, the probability that a specific link (i,j) is chosen to close the hyperedge is
Nℓ(t) being the total number of links at time t, and kij,d(t) the generalized degree of the (i,j) link, that is, the total number (at time t) of d uniform hyperedges which are incorporating the link ij. Figure 4 reports the node degree distributions P(k) (d–f) and the generalized degree distribution \(P(k_{\ell,d})\) (g–i), where \(k_{\ell,d}\) now characterizes the number of triangles (for d = 2), squares (for d = 3), or pentagons (for d = 4) supported by each link l = (i, j). The reported curves are obtained at different values of B, and the color code is visible in the legend of (a). It is possible to see that, in all cases, the grown structures display the same scaling behavior predicted by our mixed model for the case of SCs of order 2. The extension of our techniques to the growth of SC’s of higher order will be reported elsewhere.
A d hypergraph is a graph in which all hyperlinks contain d + 1 nodes, being d the order of the interaction. a–c Sketches of the processes through which uniform d hypergraphs are grown for d = 2 (a structure formed by 2-hyperlinks with triads of nodes forming triangles), d = 3 (a graph formed by 3-uniform hyperlinks, with all four nodes forming squares), and d = 4 (a uniform 4-hypergraph with all groups of five nodes forming pentagons). The procedure is such that at each time step, a chain of d − 1 new nodes (orange circles connected with solid lines) is added to the network to form a uniform d hyperedge by connecting the ends of the chain (dashed links) to an existing link (blue thick link). d–f The node degree k probability distributions P(k) and g–i the probability distribution \(P(k_{\ell,d})\) of the generalized link degree kℓ,d, that is, the number of triangles (d = 2), squares (d = 3), and pentagons (d = 4) each link participates in, obtained by growing hypergraphs of size N = 10,000 at four different values of the parameter B introduced in Eq. (16) (reported in the legend of d where NPA and PA stand for nonpreferential and preferential attachment, respectively). Data refer to an average over 100 different realizations of the growth process.
In summary, complex networks encode the basic architecture of social, biological, and technological networks, touching upon the most crucial challenges of modern science, from the spread of epidemics in social networks20,49 to the resilience of our eco-systems and critical infrastructure50.
In the context of pairwise interactions, the most natural measure of centrality is a node’s individual degree, capturing its potential dynamic impact on the system51. The discovery that most real-world networks exhibit extreme levels of degree heterogeneity was disruptive—indicating that networks are highly centralized, with a potentially disproportionate role played by a small fraction of their components42,43.
As we deepen our investigation into the interaction patterns of complex systems, it becomes increasingly clear, however, that higher order structures, beyond pairwise interactions, underlie much of the observed richness of real-world networks. Hence, we sought the fundamental rules that prescribe centrality, and govern its distribution, in an SC environment. In addition to this, the proposed models are expected to exhibit the small-world property, simultaneously featuring short average distances which increase proportionally to the logarithm of the network size (see Fig. 5) and large modularity and clustering coefficients—as new nodes enter the network closing triads. A natural measure is the number of complexes, here triangles, that a link participates in. Indeed, an SC represents a potentially functional unit, such as a collaboration of a social team17, or a trio of interacting biochemical agents20,21. A component that is part of many such complexes is, therefore, likely central in the functionality of the system.
Networks are grown with the mixed model for different values of the parameter B introduced in Eq. (13) and for the number of triangles added at each step of the growth process mtri = 1 (full symbols) and mtri = 2 (void symbols). Each point is an ensemble average over 100 network realizations and error bars represent the standard deviation. Notice the linear–log scale and that the network diameter is proportional to \(\mathrm{log}\,N\). NPA and PA stand for nonpreferential and preferential attachment, respectively.
Several growing network models have been introduced and studied, which help exposing the roots of SC heterogeneity, shedding light on the emergence of centrality beyond the degree distribution. As we seek to understand the behavior of complex systems, their resilience and dynamic functionality, we hope that our comprehension into the microscopic processes of their formation can provide meaningful macroscopic insights.
Methods
In this final section, we furnish all the details of the analytical results regarding our models. The starting point is that, as the synthesized networks have transitivity coefficient T = 1 (i.e., no links exist which do not form part of at least a triangle), the degree of each vertex is the number of triangles containing that vertex multiplied by 2, and one has that Nv(kv, t) = N(2k, t), where Nv(kv, t) is the number of nodes participating in kv triangles and N(2k, t) is the number of nodes having degree 2k at time t. As a consequence one has that P(kv) = P(k), and is therefore entitled to concentrate on either one of such distributions, depending on which one finds the simpler analytical treatment.
The nonpreferential attachment case
Let Nv(k, kv, t) be the number of nodes with k neighbors participating in kv triangles at time t. The rate equation is
where the unwritten terms account for the formation of triangles from two or more linked edges. In the sequel, we assume that the chosen edges are not linked (none of the nodes of a selected edge is linked to any of the nodes of another selected edge, which is always the case for mtri = 1). Hence, the number of edges and triangles related to a given node increases one by one. Only the new nodes entering the system have the number of edges (2mtri) double than the number of triangles in which they are participating (mtri). This way, for k big enough, one has
After summing over all values of k, one obtains an approximate equation for Nv(kv,t), the number of nodes participating in kv triangles
One then can proceed as in the case of the degree distribution P(k) (see the main text), and seek a solution of the form Nv(kv,t) = tP(kv)
The solution for kv ≥ mtri is
Therefore, one has also that P(k) ~ k−3 which coincides with Eq. (3) in the main text.
The preferential attachment case
In order to obtain P(k) and P(kℓ), one needs here to make use of the following lemma reported by Chung and Lu52:
Lemma 1 Suppose that a sequence {at} satisfies a recurrence relation
where t0 and t1 are arbitrary, positive, fixed, values. Furthermore, suppose that \({\mathrm{lim}\,}_{t\to \infty }{b}_{t}=b\,> \, 0\) and \({\mathrm{lim}\,}_{t\to \infty }{c}_{t}=c\,> \,0\).
Then, \({\mathrm{lim}\,}_{t\to \infty }\frac{{a}_{t}}{t}\) exists and one has
For instance (and indicating by \({\mathbb{E}}(\cdot )\) the expectation value), P(k) for k = 2mtri can be obtained by setting \({b}_{t}=b=\frac{2{m}_{{\rm{tri}}}}{3}\) and ct = c = 1. Application of Lemma 1 ensures that the limit \({\mathrm{lim}\,}_{t\to \infty }\frac{{\mathbb{E}}(N(2{m}_{{\rm{tri}}},t))}{t}\) exists and is equal to \(\frac{3}{3+2{m}_{{\rm{tri}}}}\), leading to \(P(1)=\frac{3}{3+2{m}_{{\rm{tri}}}}\).
On the other hand, P(k) for k > 2mtri is obtained assuming that P(k − 2) exists, and applying Lemma 1 again with \({b}_{t}=b=\frac{k}{3}\) and \({c}_{t}={\mathbb{E}}(N(k-2,t))\frac{k-2}{3t}\) (i.e., taking \(c=P(k-2)\frac{k-2}{3}\)). Such a choice, indeed, entitles one to write a recurrence relation, and to obtain an explicit formula for P(k)
which coincides with Eq. (9) of the main text. Here, Γ is the gamma function.
Furthermore, one can demonstrate that P(k) is sharp, by the use of the following theorem:
Theorem 1 For any fixed ε > 0 and δ > 0 and for any large enough t, the difference between the number of vertices with degree k at time t and P(k)t is smaller than εt with probability larger than 1 − δ.
The theorem can be proved by considering the martingale \({X}_{l}={\mathbb{E}}(N(k,t)| {{\mathcal{F}}}_{l})\) (where \({{\mathcal{F}}}_{l}\) is the σ-algebra generated by the probability space at time l). It is rather easy to show that ∣Xl+1 − Xl∣ is bounded by 4, and therefore the theorem follows from the Azuma–Hoeffding inequality53,54.
When trying to obtain P(kℓ), one can encounter a problem in the case in which the triangles added at step t have one or more common edges, so that the number of different edges added at time t might not be equal to 2mtri.
Denoting by e(t, kℓ) the number of added edges of degree kℓ at time t, the more accurate recurrence relation for kℓ > 1 is the following:
and for kℓ = 1 one has
Using Theorem 1 (and some nontrivial math, of which we omit the technical details), one can prove that \({\mathrm{lim}\,}_{t\to \infty }e(t,1)=2{m}_{{\rm{tri}}}\) and \({\mathrm{lim}\,}_{t\to \infty }e(t,{k}_{{\rm{\ell }}})=0\) for kℓ > 1, so that one has \({\mathrm{lim}\,}_{t\to \infty }{\mathbb{E}}(e(t,1))=2{m}_{{\rm{tri}}}\) and \({\mathrm{lim}\,}_{t\to \infty }{\mathbb{E}}(e(t,{k}_{{\rm{\ell }}}))=0\) for kℓ > 1.
Furthermore, one can prove another theorem:
Theorem 2 The number of different edges divided by t converges always to 2mtri.
It follows that \(P({k}_{{\rm{\ell }}})={\mathrm{lim}\,}_{t\to \infty }\frac{{\mathbb{E}}({N}_{{\rm{e}}}({k}_{{\rm{\ell }}},t))}{2{m}_{{\rm{tri}}}t}\).
Finally, one can apply Lemma 1 with \({b}_{t}=b=\frac{1}{3}\) and \(c=\mathrm{lim}\,1+O\left(\frac{1}{{t}^{2}}\right)=1\) to get that \(P(1)={\mathrm{lim}\,}_{t\to \infty }\frac{{\mathbb{E}}({N}_{{\rm{e}}}(1,t))}{2{m}_{{\rm{tri}}}t}\) exists and is equal to \(\frac{3}{4}\). For kℓ > 1, one assumes that \(P({k}_{{\rm{\ell }}}-1)={\mathrm{lim}\,}_{t\to \infty }\frac{{\mathbb{E}}({N}_{{\rm{e}}}({k}_{{\rm{\ell }}}-1,t))}{2{m}_{{\rm{tri}}}t}\) exists, and applies again Lemma 1 with \({b}_{t}=b=\frac{{k}_{{\rm{\ell }}}}{3}\) and \(c={\mathrm{lim}}\,{c}_{t}=P({k}_{{\rm{\ell }}}-1)\frac{({k}_{{\rm{\ell }}}-1)}{3t}\). Hence, \(P({k}_{{\rm{\ell }}})={\mathrm{lim}\,}_{t\to \infty }\frac{{\mathbb{E}}({N}_{{\rm{e}}}({k}_{{\rm{\ell }}},t))}{2{m}_{{\rm{tri}}}t}\) exists and is equal to \(P({k}_{{\rm{\ell }}}-1)\frac{{k}_{{\rm{\ell }}}-1}{{k}_{{\rm{\ell }}}+3}\). From such a recurrence relation, one finally obtains the explicit formula for P(kℓ)
which is identical to Eq. (4) of the main text.
Finally, since the degree of the vertex is the number of triangles containing this vertex multiplied by 2, one has Nv(kv,t) = N(2k,t) and as a consequence one obtains
The mixed case
The recurrence relation for Nv(kv,t) (the number of vertices participating in kv triangles at time t) is
For t > 0 and kv = mtri one has
Furthermore, the use of Lemma 1 gives
and
Therefore, one has
or
where \({\gamma }_{v}=1+\frac{1}{A+B}\).
In addition, one can consider \(N_{\ell}(k_{\ell}, t)\), the number of links with degree kℓ at moment t. Its recurrence equation is
For kℓ = 1 one has
One can apply again Lemma 1, and obtain
Therefore, one gets
Let us consider separately the cases B = 0 (the fully nonpreferential case) and B > 0.
For B = 0, one has
Since \(A=\frac{1}{2}\), one obtains
which fully coincide with the exponential scaling derived in the main text for the nonpreferential case.
On the other hand, when instead B > 0, one has
where \({\gamma }_{\ell}=1+\frac{2}{B}\).
Data availability
Data sharing not applicable to this article as no datasets were generated or analysed during the current study.
Code availability
All the codes used in this study can be found in Supplementary Software file.
References
Dorogovtsev, S. N. & Mendes, J. F. F. Evolution of networks. Adv. Phys. 51, 1079–1187 (2002).
Newman, M. The structure and function of complex networks. SIAM Rev. 45, 167–256 (2003).
Boccaletti, S., Latora, V., Moreno, Y., Chavez, M. & Hwang, D.-U. Complex networks: structure and dynamics. Phys. Rep. 424, 175–308 (2006).
Estrada, E. The Structure of Complex Networks: Theory and Applications (Oxford University Press, Inc., 2011).
Boccaletti, S. et al. The structure and dynamics of multilayer networks. Phys. Rep. 544, 1–122 (2014).
Latora, V., Nicosia, V. & Russo, G. Complex Networks: Principles, Methods and Applications (Cambridge University Press, 2017).
Battiston, F. et al. Networks beyond pairwise interactions: structure and dynamics. Phys. Rep. 874, 1–92 (2020).
Levine, J., Bascompte, J., Adler, P. & Allesina, S. Beyond pairwise mechanisms of species coexistence in complex communities. Nature 546, 56–64 (2017).
Petri, G., Scolamiero, M., Donato, I. & Vaccarino, F. Topological strata of weighted complex networks. PLoS ONE 8, e66506 (2013).
Sizemore, A., Giusti, C. & Bassett, D. S. Classification of weighted networks through mesoscale homological features. J. Complex Netw. 5, 245–273 (2016).
Lee, H., Kang, H., Chung, M. K., Kim, B.-N. & Lee, D. S. Persistent brain network homology from the perspective of dendrogram. IEEE Trans. Med. Imaging 31, 2267–2277 (2012).
Petri, G. et al. Homological scaffolds of brain functional networks. J. R. Soc. Interface 11, 20140873 (2014).
Lord, L.-D. et al. Insights into brain architectures from the homological scaffolds of functional connectivity networks. Front. Syst. Neurosci. 10, 85 (2016).
Sizemore, A. E. et al. Cliques and cavities in the human connectome. J. Comput. Neurosci. 44, 115–145 (2018).
Estrada, E. & Ross, G. J. Centralities in simplicial complexes. applications to protein interaction networks. J. Theor. Biol. 438, 46–60 (2018).
Sizemore, A. E., Karuza, E. A., Giusti, C. & Bassett, D. S. Knowledge gaps in the early growth of semantic feature networks. Nat. Hum. Behav. 2, 682–692 (2018).
Patania, A., Petri, G. & Vaccarino, F. The shape of collaborations. EPJ Data Sci. 6, 18 (2017).
Freeman, L. C. Q-analysis and the structure of friendship networks. Int. J. Man-Mach. Stud. 12, 367–378 (1980).
Andjelković, M., Tadić, B., Maletić, S. & Rajković, M. Hierarchical sequencing of online social graphs. Phys. A: Stat. Mech. Appl. 436, 582–595 (2015).
Harush, U. & Barzel, B. Dynamic patterns of information flow in complex networks. Nat. Commun. 8, 2181 (2017).
Kuzmin, E. et al. Systematic analysis of complex genetic interactions. Science 360, 1–9 (2018).
Centola, D. The spread of behavior in an online social network experiment. Science 329, 1194–1197 (2010).
Aleksandrov, P. Combinatorial Topology, Vol. 1 (Dover Publications, Inc, 1998).
Salnikov, V., Cassese, D. & Lambiotte, R. Simplicial complexes and complex systems. Eur. J. Phys. 40, 014001 (2018).
Sizemore, A. E., Phillips-Cremins, J. E., Ghrist, R. & Bassett, D. S. The importance of the whole: topological data analysis for the network neuroscientist. Netw. Neurosci. 3, 656–673 (2019).
Kahle, M. Topology of Random Simplicial Complexes: a Survey. AMS Contemp. Math 620, 201–222 (2014).
Costa, A. & Farber, M. Random simplicial complexes. In Configuration Spaces: Geometry, Topology and Representation Theory (eds Callegaro, F. et al.), 129–153 (Springer International Publishing, 2016).
Iacopini, I., Petri, G., Barrat, A. & Latora, V. Simplicial models of social contagion. Nat. Commun. 10, 2485 (2019).
Courtney, O. T. & Bianconi, G. Generalized network structures: the configuration model and the canonical ensemble of simplicial complexes. Phys. Rev. E 93, 062311 (2016).
Young, J.-G., Petri, G., Vaccarino, F. & Patania, A. Construction of and efficient sampling from the simplicial configuration model. Phys. Rev. E 96, 032312 (2017).
Bianconi, G., Kryven, I. & Ziff, R. M. Percolation on branching simplicial and cell complexes and its relation to interdependent percolation. Phys. Rev. E 100, 062311 (2019).
Petri, G. & Barrat, A. Simplicial activity driven model. Phys. Rev. Lett. 121, 228301 (2018).
Boguña, M. et al. Network geometry. Nat. Rev. Phys. 3, 114–135 (2021).
Bianconi, G. & Rahmede, C. Complex quantum network manifolds in dimension d > 2 are scale-free. Sci. Rep. 5, 13979 (2015).
Bianconi, G., Rahmede, C. & Wu, Z. Complex quantum network geometries: evolution and phase transitions. Phys. Rev. E 92, 022815 (2015).
Bianconi, G. & Rahmede, C. Network geometry with flavor: from complexity to quantum geometry. Phys. Rev. E 93, 032315 (2016).
Courtney, O. T. & Bianconi, G. Weighted growing simplicial complexes. Phys. Rev. E 95, 062301 (2017).
Mulder, D. & Bianconi, G. Network geometry and complexity. J. Stat. Phys. 173, 783–805 (2018).
Bianconi, G. & Rahmede, C. Emergent hyperbolic network geometry. Sci. Rep. 7, 41974 (2017).
Šuvakov, M., Andjelković, M. & Tadić, B. Hidden geometries in networks arising from cooperative self-assembly. Sci. Rep. 8, 1987 (2018).
Fountoulakis, N., Iyer, T., Mailler, C. & Sulzbach, H. Dynamical models for random simplicial complexes. arXiv: 1910.12715 (2019).
Barabási, A.-L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Albert, R. & Barabási, A.-L. Statistical mechanics of complex networks. Rev. Mod. Phys. 74, 47–97 (2002).
Gambuzza, L. V. et al. The master stability function for synchronization in simplicial complexes. Nat. Commun. 12, 1255 (2021).
Barrat, A. & Weigt, M. On the properties of small-world network models. Eur. Phys. J. B 13, 547–560 (2000).
Newman, M. E. J. Scientific collaboration networks. i. network construction and fundamental results. Phys. Rev. E 64, 016131 (2001).
Boccaletti, S., Hwang, D.-U. & Latora, V. Growing hierarchical scale-free networks by means of nonhierarchical processes. Int. J. Bifurc. Chaos 17, 2447–2452 (2007).
Dorogovtsev, S., Mendes, J. & Samukhin, A. Size-dependent degree distribution of a scale-free growing network. Phys. Rev. E 63, 062101 (2001).
Vespignani, A. Modelling dynamical processes in complex socio-technical systems. Nat. Phys. 8, 32–39 (2012).
Hacohen, A., Cohen, R., Efroni, S., Barzel, B. & Bachelet, I. Digitizable therapeutics for decentralized mitigation of global pandemics. Sci. Rep. 9, 14345 (2019).
Barzel, B. & Barabási, A.-L. Universality in network dynamics. Nat. Phys. 9, 673–681 (2013).
Chung, F. & Lu, L. Complex Graphs and Networks (American Mathematical Society, 2004).
Hoeffding, W. Probability inequalities for sums of bounded random variables. J. Am. Stat. Assoc. 58, 13–30 (1963).
Azuma, K. Weighted sums of certain dependent random variables. Tohoku Math. J. 19, 357–367 (1967).
Bonacich, P. Factoring and weighting approaches to status scores and clique identification. J. Math. Sociol. 2, 113–120 (1972).
Acknowledgements
I.S.-N. acknowledges support from the Ministerio de Economía, Industria y Competitividad, Spain, under project FIS2017-84151-P. The work of A.M.R. and D.M. was supported by the Russian Federation Government (Grant number 075-15-2019-1926).
Author information
Authors and Affiliations
Contributions
K.K., I.S.-N., N.K., S.B., and B.B. equally contributed to the manuscript. S.B., B.B., I.S.-N., and D.M. conceived the research study. K.K. and N.K. carried out all the analytical treatments. I.S.-N. wrote the codes and conducted the simulations and the analysis of the data. A.M.R. proved Theorem 2. K.K., I.S.-N., N.K., A.D., D.M., A.M.R., K.A.-B., B.B., and S.B. discussed the results, drew the conclusions, and jointly wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kovalenko, K., Sendiña-Nadal, I., Khalil, N. et al. Growing scale-free simplices. Commun Phys 4, 43 (2021). https://doi.org/10.1038/s42005-021-00538-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-021-00538-y
This article is cited by
-
Growing hypergraphs with preferential linking
Journal of the Korean Physical Society (2023)
-
The physics of higher-order interactions in complex systems
Nature Physics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.