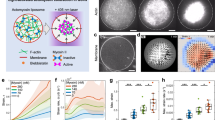
Abstract
Giant liposomes encapsulating cytoskeletons have been constructed to further understand the mechanisms of cell movement and develop cell-sized chemical machineries. Innovative studies demonstrating liposomal movements using microtubules and the molecular motors kinesin/dynein have been reported. However, no one has succeeded in generating repetitive motions controlled by external stimuli. Here we show that if the actin concentration in liposomes is comparable to that of cytoplasm of living cells, the liposomes can be deformed into spindle shapes by encapsulating only actin filaments, even without the molecular motor myosin. Furthermore, their shapes can be changed reversibly between spindle and sphere shapes by adjusting osmotic pressure or by light irradiation of fluorescent-labeled actin. In the latter case, the repetitive shape changes are accompanied with stretching and shrinking of filopodia- or acrosome projection-like extensions. Our results indicate that filamentous polymer of variable length like actin filament is a potential material for the reproduction of cell-like movement.
Similar content being viewed by others
Introduction
Molecular robotics, the development of artificial cell-like chemical machineries, has drawn much recent attention1,2,3. One such approach is to construct cell-sized giant liposomes which encapsulate artificial or native macromolecular polymers and can produce motions similar to living cells. As representative native macromolecular polymers, cytoskeletal proteins and molecular motors are frequently used. Recently, active motions of cell-sized vesicles or droplets utilizing the combination of microtubules (MTs) and kinesin as their engine have been reported4,5,6,7. Some molecular robots successfully change their shape using mechanical processes of biomolecules, including elongation/retraction of MTs controlled by hydrostatic pressure using a high-pressure microscope that was developed to acquire microscopic images while applying pressure up to 150 MPa on the sample; and sliding motions between MTs and kinesin motors switched ON/OFF controlled by a DNA-clutch system8,9,10. However, those systems require specialized equipment and/or complex components of biomolecules, which are not routinely available.
Similar to MTs, actin filaments (F-actins) are also dynamically arranged in a self-organized manner in living cells to perform various biological functions, including morphogenesis, cell motility and division, as well as the spatial arrangement and migration of organelles. It is generally accepted that complexes consisting of F-actin are involved in the constitution of backbone structures and/or skeletons inside cells through the systematic localization of networks as well as the manner of their polymerization/depolymerization11.
Living cells maintain their viability by maintaining a highly crowded cellular cytoplasm with 30–40 wt% of cellular materials12, 13. Recently, it was suggested that the high concentration and crowded environment of the cytoplasm plays an important role in the construction of cell morphology via the regulation of cytoskeletons, especially F-actin networks14, 15. Interestingly, it has become more clear that these cytoskeletons lack specific regulatory factors, suggesting a plausible scenario whereby cell morphology is generated in a self-organizing manner depending on the highly concentrated and crowded conditions in the cytoplasm.
Here, we demonstrate that liposomes encapsulating F-actin at high concentrations comparable to the cytoplasm of living cells deform to a spindle shape, which is attributable to the nematic liquid crystal formation of F-actin. Further, that deformation can be repeated by changing the external osmotic pressure or by irradiation with the excitation light of a fluorescence microscope. The aspect ratio of the liposomes can be reversibly changed by hypotonic and hypertonic treatments, due to changes in the actin concentration and the surface area/volume ratio resulting from water movement. Brief irradiation with strong excitation light can sever F-actin containing fluorescent-labeled monomers but they can subsequently repolymerize or anneal spontaneously, resulting in filament length switching and thus a reversible deformation between spindle-shaped liposomes and sphere-shaped liposomes possessing filopodia- or acrosome projection-like extensions. Interestingly, those extensions can also pull the liposome body during contraction.
These findings revealed that, not only force generating systems using cytoskeletons and molecular motors but also physicochemical phenomena, such as liquid crystal formation of cytoskeletons, are useful for actuating molecular robots. An advantage of utilizing liquid crystal formation of F-actin, as shown in this study, is that F-actin alone could deform cell-sized liposomes even without myosin, that the morphology of deformed liposomes could be further changed by external stimuli, and that the further deformation was repeatable. This concept can be applied to artificial macromolecules, and molecular robotics will be more promising by combination with macromolecular designs. So far, actin interactions with myosin and simple polymerization/depolymerization have been considered as the mechanisms exerting actin functions. Here, from the viewpoint of constructing a molecular robot as a bottom-up reconstruction approach in biology, the severing/re-elongation (repolymerization and/or annealing) of actin filaments is suggested to be another mechanism that enables the dynamic properties of living cells.
Results
Liposomes encapsulating dense F-actin acquire spindle shape
At first, we tried to encapsulate actin into cell-sized giant liposomes at high concentrations that have not been previously studied. In cases where it is necessary to experiment with repeated trials at high concentrations, a restriction is imposed even on proteins obtained with a high yield such as actin. It is important to ensure efficiency and reproducibility in the method for encapsulating high concentrations of proteins in membrane vesicles16. The amount-reduced water-in-oil (W/O) emulsion centrifugation method employed here meets those requirements8. This is a modification of the inverted emulsion method17, 18, so that the amount of protein used at a high concentration will be minimized as much as possible.
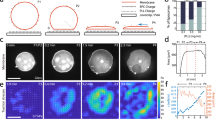
Giant liposomes prepared by this method assumed a spherical shape when no protein was encapsulated (Fig. 1a), whereas liposomes that encapsulated actin showed morphological changes depending on the polymerization state and concentration of actin encapsulated (Fig. 1b). When F-actin (polymerized filamentous actin, termed actin filaments) was encapsulated or when G-actin (monomeric actin that had not yet formed a filament) was encapsulated and subsequently polymerized to F-actin inside liposomes, the majority of liposomes tended to be spindle-shaped. The spindle-like shapes can be approximated with a morphology obtained by turning an arc around its long axis. Accordingly, the tips of some spindle-shaped liposomes were sharp, and occasionally a string-like membrane tube extended from those tips. Rarely, some liposomes encapsulating actin deformed to triangle- or cruciform- (or quadrupole-) shapes, which are complexes of three or four half-spindles, respectively (Supplementary Fig. 1). Investigating the conditions and mechanisms in which liposomes deform to these shapes is an interesting subject that will possibly lead to the induction and regulation of various morphological changes of liposomes.
Shape of liposomes encapsulating actin. a, b Phase contrast images of liposomes encapsulating without (a) or with 100 μM F-actin (b). All empty liposomes kept a spherical shape, whereas almost all liposomes encapsulating actin deformed to a spindle shape. Scale bars indicate 20 μm. c Dependence of liposome deformation on the concentration of actin encapsulated. The short axis length is plotted against the long axis length for individual liposomes when 10, 25, 44, 50, 147, or 169 μM actin were encapsulated. Note that the rightmost data point indicated by the arrow in the panel showing the results obtained at 147 μM represents a liposome with a long axis length of 48 μm and a short axis length of 10 μm, and that at 169 μM represents a liposome with a long axis length of 56 μm and a short axis length of 14 μm
First of all, it should be necessary to classify and analyze liposome morphology considering the effects of actin concentration and liposome size. Hereafter, unless otherwise noted, actin means F-actin and “liposomes encapsulating actin” means giant liposomes containing F-actin. The proportion of liposomes that deformed and the degree of their deformation depended on the concentration of actin encapsulated. The aspect ratio of individual liposomes at each actin concentration is shown in Fig. 1c (details of the results are shown in Supplementary Table 1). Points on the line with a slope of 1 that extend from the origin indicate spherical liposomes, i.e., those have not deformed. Liposomes that deformed to a spindle shape are indicated as points nearer the horizontal axis of each graph with increasing sharpness. When the concentration of actin encapsulated was higher than 50 μM, spindle-shaped liposomes were observed. With increasing actin concentrations, higher proportions of liposomes deformed to a spindle shape, which became more elongated, i.e., the aspect ratio was further away from the value of 1. It is hard to distinguish liposomes clearly into two populations of deformed and undeformed liposomes. So, if a liposome is categorized as deformed when the long axis length became equal to or more than 1.5 times the short axis length, when 169 μM actin was encapsulated, about two-thirds of the liposomes were deformed. At that concentration, liposomes with aspect ratio more than 2 were frequently observed. It is noted that approximately 200 μM actin is comparable to the actin concentration in living cells19, 20. Also, this is the upper limit concentration of F-actin that can be handled, such as by pipetting, due to its very high viscosity16. Incidentally, rare liposomal morphologies, such as triangle or cruciform shapes, could be observed when higher concentrations of actin were encapsulated. On the other hand, at least in the case of deformation to spindle shapes, liposomal size did not correlate with the deformation (Supplementary Fig. 2 and Supplementary Table 2).
The liposome shape showed some heterogeneity from spheres to sharp spindles even under the same conditions (Fig. 1c and Supplementary Fig. 2). This heterogeneity may correlate with the arrangement of F-actin confined, and indeed birefringence was strongly seen in spindle-shaped liposomes and was not seen in spherical liposomes (Supplementary Fig. 3).
Arrangement of F-actins in giant liposomes
In deformed liposomes, fluorescent signals from the encapsulated actin filled the inner space uniformly (Fig. 2a and Supplementary Fig. 4), indicating that F-actin exists in a homogeneous manner throughout the liposomes regardless of its location, e.g., the center, the tips of the spindle or the membrane portion. Moreover, deformed liposomes observed by polarization microscopy showed strong birefringence (Fig. 2a), which was observed throughout the liposomes, and was parallel to the spindle shape. Also, in the case of triangle- or cruciform-shaped liposomes, the fluorescence filled the interior uniformly and birefringence parallel to the half-spindles was obtained (Supplementary Fig. 1). On the other hand, particular structures such as bundles or backbones of F-actin did not exist. As the simplest case, inside spindle-shaped liposomes, it is assumed that straight F-actins are arranged in parallel with each other. The average distance between F-actins can be calculated as follows: Suppose that each F-actin occupies a cylindrical space with a diameter D, which is the average distance between F-actins. The volume of the cylindrical space per actin monomer is πD2/4σ, where σ is the line density of actin monomers on F-actin, 13.5 monomers/37 nm. The occupation volume by one actin monomer can be expressed as 1/(C × NA), where C is the molar concentration of actin monomer, and NA is Avogadro’s number. In the case of the highest actin concentration of our experiments (C = 169 μM), the average distance between F-actins, D, is calculated at about 70 nm, which is more than ten times the diameter of F-actin. At lower concentrations of actin, the average distance D is longer than 70 nm. F-actins can freely move relative to each other in such a sparse distribution unlike the case of bundles or networks of F-actins formed by crosslinking factors such as fascin or α-actinin21, and the collision of long F-actins (~5 μm) by thermal fluctuation keeps the nematic phase of F-actins.
Liposome morphology and arrangement of F-actin inside liposomes. a Spindle-shaped liposomes encapsulating actin observed by phase contrast polarization (direction of a polarizer and an analyzer are indicated) microscopies and fluorescence images for labeled actin and membrane stained. The actin concentration was 84 μM and 10 mol% actin in terms of the monomer was labeled with Alexa 488. This observation was done with S/G. A model of the liposome is also shown. b F-actin solution encapsulated into liposomes (100 μM) as a mixture of F-actin polymerized from fluorescently labeled monomers (50 mol%, with Alexa 488) and non-labeled F-actin with a ratio of 1/5000, in order to visualize the behavior of a single F-actin inside each liposome. Phase contrast image shows the entire shape of a spindle-shaped liposome. Model illustrates the outline of visualization of individual F-actins. The fluorescent images are still images of 1 frame and a superimposition of 440 frames (a period of about 37 s, a rate of 12 frames per second) from the movie of a liposome. All scale bars indicate 20 μm
The spindle-shaped liposomes observed by polarization microscopy show that F-actin is aligned within liposomes, perhaps along their long axial direction. In order to support this result, F-actin solutions where only some F-actin (about 1/5000 of all filaments) has been fluorescently labeled, were encapsulated in liposomes, and the behavior of the fluorescent F-actin was observed (Fig. 2b). The results show that F-actin is in a state in which it extends linearly and aligns along the long axis direction of spindle-shaped liposomes. Moreover, F-actin shows movements such as one-dimensional diffusion along the length direction of the filament, i.e., the long axis direction of the liposome (Supplementary Movie 1 and Supplementary Fig. 5). This result indicates that there is still room in deformed liposomes for F-actin to move as mentioned above, and that F-actin is not stacked. It is noted here that F-actin encapsulated in liposomes has a broad distribution of length from several up to 10 μm (depending on the conditions and the purity of the F-actin sample, median = 4–7 μm, mean = 6–10 μm), determined in fluorescence images (Figs. 2b and 3)22.
Effect of co-encapsulation of gelsolin on liposome deformation. a Fluorescence images showing F-actin in the bulk solution, phase contrast images showing liposomes encapsulating actin, and models for liposome deformation, in the absence or presence of gelsolin. A solution of rhodamine-phalloidin-labeled F-actin (76 μM) without or with a 1/100 molar ratio of gelsolin (760 nM) was diluted to 1 nM actin and then observed by fluorescence microscopy, or was encapsulated in liposomes as it is without dilution. Liposomes observed here were prepared with S/G. b, c Graphs show the distributions of F-actin length in the solutions and the aspect ratio of liposomes prepared with the solutions, respectively. Conditions are as shown in a. Red bars and red open circles indicate the results in the absence of gelsolin, and blue bars and blue filled circles indicate the results in the presence of gelsolin. d Phase contrast and polarization images of an individual liposome co-encapsulating F-actin and gelsolin. A solution of F-actin (70 μM) with a 1/1000 molar ratio of gelsolin (70 nM) was co-encapsulated in the liposome. All scale bars indicate 20 μm
Length effect of internal F-actin on the shape of liposomes
Only when F-actin, but not when G-actin, was encapsulated, liposomes deformed to a spindle shape (Figs. 1 and 2a, and Supplementary Fig. 6), indicating the necessity for the encapsulated components to be rod-like or filamentous for the deformation. We then determined how the degree of liposome deformation changes when the length of F-actin encapsulated is shortened (Fig. 3a). Gelsolin is an actin-binding protein that severs F-actin and caps its growth end in the presence of calcium ions, which results in making F-actin shorter (Fig. 3b and Supplementary Table 3)23. As expected, co-encapsulation of gelsolin reduced the liposome deformation compared with when the same concentration of F-actin was encapsulated (Fig. 3c and Supplementary Table 3). And of course, spherical liposomes co-encapsulating gelsolin showed no birefringence (Fig. 3d).
Effect of the inhibition of spontaneous alignment of F-actin
In order to examine the importance of the ability of F-actin to form a unidirectional alignment for liposome deformation from a spherical to a spindle shape, we utilized heavy meromyosin (HMM) prepared from skeletal muscle myosin. HMM has a double-headed structure and is known to be capable of crosslinking F-actins. Especially, in the absence of adenosine 5′-triphosphate (ATP), HMM forms a rigor complex between each myosin head and F-actin, resulting in an isotropic crosslinking network of F-actins24, 25. When HMM is added to the actin solution for encapsulation, ATP is depleted unless ATP is additionally supplied, because ATP that was contained in the actin solution (<0.2 mM) is sufficiently consumed by ATPase of actoHMM (μM order concentrations of F-actin and HMM) during the preparation of liposomes24, 26. Therefore, under these conditions, HMM will inhibit the nematic liquid crystallization of F-actins by its isotropic crosslinking, and thus inhibit the deformation of liposomes to a spindle shape. As expected, when F-actin was added with only HMM but no additional ATP, the liposomes kept a spherical shape (Fig. 4). Inside such spherical liposomes, the distribution of F-actin that is slightly heterogeneous and does not change at all could be observed (Fig. 4a) and, as expected, birefringence was never detected (Fig. 4b), as a result of the HMM-crosslinking (Fig. 4c). It is noted that, when liposomes did not deform, there was a possibility that actin was not encapsulated. Therefore, we counted liposomes in which the fluorescence of internal actin could be observed, as liposomes co-encapsulating F-actin and HMM. Of the 56 liposomes that could be observed, 50 were spherical and 6 were distorted, but no spindle-shaped liposomes were found.
Effect of co-encapsulation of HMM in the absence of ATP. F-actin (57 μM, 10 mol% actin labeled with Alexa 546) and HMM (5.5 μM) were co-encapsulated in liposomes. It should be noted here that, under these conditions, since ATP is depleted, HMM forms an isotropic crosslinking network as a result of rigor binding with F-actins, but does not act as a motor for F-actin. Almost all liposomes kept their shape as spherical. a Phase contrast and fluorescence images of a spherical liposome show the localization of F-actin within it. b Phase contrast and polarization images of another spherical liposome show the arrangement of F-actin inside it. c The models show that crosslinking between F-actin (red lines) and HMM (green dots) inhibits the F-actin rearrangement required for liposome deformation. Scale bars indicate 20 μm
This study demonstrated that if actin filaments are contained in cell-sized giant liposomes at high concentrations comparable to the cytoplasmic level, the liposomes spontaneously deform to spindle-like shapes even without any actin-binding proteins such as gelation or bundling factors or molecular motor myosins.
In contrast, when G-actin or gelsolin-shortened F-actin was encapsulated the deformation of liposomes did not occur (Fig. 3 and Supplementary Fig. 6), so it is indispensable that fibrous objects exist inside the liposomes. However, it did not seem that actin filaments were continuously elongating from the tip of one protrusion to the other like a beam supporting the shape. The coherence length of the nematic phase formed by encapsulated F-actins is probably longer than the liposome, and therefore it is able to deform liposomes larger than the length of F-actins (Fig. 2b). This deformation mechanism is different from that of bipolar-shaped MTs-encapsulating liposomes that are deformed by an inner pushing force generated by MTs elongating longer than the liposomes themselves. Because F-actin is uniformly distributed in the deformed liposomes and no factors that promote bundling are present, it is unlikely that F-actin is working by forming bundles and becoming rigid. In addition, because F-actin does not accumulate beneath the membrane and no factors that promote linkages between F-actin and the lipid membrane are present, it is not thought that the shape of the membrane is reinforced from inside by an actin cortex (Supplementary Notes 1 and 2, and Supplementary Fig. 4).
As important observations, F-actin, which is a representative native rod-like macromolecule, is aligned along the spindle shape, and it can move if the motion is along the long axis direction (Fig. 2b), indicating its properties as a nematic liquid crystal. It has already been shown that F-actin forms a nematic phase at high concentrations15, 27,28,29. From observations in the bulk solution, it has been reported that the nematic liquid crystallization of F-actin depends on its concentration and length like many other rod-like molecules, and the liquid crystal phase tends to appear at 50 μM or higher concentrations of F-actin29, 30. These are consistent with the results mentioned above (Figs. 1c and 2). Altogether, the unidirectional alignment of free moving F-actin, in other words, the nematic liquid crystal formation of the F-actin solution, inside liposomes is thought to be responsible for their deformation to the spindle shape. Regarding the heterogeneity seen in the degree of deformation of actin-encapsulating liposomes, it may be explained by the difference in defects or topology in active nematics5, 31, 32. If there are no or few defects, the liposome will deform to a spindle shape, whereas if there are too many defects the liposome will remain spherical without deformation.
Another finding to keep in mind is that the mechanism identified in this study is unique because the polarity of individual F-actins might be irrelevant. The two ends of actin filament have different properties from each other, and each F-actin has polarity. When F-actin is cooperating with its partner molecular motor myosin, it is the polarity of F-actin which determines the movement direction of myosin24, 26.
Reversible shape control of liposomes by osmotic pressure
Our system has only two components, lipid membranes and actin filaments. These components have flexible interactions without any static connection. Flexible interactions are a great advantage to reversibly and repeatedly control the shape of molecular robots.
As an attempt to reversibly and repeatedly deform the shape of liposomes encapsulating actin, we changed the osmolality of the external solution (Fig. 5 and Supplementary Table 4). When a hypertonic or hypotonic solution is added to an external solution of liposomes, water exits or enters through the membrane due to osmotic pressure, causing concentration changes of internal substances, i.e., the encapsulated actin and salts in this case. As a result, the alignment of encapsulated F-actin changes depending on the concentration change. As expected, the aspect ratio of liposomes varied further away from the value of 1 by the addition of a hypertonic solution to the external solution (Fig. 5a), and was reversely changed by the addition of a hypotonic solution (Fig. 5b). The deformation of individual liposomes induced by changing the osmolality finished within several seconds, indicating that the major driving force for the deformation is the realignment of F-actin, rather than slower biochemical reactions such as their polymerization or depolymerization. To be more certain, future studies should investigate how the polarization observed in liposomes changes during their deformation.
Control of the morphology of deformed cell-sized actin-encapsulating liposomes by changes in osmotic pressure of the external solution. a, b The aspect ratio of liposomes before and after the addition of a hypertonic (a) or a hypotonic solution (b) to the outer solution are plotted. The short axis length is plotted against the long axis length for individual liposomes. Blue and dark blue circles show the results before changing the osmotic pressure, and orange and cyan circles show the results after changing the osmotic pressure. Initially, the outer solution was the same as the F-actin-soluble buffer (5 mM Tris-HCl (pH 8.0), 30 mM KCl, 0.2 mM ATP, 5 mM DTT, 0.9 mM NaHCO3, 0.18 mM CaCl2). The concentration of F-actin encapsulated was 50 μM. Below each plot, the deformation of individual liposomes (phase contrast images and their models) induced by the hypertonic or the hypotonic treatment is also shown. All scale bars indicate 10 μm. Following the hypertonic treatment, liposomes deformed to a more elongated spindle shape within several seconds after the hypertonic solution was added. Following the hypotonic treatment, spindle-shaped liposomes tended to become spherical, and this shape change was completed within 5 min after the hypotonic solution was added
It is noted that if actin is not encapsulated, even when the osmotic pressure was changed, the aspect ratio of liposomes was not changed (Supplementary Fig. 7 and Supplementary Table 5), and only their size or membrane tension was affected.
Reversible shape control of liposomes by light irradiation
As another attempt to reversibly and repeatedly deform the shape of liposomes encapsulating actin, we utilized the severing of F-actin caused by irradiation with the excitation light of the fluorescence microscope (Fig. 6 and Supplementary Movie 2). It is well known that in in vitro assays, fluorescently labeled F-actin shows severing, which is attributable to radicals generated as byproducts produced by irradiation of the fluorescent dye with the excitation light33. Actually, although it was an experiment in bulk solution, we confirmed that fluorescently labeled F-actin was severed by irradiation with the excitation light (Supplementary Fig. 8a). Therefore, reducing agents and/or an oxygen removing system are added to the assay solution in order to also prevent the severing, not only to retard fluorescent bleaching34. On the other hand, it has already been reported that shortened F-actin returns to its original length under an equilibrium condition after a while by spontaneous repolymerization or annealing35. Also, under the conditions employed in this study, the spontaneous re-elongation of shortened F-actin after standing was observed (Supplementary Fig. 8b).
Repeated deformation of spindle-shaped liposomes encapsulating actin induced by light irradiation. Following brief irradiation with strong excitation light (about 20 s in this case), a spindle-shaped giant liposome that encapsulated F-actin containing fluorescent-labeled monomers deformed to spherical. In detail, the sphere elongated two very long extensions from the sites where the tips of the spindle-shaped liposome were originally. After the irradiation, if left for a while (tens of seconds to a few minutes), the liposome spontaneously returned to the original spindle shape, the central spherical part elongated, and the two extensions retracted. This repetition could be done several times. The period of no irradiation (OFF) and the timing just after the irradiation (ON) are indicated at the top, and the elapsed times (min:sec), with the first panel denoted as 0 s, are indicated at the bottom. The scale bar indicates 10 μm. A solution (5 mM Tris-HCl (pH 8.0), 30 mM KCl, 0.2 mM ATP, 10 mM DTT) containing F-actin (90 μM) polymerized from fluorescently labeled actin (10 mol%, with Alexa 546) was encapsulated in the liposomes. Models for the repeated deformation and mechanism are also shown
Following brief irradiation with the strong excitation light (10 s–1 min), spindle-shaped giant liposomes encapsulating F-actin containing fluorescent-labeled monomers deformed to spherical shapes. In other words, the spheres elongated two filopodia- or acrosome projection-like extensions from sites where the tips of the spindle shapes were originally. After the irradiation, if left for a while (tens of seconds to a few minutes), the liposomes spontaneously returned to their original spindle shape, i.e., the central spherical part elongated while the two extensions retracted (Figs. 6 and 7, Supplementary Movies 2–4). We could induce this repetition eight times at the maximum. Note that Fig. 6 and Supplementary Movies 2–4 show only the parts of the whole process that were successfully filmed. If there is a limitation to the number of repetitions, it is conceivable that actin was denatured by photoradicals, inhibiting re-elongation of F-actins.
Elongation and retraction of extensions that occur during liposome deformation. a A sequence of phase contrast and fluorescent images showing the repeats of ON/OFF of light irradiation carried out twice in succession against a liposome. The left two columns show the earlier repeat and the right two columns show the later repeat. The uppermost phase contrast images show the spindle shape of the liposome just before each light irradiation. The fluorescent images in each left column show the liposome deformation during ON of the light irradiation, and the phase contrast images in each right column show the liposome deformation during OFF of the irradiation. The number in each panel indicates the elapsed time (seconds), when the start of the first light irradiation is denoted as 0 s. Arrowheads indicate the position of the tip of the extension, and the asterisk in the panel for 101 s indicates that the extension protruded outside the observation field. In order to make the extension easy to see, the fluorescence images have been sharpened and the brightness and contrast of the phase contrast images were changed. Scale bars indicate 20 μm. b Change in the length of extensions with time measured from the above images. The white background and white circles indicate the period when the light irradiation was ON and the data that were obtained during that period. The gray background and black circles indicate the period when the irradiation was OFF and the data that were obtained during that period. As mentioned above, because the extension protruded outside the observation field at the vicinity of 100 s (asterisk), its length cannot be measured accurately. To observe the extension with fluorescence microscopy, i.e., to observe the distribution of actin, periods for the light irradiation were longer than usual. As a result, the degree of liposome deformation decreased at the later repeat. This result also indicates that the conditions of light irradiation are important for inducing the repetitive stretching of giant liposomes
Considering the application to the transportation and movement of micrometer-sized objects, it is important to reveal the processes and mechanisms of elongation and retraction of the extensions formed at both of the tips of the liposomes after light irradiation (Fig. 7 and Supplementary Movie 3). As a mechanism in which the central part occupying most of the liposome volume is deformed repeatedly between the spindle shape and the spherical shape, changes in the length distribution of the encapsulated F-actin should be involved as follows (also see Supplementary Movie 2). Radicals generated by the light irradiation cause severing of F-actin. The shortened length of F-actins resulted in the randomization of F-actin alignment. As a result, the liposomes changed to a spherical shape. It is presumed that the process occurs under the restraint condition in which the surface area of the membrane is constant. On the other hand, the volume of liposomes that had been deformed into a spindle shape is smaller than when they are spherical, because a sphere is the geometrical configuration that maximizes the ratio of volume to surface area. Therefore, only the central part will change to a spherical shape, and the remaining surplus membrane region will appear as extensions at one or both ends. It should be noted that the light irradiation carried out here does not completely depolymerize actin (Supplementary Fig. 8). Therefore, it is possible that F-actins remaining after the irradiation are within the extensions and make the extensions more stable. In fact, fluorescence observations indicate that actin is always present within the extensions. This actin may play an important role in the formation and behavior of extensions. When a protrusion is formed or disappears in the lipid bilayer membrane, a force exceeding a certain threshold (tens of pN) is necessary for its formation or disappearance, whereas almost no additional force is required for elongation or shortening of the tubular protrusions once they are formed36, 37. Regarding the extensions, such characteristics of tubular membrane protrusions may also be involved. After a while, the length of F-actins is increased by spontaneous repolymerization or annealing, resulting again in the formation of nematic liquid crystals of F-actin. As a result, the liposomes again deform to a spindle shape, and the extensions are shortened.
In order to verify the model above, when calculating the difference in surface area caused by liposome deformation between spherical and spindle shapes under the restriction of a constant liposome volume, the change is at most several percent, which seems to be too small to explain the surplus membrane surface area required for formation of the extension. The extensions contain actin inside as mentioned above. They are most likely F-actins, and we will have to consider their effect on the formation of extensions. Alternatively, some interactions between the membrane and F-actins triggered by irradiation-induced radicals may be the cause of the formation of the extensions through stabilizing tubular membrane structures in the liposome. In order to make a correct model, it is necessary to keep track of the time course of exact changes in the membrane surface area and the internal volume of the central part and extensions of individual liposomes. While it is very difficult to track in real time the detailed shapes of floating liposomes and the shapes of extensions that move in a complicated fashion like a whip, establishing a suitable model is an important issue for future research.
Discussion
The biochemical interactions between biological macromolecules, e.g., the dynamic formation of sturdy structures consisting of crosslinked cytoskeletons or the force generation by molecular motors, has been considered the mechanism for the morphogenesis of living cells. So far, the application of biochemical reactions, such as the base pairing of DNA (i.e. DNA origami) and/or the assembling of proteins (e.g. MTs or F-actin) has been expected as the engine of molecular robots1,2,3, 8, 10. The mechanism identified by this study is different from those, and sheds light on the significance of filament severing of cytoskeletons as a third mechanism23. Incidentally, whether the re-elongation of F-actin that spontaneously occurs after severing is due to repolymerization or to annealing is an issue that remains to be investigated. Previous studies have revealed that the forces generated by the cytoskeleton and molecular motors or by the polymerization of cytoskeletal filaments are sufficient to cause the deformation and/or movement in both living cells and artificial cell models4, 5. In addition, our study also suggests that the changes induced in the arrangement of F-actins that are in crowded conditions can generate forces necessary for the deformation of the cell and/or the elongation and contraction of membrane protrusions such as filopodia in living cells (Supplementary Discussion). Notably, the membrane extensions induced in spindle-shaped actin-encapsulating cell-sized liposomes by light irradiation are able to pull the liposome body, i.e., the spherical part, during the contraction (Supplementary Movies 3 and 4). Our findings will further clarify the effectiveness and future prospects of approaches already in progress to control the morphology and movement of cell-sized components, such as lipid membranes and membrane vesicles, using gelation and nematic liquid crystallization of artificial polymers and/or biopolymers including actin filaments38.
It should be noted that, in the case of the cycle of severing and repolymerization or annealing of actin, the consumption of nucleotide triphosphates, which are biological fuels, is overwhelmingly small compared with the two other mechanisms, the simple polymerization/depolymerization cycle of actin and the force generation by F-actin and myosin. This may be the reason why the liposomal deformation can be repeated many times, unlike previous studies8. Currently, the input energy of the light irradiation used may be much larger than the biochemical energy derived from nucleotide triphosphates. However, by improving this method, for example, by using the light source and irradiation method, which is capable of power output adjustment, and by regulating more spatiotemporally the light irradiation, it will be possible to sufficiently reduce energy consumption. In any case, the diversity in the method for regulating molecular robots will help to expand the possibilities of the process of evolving them. Moreover, our results show the effectiveness of physical mesoscopic phenomena, such as liquid crystal formation and phase transitions. Our findings demonstrate the possibility that a similar mechanism will work universally with rod-like and/or filamentous materials other than native polymers such as F-actin or MTs, and if so, this property will give a significant advantage for engineering.
Methods
Reagents
The materials used were obtained as follows: phospholipid, 1,2-dioleoyl-sn-glycero-3-phosphatidylcholine (DOPC), was purchased from Sigma (St. Louis, MO, USA), Avanti Polar Lipids (Alabaster, AL, USA), and Wako Pure Chemical Industries (Osaka, Japan). Liquid paraffin, squalene, chloroform, and methanol were purchased from Wako Pure Chemical Industries. Alexa 488 or 546 maleimide was purchased from Molecular Probes (Eugene, OR, USA). Rhodamine-phalloidin (R-415) was purchased from Invitrogen (Carlsbad, CA, USA). Other chemicals, including ATP, EGTA, and salts, were of analytical grade and were purchased from Wako Pure Chemical Industries or Nacalai Tesque (Kyoto, Japan). These chemicals were used without further purification.
Proteins
Actin was prepared from chicken breast muscle or rabbit skeletal muscle according to the method of Spudich and Watt39, except that the tropomyosin–troponin complex was removed before preparing the acetone powder. Labeling of actin with fluorescent dyes (Alexa 488 or 546 maleimide) was performed as described previously34, 40. Otherwise, in order to visualize actin, rhodamine-phalloidin was added to F-actin prior to the preparation of liposomes encapsulating actin16. G-actin was stored in salt-free buffer (0.2 mM ATP, 1 mM NaHCO3, 0.2 mM CaCl2 with or without 2–5 mM Tris-HCl (pH 8.0)), and F-actin was polymerized in salt-containing buffer (5 mM Tris-HCl (pH 8.0), 30 mM KCl, 0.2 mM ATP, 5 mM DTT, 0.9 mM NaHCO3, 0.18 mM CaCl2) in general. The concentration of sodium bicarbonate or calcium chloride was sometimes slightly different, but it did not affect the results if it was within 10% difference. The concentration of reducing agent was changed as appropriate from the results obtained. Actin samples were kept on ice until used. Gelsolin was prepared as follows: glutathione S-transferase-tagged human plasma gelsolin was expressed in Escherichia coli, and a soluble extract of the bacterial cells was applied to glutathione beads. The proteins adsorbed on the beads were treated with protease to separate the GST-tag, and then gelsolin was obtained in the elution. HMM was obtained from skeletal muscle myosin prepared as described previously41.
Liposomes
Giant liposomes used were prepared by the amount-reduced W/O emulsion centrifugation method performed as follows8. All procedures were performed at room temperature except where otherwise noted. Briefly, small aliquots of protein (G- or F-actin) solutions added in liquid paraffin were solubilized in phospholipid, and were shaken vigorously to make emulsions in an Eppendorf tube. Each emulsion was placed on top of other aqueous solutions in a PCR tube, and centrifuged at 18,000 × g for 10 min. Usually this was done at 4 °C, but the centrifugation speed and/or temperature was changed as appropriate from the results of liposome preparation. During the centrifugation, aqueous drops of the emulsion were pulled down and passed through the interface between the upper oil and the bottom aqueous phases. During passing the interface, the droplet was covered by a lipid bilayer and became a liposome. After centrifugation, the resulting liposomes were sedimented to the bottom of the tube. After removing the upper oil phase, the pellet was recovered and suspended in an aqueous solution, where its condition or osmolality was the same as the outer solution. The resulting suspension was kept on ice and observed for a day. The method for liposome preparation described here is applicable for trace amounts of sample and inactivation of the protein sample can be prevented by shortening the preparation time, so it has important advantages when handling biologically derived precious factors such as rare proteins or when a large number of trials are necessary. Sometimes, in order to increase the yield of liposomes encapsulating actin, an appropriate concentration of sucrose was added to the aliquot of F-actin solution and an equimolar concentration of glucose was added to the bottom aqueous solution that will become the outer solution. In cases where the addition of sucrose/glucose was performed like this, it will be stated that the experiment was done with S/G. In those experiments with sucrose and glucose, their concentrations were 25 mM for confocal microscopic observation and 500 mM for all other cases. By this method, the yield of liposomes obtained was increased several times.
Giant liposomes were observed at 25 °C by phase contrast and fluorescence microscopes (BX60/IX70; Olympus, Tokyo, Japan) attached to a polarizing unit and a camera (WAT-910HX; Watec, Tsuruoka, Japan/IR-1000; DAGE-MTI, Michigan City, IN, USA). We manually tracked the flowing liposomes in focus. The acquired images were recorded in HDD by the PC through an image capture unit (DFG/USB2aud; Imaging Source, Bremen, Germany). To adjust and analyze the contrast of images, computer software ImageJ (http://imagej.nih.gov/ij/) or plug-in View 5D (http://www.nanoimaging.de/View5D/) were used42, 43. In order to visualize fluorescent liposomal membranes, Nile-Red in dimethylsulfoxide was added to the liposome suspensions (final concentration = a few micromolar) before observation. The observation of giant liposomes by polarization microscopy was performed using two polarizers, a 1/4 phase plate and a rotation stage attached to the microscope. The direction of the slow axis of birefringence was decided by image processing using ImageJ. To prevent evaporation and condensation of sample solutions, microscopic specimens were sealed by valap sealant44. Confocal images of liposomes were obtained using a CellVoyager CV1000 512 (Yokogawa Electric Co., Musashino, Japan), which is a confocal imaging system integrated with a confocal scanner unit CSU, an excitation laser light source, and an ultra-sensitive EMCCD camera (512 × 512).
In each observation sample prepared as described above, roughly tens of liposomes could be observed. Regarding the size distribution of the liposomes obtained, the approximate range can be found from the scatter diagram of the aspect ratio of the shapes of liposomes (Figs. 1c, 3 and 5, Supplementary Fig. 7). The yield and size of liposomes were not significantly affected by the encapsulation of actin, at least under the experimental conditions carried out in this study. Incidentally, when F-actin was mixed with only HMM, but not ATP, and then encapsulated in liposomes, the yield of liposomes obtained reproducibly decreased by nearly an order of magnitude. The addition of sucrose/glucose could not overcome that decrease. That may be due to the viscosity increase caused by the rigor complex formation of F-actins and HMM, or the effect of the nature of HMM, e.g., easy to adsorb to any surface including a membrane surface16, 45, 46.
For all results reported for this study, we performed at least three independent experiments and made an effort to observe more than 20 liposomes per experiment.
Osmotic shock to giant liposomes under a microscope
Morphological controls of liposomes were as follows: for hypertonic or hypotonic treatment, that is, the application of osmotic shock, a liposome solution was mixed with a 5% volume of milli-Q water with or without 3 M KCl, under a microscope. The solution was put on a glass slide as small drops next to each other, and the drops were then covered with a cover glass with a pair of spacers 100 μm thick. The two open sides of the chamber were sealed with valap sealant to prevent evaporation and condensing of the solutions as described above. To mix each pair of drops, the upper cover glass was pushed at the middle between the two drops. Using this method allowed us to continuously observe the same liposome before and after the mixing. The indicated osmotic pressures are those after the mixing. The time range to reach a uniform concentration in the mixed drop was checked using a drop containing calcein, a water soluble fluorescent dye, and was estimated to be shorter than 1 min.
Measurement of the length distribution of F-actin
In order to assess the effect of the severing of F-actin by light irradiation, mechanical stimulation (pipetting), or the addition of gelsolin, the following procedure was used to measure the actin filament length distribution. The light irradiation, mechanical stimulation (pipetting), or addition of gelsolin was carried out using an appropriate volume (1–100 μL) of the bulk solution of F-actin (with or without fluorescence labeling) at a similar concentration as when it was encapsulated in liposomes (70–90 μM of actin concentration). After that, aliquots were mixed with equimolar amounts of phalloidin (if necessary, rhodamine-phalloidin) to prevent depolymerization, and were diluted to a final concentration of 1 nM actin to make single F-actin recognizable, and then observed in a flow chamber by fluorescence microscopy for measurement.
Light irradiation of giant liposomes
For the light irradiation, each specimen containing a solution of liposomes encapsulating actin was covered with an IR filter, and was briefly irradiated with green excitation light of the microscope (10 s –1 min) through the objective lens. It should be noted that it is essential that the actin is fluorescently labeled and that the excitation wavelength of light corresponds to the fluorescence dye used. The deformations of liposomes induced were monitored and acquired as a sequence of phase contrast images with the microscope.
Data availability
The data supporting the findings of this study are available within the paper, its Supplementary Information Files, and from the corresponding author upon reasonable request.
References
Erbas-Cakmak, S., Leigh, D. A., McTernan, C. T. & Nussbaumer, A. L. Artificial molecular machines. Chem. Rev. 115, 10081–10206 (2015).
Hagiya, M., Konagaya, A., Kobayashi, S., Saito, H. & Murata, S. Molecular robots with sensors and intelligence. Acc. Chem. Res. 47, 1681–1690 (2014).
Smith, L. M. Nanotechnology: molecular robots on the move. Nature 465, 167–168 (2010).
Keber, F. C. et al. Topology and dynamics of active nematic vesicles. Science 345, 1135–1139 (2014).
Sanchez, T., Chen, D. T. N., DeCamp, S. J., Heymann, M. & Dogic, Z. Spontaneous motion in hierarchically assembled active matter. Nature 491, 431–434 (2012).
Sanchez, T., Welch, D., Nicastro, D. & Dogic, Z. Cilia-like beating of active microtubule bundles. Science 333, 456–459 (2011).
Abkarian, M., Loiseau, E. & Massiera, G. Continuous droplet interface crossing encapsulation (cDICE) for high throughput monodisperse vesicle design. Soft Matter 7, 4610–4614 (2011).
Hayashi, M. et al. Reversible morphological control of tubulin-encapsulating giant liposomes by hydrostatic pressure. Langmuir 32, 3794–3802 (2016).
Chun, A. L. Artificial cells: pressurize and release. Nat. Nanotech. 11, 403 (2016).
Sato, Y., Hiratsuka, Y., Kawamata, I., Murata, S. & Nomura, S. M. Micrometer-sized molecular robot changes its shape in response to signal molecules. Sci. Robot. 2, eaal 3735 (2017).
Pollard, T. D. & Cooper, J. A. Actin, a central player in cell shape and movement. Science 326, 1208–1212 (2009).
Zimmerman, S. B. & Minton, A. P. Macromolecular crowding: biochemical, biophysical, and physiological consequences. Annu. Rev. Biophys. Biomol. Struct. 22, 27–65 (1993).
Ellis, R. J. Macromolecular crowding: an important but neglected aspect of the intracellular environment. Curr. Opin. Struct. Biol. 11, 114–119 (2001).
Courchaine, E. M., Lu, A. & Neugebauer, K. M. Droplet organelles? EMBO J. 35, 1603–1612 (2016).
Weirich, K. L. et al. Liquid behavior of cross-linked actin bundles. Proc. Natl. Acad. Sci. USA 114, 2131–2136 (2017).
Takiguchi, K., Yamada, A., Negishi, M., Tanaka-Takiguchi, Y. & Yoshikawa, K. Entrapping desired amounts of actin filaments and molecular motor proteins in giant liposomes. Langmuir 24, 11323–11326 (2008).
Pautot, S., Frisken, B. J. & Weitz, D. A. Production of unilamellar vesicles using an inverted emulsion. Langmuir 19, 2870–2879 (2003).
Pontani, L.-L. et al. Reconstitution of an actin cortex inside a liposome, for the encapsulation of actin with this technique. Biophys. J. 96, 192–198 (2009).
Pollard, T. D., Blanchoin, L. & Mullins, R. D. Molecular mechanisms controlling actin filament dynamics in nonmuscle cells. Annu. Rev. Biophys. Biomol. Struct. 29, 545–576 (2000).
Janson, L. W., Kolega, J. & Taylor, D. L. Modulation of contraction by gelation/solation in a reconstituted motile model. J. Cell Biol. 114, 1005–1015 (1991).
Honda, M., Takiguchi, K., Ishikawa, S. & Hotani, H. Morphogenesis of liposomes encapsulating actin depends on the type of actin-crosslinking. J. Mol. Biol. 287, 293–300 (1999).
Sept, D., Xu, J., Pollard, T. D. & McCammon, J. A. Annealing accounts for the length of actin filaments formed by spontaneous polymerization. Biophys. J. 77, 2911–2919 (1999).
Ishikawa, R., Yamashiro, S. & Matsumura, F. Annealing of gelsolin-severed actin fragments by tropomyosin in the presence of Ca2+. Potentiation of the annealing process by caldesmon. J. Biol. Chem. 264, 16764–16770 (1989).
Tanaka-Takiguchi, Y. et al. The elongation and contraction of actin bundles are induced by double-headed myosins in a motor concentration-dependent manner. J. Mol. Biol. 341, 467–476 (2004).
Tharmann, R., Claessens, M. M. A. E. & Bausch, A. R. Viscoelasticity of isotropically cross-linked actin networks. Phys. Rev. Lett. 98, 088103 (2007).
Takiguchi, K. Heavy meromyosin induces sliding movements between antiparallel actin filaments. J. Biochem. 109, 520–527 (1991).
Ott, A., Magnasco, M., Simon, A. & Libchaber, A. Measurement of the persistence length of polymerized actin using fluorescence microscopy. Phys. Rev. E 48, R1642–R1645 (1993).
Zhang, R., Kumar, N., Ross, J. L., Gardel, M. L. & de Pablo, J. J. Interplay of structure, elasticity, and dynamics in actin-based nematic materials. Proc. Natl. Acad. Sci. USA 115, E124–E133 (2018).
Viamontes, J., Oakes, P. W. & Tang, J. X. Isotropic to nematic liquid crystalline phase transition of F-actin varies from continuous to first order. Phys. Rev. Lett. 97, 118103 (2006).
Le Helfer, E., Panine, P., Carlier, M.-F. & Davidson, P. The interplay between viscoelastic and thermodynamic properties determines the birefringence of F-actin gels. Biophys. J. 89, 543–553 (2005).
Yang, Y., Barry, E., Dogic, Z. & Hagan, M. F. Self-assembly of 2D membranes from mixtures of hard rods and depleting polymers. Soft Matter 8, 707–714 (2012).
Giomi, L. Geometry and topology of turbulence in active nematics. Phys. Rev. X 5, 031003 (2015).
Golde, T. et al. Fluorescent beads disintegrate actin networks. Phys. Rev. E 88, 044601 (2013).
Fujiwara, I., Takahashi, S., Tadakuma, H., Funatsu, T. & Ishiwata, S. Microscopic analysis of polymerization dynamics with individual actin filaments. Nat. Cell Biol. 4, 666–673 (2002).
Andrianantoandro, E., Blanchoin, L., Sept, D., McCammon, J. A. & Pollard, T. D. Kinetic mechanism of end-to-end annealing of actin filaments. J. Mol. Biol. 312, 721–730 (2001).
Inaba, T. et al. Formation and maintenance of tubular membrane projections require mechanical force, but their elongation and shortening do not require additional force. J. Mol. Biol. 348, 325–333 (2005).
Umeda, T., Inaba, T., Ishijima, A., Takiguchi, K. & Hotani, H. Formation and maintenance of tubular membrane projections: experiments and numerical calculations. Biosystems 93, 115–119 (2008).
Prost, J., Jülicher, F. & Joanny, J.-F. Active gel physics. Nat. Phys. 11, 111–117 (2015).
Spudich, J. A. & Watt, S. The regulation of rabbit skeletal muscle contraction. I. Biochemical studies of the interaction of the tropomyosin-troponin complex with actin and the proteolytic fragments of myosin. J. Biol. Chem. 246, 4866–4871 (1971).
Otterbein, L. R., Graceffa, P. & Dominguez, R. The crystal structure of uncomplexed actin in the ADP state. Science 293, 708–711 (2001).
Okamoto, Y. & Sekine, T. A streamlined method of subfragment one preparation from myosin. J. Biochem. 98, 1143–1145 (1985).
Schneider, C., Rasband, W. & Eliceiri, K. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Abramoff, M., Magelhaes, P. & Ram, S. Image processing with ImageJ. Biophotonics Int. 11, 36–42 (2004).
Valap sealant. Cold Spring Harbor Protocols (Cold Spring Harbor Laboratory Press). http://cshprotocols.cshlp.org/content/2015/2/pdb.rec082917 (2015).
Kron, S. J., Toyoshima, Y., Uyeda, T. Q. P. & Spudich, J. A. Assays for actin sliding movement over myosin-coated surfaces. Methods Enzymol. 196, 399–416 (1991).
Takiguchi, K., Negishi, M., Tanaka-Takiguchi, Y., Homma, M. & Yoshikawa, K. Transformation of actoHMM assembly confined in cell-sized liposome. Langmuir 27, 11528–11535 (2011).
Acknowledgements
We thank Shuichi Takeda and Mahito Kikumoto and other members of the Structural Biology Research Center (Nagoya University), and Yoshikatsu Sato (Institute of Transformative Bio-Molecules (WPI-ITbM), Nagoya University) for their support. Observation by confocal microscopy was conducted in the WPI-ITbM and was supported by the Japan Advanced Plant Science Network. We are grateful to Kenichi Yoshikawa (Doshisha University) and Masatoshi Ichikawa (Kyoto University) for their comments. This work was supported by MEXT Japan KAKENHI Grant Number JP 24104004 and by JSPS KAKENHI Grant Numbers JP 24651134 and JP 16K07293.
Author information
Authors and Affiliations
Contributions
Experiments: S.T. and M.H.; analysis: S.T., K.T., and M.H.; manuscript preparation: K.T.; and design of the study: S.T., K.T., and M.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tanaka, S., Takiguchi, K. & Hayashi, M. Repetitive stretching of giant liposomes utilizing the nematic alignment of confined actin. Commun Phys 1, 18 (2018). https://doi.org/10.1038/s42005-018-0019-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-018-0019-2
This article is cited by
-
Spatiotemporal control of signal-driven enzymatic reaction in artificial cell-like polymersomes
Nature Communications (2022)
-
Cell-size space effects on phase separation of binary polymer blends
Biophysical Reviews (2022)
-
Towards a synthetic cell cycle
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.