Abstract
Functional materials, especially those that largely differ from known materials, are not easily discoverable because both human experts and supervised machine learning need prior knowledge and datasets. An autonomous system can evaluate various properties a priori, and thereby explore unknown extrapolation spaces in high-throughput simulations. However, high-throughput evaluations of molecular dynamics simulations are unrealistically demanding. Here, we show an autonomous search system for organic molecules implemented by a reinforcement learning algorithm, and apply it to molecular dynamics simulations of viscosity. The evaluation is dramatically accelerated (by three orders of magnitude) using a femto-second stress-tensor correlation, which underlies the glass-transition model. We experimentally examine one of 55,000 lubricant oil molecules found by the system. This study indicates that merging simulations and physical models can open a path for simulation-driven approaches to materials informatics.
Similar content being viewed by others
Introduction
The development of materials conventionally depends on human sense and trial-and-error synthesis. Such laborious developments are expected to be accelerated by materials informatics (MI)1,2, which is commonly implemented by virtual screening (see Fig. 1a). After training on existing data, a machine-learning model predicts the target properties of materials based on the features of known materials3,4,5,6,7,8,9. Rapid inference by machine learning extracts the potential candidates from hundreds of thousands of compounds in a material database. This subset of the candidates is then examined experimentally. However, the prediction ability is effective only when the target materials are within an interpolation space coordinated by a supervised dataset. To discover truly new materials, we should explore outside the scope of known materials.
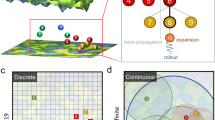
An autonomous search scheme beyond the interpolation space is called a closed-loop search1. The system configuration is illustrated in Fig. 1b. Here, a machine-learning search model accompanies robotics or simulation software. The search model receives feedback from the evaluated properties, and decides the material proposals in the next loop. This search-evaluation loop iterates until the material structure is optimized with respect to a target property. Search algorithms for this purpose are numerous and varied10,11,12,13,14. An example is the artificial neural network in the chemical language SMILES, which generates a continuous latent space of molecules, and seeks the high-scoring molecules by a gradient-based optimization procedure10,11. Elsewhere, prospective molecular structures were generated by a Bayesian approach using forward and backward predictions in the structure–property relationship12. To design synthetic strategies and uncover new organic materials, Yang et al. and Segler et al. used a reinforcement learning algorithm called Monte Carlo tree search (MCTS)13,14,15,16. This algorithm was used in the AlphaGo AI system for the Chinese board game “Go”17. The MCTS algorithm efficiently searches a tree graph whose nodes represent molecular fragments in SMILES. Its aim is to maximize the prospective reward of molecules13,14.
However, no matter what search algorithms are used, a long evaluation time is a major bottleneck in the loop. Ab initio calculations provide important material properties such as formation energies and band gaps. These static properties can be obtained at reasonable computation cost only by advanced algorithms and multicore architectures18,19,20,21. Transport-related properties, such as ion conductivity and viscosity, must be assessed in molecular dynamics (MD) calculations, which simulate the atomic dynamics of molecules. Although the evaluated transport properties are based on statistical physics, MD calculations cannot be a high-throughput evaluator22, because reliable ensemble averaging requires a huge number of MD steps23,24. Another important consideration is accuracy of the empirical force fields. This topic has been actively studied in recent years, with developments of machine-learning potentials trained on appropriate ab initio reference data25,26,27,28,29.
This paper presents an autonomous molecular-design system based on MCTS and MD simulations. As an example of transport properties, we focus on viscosity because viscosity is related to tribological properties30,31 and its reciprocal value represents a diffusion coefficient. These properties are fundamental in mechanical and chemical engineering, which use oil and electrolytes on a daily basis. Our system performs ultra-fast MD evaluations that alleviate the time-demanding bottleneck of autonomous systems.
We first explain the conventional and proposed fast viscosity evaluations by MD simulations, define the target property, and explain the rules of oil-molecule generation in MCTS. After the closed-loop search, the MI-designed oil molecule is synthesized and its viscosity performance is experimentally examined. Finally, we inductively analyze the obtained large data to guide the development of lubricants. The technical details are provided in the Methods section and Supplementary Notes.
Results
Conventional MD evaluation
One conventional schemes for obtaining transport properties is the Green–Kubo (GK) formalism32,33. Non-diagonal elements of a stress tensor Pij is observed in a MD simulation of liquid molecules. The viscosity η is obtained by dynamical fluctuations of Pij as
where kB, T, and V denote Boltzman’s constant, temperature, and volume of the simulation cell, respectively. The operator 〈〉 represents ensemble averaging in the MD calculation (see Fig. 2a), which samples the correlation Φ(t, t0) with respect to the time origin t0.
a Schematic of molecular dynamics (MD) sampling to obtain the correlation function Φ in Eq. (1). Pij, kB, and T denote the non-diagonal elements of a stress tensor, Boltzman’s constant, and temperature, respectively. The operator 〈〉 represents averaging with respect to the time origin t0. b Correlation functions of an oil molecule (molecule 13nddh shown in the Methods section) at 40 °C, and c the same correlations in the short-time range. The color bar in b represents the density of the Φ(t, t0) entries in the t0 samplings, obtained by kernel density approximation implemented in scikit-learn. The short-time correlation \(\overline {\mathrm{\Phi }}\) is central to the present fast evaluation method (Eq. (4)). The red lines are the averaged values over the samplings.
The bottleneck in the conventional MD-based evaluation is easily recognized from Φ(t, t0). Figure 2b shows the density of the sampled Φ(t, t0) entries in MD simulations of an oil molecule. After a long t, the variations among the samplings of the correlation are enlarged, meaning that the long-future state is loosely associated with its present state. Figure 2c shows the vice versa situation, in which the correlations at short times shows smaller variations. As evidenced in Eq. (1), viscosity is a long-time correlation, requiring a huge number of MD steps to obtain sufficiently many t0 samplings for accurate ensemble averaging. Based on this insight, we suggest that if the viscosity can be predicted through the short-time correlation, the number of sampling MD steps can be reduced in the viscosity evaluation. Such a strategy is sought in this paper.
Fast evaluation
To realize the above idea, we import an elastic concept of liquid viscosity called the shoving model34,35,36.
This model describes liquid from an atomic viewpoint as shown in Fig. 3. In the liquid state, a component molecule is surrounded by other liquid molecules in a caged space. Driven by thermal fluctuations, each molecule repeatedly collides with its neighbors. After a certain relaxation time, a molecule escapes from the cage by pushing its neighbors away. Through iterations of this local relaxation, all molecules are eventually rearranged and the liquid flows macroscopically. This phenomenological viewpoint suggests that the structural relaxation related to viscosity can be well represented by the energy required to push the surrounding molecules. The energy barrier is then proportional to the shear modulus of the liquid.
Combined with transition-state theory37, the shoving model provides an Arrhenius-type equation of viscosity as
where α, β, and γ are empirical parameters. Equation (2) demonstrates that viscosity is correlated with the stiffness of the liquid, which is measured under a given instantaneous force. Puosi and Leporini35 and Dyre and Wang36 improved the accuracy of viscosity calculations by a revised formula for the shear modulus \(G_\infty ^ \ast \propto {\mathrm{\Phi }}\left( {\delta t} \right)\), where δt is a short-time period of the order of molecular vibrations. In this study, we use an averaged value of Φ as follows:
and δt is set to 5.0 fs.
The shoving model was originally developed to clarify the atomic mechanism of glass transition. Here, we employ it to accelerate the MD evaluation of viscosity, as described below. Note that as Eq. (2) uses the short-time correlation, we can estimate the viscosity by \(\overline {\mathrm{\Phi }}\) instead of the conventional evaluation in Eq. (1).
To improve the accuracy of our evaluation, we modify the original Arrhenius equation in Eq. (2). Van Velzen’s model is a well-known modification of the Arrhenius form. Commonly used in lubrication engineering, this model corrects the viscosity–temperature relation with respect to the boiling point of the liquid38,39. Combining the van Velzen model with Eqs. (2) and (3), we obtain
where the boiling point Tb of the liquid is immediately estimated from a SMILES string via the Joback method40 implemented in the python library thermo. Fitting Eq. (4) to the experimental viscosities of reference organic molecules (see Methods section), the parameters A, B, and ηb were determined as 7.577 × 103, 1.607 × 107, and 0.217 cP, respectively. Interestingly, the viscosity at the boiling temperature ηb is known to be constant value 0.22 cP for typical organic molecules that contain larger than 20 carbons41. This value is consistent with the fitted value. Note that the accuracy of the proposed approach may degrade in small-molecule cases.
Target property: viscosity index
As a target property for optimization, viscosity alone is unsuitably trivial. Viscosity typically increases with number of constituent atoms of a lubricant molecule, because longer molecules become more entangled in the liquid state than short molecules39. Instead, we target the viscosity index (VI), which indicates the temperature sensitivity of viscosity42. Machinery equipment requires high-VI oil for stable mechanical operations in various environments. We use the most famous VI definition, namely the quantity VIASTM given in the American Society for Testing and Materials (ASTM) D 2270 standard42,43. The VIASTM is calculated as
where \(\eta _k^T\) is the kinematic viscosity at temperature T. In this definition, it is obtained from the kinematic viscosities L and H with VIASTM = 0 and 100, respectively, at 40 °C, and having the same kinematic viscosity as the oil of interest at 100 °C. The reference viscosities can be obtained from a viscosity conversion table42,44. We used the python library thermo to calculate VIASTM.
As a complementary measure of VI performance, we also computed the dynamic viscosity index (DVI)42,45, because the VIASTM is unsuitable for low-viscosity oils44. For example, if \(\eta _k^{40^ \circ {\mathrm{C}}}\) ≤ 2.0 mm2/s, VIASTM is undefined. Moreover, the VIASTM underestimates the viscosity susceptivity of low-viscosity oils in the range of \(\eta _k^{40^ \circ {\mathrm{C}}}\) ≤ 5.0 mm2/s44. To resolve these problems, the DVI was proposed as
where η denotes the viscosity. The kinematic viscosity and viscosity are related through ηk = η/ρ, where ρ is the density of the liquid.
An important difference between VIASTM and DVI is that the former observes the ηk variation, whereas the latter observes the η variation. Tribological properties such as oil film thickness and viscosity resistance at the sliding interface depend more on viscosity than the kinematic viscosity. Therefore, although the VIASTM is conventionally used, the DVI is also a good index of the temperature–viscosity sensitivity. These two indices are compared in the Supplementary Note 1.
Molecular fragments and rules of the Monte Carlo tree search
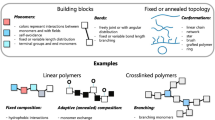
The remaining component of the autonomous design system is a search algorithm that generates molecular structures with the optimal target properties. The search algorithm should comprise both an efficient search strategy in regarding to inherent molecular representations and generation rules to meet material requirements. This study employs the MCTS as the search algorithm, which describes a molecule by a graph structure. The graph nodes describe the user-defined molecular fragments in SMILES13,14. Oil molecules synthesized and purified from crude oil generally have hydrocarbon chain structures with several branches. To represent such structures, we defined different types of molecular fragments for the main and side chains of the molecules as follows:
-
In the main chain: CC, OC, C=C, (, $, c1ccccc1$, C1CCCCC1$, =O$
-
In the side chain: CC, OC, C=C, (,), c1ccccc1), C1CCCCC1), =O)
where $ indicates the end of the molecule. These side-chain fragments can be joined only after a “(” symbol in the main chain. The c1ccccc1, C1CCCCC1, and =O fragments are terminal groups. The initial molecular fragment, called a root node, is C.
We then restricted the generated molecules to lubricants. Unbranched molecules are inappropriate because they have high freezing points, so are prone to waxing at the operating temperature. To generate molecules with one or more branches, we rejected the no-branch molecules during the rollout operation of MCTS. The branched molecules were then restricted to the allowable viscosity range. An excessively high viscosity increases the fuel consumption, whereas a very low viscosity leads to scuffing. The preferred kinematic viscosity of the base oil of automobile lubricants ranges from 3.0 to 6.0 mm2/s. As viscosity is proportional to the number of constituent atoms39, a typical oil molecule should contain 20–40 carbons46. To accord with the MCTS rules, we set an ending rule by which fragments with $ can be used only when the total number of C and O is 20 or higher. When this number is 30 or higher, fragments with $ are used mandatorily.
In summary, we define three search rules: define the molecular fragments, prohibit the unbranched molecules, and impose the ending condition. The hyperparameters of the MCTS algorithm are given in the Methods section.
Evaluations of viscosity and viscosity index
The closed-loop feasibility is mainly determined by the acceleration extent of the MD evaluations. As a baseline method, we employed the conventional Einstein–Helfand (EH) scheme33, which evaluates the viscosity by the mean-squared displacement of Pxy. We emphasize that this baseline was selected for a convenient comparison, because the EH scheme is defined to avoid erroneous negative viscosity, unlike the GK scheme. The two schemes are compared in Supplementary Note 2.
Figure 4a compares the viscosities evaluated by the fast evaluation and EH methods with an identical dataset of MD trajectories. The computational details are provided in the Methods section. Under the same sampling conditions, the root-mean-squared error (RMSE) was 3.8 cP in the proposed method, greatly reduced from 19.8 cP in the EH method. A distinctive advantage can be found in the standard deviation (STD) of each MD trajectory. In the present method, the STD is only 3.7% those of the EH method, so small that the error bars are hidden behind the points in Fig. 4a. We roughly estimated that to attain the same statistical accuracy as the EH method, the fast evaluation reduced the number of samplings in the MD steps to approximately (3.7/100)2 ∼ 1/1000. The fast evaluation is examined in detail in Supplementary Note 3.
a Comparison of the proposed fast method (left) and conventional molecular dynamics (MD) in the Einstein–Helfand (EH) scheme (right). b Plots of calculated versus experimental viscosity indices in the American Society for Testing and Materials (ASTM) D 2270 standard (VIASTM). The red circles are averaged over the MD trajectories. The reference organic molecules and MD conditions are described in the Methods section. The RMSE and STD denote the root-mean-squared error and standard deviation, respectively.
Figure 4b compares the VIASTM values of the EH and proposed methods. Because the VIASTM is very sensitive to slight deviations in kinematic viscosity, the errors in the EH method were unacceptably large for the closed-loop system. In contrast, the VIASTM values obtained by the proposed method were sufficiently accurate and efficiently obtained.
Autonomous search
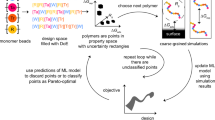
Figure 5a shows the protocol of closed-loop searching. The MCTS proposes the next molecule encoded in SMILES, and then the fast evaluation by MD simulations provides its VIASTM as feedback. The search was performed ten times with 5500 evaluation loops per search, giving 54,318 evaluated molecules. Figure 5b shows VIASTM and kinematic viscosity histograms of the molecules. Most of the viscosities ranged from 3.0 to 6.0 mm2/s as planned, and several high-VIASTM molecules were observed. As indicated by the top-ten molecules in Fig. 5c, the generated structures were very particular, unlikely to be synthesized by one or two chemical processing steps. Therefore, we investigated the candidate list for higher VIASTM molecules admitting an easy synthesis. For the easy synthesis requirement, we sought suggestions from organic chemists in our institute. Consequently, we took the 83rd-ranked molecule shown in Fig. 5d as a motif, and modified it to an easily synthesized form in Fig. 5e. The modified molecule was prepared by the etherification of farnesyl bromide with 1,5-diphenylpentan-3-ol, which is obtained by the Grignard reaction of 3-phyenylpropanal and 2-phenylethlmagnesium bromide47. As comparison molecules, we used two major high-VI base oils refined from crude oil by hydrocracking and chemical synthetic: YUBASE-4 and SpectraSyn-4 made by SK lubricants and Exxon Mobil, respectively. The viscosities of these oils were experimentally determined by a Stabinger viscometer SVMTM in Anton Paar Ltd.
a Schematic of the autonomous search system for oil-molecule design. The Monte Carlo tree search (MCTS) proposes a candidate molecule encoded in SMILES strings, for which the atomic configurations and force field are automatically generated in softwares Open Babel and Direct Force Field, respectively. The generated input files are transmitted to the ”K” super computer. The molecular dynamics (MD) simulation computes the correlation function \(\overline {\mathrm{\Phi }}\) of the fast evaluation in Eq. (4). The calculated viscosity index in the American Society for Testing and Materials (ASTM) D 2270 standard (VIASTM) updates the MCTS policy to improve the next set of candidate molecules. b Kinematic viscosity at 100 °C and VIASTM histograms of the 54,318 molecules. c The top-ten molecules. d The selected motif molecule and e its modified version that can be easily synthesized. f Molecular structure of poly-alpha oleffine (the major component of high viscosity-index base oils).
Table 1 summarizes the properties obtained in the investigation. The calculated DVIs, kinematic viscosities, viscosities, and densities deviated within 20% of the experimental values. The calculated VIASTM was overestimated because it largely responds to even slight changes in kinematic viscosity (see Supplementary Note 1). The experimental VIASTM of the present molecule was 109, smaller than those of the high-VI commercial oils, but still classifiable between the high-VI group (VIASTM = 80–110) and the very high-VI group (VIASTM > 110) according to Neale 48. In fact, when measured by another DVI metric, the obtained oil was slightly superior to the market oils.
Typically, the main components of high-VI oils are high-ration paraffin structures. For instance, poly-alpha oleffine shown in Fig. 5f is a major component of SpectraSyn. Interestingly, our molecule in Fig. 5e is quite unlike the conventional high-VI molecules. This result indicates that it extends the interpolated lubricant space. Nevertheless, engine oils in applications must not only satisfy the viscosity-index requirements but must also deliver high oxidative resistance and low freezing point at minimal production cost. These additional requirements are not considered in the present test search.
Discussion
As is often mentioned, material data are not big data, and the existing datasets of transport properties are limited. Nevertheless, experts try to deduce a design guideline from such a scarce dataset to develop better materials. For example, after observing synthesized molecules by properly controlled hydrocracking and 13C nuclear magnetic resonance (NMR), researchers deduced that high-VI molecules likely consist of long chains with few branches and rings46,49,50,51. Owing to the time-intensiveness of the experiments, the hydrocracking and NMR data constituted only several tens of entries. To our knowledge, the present dataset of 55,000 entries is the largest acquired dataset of viscosity properties. In a simple data analysis, we now extract the features from this dataset that are relevant to high-VI molecules, and compare our insights with those reported by the experts.
Figure 6a and b show the correlation heat map and the main structure–property correlations (with values exceeding 0.4), respectively. For the correlation analysis, we selected the VIASTM, kinematic viscosity ηk, density ρ, number of constituent atoms N, number of branches Nbranch, and the ring ratio Rring. The positive correlation between the kinematic viscosity and N is well known39. The VIASTM was strongly correlated with both ηk and N. To capture molecules with viscosities within the typical range of low-viscosity engine oils, we then restricted the dataset to 4.0 mm2/s ≤ \(\eta _k^{100^ \circ {\mathrm{C}}}\) ≤ 5.0 mm2/s. In Fig. 6c, the edge between VIASTM and ηk disappears because its correlation was below the threshold magnitude 0.4, but the positive correlation between N and VIASTM remained under the viscosity restriction. According to this result, VIASTM is an increasing function of N. However, as N is also positively correlated with the viscosity, it cannot be increased indefinitely, but is restricted by the upper limit of the valid viscosity range. Therefore, when increasing N, the viscosity must be simultaneously suppressed. To favor a high-VIASTM, we minimized the viscosity of molecules with constant N. Figure 6d shows the major correlations in the dataset of molecules with N = 31. The kinematic viscosities of the restricted molecules were mainly distributed over 4.0–5.0 mm2/s. The nodes Rring and Nbranch were positively correlated with the node ηk, implying that straight-chain fragments are preferable for reducing the viscosity increment.
Viscosity index in American Society for Testing and Materials (ASTM) D 2270 standard VIASTM, kinematic viscosity ηk, density ρ, number of atoms N, number of branches Nbranch, and ring ratio Rring were involved in this analysis. a Correlation heat map. b–d are graph representations that contain the edges of correlations in no restriction, c 4.0 mm2/s ≤ ηk ≤ 5.0 mm2/s, and d N = 31, respectively. The edges are presented when their correlation magnitudes are larger than or equal to 0.4. The kinematic viscosity and density are observed at 100 °C. The ring ratio refers to the number of carbon atoms in the ring bases divided by number of all elements except the hydrogens in a molecule (e.g., a SMILES ccccccC1CCCCC1 indicates Rring = 0.5).
Meanwhile, a high VI was observed for molecules with many constituent atoms, few branches, and few rings. This result is consistent with the previously reported experimental insights46,49,50,51. Note that although Nbranch and Rring negatively influenced the VIASTM, they could not describe the VI well, because they were poorly correlated with VI. The VI might be better represented by other features such as molecular configuration, dynamical entanglement, and dipole–dipole interactions. Other critical parameters of VI might be identified by mining the present dataset of 55,000 molecules; for this purpose, the dataset (see Supplementary Data 1) has been made publicly available.
In conclusion, our autonomous search confers two main advantages: (1) efficient design of a high-functioning molecule by referring to a prospective molecule selected from generated candidate molecules, and (2) acquisition of design insights and directions from the generated dataset. A major weakness of this system is the difficulty of evaluating the ease of synthesis, which has been intensively studied elsewhere14. Nevertheless, as a potentially new scheme of materials development, our MI system comprehensively explores the vast material space in high-speed evaluations. Experts can then modify the extracted prospective materials considering the required stability, safety, and production cost of the target product. Current AI systems for the “Go” game have continuously inspired professional players since demonstrating their ability to defeat the players52. This trend may also propagate into materials science, driving further technological developments through human–MI collaborations. Fast evaluation by MD simulations should be generalized to transport properties other than viscosity, such as ion conductivity. Such investigations will be undertaken in our future work.
Methods
Molecular dynamics simulation
The simulations were performed in the open-source MD solver LAMMPS with the force field TEAM_MS which is provided in the commercial software Direct Force Field (DFF). The TEAM_MS force field was constructed based on the results of ab-initio calculations of molecular fragments53. To achieve a thermal equilibrium state, we first ran an NVT calculation with time interval Δt = 0.25 fs followed by an NPT calculation with Δt = 1.0 fs. We then executed a relatively long NVT calculation with Δt = 1.0 fs to sample the non-diagonal elements of the stress tensor Pij. Table 2 summarizes the conditions of the MD simulations.
Figure 2b, c shows the distributions of Φ(t, t0) entries, calculated in MD simulations under the ”Normal” condition in Table 2. To obtain the distributions, we divided the t0 samplings into 100 domains, modifying Eq. (1) as
We employed the averaged sampling quantity as Pij ≡ (Pxy + Pyz + Pzx)/3. The MD simulations were repeated five times to increase the number of the MD samplings; therefore, Fig. 2b, c was constructed from 5 × 100 \({\mathrm{\Phi }}^\prime \left( {t,n_1} \right)\) trajectories.
Figure 4, which compares the results of the fast evaluation and conventional methods, was constructed from the same five MD trajectories under the “Normal” condition. In this case, we individually set Pxy, Pyz, and Pzx as Pij and ran the MD simulation five times, thus obtaining 5 × 3 = 15 viscosity samples for each molecule.
The traceless-symmetric part of the stress tensor Pos is known to yield good statistics. The quantity Pos consists of five independent samples Pxy, Pyz, Pzx, (Pxx − Pyy)/2, and (Pyy − Pzz)/2 collected into one MD trajectory23,24. We used Pos as the sampling quantity in the high-throughput calculations of Fig. 5. The number of molecules in the simulation cell was 120. To reduce the computational cost of the 55,000 evaluations, we decreased the cutoff length of the coulomb interaction and number of time steps (“High-throughput” row in Table 2). We confirmed that the high-throughput condition ensures acceptable accuracy for determining the order of VIASTM’s of different molecules, as shown in the Supplementary Note 4. The data in Table 1 were accurately calculated by sampling the traceless-symmetric quantity under the “Normal” condition.
Monte Carlo tree search
The reward in MCTS is defined by the upper confidence bound (UCB) score as
where n and nparent indicate the numbers of visits at a node and its parent node, respectively15,16. The quantity \(\overline {{\mathrm{VI}}} _{\mathrm{{ASTM}}}\) is obtained by averaging the VIASTMs of molecules that were randomly generated from the node called random rollout. The rollout number, which refers to the number of randomly generated molecules, was set to 10.
Because VIASTM cannot be defined when \(\eta _k^{40^ \circ {\mathrm{C}}}\) ≤ 2.0 mm2/s, we set VIASTM = 0 in such cases. If the structure of the molecule generated in the rollout phase was chemically invalid, it was automatically detected by the RDKit software and replaced with a new molecule. The bias coefficient C is an arbitrary parameter. We set C = 1, which is theoretically validated when the first term of the right-hand side of Eq. (7) ranges from 0.0 to 1.0 (refs. 15,16). We then divided \(\overline {{\mathrm{VI}}} _{\mathrm{{ASTM}}}\) by its approximately expected maximum, namely, 200.
Reference molecules
As the reference models in the MD test, we adopted typical 12e organic molecules. Their structures and abbreviated names are displayed in Fig. 7. Their formal names and viscosity properties are listed in Tables 3 and 4, respectively. In the MD calculations, the numbers of molecules in the simulation cell were 150 for 9nhhd, 9chhd, diiso_seb, and 2m4odp, 120 for 1c2mh and 13cp, and 100 for the remainder. Approximately 10,000 atoms existed in each simulation cell.
Data availability
The dataset generated during the high-throughput evaluations (54,318 SMILES of the molecules along with the VI's, viscosities, kinematic viscosities, and densities) is available in the Supplementary Data 1. The authors declare that all other data supporting the findings of this study are available within the paper and its Supplementary Notes.
References
Tabor, D. P. et al. Accelerating the discovery of materials for clean energy in the era of smart automation. Nat. Rev. Mater. 3, 5–20 (2018).
Luna, P. D. et al. Use machine learning to find energy materials. Nature 552, 23–27 (2017).
Buttler, K. T., Davies, D. W., Cartwright, H., Isayev, O. & Walsh, A. Machine learning for molecular and materials science. Nature 559, 547–555 (2018).
Sendek, A. D. et al. Machine learning-assisted discovery of solid Li-ion conducting materials. Chem. Mater. 31, 342–352 (2018).
Gomez-Bombarelli, R. et al. Design of efficient molecular organic light-emitting diodes by a high-throughput virtual screening and experimental approach. Nat. Mater. 15, 1120–1127 (2016).
Narayan, A. et al. Computational and experimental investigation for new transition metal selenides and sulfides: the importance of experimental verification for stability. Phys. Rev. B 94, 045105 (2016).
Lee, J., Ohba, N. & Asahi, R. Discovery of zirconium dioxides for the design of better oxygen-ion conductors using efficient algorithms beyond data mining. RSC Adv. 8, 25534–25545 (2018).
Ohba, N., Yokoya, T., Kajita, S. & Takechi, K. Search for high-capacity oxygen storage materials by materials informatics. RSC Adv. 9, 41811–41816 (2019).
Kajita, S., Ohba, N., Suzumura, A., Tajima, S., & Asahi, R. Discovery of superionic conductors by ensemble-scope descriptor. NPG Asia Mater. 12, 31 (2020).
Gomez-Bombarelli, R. et al. Automatic chemical design using a data-driven continuous representation of molecules. ACS Cent. Sci. 4, 268–276 (2018).
Segler, M. H., Kogej, T., Tyrchan, C. & Waller, M. P. Generating focussed molecule libraries for drug discovery with recurrent neural networks. ACS Cent. Sci. 4, 120–131 (2017).
Ikebata, H., Hongo, K., Isomura, T., Maezono, R. & Yoshida, R. Bayesian molecular design with a chemical language model. J. Comput. Aided Mol. Des. 31, 379–391 (2017).
Yang, X., Zhang, J., Yoshizoe, K., Terayama, K. & Tsuda, K. ChemTS: an efficient python library for de novo molecular generation. Sci. Technol. Adv. Mater. 18, 972–976 (2017).
Segler, M. H., Preuss, M. & Waller, M. P. Planning chemical syntheses with deep neural networks and symbolic AI. Nature 555, 604–610 (2018).
Agrawal, R. Sample mean based index policies by o (log n) regret for the multi-armed bandit problem. Adv. Appl. Probab. 27, 1054–1078 (1995).
Auer, P., Cesa-Bianchi, N. & Fischer, P. Finite-time analysis of the multiarmed bandit problem. Mach. Learn. 47, 235–256 (2002).
Silver, D. et al. Mastering the game of Go with deep neural networks and tree search. Nature 529, 484 (2016).
Hautier, G. et al. Phosphates as lithium-ion battery cathodes: an evaluation based on high-throughput ab initio calculations. Chem. Mater. 23, 3495–3508 (2011).
Studt, F. et al. CO hydrogenation to methanol on Cu-Ni catalysts: theory and experiment. J. Catal. 293, 51–61 (2012).
Nishijima, M. et al. Accelerated discovery of cathode materials with prolonged cycle life for lithium-ion battery. Nat. Commun. 5, 4553 (2014).
Hayashi, H. et al. Discovery of a novel Sn (II)-based oxide β-SnMoO4 for daylight-driven photocatalysis. Adv. Sci. 4, 1600246 (2017).
Curtarolo, S. et al. The high-throughput highway to computational materials design. Nat. Mater. 12, 191201 (2013).
Meyer, E. R., Kress, J. D., Collins, L. A. & Ticknor, C. Effect of correlation on viscosity and diffusion in molecular-dynamics simulations. Phys. Rev. E 90, 043101 (2014).
Davis, P. J. & Evans, D. J. Comparison of constant pressure and constant volume nonequilibrium simulations of sheared model decane. J. Chem. Phys. 100, 541–547 (1994).
Jinnouchi, R., Lahnsteiner, J., Karsai, F., Kresse, G. & Bokdam, M. Phase transitions of hybrid perovskites simulated by machine-learning force fields trained on the fly with Bayesian inference. Phys. Rev. Lett. 122, 225701 (2019).
Schütt, K. T., Sauceda, H. E., Kindermans, P. J., Tkatchenko, A. & Muller, K. R. SchNet-A deep learning architecture for molecules and materials. J. Chem. Phys. 148, 241722 (2018).
Chmiela, S., Sauceda, H. E., Müller, K. & Tkatchenko, A. Towards exact molecular dynamics simulations with machine-learned force fields. Nat. Commun. 9, 1–10 (2018).
Unke, T. O. & Meuwly, M. PhysNet: a neural network for predicting energies, forces, dipole moments, and partial charges. J. Chem. Theory Comput. 15, 3678–3693 (2019).
Singraber, A., Behler, J. & Dellago, C. Library-based LAMMPS implementation of high-dimensional neural network potentials. J. Chem. Theory Comput. 15, 1827–1840 (2019).
Kai, H. & Szlufarska, I. Green-Kubo relation for friction at liquid-solid interfaces. Phys. l Rev. E 89, 032119 (2014).
Washizu, H. & Ohmori, T. Molecular dynamics simulations of elastohydrodynamic lubrication oil film. Lubrication Sci. 22, 323–340 (2010).
Mondello, M. & Grest, G. S. Viscosity calculations of n-alkanes by equilibrium molecular dynamics. J. Chem. Phys. 106, 9327–9336 (1997).
Helfand, E. Transport coefficients from dissipation in a canonical ensemble. Phys. Rev. E 119, 1 (1960).
Dyre, J. C. Colloquium: The glass transition and elastic models of glass-forming liquids. Rev. Mod. Phys. 78, 953 (2006).
Puosi, F. & Leporini, D. Communication: correlation of the instantaneous and the intermediate-time elasticity with the structural relaxation in glassforming systems. J. Chem. Phys. 136, 041104 (2012).
Dyre, J. C. & Wang, W. H. The instantaneous shear modulus in the shoving model. J. Chem. Phys. 136, 224108 (2012).
Glasstone, S., Laidler, K. J. & Eyring, H. Theory of Rate Process (McGraw-Hill, New York, 1941).
Van Velzen, D., Cardozo, R. L. & Langenkamp, H. A liquid viscosity-temperature-chemical constitution relation for organic compounds. Ind. Eng. Chem. Fundam. 11, 20–25 (1972).
Viswanath, D. S., Ghosh, T. K., Prasad, D. H. L., Dutt, N. V. K. & Rani, K. Y. Viscosity of Liquids: Theory, Estimation, Experiment, and Data (Springer, Netherlands, 2007).
oback, K. G. & Reid, R. C. Estimation of pure-component properties from group-contributions. Chem. Eng. Commun. 57, 233–243 (1987).
Smith, G. J., Wilding, W. V., Oscarson, J. L., & Rowley, R. L. Correlation of liquid viscosity at the normal boiling point. Proceedings of the Fifteenth Symposium on Thermophysical Properties, Boulder, Colorado, U.S.A.
Zakarian, J. The limitations of the viscosity index and proposals for other methods to rate viscosity-temperature behavior of lubricating oils. SAE Int. J. Fuels Lubr. 5, 1123–1131 (2012).
ASTM D2270-10: Standard practice for calculating viscosity index from kinematic viscosity at 40 and 100 °C. http://ppapco.ir/wp-content/uploads/2019/07/ASTM-D2270-2016.pdf (2016).
Covitch, M. J. An improved method for calculating viscosity index (VI) of low viscosity base oils. J. Test. Eval. 46, 820–825 (2018).
Roelands, C. J. A., Blok, H., Vlugter, J. C., & Eng, M. A new viscosity-temperature criterion for lubricating oils. ASME-ASLE International Lubrication Conference, No. 64-LUB-3, Washington, D.C. (1964).
Lynch, T. R. Process Chemistry of Lubricant Base Stocks (CRC Press, Boca Raton, 2007).
Zhang, Q. C. et al. Modulating the rotation of a molecular rotor through hydrogen-bonding interactions between the rotator and stator. Angew. Chem. Int. Ed. 52, 12602–12605 (2013).
Neale, M. J. Table 2.1 in Lubrication and Reliability Handbook (Newnes, Elsevier, 2001).
Kapur, G. S., Chopra, A., Sarpal, A. S., Ramakumar, S. S. V. & Jain, S. K. Studies on competitive interactions and blending order of engine oil additives by variable temperature 31P-NMR and IR spectroscopy. Tribol. Trans. 42, 807–812 (1999).
Verdier, S., Coutinho, J. A., Silva, A. M., Alkilde, O. F. & Hansen, J. A. A critical approach to viscosity index. Fuel 88, 2199–2206 (2009).
Noh, K., Shin, J. & Lee, J. H. Change of hydrocarbon structure type in lube hydroprocessing and correlation model for viscosity index. Ind. Eng. Chem. Res. 56, 8016–8028 (2017).
Lee, C. S. et al. Human vs. computer go: review and prospect. IEEE Comput. Intell. Mag. 11, 67–72 (2016).
Sun, H. & J. COMPASS: an ab initio force-field optimized for condensed-phase applications—overview with details on alkane and benzene compounds. Phys. Chem. B 102, 7338 (1998).
Acknowledgements
S.K. thanks M. Tohyama and T. Ohmori for their useful advices regarding the specification and evaluation of lubricants. S.K. thanks H. Takeuchi and Y. Kikuzawa for assisting with the synthesis of the present oil molecules. This research used the computational resources of the K computer provided by RIKEN through the HPCI System Research project (Project ID:hp180238).
Author information
Authors and Affiliations
Contributions
S.K. developed the ultra-fast evaluation and MCTS, and operated the closed-loop search and experiment. T.K. developed the GK and EH methods, and selected the proper MD conditions. T.N. provided an idea and technical advices related to MCTS. All authors collectively wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kajita, S., Kinjo, T. & Nishi, T. Autonomous molecular design by Monte-Carlo tree search and rapid evaluations using molecular dynamics simulations. Commun Phys 3, 77 (2020). https://doi.org/10.1038/s42005-020-0338-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-020-0338-y
This article is cited by
-
Interpretability of rectangle packing solutions with Monte Carlo tree search
Journal of Heuristics (2024)
-
Human-in-the-loop assisted de novo molecular design
Journal of Cheminformatics (2022)
-
Fast evaluation technique for the shear viscosity and ionic conductivity of electrolyte solutions
Scientific Reports (2022)
-
Multi-objective goal-directed optimization of de novo stable organic radicals for aqueous redox flow batteries
Nature Machine Intelligence (2022)
-
A review of advances in tribology in 2020–2021
Friction (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.