Abstract
Photoacoustic imaging has been shown to provide high-resolution images of genetically labelled cells at depths that are inaccessible to optical microscopy. While the detection of genetic reporters, such as fluorescent proteins and pigments, has been demonstrated using multiwavelength imaging and spectral unmixing, these approaches remain challenging due to their large computational scale. In this study we report a method based on a reversibly photoswitchable phytochrome-based reporter protein (AGP1) and dual-wavelength interleaved image acquisition for obtaining difference images with unambiguous reporter-specific contrast. Detailed, full 3D images of tumours formed of cells lentivirally transduced to express AGP1 were acquired in vivo in deep tissue in a longitudinal study. This method represents a powerful new approach to studying cellular and genetic processes which, due to its experimental simplicity, can be implemented in a wide range of existing photoacoustic imaging platforms.
Similar content being viewed by others
Introduction
Biomedical photoacoustic (PA) imaging, which relies on the absorption of intensity-modulated light by tissue chromophores to generate broadband ultrasound waves1, 2, has been shown to provide exquisite high-resolution three-dimensional (3D) in vivo images of the vasculature3, 4 due to the strong contrast provided by haemoglobin in the visible to near-infrared (NIR) wavelength region. While this can be exploited using multiwavelength PA imaging to enable functional studies5, 6, most other types of tissues and cells do not exhibit sufficient endogenous contrast in this spectral region to allow detection. Exogenous contrast agents or genetically expressed absorbers are therefore used to overcome the lack of inherent optical absorption. While a wide range of exogenous contrast agents, such as dyes7 or nanoparticles8, 9, is available, their application in vivo is often hampered by the limited specificity with which cellular receptors can be targeted and their typically low systemic delivery efficiency. The use of reporter genes to provide genetically encoded PA contrast is an attractive alternative that overcomes these limitations. By incorporating reporter genes into the cells to achieve the genetic expression of molecular absorbers, this approach can be used to reveal the location or patterns of migration of otherwise transparent cells in deep tissue. More importantly, reporter genes provide the opportunity to ‘spy on cells’10 in order to obtain information on how genes and gene products affect internal cellular biochemical processes and allow the study of complex biological processes, such as cell growth, intracellular signalling, protein–protein interactions, and differentiation11, in the living organism.
While the fundamental capability of PA imaging of genetically expressed reporters in vivo has been demonstrated using fluorescent proteins or pigments, a number of limitations hamper the translation of this technology to applications in the life sciences. For example, while fluorescent proteins allow the visualisation of cells in comparatively small and transparent organisms12, 13 and in xenografts in small mammals14, they often exhibit reduced PA contrast at comparatively low fluences compared to non-fluorescent absorbers due to ground state depopulation and photobleaching15. An alternative is the genetic expression of enzymes that synthesise photostable pigments. For example, the expression of tyrosinase, which synthesises eumelanin16,17,18, has been used to provide genetically encoded PA contrast in mammalian cells to enable their detection in longitudinal tomographic imaging studies in mice16. However, while fluorescent proteins and pigments provided proof of principle of PA reporter gene imaging, their unambiguous detection in vivo and in deep tissue (>3 mm) remains challenging. Current methodologies are often based on multiwavelength image acquisition and some form of spectral unmixing and aim to exploit the differences in the absorption spectra of the tissue chromophores to recover their spatial distribution. While model-based inversion schemes are promising approaches19, recovering the spatial distribution of genetic reporters from high-resolution 3D images represents a large-scale inverse problem of considerable computational burden. In addition, it is typically assumed that the absorption spectra of the chromophores do not change during the PA signal generation. For fluorophores, this is often not valid due to ground state depopulation and fluorescence15, 20, which may adversely affect specificity.
Reversibly photoswitchable absorbing proteins, which exhibit different isomeric states with distinct absorption spectra, provide a route to overcoming these limitations. The optically induced change in absorption, which is not observed in endogenous chromophores, provides a unique contrast mechanism that can be exploited using PA difference imaging. The method relies on acquiring PA images while the protein is photoswitched between the isomeric states, thus modulating its optical absorption. From these data, PA difference images are calculated in which the otherwise overwhelming background contrast is eliminated to visualise the location of the genetic reporter with high sensitivity and specificity. This approach therefore represents a straightforward and solely experimental method for obtaining unique and highly reporter-specific image contrast.
While this principle was used to image photoswitchable fluorescent proteins in tissue phantoms21, 22, their absorption coincides with strong attenuation in blood, thus making them less suited to deep tissue PA imaging. This limitation was overcome in recent studies in which the detection of mammalian cells in deep tissue using reversibly photoswitchable phytochrome reporter proteins and PA difference imaging23, 24 was demonstrated. Phytochromes, which are light-sensitive proteins that originate from plants or bacteria, combine a number of highly advantageous properties for PA reporter gene imaging, such as strong absorption in the red to NIR wavelength region where the optical penetration depth of tissue is greatest and predominately vibrational relaxations25 for efficient PA signal generation. PA difference imaging of cells expressing phytochrome BphP1 was demonstrated by Yao et al.23 using a method that relies on acquiring two-dimensional (2D) images at a single NIR excitation wavelength and photoswitching the phytochrome absorption using the output of an additional laser source in the red wavelength region. The disadvantages of this approach are that photoswitching-induced changes in absorption at other wavelengths, which would maximise detection sensitivity, are not exploited and that the use of separate excitation lasers adds experimental complexity and cost to the imaging system.
In this paper, we report a methodology for PA reporter gene imaging based on the use of phytochrome AGP1 as a reporter protein and dual-wavelength signal acquisition, which enabled tomographic imaging of growing tumours in vivo in a longitudinal study. The advantages of this methodology are (i) dual-wavelength excitation using a single optical parametric oscillator (OPO) excitation laser to exploit the difference absorption spectrum of phytochromes, (ii) the development of a novel and biocompatible phytochrome-based reporter protein (AGP1), and (iii) full 3D tomographic imaging. Due to its conceptual and experimental simplicity, this method can be implemented in a wide range of existing in vivo PA imaging systems equipped with single 355 nm pumped OPO lasers, thus supporting translation of this technology to a broad range of preclinical applications in the life sciences.
Results
In vitro optical and PA properties of AGP1
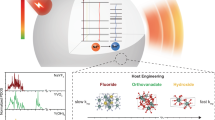
Phytochrome AGP126,27,28 (from Agrobacterium tumefaciens) is a non-fluorescent protein in which the pigment biliverdin is covalently bound to the protein barrel. AGP1 can adopt two isomeric states, Pr (red) and Pfr (far-red), which exhibit different absorption spectra in the wavelength region where the optical attenuation due to endogenous tissue absorbers, such as water29, oxy- (HbO2), and deoxyhaemoglobin (HHb)30, and optical scattering is minimal (Fig. 1a). The Pr and Pfr states can be switched using red and NIR wavelengths as shown in Fig. 1b. Photoswitching proceeds via several intermediate molecular states, such as Lumi-R and Lumi-F. A slow transition from Pfr to Pr also occurs as a result of thermal (or dark) conversion. The quantum yields of the photoconversions, ηPfr−Pr and ηPr−Pfr, differ by an order of magnitude (Fig. 1b), which results in more efficient photoswitching from Pr to Pfr31.
Optical and energetic properties of AGP1 and the fluence dependence of the PA difference signal. a Absorption spectra of AGP1 in its isomeric states Pr and Pfr, water, and oxy- (HbO2) and deoxyhaemoglobin (HHb). AGP1 provides strong absorption at wavelengths where the optical penetration depth of tissue is at a maximum. b Schematic of the electronic and vibrational energy levels of AGP1 during isomeric conversion. Illumination using red and NIR wavelengths allows photoswitching between the Pr and Pfr states. The transitions proceed via a number of intermediate states (grey dotted lines), e.g., Lumi-R and Lumi-F. The quantum yields of the transitions (ηPfr−Pr, ηPr−Pfr) differ by an order of magnitude, resulting in a more efficient Pr→Pfr transition. c Illustration of the photoswitching cycle and the PA signal acquisition during dual-wavelength excitation using a single OPO laser. The wavelength of the excitation pulses is switched to result in sequential single and dual-wavelength illumination. This modulates the AGP1 population, which adopts Pr and Pfr states during single-wavelength excitation and a mixed Pr–Pfr state during dual-wavelength excitation. After a short equilibration phase, three PA signals corresponding to each AGP1 population state, i.e., SPr, SPfr,Pr, and SPfr, were acquired. The values specified on the time axis represent those used during in vivo imaging. d Normalised PA difference signal amplitudes measured in vitro in purified solutions of AGP1 using separate and simultaneous excitation at 650 and 755 nm as a function of the relative fluences. The total fluence was kept constant at 5 mJ/cm2. The error bars represent the standard deviation of four repeated measurements
One way to exploit the photoswitching-induced change in optical absorption for PA imaging is to acquire images at an excitation wavelength where the difference in absorption between the Pr and Pfr state is at a maximum, i.e., in the NIR wavelength range. This approach, reported by Yao et al.23, was implemented by using an additional excitation source to facilitate the modulation of the isomeric state of the phytochrome. By capturing images while the phytochrome was in either the Pr or Pfr state, a difference image is obtained in which the endogenous contrast is eliminated. However, this approach does not capitalise on the differences in absorption observed in the red wavelength region, while the additional excitation laser adds experimental complexity and cost. In this work, a method was developed based on dual-wavelength excitation which, first, exploits the absorption difference spectrum of AGP1 in the red and NIR wavelength region and, second, uses a single OPO excitation laser. In principle, maximum change in phytochrome absorption, and hence maximum PA detection sensitivity, is obtained by making measurements at red and NIR excitation wavelengths while the phytochrome population is completely converted to either the Pr or the Pfr state. However, to arrive at a compromise between detection sensitivity and the image acquisition time of the raster scanning all-optical scanner, a dual-wavelength excitation photoswitching cycle as illustrated in Fig. 1c was developed. During the photoswitching cycle, the AGP1 population is converted sequentially from the Pr to the Pfr state via an intermediate state using excitation wavelengths of the signal and idler outputs of the OPO laser. After a short equilibration phase, three time-resolved PA signals corresponding to each AGP1 state population, i.e., SPr, SPfr,Pr, and SPfr, were acquired. PA difference signals, SD, were obtained by subtracting the sum of PA waveforms acquired using separate excitation pulses from the waveform acquired using simultaneous excitation pulses in the red to NIR wavelength region, i.e., SD = SPfr,Pr−SPr−SPfr. From the calculated SD, PA difference images are then obtained using image reconstruction algorithms.
To determine the experimental conditions under which this method provides maximum PA difference signal amplitude, a PA spectroscopy setup (Supplementary Figure 1) was used to record time-resolved signals in vitro in purified protein solutions (Supplementary Figure 2). By measuring the difference signal amplitude as a function of the signal and idler wavelengths, a maximum was observed at 670 and 755 nm (Supplementary Figure 3a), and these wavelengths were used throughout this study. Measurements in AGP1 solutions in the Pr state or the Pfr state at 670 and 755 nm, respectively, were used to estimate the maximum contrast-to-noise ratio (CNR) provided by dual-wavelength excitation, and was found to be twice that achieved using single-wavelength excitation (Supplementary Figure 4). Measurements of the signal amplitude as a function of the number of excitation pulses showed that the rate of photoswitching is roughly eight times greater for Pr→Pfr compared to Pfr→Pr and increases with fluence (Supplementary Figure 3b), which is in qualitative agreement with the difference in quantum yield (Fig. 1b). The difference in the photoswitching rate affects the dependence of the difference signal amplitude on the excitation fluences as shown in Fig. 1d. It is at a maximum when the ratio of the fluences at 670 and 755 nm is approximately 1:2. Importantly, AGP1 was also shown to exhibit negligible photobleaching at a fluence equivalent to the maximum permissible exposure (MPE) for skin (Supplementary Figure 3c). This is a major advantage for in vivo imaging applications where it is often necessary to minimise the effects of tissue motion using signal averaging and the acquisition of multiple images32. By contrast, a strong deterioration in PA signal amplitude was measured in solutions of a fluorescent protein with comparable NIR absorption (Supplementary Figure 3c).
Interleaved PA difference imaging of AGP1 in tissue phantoms
An all-optical PA scanner based on a Fabry–Pérot polymer film ultrasound sensor33 was used to record 3D image data sets in tissue phantoms (Fig. 2a). The phantom consisted of polymer tubes containing solutions of purified AGP1 or copper chloride immersed in a scattering lipid suspension (Fig. 2b). It was placed on the sensor and illuminated using ns excitation pulses at 670 and 755 nm. The PA field was mapped by raster scanning the interrogation beam across the sensor. A protocol for interleaved image acquisition, which was developed to minimise motion artefacts in in vivo difference images obtained using the Fabry–Pérot-based scanner32, was implemented. This involved the acquisition of three PA signals at each raster scan position according to the photoswitching cycle illustrated in Fig. 1c. To account for the difference in the photoswitching rate of AGP1 (Supplementary Figure 3b), different equilibration times preceded the acquisition of the PA signals. From the interleaved image data set, cross-sectional images of the phantom were reconstructed. Figure 2c, d and e, f show x–z and x–y maximum intensity projections (MIPs) reconstructed from the sum of SPr and SPfr, and SPfr,Pr, respectively. The MIPs show the structure of the phantom with all tubes clearly visible. In order to recover the spatial distribution of AGP1, these images are subtracted to yield a difference image, which is shown in Fig. 2g, h. The spatial distribution of AGP1 is clearly visualised while the background contrast originating from non-photoswitchable chromophores is eliminated. Using the peak-to-peak amplitude of the image background, the noise-equivalent concentration (NEC) was estimated as 3.6 µM.
Interleaved PA image acquisition protocol and experimental validation of difference imaging of AGP1 in a tissue phantom. a Schematic of the all-optical PA scanner used for phantom and in vivo imaging. PA waves are generated in the target using ns excitation pulses at 670 and 755 nm and detected by a Fabry–Pérot ultrasound sensor. The waves are mapped in x and y by scanning the interrogation beam across the sensor and by recording the acoustically induced modulation of the reflected optical power at each scan point. b Schematic of the tissue phantom. c–h The x–z and x–y maximum intensity projections (MIPs) of 3D data sets acquired in the phantom using interleaved image acquisition. c, d MIPs reconstructed from data sets of the sum of SPr and SPfr. e, f MIPs reconstructed from SPfr,Pr. g, h Difference image obtained by subtracting the images shown in (c–f), i.e., SPfr,Pr−SPr−SPfr. The location of the tubes filled with AGP1 solution are clearly visible while the background is eliminated. The scale bars represent 1 mm
In vivo longitudinal PA imaging of AGP1-expressing tumours
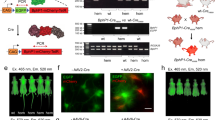
Permanent expression of AGP1 in human colorectal tumour cells (HT29) was achieved using stable lentiviral transduction to incorporate a transgene encoding AGP1 and the fluorescent protein mCherry into the chromosome (Fig. 3a–c). The co-expression of mCherry (Supplementary Figure 5a) provided a means to select cells with strong expression levels using fluorescence-activated cell sorting (FACS). The proliferation rates of transduced (HT29-AGP1-mCherry) and non-transduced cells (HT29-NT) show no significant differences (Fig. 3c), which suggests negligible cytotoxicity of AGP1 and mCherry. Stable transduction of AGP1 was validated by measuring the attenuation spectra of purified cell lysate (Supplementary Figure 6a).
Lentiviral transduction and viability of AGP1-expressing tumour cells. a Diagram of the lentiviral construct encoding long terminal repeat (LTR), AGP1, internal ribosome entry site (IRES), and the fluorescent protein mCherry, and an illustration of lentiviral transduction of a cell, which results in the integration of the reporter gene into the chromosome. b Fluorescence microscopy images of stably transduced (HT29-AGP1-mCherry) and non-transduced (HT29-NT) human colorectal cancer cells. The fluorescence emitted by the co-expressed mCherry is shown in red and the cell nuclei (stained using CytoxGreen) are shown in green. The scale bars represent 40 µm. c The proliferation of transduced and non-transduced HT29 cells over time showed no significant differences in the growth rates, demonstrating cell viability and low cytotoxicity of AGP1. The error bars represent the standard deviation of n = 3
The ability of interleaved dual-wavelength PA image acquisition to detect phytochrome-based reporter proteins in vivo was demonstrated in mice inoculated with transduced HT29 cells, which stably co-expressed AGP1 and mCherry (Supplementary Figure 5(b, c)). Tumour xenografts induced via the subcutaneous injection of 10 million cells in the flank were imaged using the Fabry–Pérot-based scanner at a number of time points post inoculation. Figure 4 shows PA images of different tumours obtained using interleaved dual-wavelength image acquisition. Figure 4a–d provides an illustration of the content of interleaved image data sets. Figure 4a, b shows cross-sectional images of the vascular morphology of a tumour and the surrounding tissue (day 24 post inoculation), which were obtained using simultaneous (SPfr,Pr) and separate (SPfr+SPr) dual-wavelength excitation. Figure 4a, b displays no visible evidence of AGP1. This is in contrast to the difference image (Fig. 4c), which clearly visualises the location of a tumour xenograft that developed two distinct nodes (see also Supplementary Figure 7a for a 3D representation). The difference image intensity, which is assumed to be proportional to AGP1 concentration, is an order of magnitude lower than that of the morphological images (Fig. 4a, b). Profiles of the difference image intensity are shown as a function of depth z in Fig. 4d for x = 5.6 mm and 10.5 mm as indicated by the green and red dashed lines in Fig. 4c. The intensity in the region corresponding to the AGP1-expressing HT29 tumour cells (0.5 mm < z < 3.5 mm) is a factor of two to three higher than that of the background intensity (z > 6 mm). Interestingly, Fig. 4d also shows negative values at z ~4.0 mm. This suggests that photoswitching of AGP1, which results in a modulation of the optical attenuation within the tumour tissue, also affects the fluence in neighbouring regions.
In vivo PA imaging of subcutaneous tumours of AGP1-expressing HT29 cells in mice. a, b Cross-sectional 2D images of the vascular morphology of a tumour reconstructed from data sets acquired on day 24 post inoculation using a simultaneous (SPfr,Pr) and b separate excitation at 670 and 755 nm (SPfr+SPr), c Difference image obtained by subtracting (a) and (b) showing the location of the tumour, which consisted of two distinct nodes (see also Supplementary Figure 7a). The scale bars indicate 1 mm and the total image acquisition time was 1.5 h. d Difference image intensity as a function of depth z for x = 5.6 mm and x = 10.5 mm corresponding to the dashed lines in (c). e, f The x–y and x–z MIPs of a different subcutaneous AGP1-expressing tumour (day 20 post inoculation) demonstrating deep tissue imaging of AGP1-expressing HT29 cells (green) to a depth of 5.5 mm. The scale bars indicate 1 mm and the colour bars represent the normalised image intensities of the background (grey) and AGP1 (green). g Volume-rendered, fused 3D image of the data set shown in (e, f) in which the vasculature is shown in red and the spatial distribution of AGP1-expressing cells in green. The colour bars represent the normalised image intensities of the background (red) and AGP1 (green). The total 3D image acquisition time was 3.5 h
In the example shown in Fig. 4d, this appears to have resulted in a shadowing effect at depths beyond the tumour margins. The reduction in the local fluence may explain the negative difference image intensities. While this is likely to have implications for a quantitative interpretation of difference images, it is unlikely to adversely affect the ability of this method to recover the spatial distribution of AGP1 since the photoswitching-induced difference image intensity is positive.
Figure 4e, f shows x–y and x–z fused colour MIPs created by combining difference images of an AGP1-expressing tumour acquired on day 20 post inoculation with high-resolution 3D images of the vascular morphology acquired conventionally. The extent of the tumour is visualised to a depth of 5.5 mm (Fig. 4f), demonstrating the capability of deep tissue imaging of reporter genes with high spatial resolution. Similar depth-to-resolution ratios are difficult to achieve using purely optical imaging modalities, such as microscopy or fluorescence molecular tomography. A further example of a 3D image of an AGP1-expressing tumour (day 28 post inoculation) is shown in Supplementary Figure 7(b, c). Figure 4g shows a volume-rendered 3D image of Fig. 4e, f in which the morphological image is rendered in red and fused with the difference image in which the location of the AGP1-expressing HT29 cells is visualised in green. Difference images obtained in xenografts of non-transduced HT29-NT cells, i.e., the control group, did not show contrast indicative of AGP1 (Supplementary Figure 8). From the peak-to-peak noise observed in the difference images of the controls (Supplementary Figure 8b), which is higher than that measured in tissue phantoms due to residual tissue motion, the CNR was estimated to be approximately 8. The NEC obtained in vivo was estimated to be 12 µM based on the NEC of the tissue phantom experiments. The concentration of AGP1 in the tumours formed by transduced HT29 cells were calculated to range from 20 to 30 µM.
The expression of AGP1 in vivo was validated independently on ex vivo tissue samples obtained from HT29-NT and HT29-AGP1-mCherry tumours. PA difference images obtained in an AGP1-expressing tumour show clear evidence of the reporter protein (Supplementary Figure 9a) while the control showed no contrast (Supplementary Figure 9b). AGP1 expression was also validated using spectrophotometry by measuring the optical attenuation spectra of filtered tumour tissue lysate (i) after light exposure and (ii) after several hours of darkness from which a difference spectrum was obtained. Difference spectra measured in lysate of AGP1-expressing tumours show the absorption changes associated with AGP1 photoswitching, while that of the control group shows no spectral features (Supplementary Figure 6b).
The images shown in Fig. 5 demonstrate in vivo longitudinal PA reporter gene imaging of growing tumours. To illustrate the dimensions and the approximate location of the tissue region over which cross-sectional images were obtained using interleaved image acquisition, a volume-rendered 3D image of the tumour on day 21 post inoculation that was acquired conventionally using a single excitation wavelength is shown in Fig. 5a. The difference image contrast observed in AGP1-expressing tumours was consistently greater than that of the background for all measured image data sets (Fig. 5b). To aid visualisation, the tumour volume was segmented manually based on the vascular morphology and the dashed lines indicate the scan region for interleaved imaging. Figure 5c, e shows x–z MIPs of the vascular morphology of the tumour and the surrounding tissue acquired on days 14, 21, and 28 post inoculation. While Fig. 5c, e allow the location of the tumour to be gleaned from the vascular features, the corresponding difference images shown in Fig. 5f, h reveal its true extent due to the unambiguous contrast provided by the AGP1-expressing cells. The images in Fig. 5f, h were thresholded to show intensities above the peak-to-peak noise of difference images acquired in vivo in non-transduced tumours (Supplementary Figure 9b). Figure 5f, h illustrates the dynamic changes over time by clearly visualising the growth of AGP1-expressing cells. The persisting difference image contrast also provides evidence of successful stable transduction, i.e., the reporter gene incorporated into the chromosome of the mother cell is passed on to daughter cells. This was also confirmed by in vivo fluorescence imaging of the tumours in mice (Supplementary Figure 5b, c) and fluorescence microscopy images of histological tumour slices (Supplementary Figure 10), which show stronger mCherry fluorescence emission in transduced cells compared to non-transduced cells but also a high degree of spatial homogeneity in the proliferation of transduced cells.
Longitudinal PA imaging of an AGP1-expressing subcutaneous tumour. a Volume-rendered 3D image of the tumour (green) surrounded by the vasculature in the skin and the underlying muscle tissue (red) acquired on day 21 post inoculation. The tumour region was segmented manually to aid visualisation. The dashed lines indicate the tissue volume over which cross-sectional image data sets were acquired using interleaved image acquisition. b Box-and-whisker plot of the PA difference image intensity of the background and the tumour regions calculated from the longitudinal images acquired in all animals (n = 5). The box shows the median, 25th and 75th percentile, the error bars represent the 10th and 90th percentile, and the dash-dot line represents the mean. A paired t-test yielded p = 0.008. c–e The x–z MIPs of the cross-sectional image data sets acquired on c day 14, d day 21, and e day 28 post inoculation showing the vascular morphology of the tumour and the surrounding tissue. f, h The x–z MIPs of difference images of the distribution of AGP1-expressing tumour cells. The scale bars indicate 1 mm and the colour bar indicates the normalised difference image intensity. The image acquisition time was 1.5 h
Discussion
This study has demonstrated full 3D PA reporter gene imaging in vivo of lentivirally transduced mammalian cells in deep tissue and in a longitudinal preclinical study. The key to these capabilities was the use of an imaging protocol based on dual-wavelength excitation and interleaved image acquisition in conjunction with phytochrome-based reporter protein AGP1.
The dual-wavelength excitation method can easily be implemented in PA imaging systems equipped with 355 nm pumped OPO laser systems. By rigorously characterising the optical properties and the photoswitching time-course of AGP1 in vitro, experimental parameters, e.g,. excitation wavelengths, relative fluences, and photoswitching equilibration times, were optimised to achieve maximium difference imaging contrast. In addition, the use of interleaved image acquisition minimised the effect of tissue motion, which manifests itself as background noise. The 12 µM NEC achieved in this work could be reduced to potentially hundreds of nM. For example, signal-to-noise ratio (SNR) could be maximised by increasing the total fluence to the MPE, by trading spatial resolution for acoustic sensitivity, i.e., by increasing the optical thickness of Fabry–Pérot sensors33, and by optimising the excitation wavelengths independently. A major advantage of dual-wavelength PA difference imaging compared to single-wavelength imaging23 is a potential reduction in NEC. This is because dual-wavelength excitation has the potential to exploit the absorption difference spectrum of phytochromes in the red and NIR region (Supplementary Figure 6), while single-wavelength excitation relies on photoswitching a single state. While the photoswitching cycle implemented in this study is a compromise between detection sensitivity and image acquisition times, in vitro measurements have demonstrated that dual-wavelength excitation offers significant increases in detection sensitivity over single-wavelength excitation (Supplementary Figure 3). Given the frame rates of recently reported highly parallelised scanners, dual-wavelength photoswitching cycles that provide maximum detection sensitivity can be implemented without adversely affecting image acquisition times.
By implementing interleaved image acquisition with optimised experimental parameters using an all-optical Fabry–Pérot PA scanner, full 3D high-resolution imaging of AGP1-expressing cells was demonstrated in vivo. While this resulted in comparatively long acquisition times (up to 3.5 h for full 3D images), this is mainly due to the specifications of the particular Fabry–Pérot-based scanner used in study and which relies on raster scanning to acquire PA signals. However, it has already been shown that acquisition times can be shortened substantially by parallelising the sensor read-out using, for example, multiple interrogation beams34, resulting in an improvement by almost an order of magnitude.
This method exploits the combination of advantageous properties provided by phytochromes. First, the isomeric states of phytochromes can be optically switched to produce a change in absorption, and hence an unambiguous PA contrast mechanism. Second, phytochromes exhibit efficient thermalisation, and therefore PA signal generation, due to primarily vibrational relaxations. Third, phytochromes provide strong absorption in the NIR wavelength region, making them highly suited to deep tissue imaging applications. Fourth, phytochromes show negligible photobleaching even at comparatively high fluences, which is vital for longitudinal studies and imaging protocols that use signal averaging and repeated photoswitching to minimise detection noise and tissue motion artefacts. Fifth, phytochromes have been shown to be biocompatible. Their genetic expression in mammals does not require the additional provision of biliverdin as it is a product of the heme metabolism and therefore readily available. AGP1 has been shown to have comparable photoswitching contrast to BphP1. The development of a lentiviral transduction method to incorporate a transgene optimised for expression in mammalian cells resulted not only in strong and consistent PA contrast, but also offers the potential for transducing slowly growing and non-dividing cells, such as stem cells.
In summary, we present a novel method for PA reporter gene imaging that overcomes the limitations of conventional multiwavelength imaging and spectral unmixing approaches. By generating unambiguous contrast in deep tissue, dual-wavelength PA difference imaging using phytochrome-based reporter proteins is a step change in PA molecular imaging capability. Due to its experimental simplicity and ease of implementation, this method enables reporter gene imaging using a wide range of existing experimental and commercially available PA imaging systems. Given their highly advantageous properties, phytochromes are a highly attractive starting point for the development of new genetic reporters for preclinical applications of PA imaging, to include functionalised expression, the study of protein–protein interaction35, and optogenetics36.
Methods
Protein expression in bacteria and purification of AGP1
Escherichia coli expression vector pET21b-agp137 encodes an AGP1-6xHis fusion protein with a C-terminal located 6xHis-tag required for affinity purification. Since functional AGP1 requires biliverdin as co-factor, a truncated sequence of human hemoxygenase 2 (deltaHOX2) was ligated into a modified E. coli protein expression vector pQE-81L (Qiagen, Germany), to enable E. coli internal synthesis of biliverdin and the use of a kanamycin resistance marker38. Both vectors, pET21b-agp1 and pQE-81L-deltahHOX2, were used for co-expression in the standard E. coli protein expression strain BL21(DE3) (Novagen, Germany). Expression was induced with 0.4 mM isopropyl-β-d-thiogalactopyranoside for 18 h at 37 °C and at 190 rpm in a rotary shaker. Cells were harvested and stored at −80 °C after shock freezing in liquid nitrogen. For AGP1 purification, frozen bacterial cells were re-suspended and sonicated in 50 ml lysis buffer (pH 8.0, 50 mM NaH2PO4, 300 mM NaCl, 0.1 mM ethylenediaminetetraacetic acid (EDTA), and 1 mM phenylmethanesulfonyl fluoride). Supernatant of centrifuged cell extract was used for purification by His-tag affinity chromatography (Protino Ni-IDA Resin) according to the instructions of the manufacturer (Macherey-Nagel, Germany). Purified AGP1 was dialysed for 18 h at 4 °C in standard phosophate-buffered saline (PBS) (pH 7.4) using Slide-A-Lyzer Dialysis Cassettes (Thermofisher Scientific, Germany). The infrared fluorescent protein iRFP713, encoded in the original expression vector pQE-81L, was co-expressed in E. coli BL21(DE3) cells together with deltaHOX2. Protein expression and purification procedures for iRFP713 are the same as described for AGP1.
Plasmids for protein expression in mammalian cells and lentiviral vector production
For protein expression in mammalian cells, a codon optimised synthetic AGP1-6xHis sequence (MWG-Biotech AG, Germany) was used. It was ligated into the BamHI and EcoRI restriction sites of the HIV-derived lentiviral expression vector LeGO-iC2 (a gift from Boris Fehse39, Addgene plasmid #27345). The sequence-verified LeGO-iC2-agp1 expression vector together with the lentiviral packaging plasmids pRSV-Rev, pMDLg-pRRE (a gift from Didier Trono40, Addgene plasmids #12253, #12251) and pCMV-VSV-G (a gift from Bob Weinberg41, Addgene plasmid #8454) were used for generation of lentivirus followed by transduction into mammalian cells as described below. Cell-free viral supernatants were generated by transient transfection of HEK293T packaging cells using Lipofectamine 3000 transfection reagent and the vectors LeGO-iC2-AGP1 (21 µg), pRSV-Rev (8 µg), pMDLg-pRRE (8 µg) and pCMV-VSV-G (8 µg) according to the instructions of the manufacturer (Thermofisher Scientific, Germany). The presence of lentivirus was confirmed five days after transfection using Lenti-X GoStix (TaKaRa Bio Europe, France). AGP1 purification procedures for mammalian cells are the same as those described for bacterial cultures.
Cell culture and lentiviral gene transfer
Human colon carcinoma cells (HT29) were purchased from Caliper Life Sciences (Rüsselsheim, Germany) and cultured in McCoy’s 5A modified medium with 2.2 g/l NaHCO3 supplemented with 10% fetal calf serum, 2 mM l-glutamine, and 10,000 IU/ml penicillin–streptomycin (Biochrom AG, Berlin, Germany) at 37 °C in a humidified incubator at 5% CO2. For transduction, the cells were incubated in 6-well plates in their respective growth media in the presence of diluted and non-diluted viral supernatants without penicillin and streptomycin at a cell density of at least 90% confluence. Cells were transduced for 48 h and prepared for cell sorting via FACS (BD FACSAria II) using the co-expressed mCherry as a fluorescent marker.
Successful transduction was evaluated by measuring the fluorescence intensity of cell lysate. In all, 10,000 and 50,000 transduced and non-transduced HT29 cells of four difference passages were seeded, grown for 24 h, and lysed with 0.1% Triton X-100 in PBS for 10 min at 37 °C. The fluorescence emitted by mCherry was measured using the EnVision 2103 Multilabel Reader (Perkin Elmer, USA). Fluorescence microscopy images of intact monolayer cell cultures were also acquired. The cells were seeded on cover slips (∅ 12 mm; R. Langenbrinck, Germany) and cultivated until confluence reached between 60 and 70%. The cells were fixed in 4% buffered formalin for 10 min and washed in tris-buffered saline (TBS). Cell nuclei were stained with 1 µM CytoxGreen in TBS (Life Technologies). After washing, the cells were briefly dehydrated in ethanol. Cover slips were air-dried and embedded with Immu-Mount (Thermo Scientific) on glass slides (Superfrost, Thermo Scientific). Images of the fluorescence emitted by mCherry were acquired using a confocal laser-scanning microscope (Axiovert 100, LSM510, Carl Zeiss, Germany).
To evaluate cell proliferation of transduced and non-transduced HT29 cells, 5000 cells of each cell line and of three different passages were seeded in 200 µl medium in 96-well plates. After 48, 72, and 96 h, half of the medium was discarded and cells were incubated in 100 µM Resazurin and culture media solution. The fluorescence emitted by the reduced Resazurin was measured after 2 h using the EnVision 2103 Multilabel Reader and the data are presented as means with standard deviation and were tested using two-way repeated measurement analysis of variance. An additional test of cell proliferation (data not shown) was conducted in which the cells were washed with PBS, fixed with 4 % buffered formalin for 10 min, and cell nuclei were stained with 0.5 µg/ml 4′,6-diamidino-2-phenylindole in PBS and 0.1% Triton-X100 for 10 min. Cells were covered with PBS and the cell nuclei were imaged using the IN Cell Analyzer 1000 (GE Healthcare, USA) using ×100 magnification. Ten images were taken per well and the cell nuclei were counted using the IN Cell Analyzer 1000 Workstation 3.5 Software (GE Healthcare, USA). GraphPad Prism software (GraphPad Software version 5.04, La Jolla, USA) was used for data analysis. AGP1 expression in transduced and non-transduced HT29 cells was confirmed by measuring the absorption spectrum of purified solutions of lysed cells.
Xenografts
Prior to inoculation, cells were grown to 80−90% confluence, detached with trypsin and EDTA. Transduced HT29-mCherry-AGP1 cells (1 × 107 cells in 100 µl 0.9% NaCl solution) were inoculated subcutaneously into the flank of adult female athymic NMRI-Foxn1nu/Foxn1nu nude mice (Janvier Labs, France) (n = 5). Non-transduced HT29 cells were injected similarly to provide a control group (n = 2). To monitor reporter protein expression in growing tumours, the fluorescence emitted by the co-expressed mCherry was imaged in vivo using the IVIS Lumina (λexc = 570 nm, Cy5.5 emission filter set) (Xenogen Corporation, USA) under isoflurane anaesthesia. The tumours were allowed to grow for up to 4 weeks and tumour growth was monitored using a digital calliper. The tumours were imaged using PA imaging once their size had reached between 100 and 1000 mm3. At the end of the PA imaging study, the animals were killed. The tumour tissue was removed immediately, snap-frozen in liquid nitrogen, and stored at −80 °C for subsequent histology. All animal experiments were in accordance with the German Animal Welfare Legislation and were approved by the Landesamt für Gesundheit und Soziales Berlin (LaGeSo approval no. G0155/15).
In vitro PA spectroscopy of purified AGP1 and iRFP solutions
PA signals were measured in purified solutions of AGP1 with concentrations ranging from 68 µm to 100 µM using the PA spectroscopy setup (Supplementary Figure 1). The solution was held in a cuvette and illuminated with the signal and idler output of an OPO laser system (Newport Spectra Physics, USA; GWU, Germany) at a pulse repetition frequency of 50 Hz. PA signals were acquired using a planar ultrasound transducer with a 10 MHz centre frequency (Precision Acoustics, UK), a 40 dB voltage preamplifier (Femto, Germany), and a digitiser card (National Instruments, USA). Difference signals were calculated by subtracting the sum of PA waveforms acquired using separate signal and idler excitation pulses from that acquired using simultaneous pulses. To determine the optimum excitation wavelength combination, difference signals were measured as a function of the signal output (630 to 680 nm) and the corresponding idler output (742 to 810 nm). To optimise the signal and idler fluences for maximum difference signal amplitudes, their relative contribution was varied using neutral density filters while keeping the total fluence constant at 5 mJ/cm2. The rate with which AGP1 can be reversibly photoswitched between the Pfr and the Pr states was measured by recording the PA signal amplitude as a function of the number of excitation pulses at a single excitation wavelength. Prior to each measurement, the solution was illuminated with the idler or the signal output for 30 s to ensure an AGP1 population in either the Pr or Pfr state. The rate of photoswitching during simultaneous signal and idler excitation was measured by varying the fluence of either signal or idler beam between 2, 3, and 5 mJ/cm2 while keeping the other constant at 5 mJ/cm2. Photobleaching was measured in solutions of purified AGP1 (68 µM) and a fluorescent protein iRFP713 (c = 100 µM) by generating PA waves using simultaneous excitation pulses at 670 and 755 nm with a combined fluence of 20 mJ/cm2 and by recording the signal amplitude as a function of the number of excitation pulses.
PA imaging and image reconstruction in phantoms and in vivo
PA image data sets were acquired in tissue phantoms and in vivo using the OPO excitation laser and an all-optical PA scanner, which is described in detail elsewhere33. Briefly, the scanner is based on a Fabry–Pérot polymer film ultrasound sensor which provides high acoustic sensitivity, broadband frequency response, and small active element sizes for high-resolution PA imaging. The sensor consists of two dichroic mirrors of high transmittance between 590 and 1200 nm and high reflectance around 1550 nm, separated by a polymer film. Its optical transparency in the visible and NIR wavelength region allows the transmission of excitation laser pulses through the sensor and into the adjacent target to generate PA waves. The optical thickness of the etalon is modulated as the waves propagate through the sensor, causing a transient change in its reflectivity. By raster scanning a focussed 1550 nm CW laser beam (Tunics T100, Yenista, France) across the surface of the sensor and by recording the time-varying reflected intensity at each point using a photodiode, the spatial–temporal distribution of the incident PA waves are mapped in 2D. The photodiode output was recorded using a digitiser card (PCI-5124, National Instruments, USA). From these data, 3D images can be obtained using image reconstruction algorithms. In this study, a Fabry–Pérot sensor of 20 µm thickness was used, which provides a uniform frequency response to 39 MHz (−3 dB point) and a peak noise-equivalent pressure of 310 Pa (over a 20 MHz detection bandwidth). The vertical line spread function of the scanner ranges from 19 µm close to the sensor to 100 µm at a depth of 10 mm, and the lateral line spread function ranges from 38 µm to around 100 µm depending on depth and lateral position33. The excitation fluence at the target was 3.5 mJ/cm2 for the signal output at 670 nm and 7.4 mJ/cm2 for the idler output at 755 nm to maximise the difference signal amplitude (Fig. 1c).
Interleaved image acquisition, which has been shown to minimise motion artefacts32, was used to obtain difference images of the spatial distribution of AGP1 in tissue phantoms and in vivo. It is based on measuring three PA signals sequentially using excitation pulses at (1) 670 nm (SPr), (2) 750 nm (SPfr), and (3) 670 and 755 nm (SPfr,Pr) at each raster scan position. The signal acquisition protocol shown in Fig. 2c was implemented by delaying the signal acquisition following a change in excitation wavelength in order to allow the AGP1 transitions between the Pr and Pfr states to complete. For the comparatively slow transition from Pfr to Pr (Supplementary Figure 3b), the delay was 1200 ms while that for the Pr to Pfr transitions was 200 ms, after which the populations of the AGP1 states were considered to have reached equilibrium. PA signals (SPr, SPfr, SPfr,Pr) were then measured using signal averaging to improve the SNR and to reduce the effect of tissue motion. Switching of the excitation wavelengths was implemented using galvanometer mirrors, which coupled the signal and idler beams into multimode fused silica fibres (1.5 mm core diameter, Thorlabs) or beam dumps (Supplementary Figure 1).
Subcutaneous tumours of transduced and non-transduced HT29 cells were imaged in vivo by anaesthetising the mice using 2% isoflurane in oxygen (2 l/min). The mice were placed on the scanner and the tumours were positioned at the centre of the scan area. Aqueous gel between the skin and the Fabry–Pérot sensor provided acoustic coupling, a thermal chamber ensured constant body temperature, and the breathing rate was monitored. Tumours of AGP1-expressing HT29 cells (n = 5) were imaged up to five times and up to day 28 post inoculation in a non-randomised, non-blinded study. Tumours formed by non-transduced HT29 cells were imaged to provide controls (n = 2). Two PA image data sets were acquired during each experiment. First, a high-resolution 3D image data set was obtained using simultaneous excitation pulses at 670 and 755 nm with a total fluence of approximately 10 mJ/cm2 to visualise the vascular morphology. The x–y scan area was 14 mm × 14 mm with increments of dx = dy = 100 µm. Time-resolved PA signals of 2000 data points were acquired with a sampling frequency of 200 MHz without signal averaging. The image acquisition time was approximately 8 min. Second, interleaved image acquisition was used to acquire cross-sectional and full 3D image data sets from which difference images could be calculated. Cross-sectional 3D image data sets of tumours were obtained using signal averaging (n = 100) over an x–y scan area of, for example, 14.0 mm × 0.6 mm with dx = 100 µm and dy = 250 µm, which resulted in an image acquisition time of 1.5 h. For full 3D interleaved image acquisition, the scan area and step increments were adjusted to match the size of the tumour and to minimise the scan duration. For example, a 7 mm × 7 mm scan area with dx = 100 µm and dy = 400 µm resulted in an acquisition time of 3.5 h.
Following data acquisition, 3D PA images of the vascular morphology were reconstructed from the data sets acquired using the standard image acquisition protocol. Prior to reconstruction, the PA image data sets were interpolated in x and y onto a three times finer spatial grid. The tissue sound speed was determined using an autofocus function42. Time-reversal image reconstruction algorithm43, 44 and time-variant filtering45 were used to obtain images of the initial pressure distribution assuming homogeneous acoustic properties. MIPs with linear or logarithmic image intensity scale were prepared using Matlab (Mathworks), and 3D fused colour volume rendering was accomplished using Amira (Visual Sciences Group). To obtain PA difference images from the interleaved data sets, the PA signals were low pass filtered (fc = 7 MHz) to improve SNR and interpolated on a finer spatial grid (2× to 4×) before image reconstruction. The intensity of the difference images was thresholded to reject values below the peak-to-peak background noise, which was obtained from the standard deviation of the image intensity of a region corresponding to tissue surrounding the tumour multiplied by a factor of three. The mean PA difference image intensity of the background, i.e., regions corresponding to normal tissue surrounding the tumour, and the tumour regions (Fig. 5b), was calculated from the longitudinal images acquired in all animals (n = 5).
The 3D image data sets of a tissue phantom were obtained using interleaved image acquisition using the Fabry–Pérot-based PA scanner as shown in Fig. 2b. The phantoms consisted of soft polymer tubes (600 µm i.d., Paradigm Optics, USA) filled with aqueous solutions of purified AGP1 (c = 27 and 55 µM) and copper chloride (c = 0.44 M). The tubes were submerged in a scattering lipid suspension with a reduced scattering coefficient of approximately 1 mm−1 as shown in Fig. 2a. PA signals were acquired using signal averaging (n = 50). The noise-equivalent concentration of AGP1 provided by interleaved image acquisition was determined from the mean difference image intensity corresponding to the AGP1-filled tubes and the peak-to-peak noise of the background.
Characterisation of tumours ex vivo
The presence of AGP1 in ex vivo tumour was confirmed by measuring the supernatant of lysed tissue using a spectrophotometer. Frozen tumour tissue (approximately 0.5 g) was ground in liquid nitrogen using a mortar and pestle followed by re-suspension of the tissue debris in 500 µl of ice-cold PBS buffer (pH 7.4, 4 °C) supplemented with 0.1% TritonX-100 (Thermofisher Scientific, Germany). This crude extract was centrifuged for 30 min at 4 °C. The supernatant was dialysed for 18 h in PBS buffer at 4 °C. Since the dialysed supernatant exhibited weak optical scattering due to residual cell debris, a difference spectrum was calculated from two spectra of the optical attenuation measured using a spectrophotometer (Perkin Elmer Lambda19). The first spectrum was measured immediately after exposing the sample to light to maximise the AGP1 concentration in the Pfr state. The second spectrum was measured after leaving the sample in the dark for 4 h to allow dark conversion of AGP1 to the Pr state. By subtracting the spectra, a difference spectrum is obtained that shows the change in absorption due to AGP1 photoswitching (Supplementary Figure 6b) while the contribution of background absorbers is largely removed.
Reporter protein expression in histological slices was demonstrated using microscopy to image the fluorescence of co-expressed mCherry. Frozen tumour tissues were embedded in Tissue-Tek O.C.T Compound (Sakura Finetek, USA) and sectioned to a thickness of 10 µm. Sections were fixed using 4% buffered formalin for 10 min. After washing with TBS, sections were dehydrated in ethanol for 2 min, air-dried, and embedded with Immu-Mount (Thermo Scientific) onto glass slides (Superfrost, Thermo Scientific). Mounted cover slides were imaged using a confocal laser-scanning microscope (Mikroskop Axiovert 100, LSM510, Carl Zeiss, Jena, Germany).
Data availability
The data sets generated and analysed during this study are available from the corresponding author on reasonable request.
References
Beard, P. Biomedical photoacoustic imaging. Interface Focus 1, 602–631 (2011).
Yao, J. & Wang, L. V. Photoacoustic tomography: fundamentals, advances and prospects. Contrast Media Mol. Imaging 6, 332–345 (2011).
Laufer, J. et al. In vivo preclinical photoacoustic imaging of tumor vasculature development and therapy. J. Biomed. Opt. 17, 56016-1–56016–8 (2012).
Xia, J., Yao, J. & Wang, L. V. Photoacoustic tomography: principles and advances. Prog. Electromagn. Res. 147, 1–22 (2014).
Krumholz, A., Wang, L. L. V. & Yao, J. Functional photoacoustic microscopy of diabetic vasculature. J. Biomed. Opt. 17, 60502 (2012).
Deán-Ben, X. L. & Razansky, D. Functional optoacoustic human angiography with handheld video rate three dimensional scanner. Photoacoustics 1, 68–73 (2013).
Weber, J., Beard, P. C. & Bohndiek, S. E. Contrast agents for molecular photoacoustic imaging. Nat. Methods 13, 639–650 (2016).
Kircher, M. F. et al A brain tumor molecular imaging strategy using a new triple-modality MRI-photoacoustic-Raman nanoparticle.Nat. Med. 18, 829–834 (2012).
De la Zerda, A. et al. Carbon nanotubes as photoacoustic molecular imaging agents in living mice. Nat. Nanotechnol. 3, 557–562 (2008).
Tsien, R. Y. Building and breeding molecules to spy on cells and tumors. FEBS Lett. 579, 927–932 (2005).
Schroeder, T. Imaging stem-cell-driven regeneration in mammals. Nature 453, 345–351 (2008).
Razansky, D. et al. Multispectral opto-acoustic tomography of deep-seated fluorescent proteins in vivo. Nat. Photonics 3, 412–417 (2009).
Liu, M. et al. In vivo three dimensional dual wavelength photoacoustic tomography imaging of the far red fluorescent protein E2-Crimson expressed in adult zebrafish. Biomed. Opt. Express 4, 1846–1855 (2013).
Krumholz, A., Shcherbakova, D. M., Xia, J., Wang, L. V. & Verkhusha, V. V. Multicontrast photoacoustic in vivo imaging using near-infrared fluorescent proteins. Sci. Rep. 4, 3939 (2014).
Laufer, J., Jathoul, A., Pule, M. & Beard, P. In vitro characterization of genetically expressed absorbing proteins using photoacoustic spectroscopy. Biomed. Opt. Express 4, 2477 (2013).
Jathoul, A. P. et al. Deep in vivo photoacoustic imaging of mammalian tissues using a tyrosinase-based genetic reporter. Nat. Photonics 9, 239–246 (2015).
Paproski, R. J., Forbrich, A. E., Wachowicz, K., Hitt, M. M. & Zemp, R. J. Tyrosinase as a dual reporter gene for both photoacoustic and magnetic resonance imaging. Biomed. Opt. Express 2, 771–780 (2011).
Paproski, R. J., Heinmiller, A., Wachowicz, K. & Zemp, R. J. Multi-wavelength photoacoustic imaging of inducible tyrosinase reporter gene expression in xenograft tumors. Sci. Rep. 4, 5329 (2014).
Cox, B., Laufer, J. G., Arridge, S. R. & Beard, P. C. Quantitative spectroscopic photoacoustic imaging: a review. J. Biomed. Opt. 17, 61202 (2012).
Märk, J. et al. Photoacoustic imaging of fluorophores using pump-probe excitation. Biomed. Opt. Express 6, 2522–2535 (2015).
Stiel, A. C. et al. High-contrast imaging of reversibly switchable fluorescent proteins via temporally unmixed multispectral optoacoustic tomography. Opt. Lett. 40, 367 (2015).
Dean-Ben, X. L. et al. Light fluence normalization in turbid tissues via temporally unmixed multispectral optoacoustic tomography. Opt. Lett. 40, 4691 (2015).
Yao, J. et al. Multiscale photoacoustic tomography using reversibly switchable bacterial phytochrome as a near-infrared photochromic probe. Nat. Methods 13, 67–73 (2015).
Dortay, H. et al. Dual-wavelength photoacoustic imaging of a photoswitchable reporter protein Proc. SPIE, Photons Plus Ultrasound: Imaging and Sensing 9708, 970820 (2016).
Zienicke, B. et al. Fluorescence of phytochrome adducts with synthetic locked chromophores. J. Biol. Chem. 286, 1103–1113 (2011).
Lamparter, T., Michael, N., Mittmann, F. & Esteban, B. Phytochrome from Agrobacterium tumefaciens has unusual spectral properties and reveals an N-terminal chromophore attachment site. Proc. Natl. Acad. Sci. USA 99, 11628–11633 (2002).
Schumann, C. et al. Subpicosecond midinfrared spectroscopy of the Pfr reaction of phytochrome Agp1 from Agrobacterium tumefaciens. Biophys. J. 94, 3189–3197 (2008).
Wagner, J. R. et al. Mutational analysis of Deinococcus radiodurans bacteriophytochrome reveals key amino acids necessary for the photochromicity and proton exchange cycle of phytochromes. J. Biol. Chem. 283, 12212–12226 (2008).
Hale, G. M. & Querry, M. R. Optical constants of water in the 200-nm to 200-microm wavelength region. Appl. Opt. 12, 555–563 (1973).
Prahl, S. Optical Absorption of Haemoglobin. Available at: http://omlc.org/spectra/hemoglobin/ (1999).
Borucki, B. et al. Light-induced proton release of phytochrome is coupled to the transient deprotonation of the tetrapyrrole chromophore. J. Biol. Chem. 280, 34358–34364 (2005).
Märk, J., Wagener, A., Zhang, E. & Laufer, J. Photoacoustic pump-probe tomography of fluorophores in vivo using interleaved image acquisition for motion suppression. Sci. Rep. 7, 40496 (2017).
Zhang, E., Laufer, J. & Beard, P. Backward-mode multiwavelength photoacoustic scanner using a planar Fabry-Perot polymer film ultrasound sensor for high-resolution three-dimensional imaging of biological tissues. Appl. Opt. 47, 561–577 (2008).
Huynh, N., Ogunlade, O., Zhang, E., Cox, B. & Beard, P. Photoacoustic imaging using an 8-beam Fabry-Perot scanner. Proc. SPIE 9708, 97082L (2016).
Shcherbakova, D. M., Baloban, M. & Verkhusha, V. V. Near-infrared fluorescent proteins engineered from bacterial phytochromes. Curr. Opin. Chem. Biol. 27, 52–63 (2015).
Kaberniuk, A. A., Shemetov, A. A. & Verkhusha, V. V. A bacterial phytochrome-based optogenetic system controllable with near-infrared light. Nat. Methods 13, 591–597 (2016).
Salewski, J. et al. Structure of the biliverdin cofactor in the Pfr state of bathy and prototypical phytochromes. J. Biol. Chem. 288, 16800–16814 (2013).
Velazquez Escobar, F. et al. Structural parameters controlling the fluorescence properties of phytochromes. Biochemistry 53, 20–29 (2014).
Weber, K., Bartsch, U., Stocking, C. & Fehse, B. A multicolor panel of novel lentiviral ‘Gene Ontology’ (LeGO) vectors for functional gene analysis. Mol. Ther. 16, 698–706 (2008).
Dull, T. et al. A third-generation lentivirus vector with a conditional packaging system. J. Virol. 72, 8463–8471 (1998).
Stewart, S. A. et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA 9, 493–501 (2003).
Treeby, B. E., Varslot, T. K., Zhang, E. Z., Laufer, J. G. & Beard, P. C. Automatic sound speed selection in photoacoustic image reconstruction using an autofocus approach. J. Biomed. Opt. 16, 90501 (2011).
Treeby, B. E. & Cox, B. T. k-Wave: MATLAB toolbox for the simulation and reconstruction of photoacoustic wave fields. J. Biomed. Opt. 15, 21314 (2010).
Treeby, B. E., Zhang, E. Z. & Cox, B. T. Photoacoustic tomography in absorbing acoustic media using time reversal. Inverse Probl. 26, 115003 (2010).
Treeby, B. E. Acoustic attenuation compensation in photoacoustic tomography using time-variant filtering. J. Biomed. Opt. 18, 36008 (2013).
Acknowledgements
This work was funded by the European Research Commission (ERC) Starting Grant 281356 and the Deutsche Forschungsgemeinschaft (DFG) project grant LA3273/1-1 (J.L.), Cluster of Excellence 'Unifying concepts in Catalysis', and the Federal Ministry of Education and Research (WTZ-RUS grant 01DJ15007). The authors would like to thank the flow cytometry unit of the Berlin Center for Regenerative Therapies for cell sorting, Norbert Michael and Peter Hildebrandt for the pET21b-AGP1 expression vector and the extinction spectra of AGP1, and Yvonne Griesecke for assisting with microscopy.
Author information
Authors and Affiliations
Contributions
J.M. conducted PA experiments in vitro and in vivo, and assisted with preparation of the manuscript. H.D. designed and constructed the plasmids and viral vectors, purified proteins, and transduced cells. A.W. carried out cell preparation and sorting, tumour inoculations, and in vitro and in vivo fluorescence tomography imaging. E.Z. developed the software for instrumentation control. J.B. conducted in vivo experiments. C.G. contributed to planning of the in vivo study and the preparation of the manuscript. T.F. contributed to the in vitro characterisation of the proteins and the preparation of the manuscript. J.L. conceived and designed the study, carried out reconstruction, processing, and rendering of in vivo images, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Märk, J., Dortay, H., Wagener, A. et al. Dual-wavelength 3D photoacoustic imaging of mammalian cells using a photoswitchable phytochrome reporter protein. Commun Phys 1, 3 (2018). https://doi.org/10.1038/s42005-017-0003-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42005-017-0003-2
This article is cited by
-
Photoacoustic Tomography with Temporal Encoding Reconstruction (PATTERN) for cross-modal individual analysis of the whole brain
Nature Communications (2024)
-
Genetically encoded photo-switchable molecular sensors for optoacoustic and super-resolution imaging
Nature Biotechnology (2022)
-
Photoacoustic in vivo 3D imaging of tumor using a highly tumor-targeting probe under high-threshold conditions
Scientific Reports (2020)
-
Small near-infrared photochromic protein for photoacoustic multi-contrast imaging and detection of protein interactions in vivo
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.